Multiple Functions of the RNA-Binding Protein HuR in Cancer Progression, Treatment Responses and Prognosis
Abstract
:1. Introduction
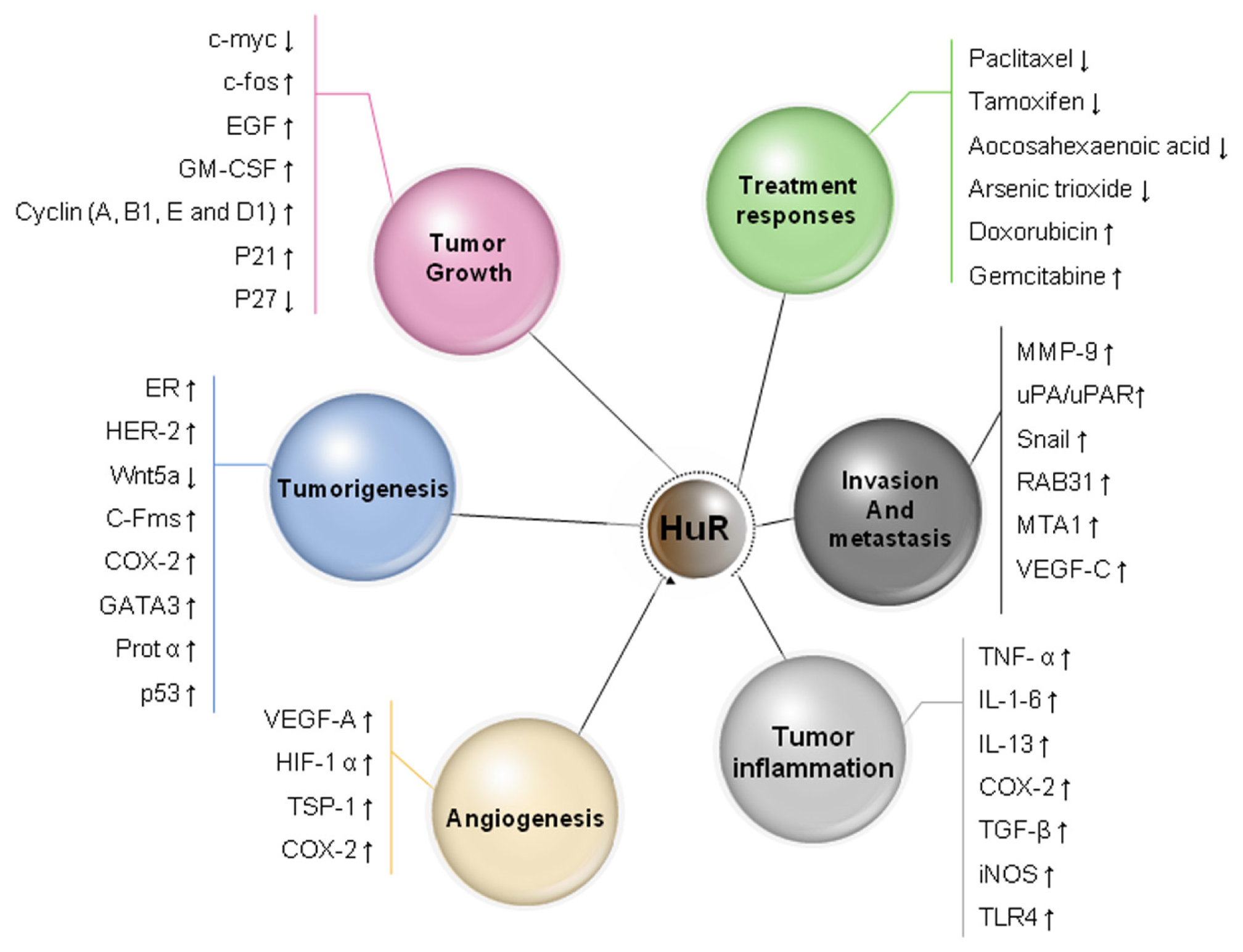
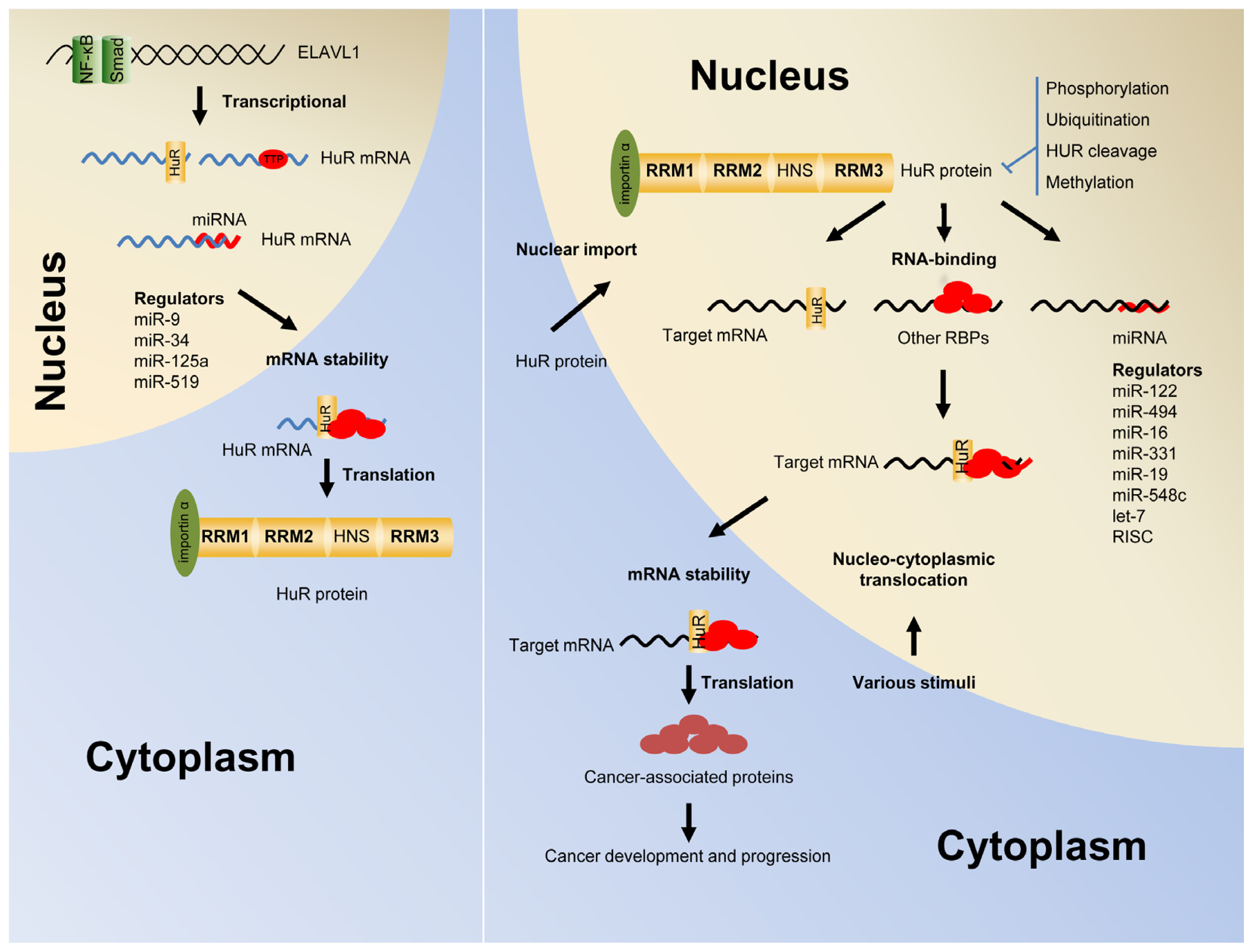
2. Post-Transcriptional Regulation of Gene Expression by HuR
3. Shuttling of HuR from the Nucleus into the Cytoplasm
4. The Autoregulation of HuR Function and Expression
5. HuR Expression in Cancer
6. HuR Expression in Pre-Malignant Lesions
7. HuR Function in Tumor Angiogenesis
8. HuR Function in Cancer Invasion and Metastasis
9. HuR and Drug Resistance and Sensitivity
10. Prognostic Significance of HuR in Human Carcinoma
11. Conclusions
Acknowledgments
Conflict of Interest
References
- Campos, A.R.; Grossman, D.; White, K. Mutant alleles at the locus elav in Drosophila melanogaster lead to nervous system defects. A developmental-genetic analysis. J. Neurogenet 1985, 2, 197–218. [Google Scholar]
- Ma, W.J.; Cheng, S.; Campbell, C.; Wright, A.; Furneaux, H. Cloning and characterization of HuR, a ubiquitously expressed Elav-like protein. J. Biol. Chem 1996, 271, 8144–8151. [Google Scholar]
- Ma, W.J.; Furneaux, H. Localization of the human HuR gene to chromosome 19p13.2. Hum. Genet 1997, 99, 32–33. [Google Scholar]
- Robinow, S.; Campos, A.R.; Yao, K.M.; White, K. The elav gene product of Drosophila, required in neurons, has three RNP consensus motifs. Science 1988, 242, 1570–1572. [Google Scholar]
- Dalmau, J.; Furneaux, H.M.; Cordon-Cardo, C.; Posner, J.B. The expression of the Hu (paraneoplastic encephalomyelitis/sensory neuronopathy) antigen in human normal and tumor tissues. Am. J. Pathol 1992, 141, 881–886. [Google Scholar]
- King, P.H.; Dropcho, E.J. Expression of Hel-N1 and Hel-N2 in small-cell lung carcinoma. Ann. Neurol 1996, 39, 679–681. [Google Scholar]
- King, P.H. Differential expression of the neuroendocrine genes Hel-N1 and HuD in small-cell lung carcinoma: Evidence for down-regulation of HuD in the variant phenotype. Int. J. Cancer 1997, 74, 378–382. [Google Scholar]
- Srikantan, S.; Gorospe, M. HuR function in disease. Front. Biosci 2012, 17, 189–205. [Google Scholar]
- Von Roretz, C.; di Marco, S.; Mazroui, R.; Gallouzi, I.E. Turnover of AU-rich-containing mRNAs during stress: A matter of survival. Wiley Interdiscip. Rev. RNA 2011, 2, 336–347. [Google Scholar]
- Abdelmohsen, K.; Gorospe, M. Posttranscriptional regulation of cancer traits by HuR. Wiley Interdiscip. Rev. RNA 2010, 1, 214–229. [Google Scholar]
- Xiao, L.; Wang, J.Y. Posttranscriptional regulation of gene expression in epithelial cells by polyamines. Methods Mol. Biol 2011, 720, 67–79. [Google Scholar]
- Carpenter, B.; MacKay, C.; Alnabulsi, A.; MacKay, M.; Telfer, C.; Melvin, W.T.; Murray, G.I. The roles of heterogeneous nuclear ribonucleoproteins in tumour development and progression. Biochim. Biophys. Acta 2006, 1765, 85–100. [Google Scholar]
- Zhou, Z.J.; Dai, Z.; Zhou, S.L.; Fu, X.T.; Zhao, Y.M.; Shi, Y.H.; Zhou, J.; Fan, J. Overexpression of HnRNP A1 promotes tumor invasion through regulating CD44v6 and indicates poor prognosis for hepatocellular carcinoma. Int. J. Cancer 2013, 132, 1080–1089. [Google Scholar]
- Wu, S.; Sato, M.; Endo, C.; Sakurada, A.; Dong, B.; Aikawa, H.; Chen, Y.; Okada, Y.; Matsumura, Y.; Sueoka, E.; et al. hnRNP B1 protein may be a possible prognostic factor in squamous cell carcinoma of the lung. Lung Cancer 2003, 41, 179–186. [Google Scholar]
- Hope, N.R.; Murray, G.I. The expression profile of RNA-binding proteins in primary and metastatic colorectal cancer: Relationship of heterogeneous nuclear ribonucleoproteins with prognosis. Hum. Pathol 2011, 42, 393–402. [Google Scholar]
- Carpenter, B.; McKay, M.; Dundas, S.R.; Lawrie, L.C.; Telfer, C.; Murray, G.I. Heterogeneous nuclear ribonucleoprotein K is over expressed, aberrantly localised and is associated with poor prognosis in colorectal cancer. Br. J. Cancer 2006, 95, 921–927. [Google Scholar]
- Papadopoulou, C.; Ganou, V.; Patrinou-Georgoula, M.; Guialis, A. HuR-hnRNP interactions and the effect of cellular stress. Mol. Cell Biochem 2013, 372, 137–147. [Google Scholar]
- Srikantan, S.; Tominaga, K.; Gorospe, M. Functional interplay between RNA-binding protein HuR and microRNAs. Curr. Protein Pept. Sci 2012, 13, 372–379. [Google Scholar]
- Bhattacharyya, S.N.; Habermacher, R.; Martine, U.; Closs, E.I.; Filipowicz, W. Relief of microRNA-mediated translational repression in human cells subjected to stress. Cell 2006, 125, 1111–1124. [Google Scholar]
- Srikantan, S.; Abdelmohsen, K.; Lee, E.K.; Tominaga, K.; Subaran, S.S.; Kuwano, Y.; Kulshrestha, R.; Panchakshari, R.; Kim, H.H.; Yang, X.; et al. Translational control of TOP2A influences doxorubicin efficacy. Mol. Cell Biol 2011, 31, 3790–3801. [Google Scholar]
- Tominaga, K.; Srikantan, S.; Lee, E.K.; Subaran, S.S.; Martindale, J.L.; Abdelmohsen, K.; Gorospe, M. Competitive regulation of nucleolin expression by HuR and miR-494. Mol. Cell Biol 2011, 31, 4219–4231. [Google Scholar]
- Wang, W.; Caldwell, M.C.; Lin, S.; Furneaux, H.; Gorospe, M. HuR regulates cyclin A and cyclin B1 mRNA stability during cell proliferation. EMBO J 2000, 19, 2340–2350. [Google Scholar]
- Epis, M.R.; Barker, A.; Giles, K.M.; Beveridge, D.J.; Leedman, P.J. The RNA-binding protein HuR opposes the repression of ERBB-2 gene expression by microRNA miR-331-3p in prostate cancer cells. J. Biol. Chem 2011, 286, 41442–41454. [Google Scholar]
- Kim, H.H.; Kuwano, Y.; Srikantan, S.; Lee, E.K.; Martindale, J.L.; Gorospe, M. HuR recruits let-7/RISC to repress c-Myc expression. Genes Dev 2009, 23, 1743–1748. [Google Scholar]
- Glorian, V.; Maillot, G.; Poles, S.; Iacovoni, J.S.; Favre, G.; Vagner, S. HuR-dependent loading of miRNA RISC to the mRNA encoding the Ras-related small GTPase RhoB controls its translation during UV-induced apoptosis. Cell Death Differ 2011, 18, 1692–1701. [Google Scholar]
- Chang, N.; Yi, J.; Guo, G.; Liu, X.; Shang, Y.; Tong, T.; Cui, Q.; Zhan, M.; Gorospe, M.; Wang, W. HuR uses AUF1 as a cofactor to promote p16INK4 mRNA decay. Mol. Cell Biol 2010, 30, 3875–3886. [Google Scholar]
- Li, B.; Si, J.; DeWille, J.W. Ultraviolet radiation (UVR) activates p38 MAP kinase and induces post-transcriptional stabilization of the C/EBPdelta mRNA in G0 growth arrested mammary epithelial cells. J. Cell Biochem 2008, 103, 1657–1669. [Google Scholar]
- Wang, W.; Furneaux, H.; Cheng, H.; Caldwell, M.C.; Hutter, D.; Liu, Y.; Holbrook, N.; Gorospe, M. HuR regulates p21 mRNA stabilization by UV light. Mol. Cell Biol 2000, 20, 760–769. [Google Scholar]
