Acyl-Homoserine Lactone Quorum Sensing in the Roseobacter Clade
Abstract
:1. Introduction
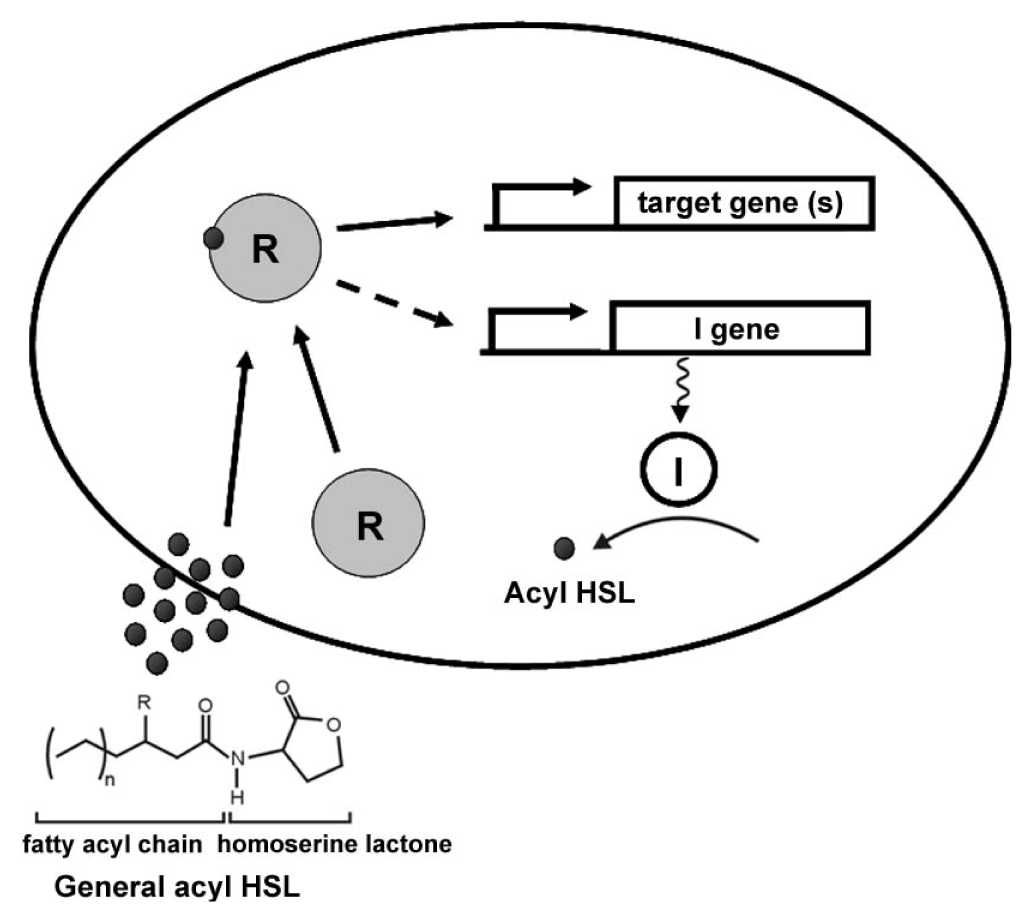
2. General Mechanisms of AHL Quorum Sensing
3. AHL Production in the Roseobacters
4. QS in Phaeobacter inhibens DSM17395
5. QS in the Marine Sponge Symbiont Ruegeria sp. KLH11
6. QS in the Dinoflagellate Symbiont D. shibae
7. LuxI-LuxR Pathways in the Roseobacter Clade: A Bioinformatics Perspective
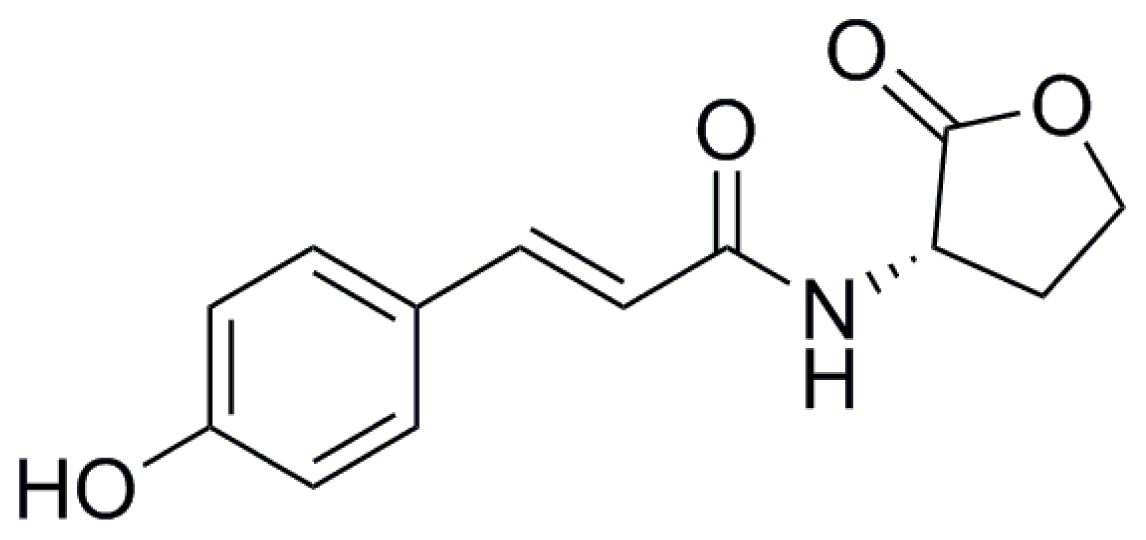
8. New Structural Variants of AHL
9. QS Control of Virulence Factors and Secondary Metabolites
10. Future Directions
11. Conclusions
Supplementary Information
ijms-15-00654-s001.pdfAcknowledgments
Conflicts of Interest
References
- Wagner-Döbler, I.; Thiel, V.; Eberl, L.; Allgaier, M.; Bodor, A.; Meyer, S.; Ebner, S.; Hennig, A.; Pukall, R.; Schulz, S. Discovery of complex mixtures of novel long-chain quorum sensing signals in free-living and host-associated marine alphaproteobacteria. ChemBioChem 2005, 6, 2195–2206. [Google Scholar]
- Buchan, A.; Gonzalez, J.M.; Moran, M.A. Overview of the marine Roseobacter lineage. Appl. Environ. Microbiol 2005, 71, 5666–5677. [Google Scholar]
- Moran, M.A.; Belas, R.; Schell, M.A.; Gonzalez, J.M.; Sun, F.; Sun, S.; Binder, B.J.; Edmonds, J.; Ye, W.; Orcutt, B.; et al. Ecological genomics of marine. Roseobacters. Appl. Environ. Microbiol 2007, 73, 4559–4569. [Google Scholar]
- Moran, M.A.; Buchan, A.; Gonzalez, J.M.; Heidelberg, J.F.; Whitman, W.B.; Kiene, R.P.; Henriksen, J.R.; King, G.M.; Belas, R.; Fuqua, C.; et al. Genome sequence of Silicibacter pomeroyi reveals adaptations to the marine environment. Nature 2004, 432, 910–913. [Google Scholar]
- Wagner-Dobler, I.; Biebl, H. Environmental biology of the marine Roseobacter lineage. Annu. Rev. Microbiol 2006, 60, 255–280. [Google Scholar]
- Mohamed, N.M.; Cicirelli, E.M.; Kan, J.; Chen, F.; Fuqua, C.; Hill, R.T. Diversity and quorum-sensing signal production of Proteobacteria associated with marine sponges. Environ. Microbiol 2008, 10, 75–86. [Google Scholar]
- Slightom, R.N.; Buchan, A. Surface colonization by marine Roseobacters: Integrating genotype and phenotype. Appl. Environ. Microbiol 2009, 75, 6027–6037. [Google Scholar]
- Taylor, M.W.; Radax, R.; Steger, D.; Wagner, M. Sponge-associated microorganisms: Evolution, ecology, and biotechnological potential. Microbiol. Mol. Biol. Rev 2007, 71, 295–347. [Google Scholar]
- Ng, W.L.; Bassler, B.L. Bacterial quorum-sensing network architectures. Annu. Rev. Genet 2009, 43, 197–222. [Google Scholar]
- Moran, M.A.; Reisch, C.R.; Kiene, R.P.; Whitman, W.B. Genomic insights into bacterial DMSP transformations. Ann. Rev. Mar. Sci 2012, 4, 523–542. [Google Scholar]
