Genetic Diversity Revealed by Single Nucleotide Polymorphism Markers in a Worldwide Germplasm Collection of Durum Wheat
Abstract
:1. Introduction
2. Results
2.1. SNP Marker Quality and Genomic Distribution
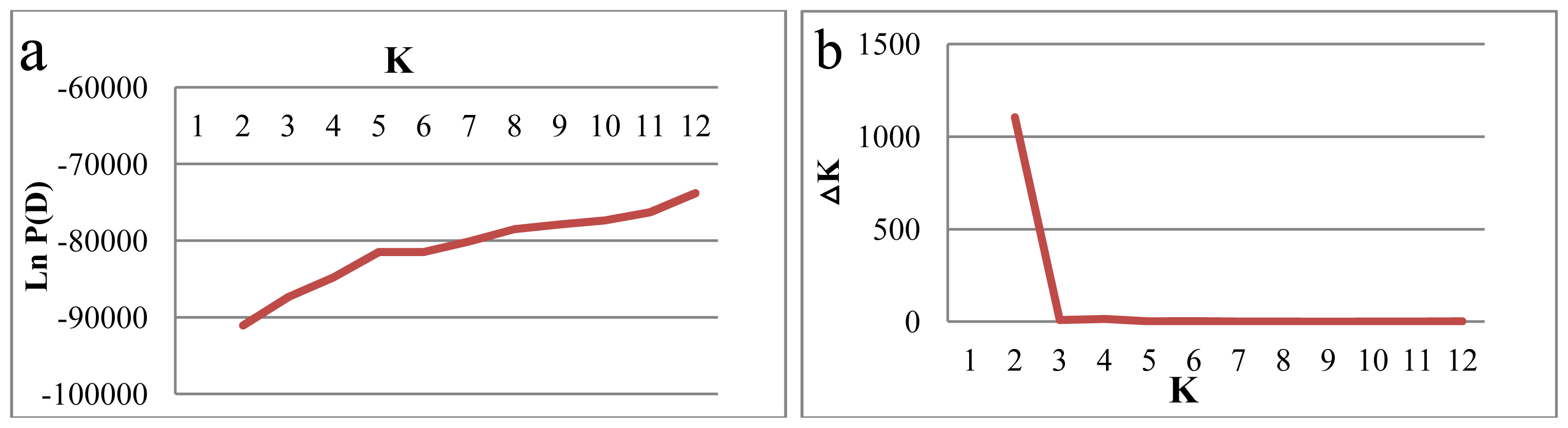
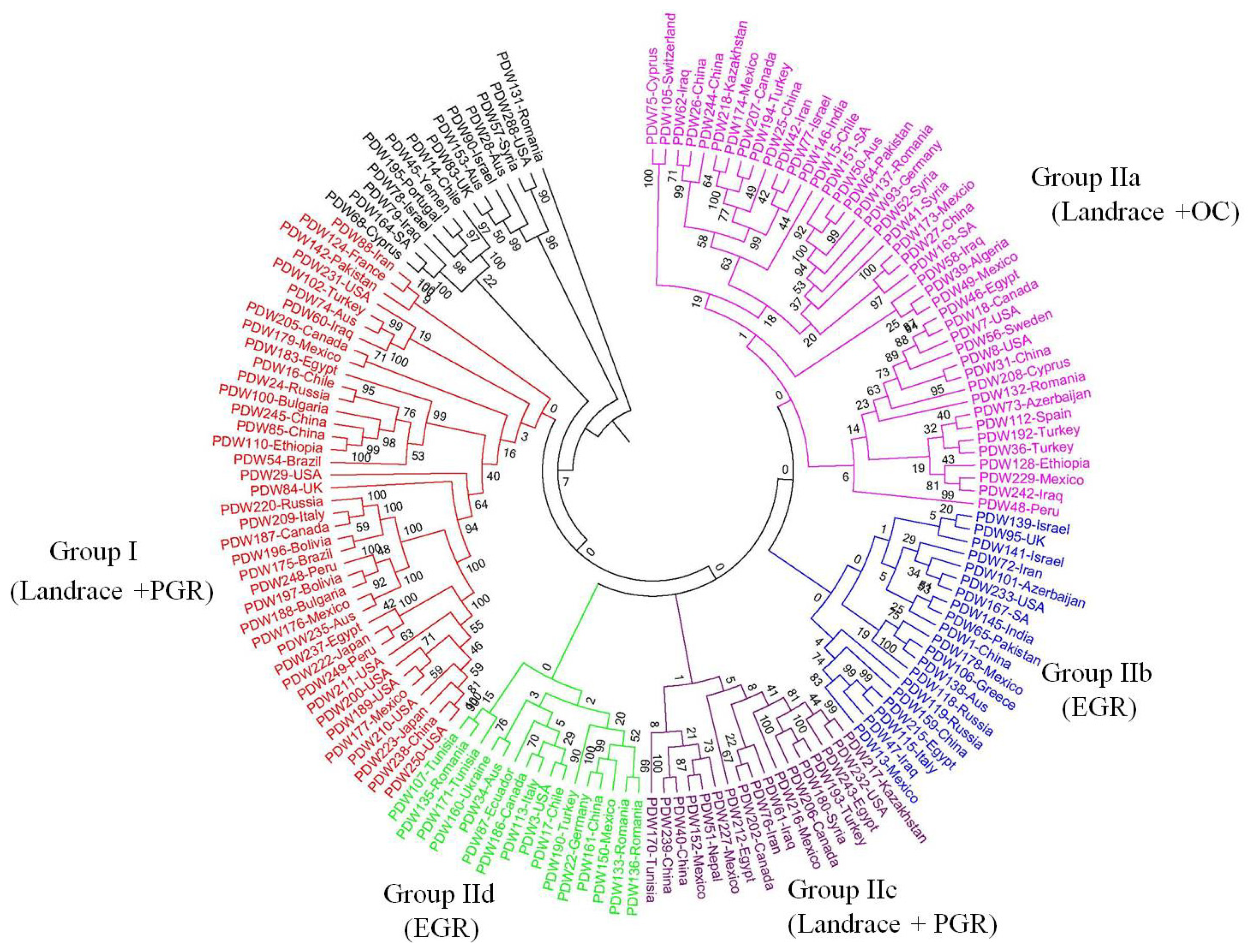
2.2. Genetic Structure
2.3. Genetic Diversity between Landraces and Cultivars
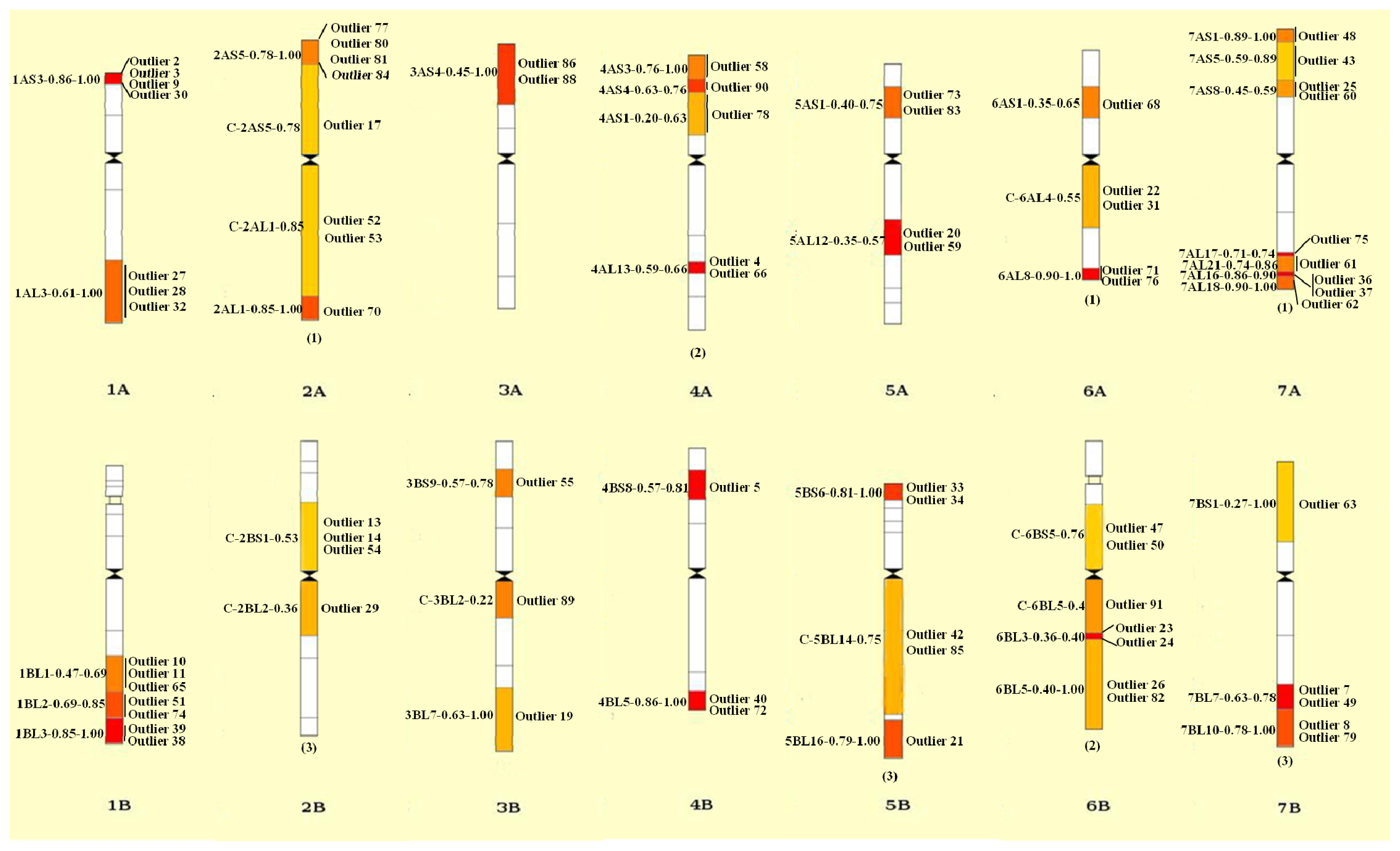
2.4. Divergence between Landraces and Cultivars
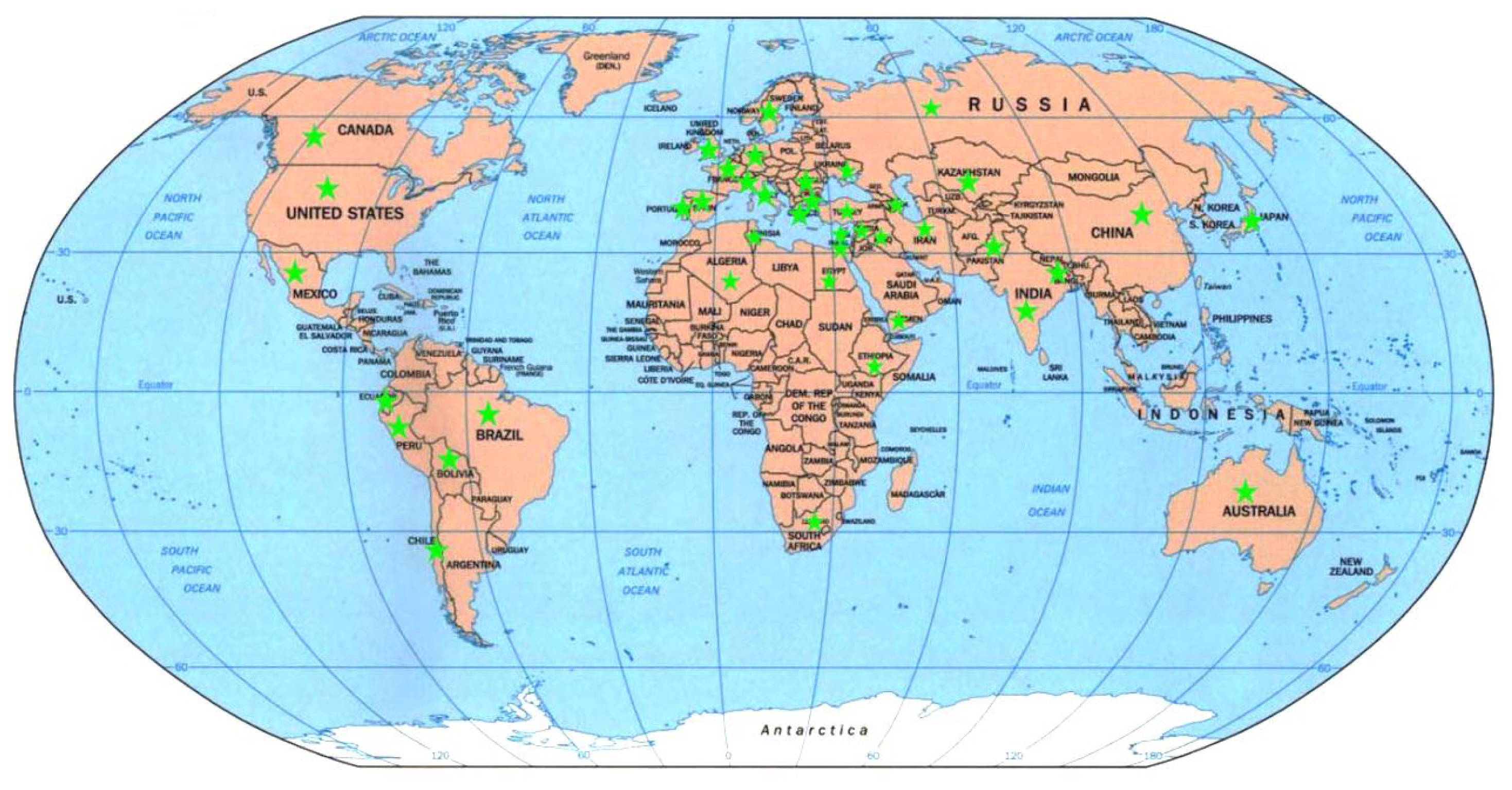
2.5. Genetic Diversity vs. Place of Origin
3. Discussion
3.1. SNP-Based Polymorphism and Genetic Diversity
3.2. Genetic Structure Raveled by SNP Markers
3.3. Genetic Diversity
3.3.1. Temporally: Genetic Diversity vs. Year of Release
3.3.2. Spatially: Genetic Diversity vs. Place of Origin
3.4. Divergence between Landraces and Cultivars Revealed by SNP Markers
4. Experimental Section
4.1. Plant Materials
4.2. Genomic DNA Extraction and SNP Genotyping
4.3. Genetic Diversity
4.4. Genetic Structure and Population Differentiation
4.5. Statistical Tests
5. Conclusions
Acknowledgments
Conflict of Interest
References
- Peng, J.H.; Sun, D.; Nevo, E. Domestication evolution, genetics and genomics in wheat. Mol. Breed 2011, 28, 281–301. [Google Scholar]
- Maccaferri, M.; Sanguineti, M.C.; Donini, P.; Tuberosa, R. Microsatellite analysis reveals a progressive widening of the genetic basis in the elite durum wheat germplasm. Theor. Appl. Genet 2003, 107, 783–797. [Google Scholar]
- Thuillet, A.C.; Bataillon, T.; Poirier, S.; Santoni, S.; David, J.L. Estimation of long-term effective population sizes through the history of durum wheat using microsatellite data. Genetics 2005, 169, 1589–1599. [Google Scholar]
- Luo, M.C.; Yang, Z.L.; You, F.M.; Kawahara, T.; Waines, J.G.; Dvorak, J. The structure of wild and domesticated emmer wheat populations, gene flow between them, and the site of emmer domestication. Theor. Appl. Genet 2007, 114, 947–959. [Google Scholar]
- Dvorak, J.; Luo, M.Ch.; Akhunov, E.D. N.I. Vavilov’s theory of centers of diversity in the light of current understanding of wheat diversity, domestication and evolution. Czech J. Genet. Plant Breed. 2011, 47, S20–S27. [Google Scholar]
- Zapata, L.; Peña-Chocarro, L.; Pérez-Jordá, G.; Stika, H.P. Early Neolithic agriculture in the Iberian Peninsula. J. World Prehist 2004, 18, 283–325. [Google Scholar]
- Crawford, D. Food: Tradition and change in Hellenistic Egypt. World Archaeol 1979, 11, 136–146. [Google Scholar]
- Moragues, M.; Moralejo, M.; Sorrells, M.E.; Royo, C. Dispersal of durum wheat [Triticum turgidum L. ssp. turgidum convar. durum (Desf.) MacKey] landraces across the Mediterranean basin assessed by AFLPs and microsatellites. Genet. Resour. Crop Evol 2007, 54, 1133–1144. [Google Scholar]
- Crosby, A.W., Jr. The Columbian Exchange: Biological and Cultural Consequences of 1492; Greenwood Press: Westport, CT, USA, 1972. [Google Scholar]
- Capparelli, A.; Lema, V.; Giovannetti, M.; Raffino, R. The introduction of Old World crops (wheat, barley and peach) in Andean Argentina during the 16th century A.D.: Archaeobotanical and ethnohistorical evidence. Veget. Hist. Archaeobot 2005, 14, 472–484. [Google Scholar]
- Soleimani, V.D.; Baum, B.R.; Johnson, D.A. AFLP and pedigree-based genetic diversity estimates in modern cultivars of durum wheat [Triticum turgidum L. subsp. durum (Desf.) Husn.]. Theor. Appl. Genet 2002, 104, 350–357. [Google Scholar]
- Autrique, E.; Nachit, M.; Monneveux, P.; Tanksley, S.D.; Sorrells, M.E. Genetic diversity in durum wheat based on RFLP, morphophysiological traits and coefficient of parentage. Crop Sci 1996, 36, 735–742. [Google Scholar]
- Peng, J.; Fahima, T.; Röder, M.S.; Li, Y.C.; Dahan, A.; Grama, A.; Ronin, Y.I.; Korol, A.B.; Nevo, E. Microsatellite tagging of the stripe-rust resistance gene YrH52 derived from wild emmer wheat, Triticum dicoccoides, and suggestive negative crossover interference on chromosome 1B. Theor. Appl. Genet 1999, 98, 862–872. [Google Scholar]
- Peng, J.; Korol, A.B.; Fahima, T.; Röder, M.S.; Ronin, Y.I.; Li, Y.C.; Nevo, E. Molecular genetic maps in wild emmer wheat, Triticum dicoccoides: Genome-wide coverage, massive negative interference, and putative quasi-linkage. Genome Res 2000, 10, 1509–1531. [Google Scholar]
- Myburg, A.A.; Cawood, M.; Wingfield, B.D.; Botha, A.M. Development of RAPD and SCAR markers linked to the Russian wheat aphid resistance gene Dn2 in wheat. Theor. Appl. Genet 1998, 96, 1162–1169. [Google Scholar]