- Nagy, L.E. Stabilization of tumor necrosis factor-alpha mRNA in macrophages in response to chronic ethanol exposure. Alcohol 2004, 33, 229–233. [Google Scholar]
- McMullen, M.R.; Cocuzzi, E.; Hatzoglou, M.; Nagy, L.E. Chronic ethanol exposure increases the binding of HuR to the TNFalpha 3′-untranslated region in macrophages. J. Biol. Chem 2003, 278, 38333–38341. [Google Scholar]
- Lin, W.N.; Lin, C.C.; Cheng, H.Y.; Yang, C.M. Regulation of cyclooxygenase-2 and cytosolic phospholipase A2 gene expression by lipopolysaccharide through the RNA-binding protein HuR: Involvement of NADPH oxidase reactive oxygen species and mitogen-activated protein kinases. Br. J. Pharmacol 2011, 163, 1691–1706. [Google Scholar]
- Lin, F.Y.; Chen, Y.H.; Lin, Y.W.; Tsai, J.S.; Chen, J.W.; Wang, H.J.; Chen, Y.L.; Li, C.Y.; Lin, S.J. The role of human antigen R: An RNA-binding protein, in mediating the stabilization of toll-like receptor 4 mRNA induced by endotoxin. a novel mechanism involved in vascular inflammation. Arterioscler Thromb Vasc. Biol 2006, 26, 2622–2629. [Google Scholar]
- Hostetter, C.; Licata, L.A.; Witkiewicz, A.; Costantino, C.L.; Yeo, C.J.; Brody, J.R.; Keen, J.C. Cytoplasmic accumulation of the RNA binding protein HuR is central to tamoxifen resistance in estrogen receptor positive breast cancer cells. Cancer Biol. Ther 2008, 7, 1496–1506. [Google Scholar]
- Costantino, C.L.; Witkiewicz, A.K.; Kuwano, Y.; Cozzitorto, J.A.; Kennedy, E.P.; Dasgupta, A.; Keen, J.C.; Yeo, C.J.; Gorospe, M.; Brody, J.R. The role of HuR in gemcitabine efficacy in pancreatic cancer. HuR Up-regulates the expression of the gemcitabine metabolizing enzyme deoxycytidine kinase. Cancer Res 2009, 69, 4567–4572. [Google Scholar]
- Chen, L.; Jarujaron, S.; Wu, X.; Sun, L.; Zha, W.; Liang, G.; Wang, X.; Gurley, E.C.; Studer, E.J.; Hylemon, P.B.; et al. HIV protease inhibitor lopinavir-induced TNF-alpha and IL-6 expression is coupled to the unfolded protein response and ERK signaling pathways in macrophages. Biochem. Pharmacol 2009, 78, 70–77. [Google Scholar]
- Bonelli, M.A.; Alfieri, R.R.; Desenzani, S.; Petronini, P.G.; Borghetti, A.F. Proteasome inhibition increases HuR level, restores heat-inducible HSP72 expression and thermotolerance in WI-38 senescent human fibroblasts. Exp. Gerontol 2004, 39, 423–432. [Google Scholar]
- Levy, N.S.; Chung, S.; Furneaux, H.; Levy, A.P. Hypoxic stabilization of vascular endothelial growth factor mRNA by the RNA-binding protein HuR. J. Biol. Chem 1998, 273, 6417–6423. [Google Scholar]
- Yaman, I.; Fernandez, J.; Sarkar, B.; Schneider, R.J.; Snider, M.D.; Nagy, L.E.; Hatzoglou, M. Nutritional control of mRNA stability is mediated by a conserved AU-rich element that binds the cytoplasmic shuttling protein HuR. J. Biol. Chem 2002, 277, 41539–41546. [Google Scholar]
- Perrone, E.E.; Liu, L.; Turner, D.J.; Strauch, E.D. Bile salts increase epithelial cell proliferation through HuR-induced c-Myc expression. J. Surg. Res 2012, 178, 155–161. [Google Scholar]
- Atasoy, U.; Watson, J.; Patel, D.; Keene, J.D. ELAV protein HuA (HuR) can redistribute between nucleus and cytoplasm and is upregulated during serum stimulation and T cell activation. J. Cell Sci 1998, 111, 3145–3156. [Google Scholar]
- Zou, T.; Rao, J.N.; Liu, L.; Xiao, L.; Yu, T.X.; Jiang, P.; Gorospe, M.; Wang, J.Y. Polyamines regulate the stability of JunD mRNA by modulating the competitive binding of its 3′ untranslated region to HuR and AUF1. Mol. Cell Biol 2010, 30, 5021–5032. [Google Scholar]
- Zou, T.; Liu, L.; Rao, J.N.; Marasa, B.S.; Chen, J.; Xiao, L.; Zhou, H.; Gorospe, M.; Wang, J.Y. Polyamines modulate the subcellular localization of RNA-binding protein HuR through AMP-activated protein kinase-regulated phosphorylation and acetylation of importin alpha1. Biochem. J 2008, 409, 389–398. [Google Scholar]
- Zou, T.; Mazan-Mamczarz, K.; Rao, J.N.; Liu, L.; Marasa, B.S.; Zhang, A.H.; Xiao, L.; Pullmann, R.; Gorospe, M.; Wang, J.Y. Polyamine depletion increases cytoplasmic levels of RNA-binding protein HuR leading to stabilization of nucleophosmin and p53 mRNAs. J. Biol. Chem 2006, 281, 19387–19394. [Google Scholar]
- Serini, S.; Fasano, E.; Piccioni, E.; Monego, G.; Cittadini, A.R.; Celleno, L.; Ranelletti, F.O.; Calviello, G. DHA induces apoptosis and differentiation in human melanoma cells in vitro. involvement of HuR-mediated COX-2 mRNA stabilization and beta-catenin nuclear translocation. Carcinogenesis 2012, 33, 164–173. [Google Scholar]
- Cao, H.; Kelly, M.A.; Kari, F.; Dawson, H.D.; Urban, J.F., Jr.; Coves, S.; Roussel, A.M.; Anderson, R.A. Green tea increases anti-inflammatory tristetraprolin and decreases pro-inflammatory tumor necrosis factor mRNA levels in rats. J. Inflamm. 2007, 4. [Google Scholar] [CrossRef]
- Zhang, J.; Ouyang, W.; Li, J.; Zhang, D.; Yu, Y.; Wang, Y.; Li, X.; Huang, C. Suberoylanilide hydroxamic acid (SAHA) inhibits EGF-induced cell transformation via reduction of cyclin D1 mRNA stability. Toxicol. Appl. Pharmacol 2012, 263, 218–224. [Google Scholar]
- Hwang, Y.S.; Park, K.K.; Chung, W.Y. Kalopanaxsaponin A inhibits the invasion of human oral squamous cell carcinoma by reducing metalloproteinase-9 mRNA stability and protein trafficking. Biol. Pharm. Bull 2012, 35, 289–300. [Google Scholar]
- Sun, L.; Zhang, S.; Jiang, Z.; Huang, X.; Wang, T.; Li, H.; Zhang, L. Triptolide inhibits COX-2 expression by regulating mRNA stability in TNF-alpha-treated A549 cells. Biochem. Biophys. Res. Commun 2011, 416, 99–105. [Google Scholar]
- Suswam, E.A.; Nabors, L.B.; Huang, Y.; Yang, X.; King, P.H. IL-1beta induces stabilization of IL-8 mRNA in malignant breast cancer cells via the 3′ untranslated region. Involvement of divergent RNA-binding factors HuR, KSRP and TIAR. Int. J. Cancer 2005, 113, 911–919. [Google Scholar]
- Nabors, L.B.; Suswam, E.; Huang, Y.; Yang, X.; Johnson, M.J.; King, P.H. Tumor necrosis factor alpha induces angiogenic factor up-regulation in malignant glioma cells. A role for RNA stabilization and HuR. Cancer Res 2003, 63, 4181–4187. [Google Scholar]
- Chae, K.S.; Kang, M.J.; Lee, J.H.; Ryu, B.K.; Lee, M.G.; Her, N.G.; Ha, T.K.; Han, J.; Kim, Y.K.; Chi, S.G. Opposite functions of HIF-alpha isoforms in VEGF induction by TGF-beta1 under non-hypoxic conditions. Oncogene 2011, 30, 1213–1228. [Google Scholar]
- Cho, N.H.; Kang, S.; Hong, S.; An, H.J.; Choi, Y.H.; Jeong, G.B.; Choi, H.K. Elevation of cyclin B1, active cdc2, and HuR in cervical neoplasia with human papillomavirus type 18 infection. Cancer Lett 2006, 232, 170–178. [Google Scholar]