- Fuqua, C.; Greenberg, E.P. Listening in on bacteria: Acyl-homoserine lactone signalling. Nat. Rev. Mol. Cell Biol 2002, 3, 685–695. [Google Scholar]
- Blackwell, H.E.; Fuqua, C. Introduction to bacterial signals and chemical communication. Chem. Rev 2011, 111, 1–3. [Google Scholar]
- Miller, M.B.; Bassler, B.L. Quorum sensing in bacteria. Annu. Rev. Microbiol 2001, 55, 165–199. [Google Scholar]
- Fuqua, C.; Parsek, M.R.; Greenberg, E.P. Regulation of gene expression by cell-to-cell communication: Acyl-homoserine lactone quorum sensing. Annu. Rev. Genet 2001, 35, 439–468. [Google Scholar]
- Moré, M.I.; Finger, L.D.; Stryker, J.L.; Fuqua, C.; Eberhard, A.; Winans, S.C. Enzymatic synthesis of a quorum-sensing autoinducer through use of defined substrates. Science 1996, 272, 1655–1658. [Google Scholar]
- Schaefer, A.L.; Hanzelka, B.L.; Eberhard, A.; Greenberg, E.P. Quorum sensing in Vibrio fischeri: Probing autoinducer—LuxR interactions with autoinducer analogs. J. Bacteriol 1996, 178, 2897–2901. [Google Scholar]
- Zan, J.; Cicirelli, E.M.; Mohamed, N.M.; Sibhatu, H.; Kroll, S.; Choi, O.; Uhlson, C.L.; Wysoczynski, C.L.; Murphy, R.C.; Churchill, M.E.; et al. A complex LuxR-LuxI type quorum sensing network in a roseobacterial marine sponge symbiont activates flagellar motility and inhibits biofilm formation. Mol. Microbiol 2012, 85, 916–933. [Google Scholar]
- Churchill, M.E.; Chen, L. Structural basis of acyl-homoserine lactone-dependent signaling. Chem. Rev 2011, 111, 68–85. [Google Scholar]
- Subramoni, S.; Venturi, V. LuxR-family “solos”: Bachelor sensors/regulators of signalling molecules. Microbiology 2009, 155, 1377–1385. [Google Scholar]
- Zhu, J.; Winans, S.C. The quorum-sensing transcriptional regulator TraR requires its cognate signaling ligand for protein folding, protease resistance, and dimerization. Proc. Natl. Acad. Sci. USA 2001, 98, 1507–1512. [Google Scholar]
- Choi, S.H.; Greenberg, E.P. Genetic dissection of DNA binding and luminescence gene activation by the Vibrio fischeri LuxR protein. J. Bacteriol 1992, 174, 4064–4069. [Google Scholar]
- Kiratisin, P.; Tucker, K.D.; Passador, L. LasR, a transcriptional activator of Pseudomonas aeruginosa virulence genes, functions as a multimer. J. Bacteriol 2002, 184, 4912–4919. [Google Scholar]
- Stevens, A.M.; Greenberg, E.P. Transcriptional activation by LuxR. In Cell-Cell Signaling in Bacteria; Dunny, G.M., Winans, S.C., Eds.; ASM Press: Washington, DC, USA, 1999; p. 231. [Google Scholar]
- Tsai, C.S.; Winans, S.C. LuxR-type quorum-sensing regulators that are detached from common scents. Mol. Microbiol 2010, 77, 1072–1082. [Google Scholar]