- Vierling, R.A.; Nguyen, H.T. Use of RAPD markers to determine the genetic diversity of diploid, wheat genotypes. Theor. Appl. Genet 1992, 84, 835–838. [Google Scholar]
- Incirli, A.; Akkaya, M.S. Assessment of genetic relationships in durum wheat cultivars using AFLP markers. Genet. Resour. Crop Evol 2001, 48, 233–238. [Google Scholar]
- Medini, M.; Hamza1, S.; Rebai, A.; Baum, M. Analysis of genetic diversity in Tunisian durum wheat cultivars and related wild species by SSR and AFLP markers. Genet. Resour. Crop Evol 2005, 52, 21–31. [Google Scholar]
- Shoaib, A.; Arabi, M.I.E. Genetic diversity among Syrian cultivated and landraces wheat revealed by AFLP markers. Genet. Resour. Crop Evol 2006, 53, 901–906. [Google Scholar]
- Altintas, S.; Toklu, F.; Kafkas, S.; Kilian, B.; Brandolini, A.; Ozkan, H. Estimating genetic diversity in durum and bread wheat cultivars from Turkey using AFLP and SAMPL markers. Plant Breed 2008, 127, 9–14. [Google Scholar]
- Collard, B.C.; Mackill, D.J. Marker-Assisted selection: an approach for precision plant breeding in the twenty-first century. Philos. Trans. R. Soc. Lond. B. Biol. Sci 2008, 363, 557–572. [Google Scholar]
- Paux, E.; Sourdille, P.; Mackay, I.; Feuillet, C. Sequence-Based marker development in wheat: Advances and applications to breeding. Biotechnol. Adv 2012, 30, 1071–1088. [Google Scholar]
- Noli, E.; Teriaca, M.S.; Sanguineti, M.C.; Conti, S. Utilization of SSR and AFLP markers for the assessment of distinctness in durum wheat. Mol. Breed 2008, 22, 301–313. [Google Scholar]
- Van Inghelandt, D.; Melchinger, A.E.; Lebreton, C.; Stich, B. Population structure and genetic diversity in a commercial maize breeding program assessed with SSR and SNP markers. Theor. Appl. Genet 2010, 120, 1289–1299. [Google Scholar]
- GrainGenes: A Database for Triticeae and Avena. Available online: http://wheat.pw.usda.gov/GG2/index.shtml (assessed on 15 January 2013).
- Jones, E.S.; Sullivan, H.; Bhattramakki, D.; Smith, J.S. A comparison of simple sequence repeat and single nucleotide polymorphism marker technologies for the genotypic analysis of maize (Zea mays L.). Theor. Appl. Genet 2007, 115, 361–371. [Google Scholar]
- Rafalski, J.A. Novel genetic mapping tools in plant: SNPs and LD-based approaches. Plant Sci 2002, 162, 329–333. [Google Scholar]
- Mackay, I.; Powell, W. Methods for linkage disequilibrium mapping in crops. Trends Plant Sci 2007, 12, 57–63. [Google Scholar]
- Jannink, J.L.; Lorenz, A.J. Iwata H: Genomic selection in plant breeding: From theory to practice. Brief Funct. Genomics 2010, 9, 166–177. [Google Scholar]
- Bhattramakki, D.; Dolan, M.; Hanafey, M.; Wineland, R.; Vaske, D.; Register, J.C.; Tingey, S.V.; Rafalski, A. Insertion-Deletion polymorphisms in 3′ regions of maize genes occur frequently and can be used as highly informative genetic markers. Plant Mol. Biol 2002, 48, 539–547. [Google Scholar]
- Cho, R.J.; Mindrinos, M.; Richards, D.R.; Sapolsky, R.J.; Anderson, M.; Drenkard, E.; Dewdney, J.; Reuber, T.L.; Stammers, M.; Federspiel, N.; et al. Genome-Wide mapping with biallelic markers in Arabidopsis thaliana. Nat. Genet 1999, 23, 203–207. [Google Scholar]
- Nasu, S.; Suzuki, J.; Ohta, R.; Hasegawa, K.; Yui, R.; Kitazawa, N.; Monna, L.; Minobe, Y. Search for and analysis of single nucleotide polymorphisms (SNPs) in rice (Oryza sativa, Oryza rufipogon) and establishment of SNP markers. DNA Res 2002, 9, 163–171. [Google Scholar]
- Choi, I.Y.; Hyten, D.L.; Matukumalli, L.K.; Song, Q.; Chaky, J.M.; Quigley, C.V.; Chase, K.; Lark, K.G.; Reiter, R.S.; Yoon, M.S.; et al. A soybean transcript map: Gene distribution, haplotype and single-nucleotide polymorphism analysis. Genetics 2007, 176, 685–696. [Google Scholar]
- Kota, R.; Varshney, R.K.; Prasad, M.; Zhang, H.; Stein, N.; Graner, A. EST-Derived single nucleotide polymorphism markers for assembling genetic and physical maps of the barley genome. Funct. Integr. Genomics 2007, 8, 223–233. [Google Scholar]
- Akhunov, E.; Nicolet, C.; Dvorak, J. Single nucleotide polymorphism genotyping in polyploid wheat with the illumine Golden Gate assay. Theor. Appl. Genet 2009, 119, 507–517. [Google Scholar]