- Cumming, S.A.; Chuen-Im, T.; Zhang, J.; Graham, S.V. The RNA stability regulator HuR regulates L1 protein expression in vivo in differentiating cervical epithelial cells. Virology 2009, 383, 142–149. [Google Scholar]
- Dickson, A.M.; Anderson, J.R.; Barnhart, M.D.; Sokoloski, K.J.; Oko, L.; Opyrchal, M.; Galanis, E.; Wilusz, C.J.; Morrison, T.E.; Wilusz, J. Dephosphorylation of HuR Protein during alphavirus infection is associated with HuR relocalization to the cytoplasm. J. Biol. Chem 2012, 287, 36229–36238. [Google Scholar]
- Cherradi, N.; Lejczak, C.; Desroches-Castan, A.; Feige, J.J. Antagonistic functions of tetradecanoyl phorbol acetate-inducible-sequence 11b and HuR in the hormonal regulation of vascular endothelial growth factor messenger ribonucleic acid stability by adrenocorticotropin. Mol. Endocrinol 2006, 20, 916–930. [Google Scholar]
- Sheflin, L.G.; Zou, A.P.; Spaulding, S.W. Androgens regulate the binding of endogenous HuR to the AU-rich 3′UTRs of HIF-1alpha and EGF mRNA. Biochem. Biophys. Res. Commun 2004, 322, 644–651. [Google Scholar]
- Sheflin, L.G.; Zhang, W.; Spaulding, S.W. Androgen regulates the level and subcellular distribution of the AU-rich ribonucleic acid-binding protein HuR both in vitro and in vivo. Endocrinology 2001, 142, 2361–2368. [Google Scholar]
- Karipcin, F.S.; Ensari, T.A.; Kayisli, U.A.; Guzel, E.; Kallen, C.B.; Seli, E. The mRNA-binding protein HuR is regulated in the menstrual cycle and repressed in ectopic endometrium. Reprod. Sci 2011, 18, 145–155. [Google Scholar]
- Guttinger, S.; Muhlhausser, P.; Koller-Eichhorn, R.; Brennecke, J.; Kutay, U. Transportin2 functions as importin and mediates nuclear import of HuR. Proc. Natl. Acad. Sci. USA 2004, 101, 2918–2923. [Google Scholar]
- Fan, X.C.; Steitz, J.A. HNS: A nuclear-cytoplasmic shuttling sequence in HuR. Proc. Natl. Acad. Sci. USA 1998, 95, 15293–15298. [Google Scholar]
- Doller, A.; Huwiler, A.; Muller, R.; Radeke, H.H.; Pfeilschifter, J.; Eberhardt, W. Protein kinase C alpha-dependent phosphorylation of the mRNA-stabilizing factor HuR. implications for posttranscriptional regulation of cyclooxygenase-2. Mol. Biol. Cell 2007, 18, 2137–2148. [Google Scholar]
- Doller, A.; Akool, el-S.; Huwiler, A.; Muller, R.; Radeke, H.H.; Pfeilschifter, J.; Eberhardt, W. Posttranslational modification of the AU-rich element binding protein HuR by protein kinase Cdelta elicits angiotensin II-induced stabilization and nuclear export of cyclooxygenase 2 mRNA. Mol. Cell Biol 2008, 28, 2608–2625. [Google Scholar]
- Kim, H.H.; Abdelmohsen, K.; Lal, A.; Pullmann, R., Jr; Yang, X.; Galban, S.; Srikantan, S.; Martindale, J.L.; Blethrow, J.; Shokat, K.M.; et al. Nuclear HuR accumulation through phosphorylation by Cdk1. Genes Dev 2008, 22, 1804–1815. [Google Scholar]
- Abdelmohsen, K.; Pullmann, R., Jr; Lal, A.; Kim, H.H.; Galban, S.; Yang, X.; Blethrow, J.D.; Walker, M.; Shubert, J.; Gillespie, D.A.; et al. Phosphorylation of HuR by Chk2 regulates SIRT1 expression. Mol. Cell 2007, 25, 543–557. [Google Scholar]
- Lafarga, V.; Cuadrado, A.; Lopez de Silanes, I.; Bengoechea, R.; Fernandez-Capetillo, O.; Nebreda, A.R. p38 Mitogen-activated protein kinase- and HuR-dependent stabilization of p21(Cip1) mRNA mediates the G(1)/S checkpoint. Mol. Cell Biol 2009, 29, 4341–4351. [Google Scholar]
- Li, H.; Park, S.; Kilburn, B.; Jelinek, M.A.; Henschen-Edman, A.; Aswad, D.W.; Stallcup, M.R.; Laird-Offringa, I.A. Lipopolysaccharide-induced methylation of HuR, an mRNA-stabilizing protein, by CARM1. Coactivator-associated arginine methyltransferase. J. Biol. Chem 2002, 277, 44623–44630. [Google Scholar]
- Abdelmohsen, K.; Srikantan, S.; Yang, X.; Lal, A.; Kim, H.H.; Kuwano, Y.; Galban, S.; Becker, K.G.; Kamara, D.; de Cabo, R.; et al. Ubiquitin-mediated proteolysis of HuR by heat shock. EMBO J 2009, 28, 1271–1282. [Google Scholar]
- Wang, W.; Yang, X.; Kawai, T.; Lopez de Silanes, I.; Mazan-Mamczarz, K.; Chen, P.; Chook, Y.M.; Quensel, C.; Kohler, M.; Gorospe, M. AMP-activated protein kinase-regulated phosphorylation and acetylation of importin alpha1. Involvement in the nuclear import of RNA-binding protein HuR. J. Biol. Chem 2004, 279, 48376–48388. [Google Scholar]
- Wang, W.; Yang, X.; Lopez de Silanes, I.; Carling, D.; Gorospe, M. Increased AMP:ATP ratio and AMP-activated protein kinase activity during cellular senescence linked to reduced HuR function. J. Biol. Chem 2003, 278, 27016–27023. [Google Scholar]
- Annabi, B.; Currie, J.C.; Moghrabi, A.; Beliveau, R. Inhibition of HuR and MMP-9 expression in macrophage-differentiated HL-60 myeloid leukemia cells by green tea polyphenol EGCg. Leuk. Res 2007, 31, 1277–1284. [Google Scholar]
- Lin, F.Y.; Chen, Y.H.; Chen, Y.L.; Wu, T.C.; Li, C.Y.; Chen, J.W.; Lin, S.J. Ginkgo biloba extract inhibits endotoxin-induced human aortic smooth muscle cell proliferation via suppression of toll-like receptor 4 expression and NADPH oxidase activation. J. Agric. Food Chem 2007, 55, 1977–1984. [Google Scholar]
- Kang, M.J.; Ryu, B.K.; Lee, M.G.; Han, J.; Lee, J.H.; Ha, T.K.; Byun, D.S.; Chae, K.S.; Lee, B.H.; Chun, H.S.; et al. NF-kappaB activates transcription of the RNA-binding factor HuR, via PI3K-AKT signaling, to promote gastric tumorigenesis. Gastroenterology 2008, 135, 2030–2042. [Google Scholar]
- Jeyaraj, S.C.; Singh, M.; Ayupova, D.A.; Govindaraju, S.; Lee, B.S. Transcriptional control of human antigen R by bone morphogenetic protein. J. Biol. Chem 2010, 285, 4432–4440. [Google Scholar]
- Al-Ahmadi, W.; Al-Ghamdi, M.; Al-Souhibani, N.; Khabar, K.S. miR-29a inhibition normalizes HuR over-expression and aberrant AU-rich mRNA stability in invasive cancer. J. Pathol 2013, 230, 28–38. [Google Scholar]
- Prislei, S.; Martinelli, E.; Mariani, M.; Raspaglio, G.; Sieber, S.; Ferrandina, G.; Shahabi, S.; Scambia, G.; Ferlini, C. MiR-200c and HuR in ovarian cancer. BMC Cancer 2013, 13, 72. [Google Scholar]
- Leucci, E.; Zriwil, A.; Gregersen, L.H.; Jensen, K.T.; Obad, S.; Bellan, C.; Leoncini, L.; Kauppinen, S.; Lund, A.H. Inhibition of miR-9 de-represses HuR and DICER1 and impairs Hodgkin lymphoma tumour outgrowth in vivo. Oncogene 2012, 31, 5081–5089. [Google Scholar]
- Kojima, K.; Fujita, Y.; Nozawa, Y.; Deguchi, T.; Ito, M. MiR-34a attenuates paclitaxel-resistance of hormone-refractory prostate cancer PC3 cells through direct and indirect mechanisms. Prostate 2010, 70, 1501–1512. [Google Scholar]
- Xu, F.; Zhang, X.; Lei, Y.; Liu, X.; Liu, Z.; Tong, T.; Wang, W. Loss of repression of HuR translation by miR-16 may be responsible for the elevation of HuR in human breast carcinoma. J. Cell Biochem 2010, 111, 727–734. [Google Scholar]