- Thole, S.; Kalhoefer, D.; Voget, S.; Berger, M.; Engelhardt, T.; Liesegang, H.; Wollherr, A.; Kjelleberg, S.; Daniel, R.; Simon, M.; et al. Phaeobacter gallaeciensis genomes from globally opposite locations reveal high similarity of adaptation to surface life. ISME J 2012, 6, 2229–2244. [Google Scholar]
- Berger, M.; Neumann, A.; Schulz, S.; Simon, M.; Brinkhoff, T. Tropodithietic acid production in Phaeobacter gallaeciensis is regulated by N-acyl homoserine lactone-mediated quorum sensing. J. Bacteriol 2011, 193, 6576–6585. [Google Scholar]
- Prol Garcia, M.J.; D’Alvise, P.W.; Gram, L. Disruption of cell-to-cell signaling does not abolish the antagonism of Phaeobacter gallaeciensis toward the fish pathogen Vibrio anguillarum in algal systems. Appl. Environ. Microbiol 2013, 79, 5414–5417. [Google Scholar]
- Berger, M.; Brock, N.L.; Liesegang, H.; Dogs, M.; Preuth, I.; Simon, M.; Dickschat, J.S.; Brinkhoff, T. Genetic analysis of the upper phenylacetate catabolic pathway in the production of tropodithietic acid by Phaeobacter gallaeciensis. Appl. Environ. Microbiol 2012, 78, 3539–3551. [Google Scholar]
- Zan, J.; Heindl, J.E.; Liu, Y.; Fuqua, C.; Hill, R.T. The CckA-ChpT-CtrA phosphorelay system is regulated by quorum sensing and controls flagellar motility in the marine sponge symbiont Ruegeria sp. KLH11. PLoS One 2013, 8, e66346. [Google Scholar]
- Zan, J.; Fricke, W.F.; Fuqua, C.; Ravel, J.; Hill, R.T. Genome sequence of Ruegeria sp. strain KLH11, an N-acylhomoserine lactone-producing bacterium isolated from the marine sponge Mycale laxissima. J. Bacteriol 2011, 193, 5011–5012. [Google Scholar]
- Biebl, H.; Allgaier, M.; Tindall, B.J.; Koblizek, M.; Lunsdorf, H.; Pukall, R.; Wagner-Dobler, I. Dinoroseobacter shibae gen. nov., sp. nov., a new aerobic phototrophic bacterium isolated from dinoflagellates. Int. J. Syst. Evol. Microbiol 2005, 55, 1089–1096. [Google Scholar]
- Wagner-Döbler, I.; Ballhausen, B.; Berger, M.; Brinkhoff, T.; Buchholz, I.; Bunk, B.; Cypionka, H.; Daniel, R.; Drepper, T.; Gerdts, G.; et al. The complete genome sequence of the algal symbiont Dinoroseobacter shibae: A hitchhiker’s guide to life in the sea. ISME J 2010, 4, 61–77. [Google Scholar]
- Patzelt, D.; Wang, H.; Buchholz, I.; Rohde, M.; Grobe, L.; Pradella, S.; Neumann, A.; Schulz, S.; Heyber, S.; Munch, K.; et al. You are what you talk: Quorum sensing induces individual morphologies and cell division modes in Dinoroseobacter shibae. ISME J 2013, 7, 2274–2286. [Google Scholar]
- Buddruhs, N.; Pradella, S.; Goker, M.; Pauker, O.; Pukall, R.; Sproer, C.; Schumann, P.; Petersen, J.; Brinkhoff, T. Molecular and phenotypic analyses reveal the non-identity of the Phaeobacter gallaeciensis type strain deposits CIP 105210T and DSM 17395. Int. J. Syst. Evol. Microbiol 2013, 63, 4340–4349. [Google Scholar]