- Akhunov, E.D.; Akhunova, A.R.; Anderson, O.D.; Anderson, J.A.; Blake, N.; Clegg, M.T.; Coleman-Derr, D.; Conley, E.J.; Crossman, C.C.; Deal, K.R.; et al. Nucleotide diversity maps reveal variation in diversity among wheat genomes and chromosomes. BMC Genomics 2010, 11, 702. [Google Scholar]
- Bérard, A.; Le Paslier, M.C.; Dardevet, M.; Exbrayat-Vinson, F.; Bonnin, I.; Cenci, A.; Haudry, A.; Brunel, D.; Ravel, C. High-Throughput single nucleotide polymorphism genotyping in wheat (Triticum spp.). Plant Biotechnol. J 2009, 7, 364–374. [Google Scholar]
- Chao, S.; Zhang, W.; Akhunov, E.; Sherman, J.; Ma, Y.; Luo, M.C.; Dubcovsky, J. Analysis of gene-derived SNP marker polymorphism in US wheat (Triticum aestivum L.) cultivars. Mol. Breed 2009, 23, 23–33. [Google Scholar]
- Edwards, K.J.; Reid, A.L.; Coghill, J.A.; Berry, S.T.; Barker, G.L. Multiplex single nucleotide polymorphism (SNP)-based genotyping in allohexaploid wheat using padlock probes. Plant Biotechnol. J 2009, 7, 375–390. [Google Scholar]
- Kozlova, S.A.; Khlestkina, E.K.; Salina, E.A. Specific features in using SNP markers developed for allopolyploid wheat. Russ. J. Genet 2009, 45, 81–84. [Google Scholar]
- Somers, D.J.; Kirkpatrick, R.; Moniwa, M.; Walsh, A. Mining single-nucleotide polymorphisms from hexaploid wheat ESTs. Genome 2003, 46, 431–437. [Google Scholar]
- Hoisington, D.; Khairallah, M.; Reeves, T.; Ribaut, J.M.; Skovmand, B.; Taba, S.; Warburton, M. Plant genetic resources: What can they contribute toward increased crop productivity? Proc. Natl. Acad. Sci. USA 1999, 96, 5937–5943. [Google Scholar]
- Donini, P.; Law, J.R.; Koebner, R.M.D.; Reeves, J.C.; Cooke, R.J. Temporal trends in the diversity of UK wheats. Theor. Appl. Genet 2000, 100, 912–917. [Google Scholar]
- Martos, V.; Royo, C.; Rharrabti, Y.; Garcia del Morala, L.F. Using AFLPs to determine phylogenetic relationships and genetic erosion in durum wheat cultivars released in Italy and Spain throughout the 20th century. Field Crop Res 2005, 91, 107–116. [Google Scholar]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol 2005, 14, 2611–2620. [Google Scholar]
- Karagöz, A.; Zencirci, N. Variation in wheat (Triticum spp.) landraces from different altitudes of three regions of Turkey. Genet. Resour. Crop Evol 2005, 52, 75–785. [Google Scholar]
- Zencirci, N.; Karagoz, A. Effect of developmental stages length on yield and some quality traits of Turkish durum wheat (Triticum turgidum L. convar. durum (Desf.) Mackey) landraces: Influence of developmental stages length on yield and quality of durum wheat. Genet. Resour. Crop Evol 2005, 52, 765–774. [Google Scholar]
- Hedden, P. The genes of the green revolution. Trends Genet 2003, 19, 5–9. [Google Scholar]
- Sim, S.C.; Robbins, M.D.; van Deynze, A.; Michel, A.P.; Francis, D.M. Population structure and genetic differentiation associated with breeding history and selection in tomato (Solanumly copersicum L.). Heredity 2011, 106, 927–935. [Google Scholar]
- Antao, T.; Lopes, A.; Lopes, R.; Beja-Pereira, A.; Luikart, G. LOSITAN: A workbench to detect molecular adaptation based on a Fst-outlier method. BMC Bioinforma 2008, 9, 323. [Google Scholar]
- Liu, C.J.; Atkinson, M.D.; Chinoy, C.N.; Devos, K.M.; Gale, M.D. Nonhomoeologous translocations between group 4, 5 and 7 chromosomes within wheat and rye. Theor. Appl. Genet 1992, 83, 305–312. [Google Scholar]
- Devos, K.M.; Dubcovsky, J.; Dvorak, J.; Chinoy, C.N. Structural evolution of wheat chromosomes 4A, 5A and 7B and on their recombination. Theor. Appl. Genet 1995, 91, 282–288. [Google Scholar]
- Röder, M.S.; Korzun, V.; Wendehake, K.; Plaschke, J.; Tixier, M.H.; Leroy, P.; Ganal, M.W. A microsatellite map of wheat. Genetics 1998, 149, 2007–2023. [Google Scholar]
- Liu, Y.G.; Tsunewaki, K. Restriction fragment length polymorphism analysis of wheat. II. Linkage maps of the RFLP sites in common wheat. Jpn. J. Genet 1991, 66, 617–633. [Google Scholar]
- Li, Y.; Fahima, T.; Korol, A.B.; Peng, J.; Röder, M.S.; Kirzhner, V.; Beiles, A.; Nevo, E. Microsatellite diversity correlated with ecological-edaphic and genetic factors in three microsites of wild emmer wheat in North Israel. Mol. Biol. Evol 2000, 17, 851–862. [Google Scholar]
- Fu, Y.B.; Peterson, G.W.; Richards, K.W.; Somers, D.; de Pauw, R.M.; Clarke, J.M. Allelic reduction and genetic shift in the Canadian hard red spring wheat germplasm released from 1845 to 2004. Theor. Appl. Genet 2005, 110, 1505–1516. [Google Scholar]
