Comparative Proteomic Analysis of Lactobacillus plantarum WCFS1 and ΔctsR Mutant Strains Under Physiological and Heat Stress Conditions
Abstract
:1. Introduction
2. Results and Discussion
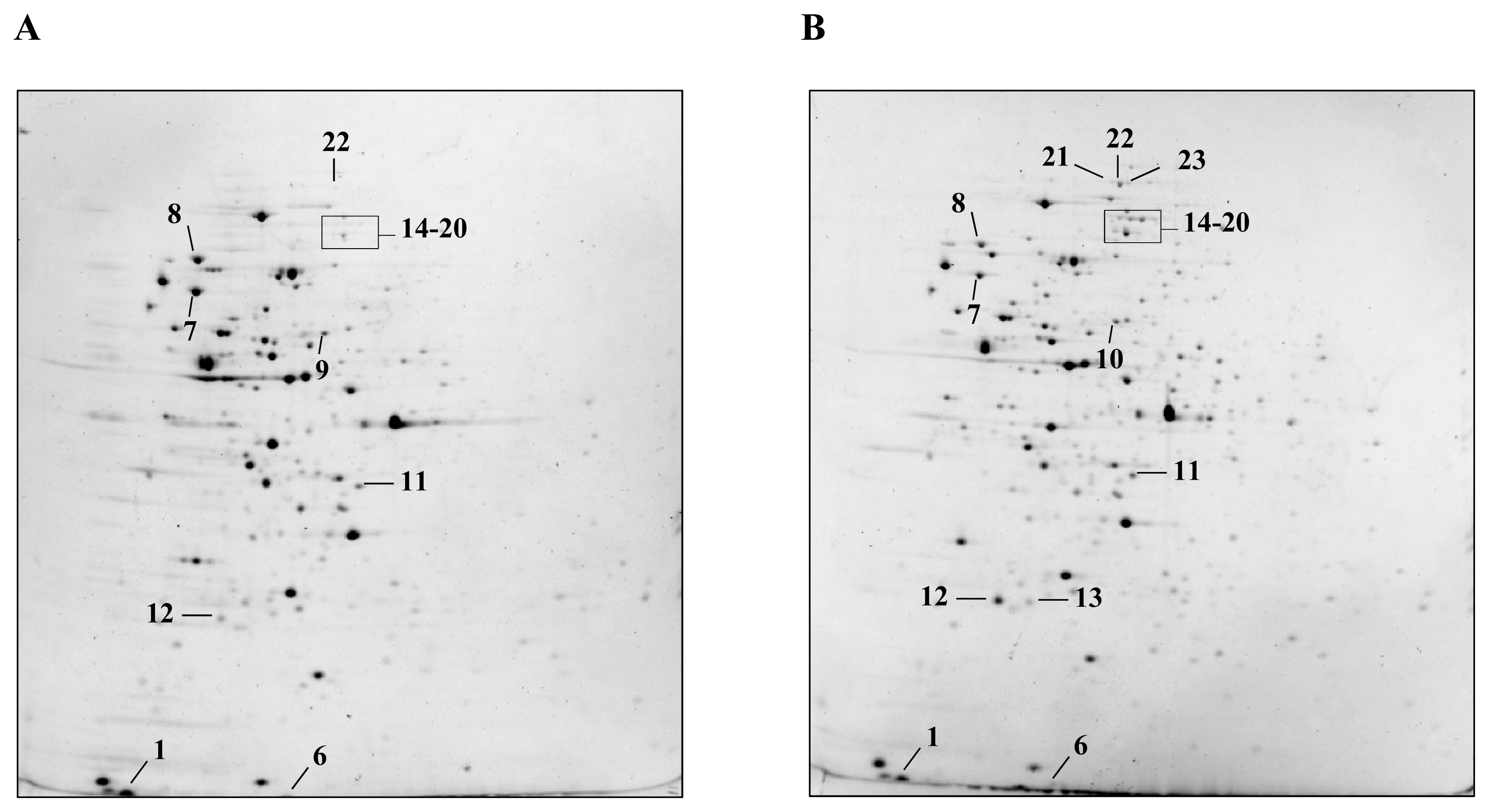
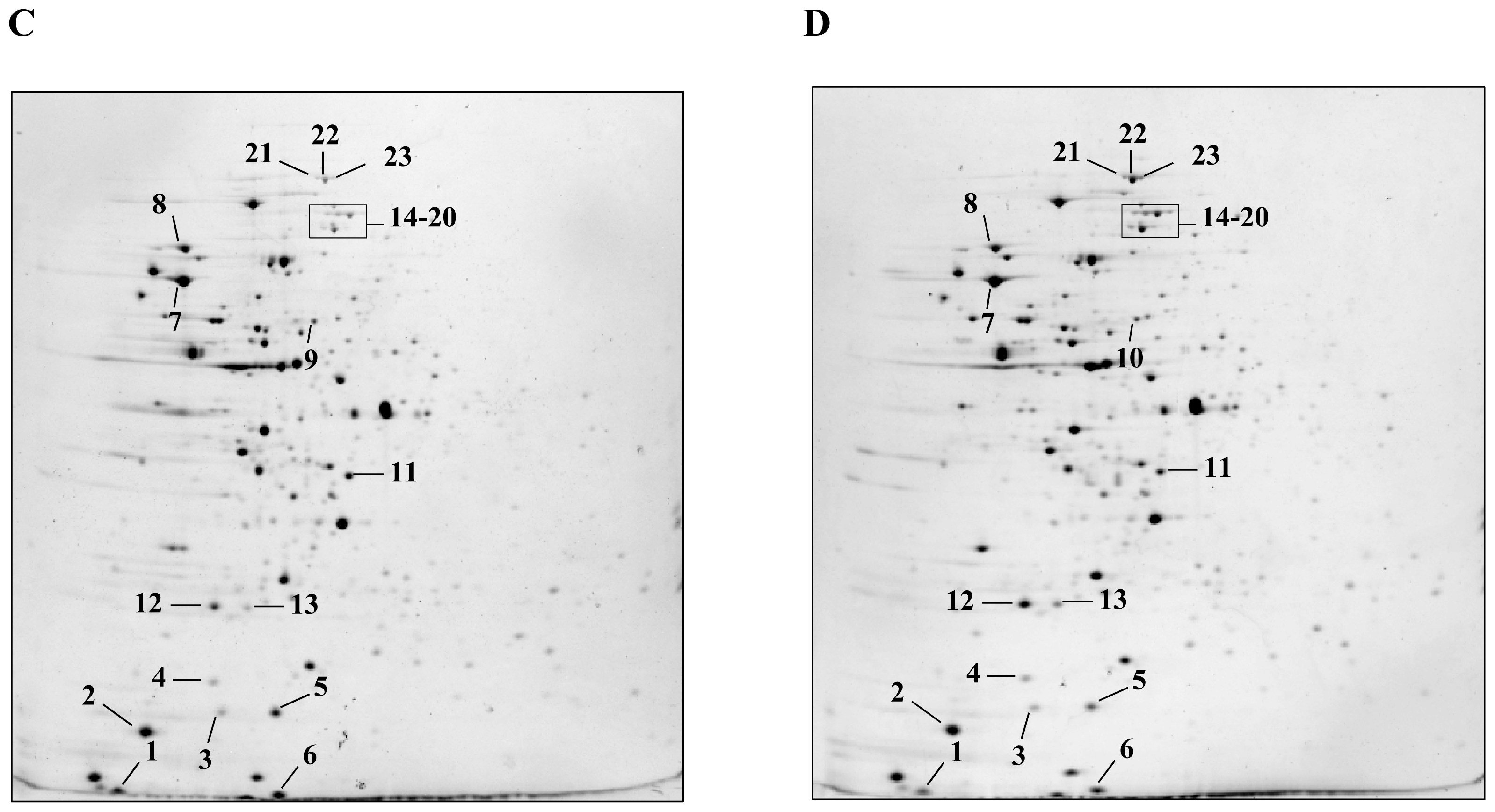
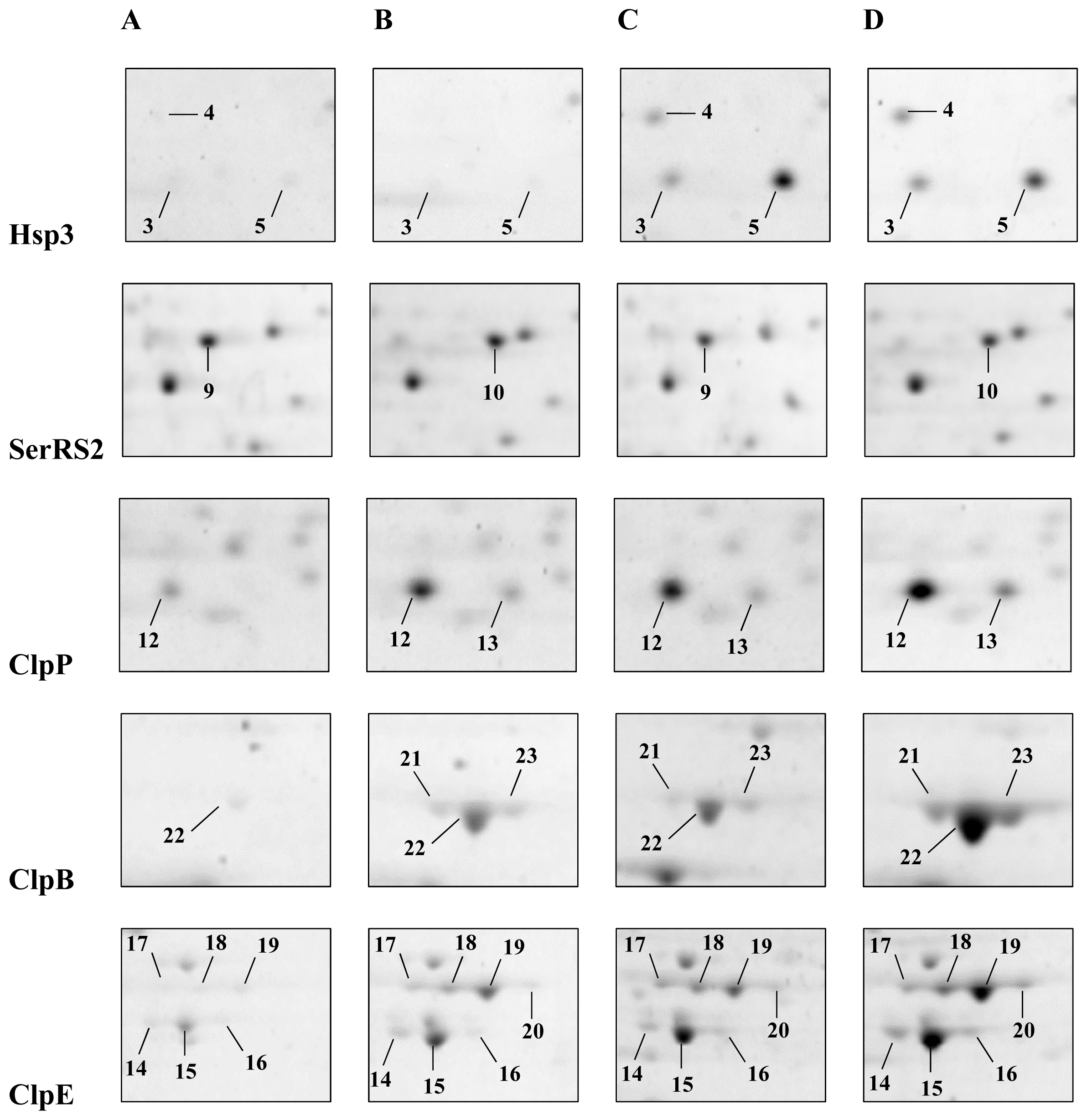
2.1. Detection and Identification of Spots with Altered Expression Levels
2.2. Influence of Heat Shock on Protein Levels