- Gram, L.; Grossart, H.P.; Schlingloff, A.; Kiorboe, T. Possible quorum sensing in marine snow bacteria: Production of acylated homoserine lactones by Roseobacter strains isolated from marine snow. Appl. Environ. Microbiol 2002, 68, 4111–4116. [Google Scholar]
- Bruhn, J.B.; Gram, L.; Belas, R. Production of antibacterial compounds and biofilm formation by Roseobacter species are influenced by culture conditions. Appl. Environ. Microbiol 2007, 73, 442–450. [Google Scholar]
- Taylor, M.W.; Schupp, P.J.; Baillie, H.J.; Charlton, T.S.; de Nys, R.; Kjelleberg, S.; Steinberg, P.D. Evidence for acyl homoserine lactone signal production in bacteria associated with marine sponges. Appl. Environ. Microbiol 2004, 70, 4387–4389. [Google Scholar]
- Zan, J. Quorum sensing in bacteria associated with marine sponges Mycale laxissima and Ircinia strobilina. Ph.D Dissertation, University of Maryland, College Park, MD, USA, May 2013. [Google Scholar]
- Geng, H.; Belas, R. TdaA regulates tropodithietic acid synthesis by binding to the tdaC promoter region. J. Bacteriol 2011, 193, 4002–4005. [Google Scholar]
- Zan, J.; Choi, O.; Meharena, H.; Uhlson, C.L.; Churchill, M.E.A.; Hill, R.T.; Fuqua, C. A solo luxI-type gene directs acylhomoserine lactone synthesis and contributes to motility control in the marine sponge symbiont Ruegeria sp. KLH11. Microbiology 2013. to be submitted for publication. [Google Scholar]
- Yates, E.A.; Philipp, B.; Buckley, C.; Atkinson, S.; Chhabra, S.R.; Sockett, R.E.; Goldner, M.; Dessaux, Y.; Camara, M.; Smith, H.; et al. N-acylhomoserine lactones undergo lactonolysis in a pH-, temperature-, and acyl chain length-dependent manner during growth of Yersinia pseudotuberculosis and Pseudomonas aeruginosa. Infect. Immun 2002, 70, 5635–5646. [Google Scholar]
- Riebesell, U.; Zondervan, I.; Rost, B.; Tortell, P.D.; Zeebe, R.E.; Morel, F.M. Reduced calcification of marine plankton in response to increased atmospheric CO2. Nature 2000, 407, 364–367. [Google Scholar]
- Capra, E.J.; Laub, M.T. Evolution of two-component signal transduction systems. Annu. Rev. Microbiol 2012, 66, 325–347. [Google Scholar]
- Kim, J.; Heindl, J.E.; Fuqua, C. Coordination of division and development influences complex multicellular behavior in Agrobacterium tumefaciens. PLoS One 2013, 8, e56682. [Google Scholar]
- Miller, T.R.; Hnilicka, K.; Dziedzic, A.; Desplats, P.; Belas, R. Chemotaxis of Silicibacter sp. strain TM1040 toward dinoflagellate products. Appl. Environ. Microbiol 2004, 70, 4692–4701. [Google Scholar]
- Hammer, B.K.; Bassler, B.L. Quorum sensing controls biofilm formation in Vibrio cholerae. Mol. Microbiol 2003, 50, 101–104. [Google Scholar]