- Fu, Y.B.; Peterson, G.W.; Yu, J.K.; Gao, L.F.; Jia, J.Z.; Richards, K.W. Impact of plant breeding on genetic diversity of the Canadian hard red spring wheat germplasm as revealed by EST-derived SSR markers. Theor. Appl. Genet 2006, 112, 1239–1247. [Google Scholar]
- Fu, Y.B.; Somers, D.J. Genome-Wide reduction of genetic diversity in wheat breeding. Crop Sci 2009, 49, 161–168. [Google Scholar]
- Nevo, E.; Fu, Y.B.; Pavlicek, T.; Khalifa, S.; Tavasi, M.; Beiles, A. Evolution of wild cereals during 28 years of global warming in Israel. Proc. Natl. Acad. Sci. USA 2012, 109, 3412–3415. [Google Scholar]
- Evenson, R.E.; Gollin, D. Assessing the impact of the Green Revolution, 1960 to 2000. Science 2003, 300, 758. [Google Scholar]
- Reif, J.C.; Zhang, P.; Dreisigacker, S.; Warburton, M.L.; van Ginkel, M.; Hoisington, D.; Bohn, M.; Melchinger, A.E. Wheat genetic diversity trends during domestication and breeding. Theor. Appl. Genet 2005, 110, 859–864. [Google Scholar]
- Rajaram, S.; Saari, E.E.; Hettel, G.P. Durum Wheats: Challenges and Opportunities; Wheat Special Report No. 9; International Maize and Wheat Improvement Center (CIMMYT): Mexico City, Mexico, 1992. [Google Scholar]
- Rajaram, S.; van Ginkel, M. Mexico: 50 Years of International Wheat Breeding. In The World Wheat Book: A history of wheat breading; Bonjean, A.P., Angus, W.J., Eds.; Lavoisier: Paris, France, 2001; pp. 579–610. [Google Scholar]
- Reeves, T.; Rajaram, S.; van Ginkel, M.; Trethowan, R.; Braun, H.; Cassaday, K. New Wheats for a Secure, Sustainable Future; International Maize and Wheat Improvement Center (CIMMYT): Mexico City, Mexico, 1999. [Google Scholar]
- Germplasm Resources Information Network (GRIN). Available online: http://www.ars-grin.gov/npgs/acc/acc_queries.html (assessed on 15 January 2013).
- Vavilov, N.I. Phytogeographic basis of plant breeding: The origin, variation, immunity and breeding of cultivated plants. Chronica Bot 1951, 13, 1–366. [Google Scholar]
- Teklu, Y.; Hammer, K.; Röder, M.S. Simple sequence repeats marker polymorphism in emmer wheat (Triticum dicoccon Schrank): Analysis of genetic diversity and differentiation. Genet. Resour. Crop Evol 2007, 54, 543–554. [Google Scholar]
- Harlan, J.R. The great plains region (Part 4). Agric. Food Chem 1955, 3, 29–31. [Google Scholar]
- Harlan, J.R. Agricultural origins: Centers and noncenters. Science 1971, 174, 468–474. [Google Scholar]
- Byerlee, D.; Moya, P. Impacts of International Wheat Breeding Research in the Developing World: 1966–90; International Maize and Wheat Improvement Center (CIMMYT): Mexico City, Mexico, 1993. [Google Scholar]
- Rajaram, S. Wheat Breeding at CIMMYT: Commemorating 50 Years of Research in Mexico for Global Wheat Improvement; Wheat Special Report No 29; International Maize and Wheat Improvement Center (CIMMYT): Mexico City, Mexico, 1994. [Google Scholar]
- Peng, J.; Wang, H.; Haley, S.D.; Peairs, F.B.; Lapitan, N.L.V. Molecular mapping of the Russian wheat aphid resistance gene Dn2414 in wheat. Crop Sci 2007, 47, 2418–2429. [Google Scholar]
- Wheat SNP Database. Available online: http://probes.pw.usda.gov:8080/snpworld/Search (assessed on 15 January 2013).
- Chao, S.; Dubcovsky, J.; Dvorak, J.; Luo, M.C.; Baenziger, S.P.; Matnyazov, R.; Clark, D.R.; Talbert, L.E.; Anderson, J.A.; Dreisigacker, S.; et al. Population- and genome-specific patterns of linkage disequilibrium and SNP variation in spring and winter wheat (Triticum aestivum L.). BMC Genomics 2010, 11, 727. [Google Scholar]
- Luo, M.C.; Deal, K.R.; Akhunov, E.D.; Akhunova, A.R.; Anderson, O.D.; Anderson, J.A.; Blake, N.; Clegg, M.T.; Coleman-Derr, D.; Conley, E.J.; et al. Genome comparisons reveal a dominant mechanism of chromosome number reduction in grasses and accelerated genome evolution in Triticeae. Proc. Natl. Acad. Sci. USA 2009, 106, 15780–15785. [Google Scholar]
- Liu, K.; Muse, S.V. Powermarker: An integrated analysis environment for genetic marker analysis. Bioinformatics 2005, 21, 2128–2129. [Google Scholar]
- Weir, B.S. Genetic Data Analysis II; Sinauer Associates, Inc: Sunderland, MA, USA, 1996. [Google Scholar]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar]
- Felsenstein, J. PHYLIP (Phylogeny Inference Package) Version 3.66; Department of Genome Sciences, University of Washington: Seattle, WA, USA, 2006. [Google Scholar]
- Rambaut, A. FigTree, version 1.3.1. Available online: http://tree.bio.ed.ac.uk/software/figtree/ (assessed on 15 January 2013).