- Guo, X.; Wu, Y.; Hartley, R.S. MicroRNA-125a represses cell growth by targeting HuR in breast cancer. RNA Biol 2009, 6, 575–583. [Google Scholar]
- Marasa, B.S.; Srikantan, S.; Martindale, J.L.; Kim, M.M.; Lee, E.K.; Gorospe, M.; Abdelmohsen, K. MicroRNA profiling in human diploid fibroblasts uncovers miR-519 role in replicative senescence. Aging 2010, 2, 333–343. [Google Scholar]
- Abdelmohsen, K.; Srikantan, S.; Kuwano, Y.; Gorospe, M. miR-519 reduces cell proliferation by lowering RNA-binding protein HuR levels. Proc. Natl. Acad. Sci. USA 2008, 105, 20297–20302. [Google Scholar]
- Abdelmohsen, K.; Kim, M.M.; Srikantan, S.; Mercken, E.M.; Brennan, S.E.; Wilson, G.M.; Cabo, R.; Gorospe, M. miR-519 suppresses tumor growth by reducing HuR levels. Cell Cycle 2010, 9, 1354–1359. [Google Scholar]
- Dai, W.; Zhang, G.; Makeyev, E.V. RNA-binding protein HuR autoregulates its expression by promoting alternative polyadenylation site usage. Nucleic Acids Res 2012, 40, 787–800. [Google Scholar]
- Mansfield, K.D.; Keene, J.D. Neuron-specific ELAV/Hu proteins suppress HuR mRNA during neuronal differentiation by alternative polyadenylation. Nucleic Acids Res 2012, 40, 2734–2746. [Google Scholar]
- Williams, T.K.; Costantino, C.L.; Bildzukewicz, N.A.; Richards, N.G.; Rittenhouse, D.W.; Einstein, L.; Cozzitorto, J.A.; Keen, J.C.; Dasgupta, A.; Gorospe, M.; et al. pp32 (ANP32A) expression inhibits pancreatic cancer cell growth and induces gemcitabine resistance by disrupting HuR binding to mRNAs. PLoS One 2010, 5, e15455. [Google Scholar]
- Cho, S.J.; Zhang, J.; Chen, X. RNPC1 modulates the RNA-binding activity of, and cooperates with, HuR to regulate p21 mRNA stability. Nucleic Acids Res 2010, 38, 2256–2267. [Google Scholar]
- Cho, S.J.; Jung, Y.S.; Zhang, J.; Chen, X. The RNA-binding protein RNPC1 stabilizes the mRNA encoding the RNA-binding protein HuR and cooperates with HuR to suppress cell proliferation. J. Biol. Chem 2012, 287, 14535–14544. [Google Scholar]
- Embade, N.; Fernández-Ramos, D.; Varela-Rey, M.; Beraza, N.; Sini, M.; Gutiérrez de Juan, V.; Woodhoo, A.; Martínez-López, N.; Rodríguez-Iruretagoyena, B.; Bustamante, F.J.; et al. Murine double minute 2 regulates Hu antigen R stability in human liver and colon cancer through NEDDylation. Hepatology 2012, 55, 1237–1248. [Google Scholar]
- Gabai, V.L.; Meng, L.; Kim, G.; Mills, T.A.; Benjamin, I.J.; Sherman, M.Y. Heat shock transcription factor Hsf1 is involved in tumor progression via regulation of hypoxia-inducible factor 1 and RNA-binding protein HuR. Mol. Cell Biol 2012, 32, 929–940. [Google Scholar]
- Al-Ahmadi, W.; Al-Ghamdi, M.; Al-Haj, L.; Al-Saif, M.; Khabar, K.S. Alternative polyadenylation variants of the RNA binding protein., HuR. Abundance, role of AU-rich elements and auto-Regulation. Nucleic Acids Res 2009, 37, 3612–3624. [Google Scholar]
- Talwar, S.; Jin, J.; Carroll, B.; Liu, A.; Gillespie, M.B.; Palanisamy, V. Caspase-mediated cleavage of RNA-binding protein HuR regulates c-Myc protein expression after hypoxic stress. J. Biol. Chem 2011, 286, 32333–32343. [Google Scholar]
- Yi, J.; Chang, N.; Liu, X.; Guo, G.; Xue, L.; Tong, T.; Gorospe, M.; Wang, W. Reduced nuclear export of HuR mRNA by HuR is linked to the loss of HuR in replicative senescence. Nucleic Acids Res 2010, 38, 1547–1558. [Google Scholar]
- Upadhyay, R.; Sanduja, S.; Kaza, V.; Dixon, D.A. Genetic polymorphisms in RNA binding proteins contribute to breast cancer survival. Int. J. Cancer 2012, 132, E128–E138. [Google Scholar]
- Cleary, M.L.; Mellentin, J.D.; Spies, J.; Smith, S.D. Chromosomal translocation involving the beta T cell receptor gene in acute leukemia. J. Exp. Med 1998, 167, 682–687. [Google Scholar]
- Bitoun, M.; Maugenre, S.; Jeannet, P.Y.; Lacène, E.; Ferrer, X.; Laforêt, P.; Martin, J.J.; Laporte, J.; Lochmüller, H.; Beggs, A.H.; et al. Mutations in dynamin 2 cause dominant centronuclear myopathy. Nat. Genet 2005, 37, 1207–1209. [Google Scholar]
- Rosette, C.; Roth, R.B.; Oeth, P.; Braun, A.; Kammerer, S.; Ekblom, J.; Denissenko, M.F. Role of ICAM1 in invasion of human breast cancer cells. Carcinogenesis 2005, 26, 943–950. [Google Scholar]
- Koljonen, V.; Bohling, T.; Haglund, C.; Ristimaki, A. Expression of HuR in Merkel cell carcinoma and in normal skin. J. Cutan Pathol 2008, 35, 10–14. [Google Scholar]
- Kim, K.Y.; Li, S.; Cha, J.D.; Zhang, X.; Cha, I.H. Significance of molecular markers in survival prediction of oral squamous cell carcinoma. Head Neck 2012, 34, 929–936. [Google Scholar]
- Denkert, C.; Koch, I.; von Keyserlingk, N.; Noske, A.; Niesporek, S.; Dietel, M.; Weichert, W. Expression of the ELAV-like protein HuR in human colon cancer. Association with tumor stage and cyclooxygenase-2. Mod Pathol 2006, 19, 1261–1269. [Google Scholar]
- Mrena, J.; Wiksten, J.P.; Thiel, A.; Kokkola, A.; Pohjola, L.; Lundin, J.; Nordling, S.; Ristimaki, A.; Haglund, C. Cyclooxygenase-2 is an independent prognostic factor in gastric cancer and its expression is regulated by the messenger RNA stability factor HuR. Clin. Cancer Res 2005, 11, 7362–7368. [Google Scholar]
- Wang, J.; Zhao, W.; Guo, Y.; Zhang, B.; Xie, Q.; Xiang, D.; Gao, J.; Wang, B.; Chen, Z. The expression of RNA-binding protein HuR in non-small cell lung cancer correlates with vascular endothelial growth factor-C expression and lymph node metastasis. Oncology 2009, 76, 420–429. [Google Scholar]
- Wang, J.; Wang, B.; Bi, J.; Zhang, C. Cytoplasmic HuR expression correlates with angiogenesis, lymphangiogenesis, and poor outcome in lung cancer. Med. Oncol 2011, 28, S577–S585. [Google Scholar]
- Denkert, C.; Weichert, W.; Winzer, K.J.; Muller, B.M.; Noske, A.; Niesporek, S.; Kristiansen, G.; Guski, H.; Dietel, M.; Hauptmann, S. Expression of the ELAV-like protein HuR is associated with higher tumor grade and increased cyclooxygenase-2 expression in human breast carcinoma. Clin. Cancer Res 2004, 10, 5580–5586. [Google Scholar]
- Denkert, C.; Weichert, W.; Pest, S.; Koch, I.; Licht, D.; Kobel, M.; Reles, A.; Sehouli, J.; Dietel, M.; Hauptmann, S. Overexpression of the embryonic-lethal abnormal vision-like protein HuR in ovarian carcinoma is a prognostic factor and is associated with increased cyclooxygenase 2 expression. Cancer Res 2004, 64, 189–195. [Google Scholar]
- Danilin, S.; Sourbier, C.; Thomas, L.; Lindner, V.; Rothhut, S.; Dormoy, V.; Helwig, J.J.; Jacqmin, D.; Lang, H.; Massfelder, T. Role of the RNA-binding protein HuR in human renal cell carcinoma. Carcinogenesis 2010, 31, 1018–1026. [Google Scholar]
- Kim, G.Y.; Lim, S.J.; Kim, Y.W. Expression of HuR, COX-2, and survivin in lung cancers, cytoplasmic HuR stabilizes cyclooxygenase-2 in squamous cell carcinomas. Mod Pathol 2011, 24, 1336–1347. [Google Scholar]