- Zhu, J.; Mekalanos, J.J. Quorum sensing-dependent biofilms enhance colonization in Vibrio cholerae. Dev. Cell 2003, 5, 647–656. [Google Scholar]
- Puskas, A.; Greenberg, E.P.; Kaplan, S.; Schaefer, A.L. A quorum-sensing system in the free-living photosynthetic bacterium Rhodobacter sphaeroides. J. Bacteriol 1997, 179, 7530–7537. [Google Scholar]
- Veening, J.W.; Smits, W.K.; Kuipers, O.P. Bistability, epigenetics, and bet-hedging in bacteria. Annu. Rev. Microbiol 2008, 62, 193–210. [Google Scholar]
- Gould, T.A.; Schweizer, H.P.; Churchill, M.E. Structure of the Pseudomonas aeruginosa acyl-homoserinelactone synthase LasI. Mol. Microbiol 2004, 53, 1135–1146. [Google Scholar]
- Watson, W.T.; Minogue, T.D.; Val, D.L.; von Bodman, S.B.; Churchill, M.E. Structural basis and specificity of acyl-homoserine lactone signal production in bacterial quorum sensing. Mol. Cell 2002, 9, 685–694. [Google Scholar]
- Schaefer, A.L.; Greenberg, E.P.; Oliver, C.M.; Oda, Y.; Huang, J.J.; Bittan-Banin, G.; Peres, C.M.; Schmidt, S.; Juhaszova, K.; Sufrin, J.R.; et al. A new class of homoserine lactone quorum-sensing signals. Nature 2008, 454, 595–599. [Google Scholar]
- Ahlgren, N.A.; Harwood, C.S.; Schaefer, A.L.; Giraud, E.; Greenberg, E.P. Aryl-homoserine lactone quorum sensing in stem-nodulating photosynthetic bradyrhizobia. Proc. Natl. Acad. Sci. USA 2011, 108, 7183–7188. [Google Scholar]
- Lindemann, A.; Pessi, G.; Schaefer, A.L.; Mattmann, M.E.; Christensen, Q.H.; Kessler, A.; Hennecke, H.; Blackwell, H.E.; Greenberg, E.P.; Harwood, C.S. Isovaleryl-homoserine lactone, an unusual branched-chain quorum-sensing signal from the soybean symbiont Bradyrhizobium japonicum. Proc. Natl. Acad. Sci. USA 2011, 108, 16765–16770. [Google Scholar]
- Seyedsayamdost, M.R.; Case, R.J.; Kolter, R.; Clardy, J. The Jekyll-and-Hyde chemistry of Phaeobacter gallaeciensis. Nat. Chem. Biol 2011, 4, 331–335. [Google Scholar]
- Sule, P.; Belas, R. A novel inducer of Roseobacter motility is also a disruptor of algal symbiosis. J. Bacteriol 2013, 195, 637–646. [Google Scholar]
- Boettcher, K.J.; Barber, B.J.; Singer, J.T. Use of antibacterial agents to elucidate the etiology of juvenile oyster disease (JOD) in Crassostrea virginica and numberical dominance of an alpha-proteobacterium in JOD-affected animals. Appl. Environ. Microbiol 1999, 65, 2534–2539. [Google Scholar]
- Boettcher, K.J.; Barber, B.J.; Singer, J.T. Additional evidence that juvenile oyster diease is caused by a member of the Roseobacter group and colonization of nonaffected animals by Stappia stellulata-like strains. Appl. Environ. Microbiol 2000, 66, 3924–3930. [Google Scholar]