- Excoffier, L.; Laval, G.; Schneider, S. Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evol. Bioinform 2005, 1, 47–50. [Google Scholar]
- SPSS Web site. Available online: http://www.spss.com (assessed on 15 January 2013).
| Chromosome | No. of SNP Markers | No. of Polymorphic Markers | Gene Diversity | PIC |
|---|---|---|---|---|
| A Genome | ||||
| 1A | 114 | 75 | 0.2319 | 0.1905 |
| 2A | 96 | 65 | 0.2180 | 0.1840 |
| 3A | 98 | 67 | 0.2036 | 0.1697 |
| 4A | 124 | 86 | 0.1899 * | 0.1576 * |
| 5A | 85 | 59 | 0.2179 | 0.1798 |
| 6A | 125 | 78 | 0.2526 * | 0.2072 * |
| 7A | 135 | 88 | 0.2249 | 0.1884 |
| Subtotal/Mean | 767 | 516 | 0.2193 | 0.1819 |
| B Genome | ||||
| 1B | 99 | 76 | 0.2695 * | 0.2225 * |
| 2B | 87 | 64 | 0.2553 | 0.2097 |
| 3B | 67 | 49 | 0.2180 * | 0.1832 |
| 4B | 75 | 46 | 0.2200 * | 0.1804 * |
| 5B | 76 | 49 | 0.2120 * | 0.1747 * |
| 6B | 105 | 83 | 0.2211 * | 0.1842 |
| 7B | 101 | 70 | 0.2404 | 0.1982 |
| Subtotal/Mean | 599 | 430 | 0.2384 | 0.1970 |
| Hemoeologous | ||||
| 1 | 213 | 151 | 0.2508 * | 0.2066 * |
| 2 | 183 | 129 | 0.2365 | 0.1967 |
| 3 | 165 | 116 | 0.2097 * | 0.1754 * |
| 4 | 199 | 132 | 0.2004 * | 0.1656 * |
| 5 | 161 | 108 | 0.2153 * | 0.1775 * |
| 6 | 230 | 161 | 0.2364 | 0.1953 |
| 7 | 236 | 158 | 0.2318 | 0.1927 |
| Total/Grand mean | 1366 | 946 | 0.2280 | 0.1888 |
| Sample Size | No. of Polymorphic Marker | Polymorphic Rate (%) | Gene Diversity * | PIC * | |
|---|---|---|---|---|---|
| Improvement status | |||||
| Landrace | 53 | 756 | 79.9% | 0.2192 b | 0.1800 b |
| Cultivar | 97 | 933 | 98.6% | 0.2310 a | 0.1919 a |
| Time group† | |||||
| Landrace | 53 | 756 | 79.9% | 0.2192 b | 0.1800 b |
| OC | 32 | 757 | 80.0% | 0.2192 b | 0.1807 b |
| EGR | 35 | 728 | 77.0% | 0.2034 c | 0.1680 c |
| PGR | 30 | 825 | 87.2% | 0.2474 a | 0.2039 a |
| Source of Variation | Sum of Squares | Percentage of Variation (%) |
|---|---|---|
| Among Populations | 321.84 | 0.50 |
| Within Population (Cultivar) | 42,400.65 | 65.54 |
| Within Population (landrace) | 21,977.11 | 33.97 |
| Total | 64,699.60 | 100.00 |
| Group | Sample Size | Mean Plant Height, cm (SE) |
|---|---|---|
| Landrace | 53 | 132.46 (1.91) a |
| OC | 32 | 130.72 (2.48) a |
| EGR | 35 | 119.13 (4.05) b |
| PGR | 30 | 101.91 (4.27) c |
| SNP marker and the EST | Gene function and the homologous EST | ||||||
|---|---|---|---|---|---|---|---|
| Code | SNP Marker | Accession No. | Map position (Bin) | Function | Accession No. | Identity (%) | E-value |
| Outlier 1 | AY244508_5_B_Y_26 | AY244508 | 5B | G1777 MADS-box transcriptional factor (AP1) gene, T. monococcum | AY244508.1 | ||
| Outlier 2 | BE405518_1_A_95 | BE405518 | 1AS3-0.86–1.00 | Alternative splicing regulator (RSZ38), T. aestivum | DQ019628.1 | 93% | 0 |
| Outlier 3 | BE405518_1_A_Y_106 | BE405518 | 1AS3-0.86–1.00 | Alternative splicing regulator (RSZ38), T. aestivum | DQ019628.1 | 93% | 0 |
| Outlier 4 | BE442666_4_A_269 | BE442666 | 4AL13-0.59–0.66 | Lipoxygenase 3 (LOX3), T. aestivum | HQ913602.1 | 99% | 0 |
| Outlier 5 | BE442666_4_B_Y_327 | BE442666 | 4BS8-0.57–0.81 | Lipoxygenase 3 (LOX3), T. aestivum | HQ913602.1 | 99% | 0 |
| Outlier 6 | BE404341_5_B_Y_124 | BE404341 | 5B | Phytochelatin synthetase, T. aestivum | AY442329.1 | 98% | 0 |
| Outlier 7 | BE406148_7_B_Y_647 | BE406148 | 7BL7-0.63–0.78 | Cyclophilin B-B gene, T. aestivum | EU627095.1 | 100% | 9 × 10−101 |
| Outlier 8 | BE445506_7_B_Y_355 | BE445506 | 7BL10-0.78–1.00 | Unknown | |||
| Outlier 9 | BE405834_1_A_N_641 | BE405834 | 1AS3-0.86–1.00 | Soluble inorganic pyrophosphatase-like, B. distachyon | XM_003568957.1 | 91% | 0 |
| Outlier 10 | BE405834_1_B_Y_216 | BE405834 | 1BL1-0.47–0.69 | Soluble inorganic pyrophosphatase-like, B. distachyon | XM_003568957.1 | 91% | 0 |
| Outlier 11 | BE446240_1_B_131 | BE446240 | 1BL1-0.47–0.69 | Rab GDP dissociation inhibitor, B. distachyon | XM_003568390.1 | 93% | 0 |
| Outlier 12 | BE403177_2_B_409 | BE403177 | 2B | F-box protein 7-like, B. distachyon | XM_003579715.1 | 90% | 3 × 10−136 |
| Outlier 13 | BE404332_2_B_29 | BE404332 | C-2BS4-0.75 * | Ribosomal protein S12 (rps12), H. vulgare | AF067732.1 | 94% | 0 |
| Outlier 14 | BE444144_2_B_92 | BE444144 | 2BS | Unknown | |||
| Outlier 15 | BE445278_2_B_143 | BE445278 | 2B | RuvB-like 2-like, B. distachyon | XM_003562775.1 | 92% | 0 |
| Outlier 16 | BE445278_2_B_243 | BE445278 | 2B | RuvB-like 3-like, B. distachyon | XM_003562775.1 | 92% | 0 |
| Outlier 17 | BE445242_2_A_362 | BE445242 | C-2AS5-0.78 | Unknown | |||
| Outlier 18 | BE444579_3_B_Y_375 | BE444579 | 3B | Unknown | |||
| Outlier 19 | BE444864_3_B_373 | BE444864 | 3BL7-0.63–1.00 | C2 domain-containing protein C31G5.15-like, B. distachyon | XR_138068.1 | 91% | 0 |
| Outlier 20 | BE443187_5_A_511 | BE443187 | 5AL12-0.35–0.57 | 65-kDa microtubule-associated protein 7-like, B. distachyon | XM_003578156.1 | 88% | 0 |
| Outlier 21 | CD373602_5_B_Y_310 | CD373602 | 5BL16-0.79–1.00 | Unknown | |||
| Outlier 22 | BE444256_6_A_N_1118 | BE444256 | C-6AL4-0.55 | Alcohol dehydrogenase-like 6-like, B. distachyon | XM_003569903.1 | 93% | 0 |
| Outlier 23 | CD452643_6_B_111 | CD452643 | 6BL3 | Alcohol dehydrogenase-like 6-like, B. distachyon | XM_003569903.1 | 92% | 1 × 10−117 |
| Outlier 24 | CD452643_6_B_Y_113 | CD452643 | 6BL3 | Alcohol dehydrogenase-like 6-like, B. distachyon | XM_003569903.1 | 92% | 1 × 10−117 |
| Outlier 25 | BE446380_7_A_577 | BE446380 | 7AS8-0.45–0.59 | Putative phospholipid-transporting ATPase 9-like, B. distachyon | XM_003563827.1 | 91% | 0 |
| Outlier 26 | BE403950_6_B_Y_325 | BE403950 | 6BL5-0.40–1.00 | ABC transporter F family member 3-like, B. distachyon | XM_003570443.1 | 93% | 0 |
| Outlier 27 | BE517729_1_A_116 | BE517729 | 1AL3-0.61–1.00 | Putative prolyl aminopeptidase 1 (PAP1), T. durum × Secalecereale | JN808306.2 | 97% | 0 |
| Outlier 28 | BE517729_1_A_Y_117 | BE517729 | 1AL3-0.61–1.00 | Putative prolyl aminopeptidase 1 (PAP1), T. durum × Secalecereale | JN808306.2 | 97% | 0 |
| Outlier 29 | BE517831_2_B_70 | BE517831 | C-2BL2-0.36 | Phosphoinositide-specific phospholipase C1, T. aestivum | HM754654.1 | 95% | 0 |
| Outlier 30 | BF200531_1_A_N_573 | BF200531 | 1AS3-0.86–1.00 | Protein notum homolog, B. distachyon | XM_003566643.1 | 94% | 4 × 10−169 |