- Mrena, J.; Wiksten, J.P.; Kokkola, A.; Nordling, S.; Ristimaki, A.; Haglund, C. COX-2 is associated with proliferation and apoptosis markers and serves as an independent prognostic factor in gastric cancer. Tumour Biol 2010, 31, 1–7. [Google Scholar]
- Stoppoloni, D.; Cardillo, I.; Verdina, A.; Vincenzi, B.; Menegozzo, S.; Santini, M.; Sacchi, A.; Baldi, A.; Galati, R. Expression of the embryonic lethal abnormal vision-like protein HuR in human mesothelioma: Association with cyclooxygenase-2 and prognosis. Cancer 2008, 113, 2761–2769. [Google Scholar]
- Miyata, Y.; Watanabe, S.; Sagara, Y.; Mitsunari, K.; Matsuo, T.; Ohba, K.; Sakai, H. High expression of HuR in cytoplasm, but not nuclei, is associated with malignant aggressiveness and prognosis in bladder cancer. PLoS One 2013, 8, e59095. [Google Scholar]
- Heinonen, M.; Fagerholm, R.; Aaltonen, K.; Kilpivaara, O.; Aittomaki, K.; Blomqvist, C.; Heikkila, P.; Haglund, C.; Nevanlinna, H.; Ristimaki, A. Prognostic role of HuR in hereditary breast cancer. Clin. Cancer Res 2007, 13, 6959–6963. [Google Scholar]
- Heinonen, M.; Bono, P.; Narko, K.; Chang, S.H.; Lundin, J.; Joensuu, H.; Furneaux, H.; Hla, T.; Haglund, C.; Ristimaki, A. Cytoplasmic HuR expression is a prognostic factor in invasive ductal breast carcinoma. Cancer Res 2005, 65, 2157–2161. [Google Scholar]
- Lim, S.J.; Kim, H.J.; Kim, J.Y.; Park, K.; Lee, C.M. Expression of HuR is associated with increased cyclooxygenase-2 expression in uterine cervical carcinoma. Int. J. Gynecol. Pathol 2007, 26, 229–234. [Google Scholar]
- Blaxall, B.C.; Dwyer-Nield, L.D.; Bauer, A.K.; Bohlmeyer, T.J.; Malkinson, A.M.; Port, J.D. Differential expression and localization of the mRNA binding proteins, AU-rich element mRNA binding protein (AUF1) and Hu antigen R (HuR), in neoplastic lung tissue. Mol. Carcinog 2000, 28, 76–83. [Google Scholar]
- Barbisan, F.; Mazzucchelli, R.; Santinelli, A.; Lopez-Beltran, A.; Cheng, L.; Scarpelli, M.; Montorsi, F.; Montironi, R. Overexpression of ELAV-like protein HuR is associated with increased COX-2 expression in atrophy, high-grade prostatic intraepithelial neoplasia, and incidental prostate cancer in cystoprostatectomies. Eur. Urol 2009, 56, 105–112. [Google Scholar]
- Brosens, L.A.; Keller, J.J.; Pohjola, L.; Haglund, C.; Morsink, F.H.; Iacobuzio-Donahue, C.; Goggins, M.; Giardiello, F.M.; Ristimaki, A.; Offerhaus, G.J. Increased expression of cytoplasmic HuR in familial adenomatous polyposis. Cancer Biol. Ther 2008, 7, 424–427. [Google Scholar]
- Fay, J.; Kelehan, P.; Lambkin, H.; Schwartz, S. Increased expression of cellular RNA-binding proteins in HPV-induced neoplasia and cervical cancer. J. Med. Virol 2009, 81, 897–907. [Google Scholar]
- Lopez de Silanes, I.; Fan, J.; Yang, X.; Zonderman, A.B.; Potapova, O.; Pizer, E.S.; Gorospe, M. Role of the RNA-binding protein HuR in colon carcinogenesis. Oncogene 2003, 22, 7146–7154. [Google Scholar]
- Lopez de Silanes, I.; Fan, J.; Galban, C.J.; Spencer, R.G.; Becker, K.G.; Gorospe, M. Global analysis of HuR-regulated gene expression in colon cancer systems of reducing complexity. Gene Expr 2004, 12, 49–59. [Google Scholar]
- Milne, A.N.; Carvalho, R.; Morsink, F.M.; Musler, A.R.; de Leng, W.W.; Ristimaki, A.; Offerhaus, G.J. Early-onset gastric cancers have a different molecular expression profile than conventional gastric cancers. Mod. Pathol 2006, 19, 564–572. [Google Scholar]
- Young, L.E.; Sanduja, S.; Bemis-Standoli, K.; Pena, E.A.; Price, R.L.; Dixon, D.A. The mRNA binding proteins HuR and tristetraprolin regulate cyclooxygenase 2 expression during colon carcinogenesis. Gastroenterology 2009, 136, 1669–1679. [Google Scholar]
- Cho, N.P.; Han, H.S.; Soh, Y.; Son, H.J. Overexpression of cyclooxygenase-2 correlates with cytoplasmic HuR expression in salivary mucoepidermoid carcinoma but not in pleomorphic adenoma. J. Oral. Pathol. Med 2007, 36, 297–303. [Google Scholar]
- Saunus, J.M.; French, J.D.; Edwards, S.L.; Beveridge, D.J.; Hatchell, E.C.; Wagner, S.A.; Stein, S.R.; Davidson, A.; Simpson, K.J.; Francis, G.D.; et al. Posttranscriptional regulation of the breast cancer susceptibility gene BRCA1 by the RNA binding protein HuR. Cancer Res 2008, 68, 9469–9478. [Google Scholar]
- Woo, H.H.; Yi, X.; Lamb, T.; Menzl, I.; Baker, T.; Shapiro, D.J.; Chambers, S.K. Posttranscriptional suppression of proto-oncogene c-fms expression by vigilin in breast cancer. Mol. Cell Biol 2011, 31, 215–225. [Google Scholar]
- Guo, X.; Hartley, R.S. HuR contributes to cyclin E1 deregulation in MCF-7 breast cancer cells. Cancer Res 2006, 66, 7948–7956. [Google Scholar]
- Yuan, Z.; Sanders, A.J.; Ye, L.; Wang, Y.; Jiang, W.G. Knockdown of human antigen R reduces the growth and invasion of breast cancer cells in vitro and affects expression of cyclin D1 and MMP-9. Oncol. Rep 2011, 26, 237–245. [Google Scholar]
- López de Silanes, I.; Gorospe, M.; Taniguchi, H.; Abdelmohsen, K.; Srikantan, S.; Alaminos, M.; Berdasco, M.; Urdinguio, R.G.; Fraga, M.F.; Jacinto, F.V.; et al. The RNA-binding protein HuR regulates DNA methylation through stabilization of DNMT3b mRNA. Nucleic Acids Res 2009, 37, 2658–2671. [Google Scholar]
- Kakuguchi, W.; Kitamura, T.; Kuroshima, T.; Ishikawa, M.; Kitagawa, Y.; Totsuka, Y.; Shindoh, M.; Higashino, F. HuR knockdown changes the oncogenic potential of oral cancer cells. Mol. Cancer Res 2010, 8, 520–528. [Google Scholar]
- Linker, K.; Pautz, A.; Fechir, M.; Hubrich, T.; Greeve, J.; Kleinert, H. Involvement of KSRP in the post-transcriptional regulation of human iNOS expression-complex interplay of KSRP with TTP and HuR. Nucleic Acids Res 2005, 33, 4813–4827. [Google Scholar]
- Suzuki, E.; Tsutsumi, A.; Sugihara, M.; Mamura, M.; Goto, D.; Matsumoto, I.; Ito, S.; Ikeda, K.; Ochiai, N.; Sato, Y.; et al. Expression of TNF-alpha., tristetraprolin., T-cell intracellular antigen-1 and Hu antigen R genes in synovium of patients with rheumatoid arthritis. Int. J. Mol. Med 2006, 18, 273–278. [Google Scholar]
- Nabors, L.B.; Gillespie, G.Y.; Harkins, L.; King, P.H. HuR, a RNA stability factor, is expressed in malignant brain tumors and binds to adenine- and uridine-rich elements within the 3′ untranslated regions of cytokine and angiogenic factor mRNAs. Cancer Res 2001, 61, 2154–2161. [Google Scholar]
- Ishimaru, D.; Ramalingam, S.; Sengupta, T.K.; Bandyopadhyay, S.; Dellis, S.; Tholanikunnel, B.G.; Fernandes, D.J.; Spicer, E.K. Regulation of Bcl-2 expression by HuR in HL60 leukemia cells and A431 carcinoma cells. Mol. Cancer Res 2009, 7, 1354–1366. [Google Scholar]