- Fernandes, N.; Case, R.J.; Longford, S.R.; Seyedsayamdost, M.R.; Steinberg, P.D.; Kjelleberg, S.; Thomas, T. Genomes and virulence factors of novel bacterial pathogens causing bleaching disease in the marine red alga Delisea pulchra. PLoS One 2011, 6, e27387. [Google Scholar]
- Martens, T.; Gram, L.; Grossart, H.P.; Kessler, D.; Muller, R.; Simon, M.; Wenzel, S.C.; Brinkoff, T. Bacteria of the Roseobacter clade show potential for secondary metabolite production. Mol. Ecol 2007, 54, 31–42. [Google Scholar]
- Cude, W.N.; Mooney, J.; Tavanaei, A.A.; Hadden, M.K.; Frank, A.M.; Gulvik, C.A.; May, A.L.; Buchan, A. Production of the antimicrobial secondary metabolite indigoidine contributes to competitive surface colonization by the marine roseobacter Phaeobacter sp. strain Y4I. Appl. Environ. Microbiol 2012, 78, 4771–4780. [Google Scholar]
- Ruby, E.G. Symbiotic conversations are revealed under genetic interrogation. Nat. Rev. Microbiol 2008, 6, 752–762. [Google Scholar]
- Cude, W.N.; Buchan, A. Acyl-homoserine lactone-based quorum sensing in the Roseobacter clade: Complex cell-to-cell communication controls multiple physiologies. Front Microbiol 2013, 4, 336. [Google Scholar]

| Species | LuxI-LuxR | Isolation source | Signal profile | Function | Ref |
|---|---|---|---|---|---|
| P. inhibens DSM17395 | pgaRI | coastal water | 3-OH-C10-HSL | Temporal regulation of TDA production | [25–28] |
| Ruegeria sp. KLH11 | ssaRI, ssbRI sscI | marine sponge Mycale laxissima | 3-OH-C14-HSL, 3-OH-C14:1-HSL, 3-OH-C12-HSL | Activation of flagellar synthesis and swimming motility and inhibition of biofilm formation | [17,29,30] |
| D. shibae DFL12T | luxR1I1, luxR2I2, luxI3 | Dinoflagellate Prorocentrum lima | 3-C18en-HSL | Cell morphology and flagellar biosynthesis | [31–33] |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zan, J.; Liu, Y.; Fuqua, C.; Hill, R.T. Acyl-Homoserine Lactone Quorum Sensing in the Roseobacter Clade. Int. J. Mol. Sci. 2014, 15, 654-669. https://doi.org/10.3390/ijms15010654
Zan J, Liu Y, Fuqua C, Hill RT. Acyl-Homoserine Lactone Quorum Sensing in the Roseobacter Clade. International Journal of Molecular Sciences. 2014; 15(1):654-669. https://doi.org/10.3390/ijms15010654
Chicago/Turabian StyleZan, Jindong, Yue Liu, Clay Fuqua, and Russell T. Hill. 2014. "Acyl-Homoserine Lactone Quorum Sensing in the Roseobacter Clade" International Journal of Molecular Sciences 15, no. 1: 654-669. https://doi.org/10.3390/ijms15010654
APA StyleZan, J., Liu, Y., Fuqua, C., & Hill, R. T. (2014). Acyl-Homoserine Lactone Quorum Sensing in the Roseobacter Clade. International Journal of Molecular Sciences, 15(1), 654-669. https://doi.org/10.3390/ijms15010654