| Outlier 31 | BF474493_6_A_N_40 | BF474493 | C-6AL4-0.55 | Pescadillo homolog, B. distachyon | XM_003560899.1 | 91% | 0 |
| Outlier 32 | BF474139_1_A_144 | BF474139 | 1AL3-0.61–1.00 | 6 phosphofructo kinase 3-like, B. distachyon | XM_003568020.1 | 95% | 6 × 10−157 |
| Outlier 33 | BF201102_5_B_444 | BF201102 | 5BS6-0.81–1.00 | Methionine synthase 1 enzyme (ms1 gene), Hordeum vulgare | AM039904.1 | 93% | 2 × 10−168 |
| Outlier 34 | BF201102_5_B_Y_373 | BF201102 | 5BS6-0.81–1.00 | Methionine synthase 1 enzyme (ms1 gene), Hordeum vulgare | AM039904.1 | 93% | 2 × 10−168 |
| Outlier 35 | CD453605_6_B_427 | CD453605 | 6B | Putative nitric oxide synthase-like, B. distachyon | XM_003570728.1 | 89% | 2 × 10−179 |
| Outlier 36 | BF474379_7_A_83 | BF474379 | 7AL16-0.86–0.90 | Protein N-terminal asparagine amidohydrolase-like, B. distachyon | XM_003563571.1 | 90% | 0 |
| Outlier 37 | BF474379_7_A_Y_253 | BF474379 | 7AL16-0.86–0.90 | Protein N-terminal asparagine amidohydrolase-like, B. distachyon | XM_003563571.1 | 90% | 0 |
| Outlier 38 | BE494527_1_B_77 | BE494527 | 1BL2-0.0.69–0.85 | Phosphoethanolamine methyltransferase, T. aestivum | AY065971.1 | 96% | 3 × 10−86 |
| Outlier 39 | BE494527_1_B_Y_438 | BE494527 | 1BL2-0.0.69–0.85 | Phosphoethanolamine methyltransferase, T. aestivum | AY065971.1 | 96% | 3 × 10−86 |
| Outlier 40 | BE494765_4_B_Y_426 | BE494765 | 4BL5-0.86–1.00 | Unknown | |||
| Outlier 41 | BE636872_6_A_119 | BE636872 | 6A | Unknown | |||
| Outlier 42 | BE495277_5_B_336 | BE495277 | C-5BL14-0.75 * | UPF0664 stress-induced protein C29B12.11c-like, B. distachyon | XM_003578371.1 | 91% | 2 × 10−137 |
| Outlier 43 | BE493868_7_A_Y_93 | BE493868 | 7AS5-0.59–0.89 | Probable protein phosphatase 2C 54-like, B. distachyon | XM_003564166.1 | 91% | 0 |
| Outlier 44 | BE494482_7_B_Y_29 | BE494482 | 7B | Zuxin response factor 21 (ARF21) gene, Zea mays | HM004536.1 | 92% | 3 × 10−67 |
| Outlier 45 | CD491758_6_A_Y_81 | CD491758 | 6A | Calcium-dependent protein kinase-like (CPK10), T. aestivum | EU181189.1 | 92% | 0 |
| Outlier 46 | BQ159615_6_B_Y_336 | BQ159615 | 6B | Leucine-rich repeat protein (LRR2), T. aestivum | EF555120.1 | 98% | 0 |
| Outlier 47 | BF291774_6_B_181 | BF291774 | 6BSc | Putative vacuolar cation/proton exchanger 4-like, B. distachyon | XM_003570864.1 | 83% | 0 |
| Outlier 48 | BF292264_7_A_712 | BF292264 | 7AS1-0.89–1.00 | Unknown | |||
| Outlier 49 | BF292193_7_B_N_78 | BF292193 | 7BL7-0.63–0.78 | Cytochrome b5 (cb5-1 gene), Oryza sativa | AJ429043.1 | 84% | 8 × 10−103 |
| Outlier 50 | BF291774_6_B_519 | BF291774 | 6BSc | Putative vacuolar cation/proton exchanger 4-like, B. distachyon | XM_003570864.1 | 83% | 0 |
| Outlier 51 | BG263233_1_B_825 | BG263233 | 1BL2-0.0.69–0.85 | Flap endonuclease 1-A-like, B. distachyon | XM_003567949.1 | 91% | 0 |
| Outlier 52 | BG605368_2_A_156 | BG605368 | C-2AL1-0.85 | Exopolygalacturonase-like, B. distachyon | XM_003571584.1 | 86% | 4 × 10−136 |
| Outlier 53 | BG605368_2_A_Y_310 | BG605368 | C-2AL1-0.85 | Exopolygalacturonase-like, B. distachyon | XM_003571584.1 | 86% | 4 × 10−136 |
| Outlier 54 | BG263521_2_B_Y_261 | BG263521 | C-2BS1-0.53 | Mitogen activated protein kinase (MEK1), O, sativa | AF080436.1 | 83% | 4 × 10−141 |
| Outlier 55 | BF203070_3_B_Y_52 | BF203070 | 3BS9-0.57–0.78 | Unknown | |||
| Outlier 56 | BE637808_4_A_Y_332 | BE637808 | 4A | DEAD-box ATP-dependent RNA helicase 16-like, B. distachyon | XM_003559423.1 | 90% | 4 × 10−165 |
| Outlier 57 | BF482950_4_A_Y_272 | BF482950 | 4A | Lariat debranching enzyme-like, B. distachyon | XM_003559432.1 | 90% | 7 × 10−117 |
| Outlier 58 | BF483551_4_A_N_203 | BF483551 | 4AS3-0.76–1.00 | Unknown | |||
| Outlier 59 | BE497820_5_A_Y_664 | BE497820 | C-5AL10-0.57 * | Probable thylakoidal processing peptidase 2, chloroplastic-like, B. distachyon | XM_003578166.1 | 89% | 0 |
| Outlier 60 | BE498662_7_A_Y_513 | BE498662 | 7AS8-0.45–0.59 | Unknown | |||
| Outlier 61 | BF482403_7_A_126 | BF482403 | 7AL21-0.74–0.86 | Unknown | |||
| Outlier 62 | BQ169669_7_A_Y_378 | BQ169669 | 7AL18 | Unknown | |||
| Outlier 63 | BE499248_7_B_Y_63 | BE499248 | 7BS1-0.27–1.00 | Caffeoyl-CoA O-methyltransferase 2, B. distachyon | XM_003564219.1 | 95% | 6 × 10−153 |
| Outlier 64 | BF485380_7_B_Y_479 | BF485380 | 7B | Unknown | |||
| Outlier 65 | BM140362_1_B_432 | BM140362 | 1BL1-0.47–0.69 | Glyoxysomal processing protease, glyoxysomal-like, B. distachyon | XM_003568135.1 | 89% | 0 |
| Outlier 66 | BG604678_4_A_Y_256 | BG604678 | 4AL13-0.59–0.66 | Phytanoyl-CoA dioxygenase domain-containing protein 1-like, B. distachyon | XM_003560712.1 | 92% | 0 |
| Outlier 67 | CD453913_7_A_105 | CD453913 | 7A | Phosphoserine phosphatase, chloroplastic-like, B. distachyon | XM_003577403.1 | 89% | 2 × 10−179 |
| Outlier 68 | BG262421_6_A_87 | BG262421 | 6AS1-0.35–0.65 | Purple acid phosphatase 18-like, B. distachyon | XM_003562305.1 | 91% | 0 |
| Outlier 69 | BG262287_7_B_Y_175 | BG262287 | 7B | Vacuolar proton-ATPase subunit A, T. aestivum | DQ432014.1 | 99% | 0 |
| Outlier 70 | BE490763_2_A_1462 | BE490763 | 2AL1-0.85–1.00 | Endoplasmic reticulum metallopeptidase 1-like, B. distachyon | XM_003580100.1 | 88% | 0 |
| Outlier 71 | BE471213_6_A_N_28 | BE471213 | 6AL8-0.90–1.00 | Metal tolerance protein C2-like, B. distachyon | XM_003570688.1 | 92% | 6 × 10−178 |
| Outlier 72 | BE591172_4_B_Y_148 | BE591172 | 4BL5-0.86–1.00 | Phytoenedesaturase (PDS), T. aestivum | FJ517553.1 | 98% | 0 |
| Outlier 73 | BE591974_5_A_1534 | BE591974 | 5AS1-0.40–0.75 | Unknown | |||
| Outlier 74 | BE591290_1_B_Y_289 | BE591290 | 1BL2-0.0.69–0.85 | B73 WTF1 gene, Zea mays cultivar | FJ264201.1 | 82% | 2 × 10−134 |
| Outlier 75 | BE591002_7_A_244 | BE591002 | 7AL17-0.71–0.74 | Probable alanyl-t RNA synthetase, chloroplastic-like, transcript variant 2, B. distachyon | XM_003563964.1 | 85% | 2 × 10−108 |
| Outlier 76 | BE591777_6_A_Y_394 | BE591777 | 6AL8-0.90–1.00 | PAP-specific phosphatase HAL2-like, B. distachyon | XM_003570307.1 | 89% | 1 × 10−128 |
| Outlier 77 | BE497494_2_A_Y_475 | BE497494 | 2AS5-0.78–1.00 | GLU gene for ferredoxin-dependent glutamate synthase precursor, O. sativa | AB061357.1 | 96% | 0 |
| Outlier 78 | BE497224_4_A_Y_41 | BE497224 | 4AS1-0.20–0.63 | Unknown | |||
| Outlier 79 | BE605194_7_B_Y_583 | BE605194 | 7BL10-0.78–1.00 | Serine/threonine-protein kinase At5g01020-like, B. distachyon | XM_003563310.1 | 92% | 2 × 10−131 |