- Pineda, D.M.; Rittenhouse, D.W.; Valley, C.C.; Cozzitorto, J.A.; Burkhart, R.A.; Leiby, B.; Winter, J.M.; Weber, M.C.; Londin, E.R.; Rigoutsos, I.; et al. HuR’s post-transcriptional regulation of Death Receptor 5 in pancreatic cancer cells. Cancer Biol. Ther 2012, 13, 946–955. [Google Scholar]
- Giles, K.M.; Daly, J.M.; Beveridge, D.J.; Thomson, A.M.; Voon, D.C.; Furneaux, H.M.; Jazayeri, J.A.; Leedman, P.J. The 3′-untranslated region of p21WAF1 mRNA is a composite cis-acting sequence bound by RNA-binding proteins from breast cancer cells, including HuR and poly(C)-binding protein. J. Biol. Chem 2003, 278, 2937–2946. [Google Scholar]
- Leandersson, K.; Riesbeck, K.; Andersson, T. Wnt-5a mRNA translation is suppressed by the Elav-like protein HuR in human breast epithelial cells. Nucleic Acids Res 2006, 34, 3988–3999. [Google Scholar]
- Yoo, P.S.; Mulkeen, A.L.; Cha, C.H. Post-transcriptional regulation of vascular endothelial growth factor. Implications for tumor angiogenesis. World J. Gastroenterol 2006, 12, 4937–4942. [Google Scholar]
- Ido, K.; Nakagawa, T.; Sakuma, T.; Takeuchi, H.; Sato, K.; Kubota, T. Expression of vascular endothelial growth factor-A and mRNA stability factor HuR in human astrocytic tumors. Neuropathology 2008, 28, 604–611. [Google Scholar]
- Sakuma, T.; Nakagawa, T.; Ido, K.; Takeuchi, H.; Sato, K.; Kubota, T. Expression of vascular endothelial growth factor-A and mRNA stability factor HuR in human meningiomas. J. Neurooncol 2008, 88, 143–155. [Google Scholar]
- Niesporek, S.; Kristiansen, G.; Thoma, A.; Weichert, W.; Noske, A.; Buckendahl, A.C.; Jung, K.; Stephan, C.; Dietel, M.; Denkert, C. Expression of the ELAV-like protein HuR in human prostate carcinoma is an indicator of disease relapse and linked to COX-2 expression. Int. J. Oncol 2008, 32, 341–347. [Google Scholar]
- Dixon, D.A.; Tolley, N.D.; King, P.H.; Nabors, L.B.; McIntyre, T.M.; Zimmerman, G.A.; Prescott, S.M. Altered expression of the mRNA stability factor HuR promotes cyclooxygenase-2 expression in colon cancer cells. J. Clin. Invest 2001, 108, 1657–1665. [Google Scholar]
- Erkinheimo, T.L.; Lassus, H.; Sivula, A.; Sengupta, S.; Furneaux, H.; Hla, T.; Haglund, C.; Butzow, R.; Ristimaki, A. Cytoplasmic HuR expression correlates with poor outcome and with cyclooxygenase 2 expression in serous ovarian carcinoma. Cancer Res 2003, 63, 7591–7594. [Google Scholar]
- Yoo, P.S.; Sullivan, C.A.; Kiang, S.; Gao, W.; Uchio, E.M.; Chung, G.G.; Cha, C.H. Tissue microarray analysis of 560 patients with colorectal adenocarcinoma. high expression of HuR predicts poor survival. Ann. Surg. Oncol 2009, 16, 200–207. [Google Scholar]
- Richards, N.G.; Rittenhouse, D.W.; Freydin, B.; Cozzitorto, J.A.; Grenda, D.; Rui, H.; Gonye, G.; Kennedy, E.P.; Yeo, C.J.; Brody, J.R.; et al. HuR status is a powerful marker for prognosis and response to gemcitabine-based chemotherapy for resected pancreatic ductal adenocarcinoma patients. Ann. Surg 2010, 252, 499–505. [Google Scholar]
- Kurosu, T.; Ohga, N.; Hida, Y.; Maishi, N.; Akiyama, K.; Kakuguchi, W.; Kuroshima, T.; Kondo, M.; Akino, T.; Totsuka, Y.; et al. HuR keeps an angiogenic switch on by stabilising mRNA of VEGF and COX-2 in tumour endothelium. Br. J. Cancer 2011, 104, 819–829. [Google Scholar]
- Gubin, M.M.; Calaluce, R.; Davis, J.W.; Magee, J.D.; Strouse, C.S.; Shaw, D.P.; Ma, L.; Brown, A.; Hoffman, T.; Rold, T.L.; et al. Overexpression of the RNA binding protein HuR impairs tumor growth in triple negative breast cancer associated with deficient angiogenesis. Cell Cycle 2010, 9, 3337–3346. [Google Scholar]
- Calaluce, R.; Gubin, M.M.; Davis, J.W.; Magee, J.D.; Chen, J.; Kuwano, Y.; Gorospe, M.; Atasoy, U. The RNA binding protein HuR differentially regulates unique subsets of mRNAs in estrogen receptor negative and estrogen receptor positive breast cancer. BMC Cancer 2010, 10, 126. [Google Scholar]
- Ale-Agha, N.; Galban, S.; Sobieroy, C.; Abdelmohsen, K.; Gorospe, M.; Sies, H.; Klotz, L.O. HuR regulates gap junctional intercellular communication by controlling beta-catenin levels and adherens junction integrity. Hepatology 2009, 50, 1567–1576. [Google Scholar]
- Lim, S.J.; Lee, S.H.; Joo, S.H.; Song, J.Y.; Choi, S.I. Cytoplasmic expression of HuR is related to cyclooxygenase-2 expression in colon cancer. Cancer Res. Treat 2009, 41, 87–92. [Google Scholar]
- Liang, P.I.; Li, W.M.; Wang, Y.H.; Wu, T.F.; Wu, W.R.; Liao, A.C.; Shen, K.H.; Wei, Y.C.; Hsing, C.H.; Shiue, Y.L.; et al. HuR cytoplasmic expression is associated with increased cyclin A expression and poor outcome with upper urinary tract urothelial carcinoma. BMC Cancer 2012, 12, 611. [Google Scholar]
- Heinonen, M.; Hemmes, A.; Salmenkivi, K.; Abdelmohsen, K.; Vilén, S.T.; Laakso, M.; Leidenius, M.; Salo, T.; Hautaniemi, S.; Gorospe, M.; et al. Role of RNA binding protein HuR in ductal carcinoma in situ of the breast. J. Pathol 2011, 224, 529–539. [Google Scholar]
- Lauriola, L.; Granone, P.; Ramella, S.; Lanza, P.; Ranelletti, F.O. Expression of the RNA-binding protein HuR and its clinical significance in human stage I and II lung adenocarcinoma. Histol. Histopathol 2012, 27, 617–626. [Google Scholar]
- Tran, H.; Maurer, F.; Nagamine, Y. Stabilization of urokinase and urokinase receptor mRNAs by HuR is linked to its cytoplasmic accumulation induced by activated mitogen-activated protein kinase-activated protein kinase 2. Mol. Cell Biol 2003, 23, 7177–7188. [Google Scholar]
- Dong, R.; Lu, J.G.; Wang, Q.; He, X.L.; Chu, Y.K.; Ma, Q.J. Stabilization of Snail by HuR in the process of hydrogen peroxide induced cell migration. Biochem. Biophys. Res. Commun 2007, 356, 318–321. [Google Scholar]
- Lee, Y.R.; Noh, E.M.; Oh, H.J.; Hur, H.; Kim, J.M.; Han, J.H.; Hwang, J.K.; Park, B.H.; Park, J.W.; Youn, H.J.; et al. Dihydroavenanthramide D inhibits human breast cancer cell invasion through suppression of MMP-9 expression. Biochem. Biophys. Res. Commun 2011, 405, 552–557. [Google Scholar]
- Noh, E.M.; Lee, Y.R.; Hur, H.; Kim, J.S. Radix clematidis extract inhibits TPA-induced MMP-9 expression by suppressing NF-kappaB activation in MCF-7 human breast cancer cells. Mol. Med. Report 2011, 4, 879–883. [Google Scholar]
- Hsia, T.C.; Tu, C.Y.; Chen, Y.J.; Wei, Y.L.; Yu, M.C.; Hsu, S.C.; Tsai, S.L.; Chen, W.S.; Yeh, M.H.; Yen, C.J.; et al. Lapatinib-mediated COX-2 Expression Via EGFR/HuR interaction enhances the aggressiveness of triple-negative breast cancer cells. Mol. Pharmacol 2013, 83, 857–869. [Google Scholar]