| Outlier 80 | BG275030_2_A_96 | BG275030 | 2AS5-0.78–1.00 | Symplekin-like, B. distachyon | XM_003559695.1 | 91% | 4 × 10−144 |
| Outlier 81 | BG275030_2_A_Y_103 | BG275030 | 2AS5-0.78–1.00 | Symplekin-like, B. distachyon | XM_003559695.1 | 91% | 4 × 10−144 |
| Outlier 82 | BF475120_6_B_Y_75 | BF475120 | 6BL5-0.40–1.00 | Unknown | |||
| Outlier 83 | BG313707_5_A_Y_547 | BG313707 | 5AS1-0.40–0.75 | 2 oxoglutarate/malate translocator, chloroplastic-like, B. distachyon | XM_003575906.1 | 93% | 3 × 10−160 |
| Outlier 84 | BG314532_2_A_Y_446 | BG314532 | 2AS5-0.78–1.00 | Unknown | |||
| Outlier 85 | BQ168780_5_B_995 | BQ168780 | C-5BL14–0.75 * | Actin-related protein 2/3 complex subunit 5-like, B. distachyon | XM_003577407.1 | 92% | 1 × 10−145 |
| Outlier 86 | BG314551_3_A_Y_162 | BG314551 | 3AS4-0.45–1.00 | 66 kDa stress protein-like, B. distachyon | XM_003567837.1 | 87% | 4 × 10−176 |
| Outlier 87 | BQ168329_2_A_Y_198 | BQ168329 | 2A | Protoporphyrin IX Mg-chelatase subunit precursor (Xantha-f) gene, H. vulgare | U26916.1 | 97% | 0 |
| Outlier 88 | BE426222_3_A_68 | BE426222 | C-3AS2-0.23 | Topless-related protein 2-like, transcript variant 1, B. distachyon | XM_003566383.1 | 91% | 0 |
| Outlier 89 | BE489326_3_B_Y_300 | BE489326 | C-3BL2-0.22 | CTD-phosphatase-like protein, Zea mays | NM_001155943.1 | 80% | 1 × 10−115 |
| Outlier 90 | BE425301_4_A_Y_160 | BE425301 | 4AS4-0.63–0.76 | 40S ribosomal protein gene, T. aestivum | AF479043.1 | 99 | 5 × 10−175 |
| Outlier 91 | BE426413_6_B_286 | BE426413 | C-6BL5-0.40 * | Adenosine kinase 2-like, B. distachyon | XM_003575347.1 | 94% | 0 |
| Outlier 92 | BJ291318_5_B_Y_120 | BJ291318 | 5B | 60S ribosomal protein L23a-like, B. distachyon | XM_003557882.1 | 87% | 2 × 10−179 |
| Origin | Sample Size | Gene Diversity | PIC |
|---|---|---|---|
| East-Asia | 15 | 0.2220 | 0.1798 |
| Eastern-Europe | 15 | 0.2183 | 0.1792 |
| Latin-America | 12 | 0.2518 | 0.2044 |
| Middle-East | 32 | 0.1906 | 0.1549 |
| North-Africa | 12 | 0.2054 | 0.1682 |
| North-America | 33 | 0.2351 | 0.1937 |
| Oceania | 7 | 0.2179 | 0.1747 |
| South-Africa | 4 | 0.1591 | 0.1252 |
| South-Asia | 6 | 0.1575 | 0.1258 |
| Western-Europe | 14 | 0.2299 | 0.1902 |
| Geographical Region of Origin | Country | Region within Country | Code | Accession Identifier# | Collection Year | Latitude | Longitude | Elevation |
|---|---|---|---|---|---|---|---|---|
| East Asia (15) | China | Heilongjiang | PDW1 | CItr 11495 | 1932 | 48.00N | 128.00E | |
| Heilongjiang | PDW238 * | PI 70658 | 1926 | 45.75N | 126.65E | 140 | ||
| Heilongjiang | PDW239 * | PI 70662 | 1926 | 45.76N | 126.66E | 140 | ||
| Heilongjiang | PDW245 * | PI 79900 | 1929 | |||||
| Xinjiang | PDW161 | PI 447421 | 1980 | |||||
| Jiangsu | PDW40 * | PI 124292 | 1937 | 31.75N | 120.25E | |||
| Jiangsu | PDW244 * | PI 74830 | 1927 | 33N | 120E | |||
| Beijing | PDW27 * | CItr 5094 | 1916 | 39.93N | 116.40E | 62 | ||
| Sichuan | PDW31 * | CItr 8327 | 1924 | 28.83N | 104.58E | 452 | ||
| unknown | PDW25 * | CItr 5077 | 1916 | |||||
| unknown | PDW26 * | CItr 5083 | 1916 | |||||
| unknown | PDW85 | PI 283853 | 1962 | |||||
| unknown | PDW159 | PI 435100 | 1979 | |||||
| Japan | Hokkaido | PDW222 * | PI 61351 | 1924 | 40.71N | 142.50E | ||
| Hokkaido | PDW223 * | PI 61352 | 1924 | 40.72N | 142.51E | |||
| Central Asia (2) | Kazakhstan | Kazakhstan | PDW217 * | PI 61112 | 1924 | 50.47N | 80.22E | 220 |
| Kazakhstan | PDW218 * | PI 61123 | 1924 | 50.48N | 80.23E | 220 | ||
| South Asia (6) | Nepal | Sonsera | PDW51 * | PI 176228 | 1949 | 2128 | ||
| Pakistan | Punjab | PDW64 | PI 210910 | 1953 | 31.00N | 72.00E | ||
| Punjab | PDW65 | PI 210911 | 1953 | 31.01N | 72.01E | |||
| Punjab | PDW142 * | PI 388132 | 1974 | 31.02N | 72.02E | |||
| India | Madhya Pradesh, | PDW145 * | PI 41015 | 1915 | 22.00N | 79.00E | ||
| Gujarat | PDW146 * | PI 41342 | 1915 | 21.70N | 72.97E | |||
| Middle East (32) | Turkey | Ankara | PDW36 | PI 109588 | 1935 | 39.53N | 32.63E | 938 |
| Bitlis | PDW192 * | PI 560717 | 1986 | 38.77N | 42.37E | 1770 | ||
| Bitlis | PDW193 * | PI 560718 | 1986 | 38.78N | 42.38E | 1770 | ||
| Siirt | PDW190 * | PI 560702 | 1986 | 37.82N | 41.87E | 560 | ||
| Siirt | PDW194 * | PI 560889 | 1989 | 37.75N | 42.20E | 1070 | ||
| unknown | PDW102 | PI 346985 | 1970 | |||||
| Syria | Dimashq | PDW52 * | PI 182697 | 1949 | 33.5N | 36.30E | 690 | |
| Halab | PDW57 * | PI 193391 | 1951 | 36.2N | 37.17E | 410 | ||
| Unknown | PDW180 | PI 520415 | 1987 | |||||
| Unknown | PDW41 * | PI 134596 | 1939 | |||||
| Iran | Khuzestan, | PDW42 * | PI 140184 | 1941 | 32.38N | 48.40E | 126 | |
| East Azerbaijan | PDW72 * | PI 222675 | 1954 | 38.08N | 46.30E | 1399 | ||
| Tehran | PDW76 * | PI 243790 | 1957 | 35.27N | 49.28E | 1866 | ||
| Fars | PDW88 * | PI 289821 | 1963 | 30.33N | 51.52E | 1130 | ||
| Iraq | Ninawa | PDW79 * | PI 253801 | 1958 | 36.33N | 43.13E | 223 | |
| Unknown | PDW47 | PI 165846 | 1948 | |||||
| Unknown | PDW58 * | PI 208903 | 1953 | |||||
| Unknown | PDW60 * | PI 208907 | 1953 | |||||
| Unknown | PDW61 * | PI 208908 | 1953 | |||||
| Unknown | PDW62 * | PI 208910 | 1953 | |||||
| Unknown | PDW242 * | PI 70736 | 1926 | |||||
| Israel | Unknown | PDW77 | PI 249816 | 1958 | ||||
| Unknown | PDW78 | PI 249820 | 1958 | |||||
| Unknown | PDW90 | PI 292035 | 1963 | |||||
| Unknown | PDW139 | PI 384043 | 1973 | |||||
| Unknown | PDW141 | PI 388035 | 1974 | |||||
| Cyprus | Unknown | PDW68 * | PI 210952 | 1953 | ||||
| Unknown | PDW75 | PI 237632 | 1957 | |||||
| Unknown | PDW208 | PI 591959 | 1994 | |||||
| Yemen | Aden | PDW45 | PI 152567 | 1945 | 12.77N | 45.01E | 79 | |
| Azerbaijan | Unknown | PDW73 | PI 233213 | 1956 | ||||
| Unknown | PDW101 | PI 345707 | 1950 | |||||
| North America (33) | USA | North Dakota | PDW3 | Citr 12068 | 1940 | |||
| North Dakota | PDW7 | Citr 13246 | 1955 | |||||
| North Dakota | PDW8 | Citr 13333 | 1957 | |||||
| North Dakota | PDW288 | Ldn 16 | ||||||
| Colorado | PDW29 | Citr 6881 | 1923 | |||||
| Kansas | PDW189 | PI 560335 | 1992 | |||||
| Arizona | PDW200 | PI 573005 | 1988 | |||||
| Arizona | PDW211 | PI 601250 | 1985 | |||||
| California | PDW210 | PI 600931 | 1982 | |||||
| California | PDW231 | PI 656793 | 2009 | |||||