- Filippova, N.; Yang, X.; Wang, Y.; Gillespie, G.Y.; Langford, C.; King, P.H.; Wheeler, C.; Nabors, L.B. The RNA-binding protein HuR promotes glioma growth and treatment resistance. Mol. Cancer Res 2011, 9, 648–659. [Google Scholar]
- Urano, N.; Fujiwara, Y.; Doki, Y.; Kim, S.J.; Miyoshi, Y.; Noguchi, S.; Miyata, H.; Takiguchi, S.; Yasuda, T.; Yano, M.; et al. Clinical significance of class III beta-tubulin expression and its predictive value for resistance to docetaxel-based chemotherapy in gastric cancer. Int. J. Oncol 2006, 28, 375–381. [Google Scholar]
- Lee, K.M.; Cao, D.; Itami, A.; Pour, P.M.; Hruban, R.H.; Maitra, A.; Ouellette, M.M. Class III beta-tubulin, a marker of resistance to paclitaxel, is overexpressed in pancreatic ductal adenocarcinoma and intraepithelial neoplasia. Histopathology 2007, 51, 539–546. [Google Scholar]
- Seve, P.; Isaac, S.; Tredan, O.; Souquet, P.J.; Pacheco, Y.; Perol, M.; Lafanechere, L.; Penet, A.; Peiller, E.L.; Dumontet, C. Expression of class III β-tubulin is predictive of patient outcome in patients with non-small cell lung cancer receiving vinorelbine-based chemotherapy. Clin. Cancer Res 2005, 11, 5481–5486. [Google Scholar]
- Raspaglio, G.; de Maria, I.; Filippetti, F.; Martinelli, E.; Zannoni, G.F.; Prislei, S.; Ferrandina, G.; Shahabi, S.; Scambia, G.; Ferlini, C. HuR regulates beta-tubulin isotype expression in ovarian cancer. Cancer Res 2010, 70, 5891–5900. [Google Scholar]
- Liu, Z.M.; Tseng, J.T.; Hong, D.Y.; Huang, H.S. Suppression of TG-interacting factor sensitizes arsenic trioxide-induced apoptosis in human hepatocellular carcinoma cells. Biochem. J 2011, 438, 349–358. [Google Scholar]
- Serini, S.; Donato, V.; Piccioni, E.; Trombino, S.; Monego, G.; Toesca, A.; Innocenti, I.; Missori, M.; de Spirito, M.; Celleno, L.; et al. Docosahexaenoic acid reverts resistance to UV-induced apoptosis in human keratinocytes. involvement of COX-2 and HuR. J. Nutr. Biochem 2011, 22, 874–885. [Google Scholar]
- Latorre, E.; Tebaldi, T.; Viero, G.; Sparta, A.M.; Quattrone, A.; Provenzani, A. Downregulation of HuR as a new mechanism of doxorubicin resistance in breast cancer cells. Mol. Cancer 2012. [Google Scholar] [CrossRef]
- Griseri, P.; Bourcier, C.; Hieblot, C.; Essafi-Benkhadir, K.; Chamorey, E.; Touriol, C.; Pages, G. A synonymous polymorphism of the Tristetraprolin (TTP) gene, an AU-rich mRNA-binding protein, affects translation efficiency and response to Herceptin treatment in breast cancer patients. Hum. Mol. Genet 2011, 20, 4556–4568. [Google Scholar]
- Ronkainen, H.; Vaarala, M.H.; Hirvikoski, P.; Ristimaki, A. HuR expression is a marker of poor prognosis in renal cell carcinoma. Tumour. Biol 2011, 32, 481–487. [Google Scholar]
- Cha, J.D.; Li, S.; Cha, I.H. Association between expression of embryonic lethal abnormal vision-like protein HuR and cyclooxygenase-2 in oral squamous cell carcinoma. Head Neck 2011, 33, 627–637. [Google Scholar]
- Zhu, Z.; Wang, B.; Bi, J.; Zhang, C.; Guo, Y.; Chu, H.; Liang, X.; Zhong, C.; Wang, J. Cytoplasmic HuR expression correlates with P-gp, HER-2 positivity, and poor outcome in breast cancer. Tumor. Biol 2013. [Google Scholar] [CrossRef]
- Yi, X.; Zhou, Y.; Zheng, W.; Chambers, S.K. HuR expression in the nucleus correlates with high histological grade and poor disease-free survival in ovarian cancer. Aust. N. Z. J. Obstet. Gynaecol 2009, 49, 93–98. [Google Scholar]
- Ortega, A.D.; Sala, S.; Espinosa, E.; Gonzalez-Baron, M.; Cuezva, J.M. HuR and the bioenergetic signature of breast cancer, a low tumor expression of the RNA-binding protein predicts a higher risk of disease recurrence. Carcinogenesis 2008, 29, 2053–2061. [Google Scholar]
- Yuan, Z.; Sanders, A.J.; Ye, L.; Wang, Y.; Jiang, W.G. Prognostic value of the human antigen R (HuR) in human breast cancer. high level predicts a favourable prognosis. Anticancer Res 2011, 31, 303–310. [Google Scholar]
- Lopez de Silanes, I.; Zhan, M.; Lal, A.; Yang, X.; Gorospe, M. Identification of a target RNA motif for RNA-binding protein HuR. Proc. Natl. Acad. Sci. USA 2004, 101, 2987–2992. [Google Scholar]
- Lopez de Silanes, I.; Lal, A.; Gorospe, M. HuR post-transcriptional paths to malignancy. RNA Biol 2005, 2, 11–13. [Google Scholar]
- Lebedeva, S.; Jens, M.; Theil, K.; Schwanhäusser, B.; Selbach, M.; Landthaler, M.; Rajewsky, N. Transcriptome-wide analysis of regulatory interactions of the RNA-binding protein HuR. Mol. Cell 2011, 43, 340–352. [Google Scholar]
- Mukherjee, N.; Corcoran, D.L.; Nusbaum, J.D.; Reid, D.W.; Georgiev, S.; Hafner, M.; Ascano, M., Jr; Tuschl, T.; Ohler, U.; Keene, J.D. Integrative regulatory mapping indicates that the RNA-binding protein HuR couples pre-mRNA processing and mRNA stability. Mol. Cell 2011, 43, 327–339. [Google Scholar]
- Kishore, S.; Jaskiewicz, L.; Burger, L.; Hausser, J.; Khorshid, M.; Zavolan, M. A quantitative analysis of CLIP methods for identifying binding sites of RNA-binding proteins. Nat. Methods 2011, 8, 559–564. [Google Scholar]
- Akool el, S.; Kleinert, H.; Hamada, F.M.; Abdelwahab, M.H.; Forstermann, U.; Pfeilschifter, J.; Eberhardt, W. Nitric oxide increases the decay of matrix metalloproteinase 9 mRNA by inhibiting the expression of mRNA-stabilizing factor HuR. Mol. Cell Biol 2003, 23, 4901–4916. [Google Scholar]
| Regulators | Effect on HuR | References |
|---|---|---|
| UVR | Cytoplasmic accumulation ↑ | [27,28] |
| Compound | ||
| Ethanol | Cytoplasmic accumulation ↑ | [29,30] |
| LPS | Cytoplasmic accumulation ↑ | [31,32] |
| SAHA | Protein ↓ | [44] |
| Tamoxifen | Cytoplasmic accumulation ↑ | [33] |
| Gemcitabine | Cytoplasmic accumulation ↑ | [34] |
| Nitric oxide | mRNA ↓, protein ↓ | [23] |
| HIV protease inhibitor | Cytoplasmic accumulation ↑ | [35] |
| Proteasome inhibitor MG132 | Cytoplasmic accumulation ↑ | [36] |
| Microenvironment change | ||
| Hypoxia | Cytoplasmic accumulation ↑ | [37] |
| Amino acid limitation | Cytoplasmic accumulation ↑ | [38] |
| Bile salts | Cytoplasmic accumulation ↑ | [39] |
| Serum | Cytoplasmic accumulation ↑ | [40] |
| Polyamines depletion | Cytoplasmic accumulation ↑ | [41–43] |
| DHA | Cytoplasmic accumulation ↑ | [44] |
| Nature reagent | ||
| Green tea | Cytoplasmic accumulation ↓ | [45,70] |
| Ginkgo biloba extract | Cytoplasmic accumulation ↓ | [71] |
| KPS-A | Cytoplasmic accumulation ↓ | [47] |
| Triptolide | Cytoplasmic accumulation ↓ | [48] |
| Cytokine | ||
| IL-1β | Cytoplasmic accumulation ↑ | [49] |
| TNF-α | Cytoplasmic accumulation ↑ | [50] |
| TGF-β1 | Cytoplasmic accumulation ↑ | [51] |
| Virus infection | ||
| HPV | Cytoplasmic accumulation ↑ | [52,53] |
| Alphavirus | Cytoplasmic accumulation ↑ | [54] |
| Hormone | ||
| ACTH | Cytoplasmic accumulation ↑ | [55] |
| Androgens | Cytoplasmic accumulation ↑ | [56,57] |
| 17β-estradiol | Cytoplasmic accumulation ↑ | [58] |
| Regulators | Mechanism | Effect sites | Effect on HuR | References |
|---|---|---|---|---|
| NF-κB | Transcriptional | Promoter | HuR mRNA ↑ | [72] |
| Smad | Transcriptional | Promoter | HuR mRNA ↑ | [73] |
| Kinases | ||||
| PKCα | Phosphorylation | S158, S221 | RNA-binding ↑, cytoplasmic accumulation ↑ | [61] |
| PKCδ | Phosphorylation | S318, S221 | RNA-binding ↑, cytoplasmic accumulation ↑ | [62] |
| Cdk1 | Phosphorylation | S202 | Cytoplasmic accumulation ↑ | [63] |
| Chk2 | Phosphorylation | S88, S100, T118 | RNA-binding ↑ | [64] |
| p38 | MAPK Phosphorylation | T118 | Cytoplasmic accumulation | [65] |
| PI3K-AKT | Transcriptional. | Promoter | p65/RelA binding to a putative NF-κB binding site in the HuR promoter ↑ | [72] |
| AMPK | Transcriptional | K22 and S105 of importin α | Nuclear import via phosphorylation and acetylation of importin α ↑ | [41,68,69] |
| miRNAs | ||||
| miR-9 | Transcriptional | Unknown | HuR mRNA↓; HuR protein ↓ | [74] |
| miR-200c | Unknown | Unknown | Interaction of HuR and mRNA ↓ | [75] |
| miR-9 | Post-transcriptional | 3′UTR | HuR mRNA↓; HuR protein ↓ | [76] |
| miR-34a | Post-transcriptional | 3′UTR | HuR mRNA↓; HuR protein ↓ | [77] |
| miR-16 | Translational | 3′UTR | HuR protein ↓ | [78] |
| miR-125a | Translational | 3′UTR | HuR protein ↓ | [79] |
| miR-519 | Translational | 3′UTR | HuR protein ↓ | [80–82] |
| Proteins | ||||
| CARM1 | Methylation | R217 | RNA-binding ↑, cytoplasmic accumulation ↑ | [66] |
| HuR | Ubiquitinylation | K182 | Protein stability ↑ | [67] |
| HuR | Post-transcriptional | polyadenylation site | mRNA stability ↑ | [83] |
| Hu (B-D) | Post-transcriptional | polyadenylation site | mRNA stability ↑ | [84] |
| TTP | Post-transcriptional | 3′UTR | mRNA stability ↓ | [83] |
| pp32 | Interaction | Not indicated | RNA-binding ↓ | [85] |
| RNP C1 | Post-transcriptional | RRM, 3′UTR | RNA-binding ↑, mRNA stability ↓ | [86,87] |
| Mdm2 | Ubiquitinylation | K283, K313, K326 | Protein stability ↑ | [88] |
| Hsf1 | Not indicated | Not indicated | HuR protein ↑ | [89] |
| First author | Year | Country | Method | N | Prognostic effect of HuR | Type of cancer | |
|---|---|---|---|---|---|---|---|
| Unadjusted | Adjusted | ||||||
| Miyata et al. [109] | 2013 | Japan | IHC | 122 | C b,f↓ | C b,f↓ | Bladder Cancer, pTa-3 |
| Zhu et al. [167] | 2013 | China | IHC | 82 | C a,c↓, N a,c (NS) | C a,c↓, N a,c (NS) | Breast cancer, stage I–III |
| Lauriola et al. [150] | 2012 | Italy | IHC | 54 | C b,c↓, NCR b,c↓, N b,c (NS) | C b,d↓, NCR b,c | Lung adenocarcinoma, stage I–II |
| Kim et al. [98] | 2012 | South Korea | IHC | 96 | C c (NS), N c (NS) | C c (NS), N c (NS) | Oral squamous cell carcinoma, stage I–IV |
| Liang et al.[148] | 2012 | China | IHC | 340 | C a,b,f↓ | C f (NS) | Upper urinary tract urothelial carcinoma |
| Kim et al. [106] | 2011 | South Korea | IHC | 244 | C c (NS), N c (NS), (C + N) c (NS), (C − N) c (NS) | C c (NS), N c (NS), (C + N) c (NS), (C − N) c (NS) | Lung adenocarcinoma and squamous cell carcinomas, I–IV |
| Yuan et al. [170] | 2011 | UK | RT-PCR | 109 | HuR mRNA a,c (NS) | HuR mRNA a,c (NS) | Invasive breast carcinoma, stage I–IV |
| Ronkainen et al. [165] | 2011 | Finland | IHC | 152 | C c↓ | C c (NS) | Renal cell carcinoma, stage I–IV |
| Wang et al. [102] | 2011 | China | IHC | 132 | C a,c↓, N a,c (NS) | C a,d↓, N a,c (NS) | Non-small cell lung carcinoma, I–IIIB |
| Cha et al. [166] | 2011 | South Korea | IHC | 103 | C c↓, N a,c (NS) | C d↓, N a,c (NS) | Oral squamous cell carcinoma, I–IV |
| Richards et al. [142] | 2010 | USA | IHC | 52 | C d↑ | C d↑ | Pancreatic ductal adenocarcinoma |
| Mrena et al. [107] | 2010 | Finland | IHC | 316 | C c↓ | C c (NS) | Gastric carcinoma, stage I–IV |
| Costantino et al. [34] | 2009 | USA | IHC | 32 | C d↑ | C d↑ | Pancreatic ductal adenocarcinoma |
| Yi et al. [168] | 2009 | USA | IHC | 113 | N a↓ | N a↓ | Ovarian carcinoma, stage I–IV |
| Yoo et al. [141] | 2009 | USA | Immunofluorescence | 560 | (C + N) c↓, NCR c↑ | (C + N) c↓ | Colorectal carcinoma, stage I–IV |
| Stoppoloni et al. [108] | 2009 | Italy | IHC | 29 | C c↓ | Not indicated | Mesothelioma |
| Ortega et al. [169] | 2008 | Spain | Western blotting | 89 | (C + N) a↑ | Not indicated | Invasive breast carcinoma |
| Niesporek et al. [138] | 2008 | Germany | IHC | 104 | C a (NS), N a↑ | N a↑ | Prostate carcinoma |
| Heinonen et al. [110] | 2007 | Finland | IHC | 641 | C c,e↓ | C c↓ | Invasive breast carcinoma |
| Lim et al. [112] | 2007 | South Korea | IHC | 308 | C c (NS), N c (NS) | C c (NS), N c (NS) | Cervical carcinoma, carcinoma in situ and stage I–II |
| Denkert et al. [99] | 2006 | Germany | IHC | 87 | C c (NS) | C c (NS) | Colorectal carcinoma, |
| Mrena et al. [100] | 2005 | Finland | IHC | 316 | C c↓ | C c (NS) | Gastric carcinoma, stage I–IV |
| Heinonen et al. [111] | 2005 | USA | IHC | 133 | C a↓ | C a↓ | Invasive breast carcinoma, stage I–III |
| Denkert et al. [103] | 2004 | Germany | IHC | 208 | C a,c (NS), N a,c (NS) | C a,c (NS), N a,c (NS) | Invasive breast carcinoma, stage I–III |
| Denkert et al. [104] | 2004 | Germany | IHC | 83 | C a,c↓, N a,c (NS) | C a,c↓ | Ovarian carcinoma, stage I–IV |
| Erkinheimo et al. [140] | 2003 | Finland | IHC | 445 | C c↓, N c (NS) | C c (NS) | Ovarian carcinoma, stage I–IV |
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wang, J.; Guo, Y.; Chu, H.; Guan, Y.; Bi, J.; Wang, B. Multiple Functions of the RNA-Binding Protein HuR in Cancer Progression, Treatment Responses and Prognosis. Int. J. Mol. Sci. 2013, 14, 10015-10041. https://doi.org/10.3390/ijms140510015
Wang J, Guo Y, Chu H, Guan Y, Bi J, Wang B. Multiple Functions of the RNA-Binding Protein HuR in Cancer Progression, Treatment Responses and Prognosis. International Journal of Molecular Sciences. 2013; 14(5):10015-10041. https://doi.org/10.3390/ijms140510015
Chicago/Turabian StyleWang, Jun, Yan Guo, Huili Chu, Yaping Guan, Jingwang Bi, and Baocheng Wang. 2013. "Multiple Functions of the RNA-Binding Protein HuR in Cancer Progression, Treatment Responses and Prognosis" International Journal of Molecular Sciences 14, no. 5: 10015-10041. https://doi.org/10.3390/ijms140510015
APA StyleWang, J., Guo, Y., Chu, H., Guan, Y., Bi, J., & Wang, B. (2013). Multiple Functions of the RNA-Binding Protein HuR in Cancer Progression, Treatment Responses and Prognosis. International Journal of Molecular Sciences, 14(5), 10015-10041. https://doi.org/10.3390/ijms140510015