| California | PDW232 | PI 656794 | 2009 | |||||
| California | PDW233 | PI 656795 | 2009 | |||||
| Erevan | PDW250 | PI 9872 | 1903 | 40.18N | 44.50E | 1120 | ||
| Mexico | Federal District | PDW152 | PI 428453 | 1978 | ||||
| Federal District | PDW173 | PI 519751 | 1987 | |||||
| Federal District | PDW174 | PI 519752 | 1987 | |||||
| Federal District | PDW176 | PI 519761 | 1987 | |||||
| Federal District | PDW177 | PI 519866 | 1987 | |||||
| Federal District | PDW178 | PI 520053 | 1987 | |||||
| Federal District | PDW216 | PI 610765 | 1999 | |||||
| Federal District | PDW227 | PI 634315 | 2001 | |||||
| Federal District | PDW229 | PI 634318 | 2001 | |||||
| Unknown | PDW179 | PI 520173 | 1987 | |||||
| Unknown | PDW49 | PI 168708 | 1948 | |||||
| Unknown | PDW150 | PI 422289 | 1978 | |||||
| Unknown | PDW13 | Citr 15874 | 1972 | |||||
| Canada | Saskatchewan | PDW18 | Citr 17337 | 1974 | ||||
| Saskatchewan | PDW186 | PI 546060 | 1990 | |||||
| Saskatchewan | PDW187 | PI 546362 | 1991 | |||||
| Saskatchewan | PDW202 | PI 583724 | 1994 | |||||
| Saskatchewan | PDW205 | PI 583731 | 1994 | |||||
| Saskatchewan | PDW206 | PI 583732 | 1994 | |||||
| Saskatchewan | PDW207 | PI 583733 | 1994 | |||||
| Latin America (12) | Chile | La Araucania | PDW14 | Citr 17057 | 1972 | |||
| La Araucania | PDW15 | Citr 17058 | 1972 | |||||
| La Araucania | PDW16 | Citr 17157 | 1972 | |||||
| La Araucania | PDW17 | Citr 17159 | 1972 | |||||
| Peru | Junin | PDW248 | PI 91956 | 1931 | 12.03S | 75.28W | 3252 | |
| Cajamarca | PDW249 | PI 92024 | 1931 | 7.60S | 78.47W | 3050 | ||
| Unknown | PDW48 | PI 168692 | 1948 | |||||
| Brazil | Sao Paulo | PDW54 | PI 191645 | 1950 | 22.00S | 49.00W | ||
| Unknown | PDW175 | PI 519759 | 1987 | |||||
| Bolivia | Cochabamba | PDW196 * | PI 565259 | 1991 | 17.40S | 66.23W | 3245 | |
| Cochabamba | PDW197 * | PI 565266 | 1991 | 17.57S | 65.83W | 2730 | ||
| Ecuador | Pichincha | PDW87 | PI 286546 | 1963 | ||||
| Oceania (7) | Australia | Victoria | PDW28 * | Citr 5136 | 1916 | 34.25S | 141.50E | |
| Western Australia | PDW50 | PI 174645 | 1949 | |||||
| Western Australia | PDW235 | PI 67341 | 1926 | |||||
| New South Wales | PDW74 | PI 235159 | 1956 | 33.00S | 146.00E | |||
| Unknown | PDW34 | PI 107606 | 1934 | |||||
| Unknown | PDW138 | PI 377882 | 1973 | |||||
| Unknown | PDW153 | PI 428701 | 1978 | |||||
| Western Europe (14) | Portugal | Lisboa | PDW195 | PI 56233 | 1923 | |||
| France | Unknown | PDW124 | PI 352450 | 1969 | ||||
| Greece | Unknown | PDW106 | PI 352389 | 1969 | ||||
| Sweden | Gotland | PDW56 | PI 192711 | 1950 | ||||
| Switzerland | Switzerland | PDW105 | PI 352377 | 1969 | ||||
| Spain | Unknown | PDW112 | PI 352404 | 1969 | ||||
| Germany | Unknown | PDW22 * | Citr 2468 | 1904 | ||||
| Germany | Lower Saxony | PDW93 | PI 306664 | 1965 | ||||
| Bulgaria | Unknown | PDW100 | PI 344743 | 1969 | ||||
| Bulgaria | Khaskovo | PDW188 | PI 546462 | 1990 | ||||
| Italy | Unknown | PDW113 | PI 352408 | 1969 | ||||
| Latium | PDW115 | PI 352415 | 1969 | |||||
| Latium | PDW209 | PI 593005 | 1996 | |||||
| England | Unknown | PDW83 | PI 278223 | 1962 | ||||
| Unknown | PDW84 | PI 278648 | 1962 | 53.00N | 2.00W | |||
| Unknown | PDW95 | PI 321702 | 1967 | |||||
| Romania | Unknown | PDW131 | PI 376498 | 1972 | ||||
| Unknown | PDW132 | PI 376500 | 1972 | |||||
| Unknown | PDW133 | PI 376501 | 1972 | |||||
| Unknown | PDW135 | PI 376509 | 1972 | |||||
| Unknown | PDW136 | PI 376511 | 1972 | |||||
| Unknown | PDW137 | PI 376512 | 1972 | |||||
| Eastern Europe (5) | Ukraine | Kharkiv | PDW160 | PI 438973 | 1980 | |||
| Russian | Altay | PDW24 * | Citr 3267 | 1911 | 52.68N | 83.21E | 152 | |
| Former Soviet | PDW118 | PI 352436 | 1969 | |||||
| Union | ||||||||
| Former Soviet | PDW119 | PI 352437 | 1969 | |||||
| Union | ||||||||
| Krasnoyarsk | PDW220 * | PI 61189 | 1924 | 58.45N | 92.17E | 79 | ||
| South Africa (4) | South Africa | Unknown | PDW151 * | PI 42425 | 1916 | |||
| Free State | PDW163 * | PI 45442 | 1917 | 29.17S | 24.75E | 1123 | ||
| Cape Province | PDW164 * | PI 45443 | 1917 | 30.98S | 27.33E | 1703 | ||
| Cape Province | PDW167 | PI 46766 | 1918 | 31.47S | 19.78E | 994 | ||
| North Africa (12) | Algeria | Mascara | PDW39 * | PI 11715 | 1904 | 35.74N | 0.55E | 104 |
| Tunisia | Unknown | PDW107 | PI 352390 | 1969 | ||||
| Unknown | PDW170 * | PI 51210 | 1920 | 33.02N | 35.57E | |||
| Unknown | PDW171 | PI 519380 | 1987 | |||||
| Egypt | Giza | PDW46 | PI 153774 | 1946 | 29.77N | 31.30E | ||
| Minufiya | PDW183 | PI 532119 | 1988 | 30.47N | 30.93E | 12 | ||
| Unknown | PDW212 * | PI 60712 | 1924 | |||||
| Sinai | PDW215 * | PI 60742 | 1924 | 29.50N | 34.00E | |||
| Alexandria | PDW237 * | PI 7016 | 1901 | 31.17N | 29.87E | |||
| Sawhaj | PDW243 * | PI 7422 | 1901 | 26.35N | 31.89E | 65 | ||
| Ethiopia | Unknown | PDW110 | PI 352395 | 1969 | ||||
| Unknown | PDW128 * | PI 352551 | 1969 | |||||
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Ren, J.; Sun, D.; Chen, L.; You, F.M.; Wang, J.; Peng, Y.; Nevo, E.; Sun, D.; Luo, M.-C.; Peng, J. Genetic Diversity Revealed by Single Nucleotide Polymorphism Markers in a Worldwide Germplasm Collection of Durum Wheat. Int. J. Mol. Sci. 2013, 14, 7061-7088. https://doi.org/10.3390/ijms14047061
Ren J, Sun D, Chen L, You FM, Wang J, Peng Y, Nevo E, Sun D, Luo M-C, Peng J. Genetic Diversity Revealed by Single Nucleotide Polymorphism Markers in a Worldwide Germplasm Collection of Durum Wheat. International Journal of Molecular Sciences. 2013; 14(4):7061-7088. https://doi.org/10.3390/ijms14047061
Chicago/Turabian StyleRen, Jing, Daokun Sun, Liang Chen, Frank M. You, Jirui Wang, Yunliang Peng, Eviatar Nevo, Dongfa Sun, Ming-Cheng Luo, and Junhua Peng. 2013. "Genetic Diversity Revealed by Single Nucleotide Polymorphism Markers in a Worldwide Germplasm Collection of Durum Wheat" International Journal of Molecular Sciences 14, no. 4: 7061-7088. https://doi.org/10.3390/ijms14047061
APA StyleRen, J., Sun, D., Chen, L., You, F. M., Wang, J., Peng, Y., Nevo, E., Sun, D., Luo, M.-C., & Peng, J. (2013). Genetic Diversity Revealed by Single Nucleotide Polymorphism Markers in a Worldwide Germplasm Collection of Durum Wheat. International Journal of Molecular Sciences, 14(4), 7061-7088. https://doi.org/10.3390/ijms14047061