2.3. Influence of ΔctsR Mutation in Protein Expression Levels
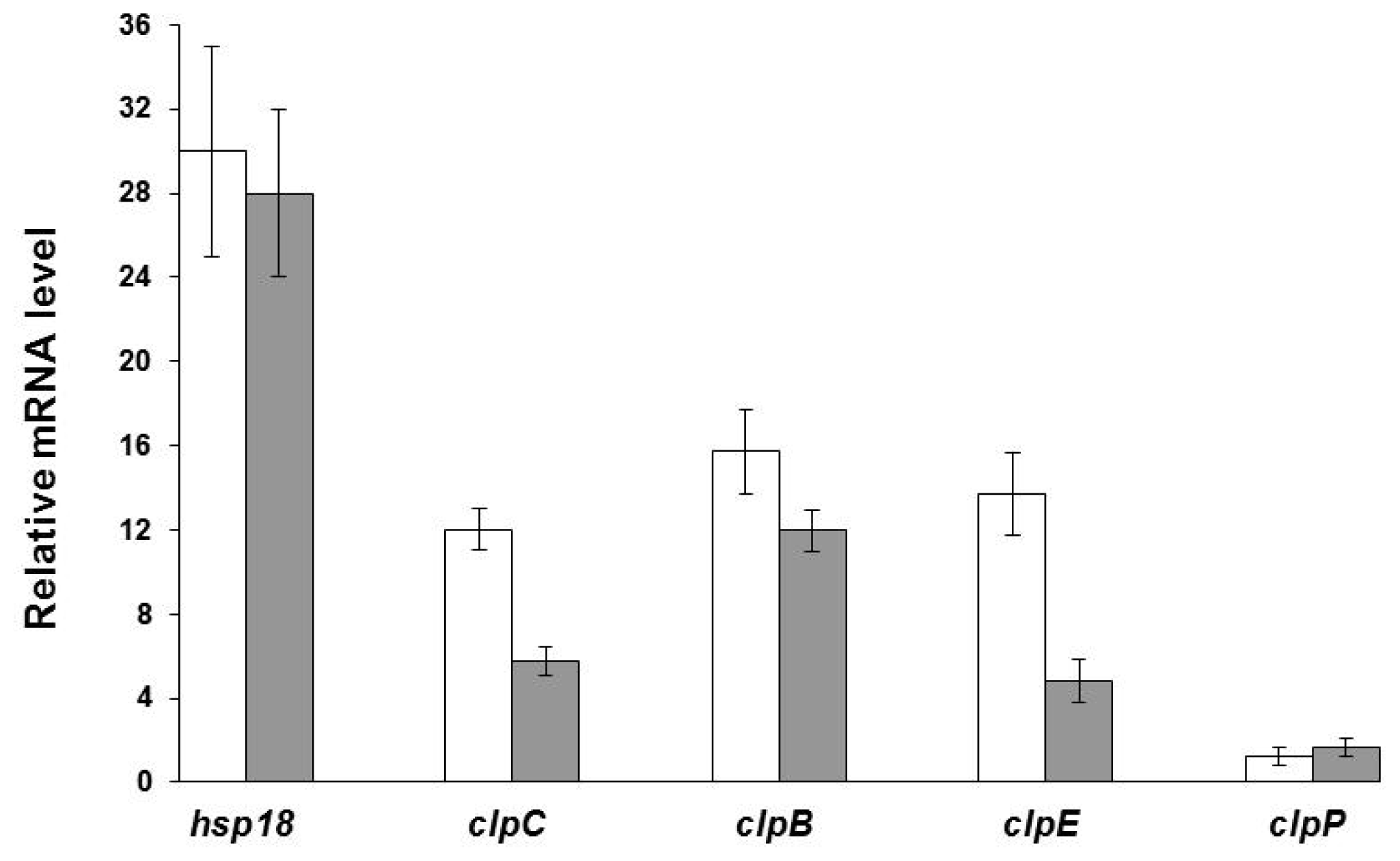
2.4. Transcriptional Analysis of Relevant Genes
3. Experimental Section
3.1. Bacterial Strains and Experimental Growth Conditions
3.2. Preparation of Protein Extracts
3.3. Two-Dimensional (2D) Gel Analysis and Image Acquisition
3.4. 2D-Gel Image Analysis
3.5. Identification of Proteins by MALDI-TOF-TOF Peptide Mass Fingerprinting
3.6. Quantitative RT-PCR Analysis
4. Conclusions
Acknowledgments
References
- Van de Guchte, M.; Serror, P.; Chervaux, C.; Smokvina, T.; Ehrlich, S.D.; Maguin, E. Stress responses in lactic acid bacteria. Antonie Van Leeuwenhoek 2002, 82, 187–216. [Google Scholar]
- Gottesman, S.; Wickner, S.; Maurizi, M.R. Protein quality control: Triage by chaperones and proteases. Genes Dev 1997, 11, 815–823. [Google Scholar]
- Schumann, W. The Bacillus subtilis heat shock stimulon. Cell Stress Chaperones 2003, 8, 207–217. [Google Scholar]
- Krüger, E.; Hecker, M. The first gene of the Bacillus subtilis clpC operon, ctsR, encodes a negative regulator of its own operon and other class III heat shock genes. J. Bacteriol 1998, 180, 6681–6688. [Google Scholar]
- Derré, I.; Rapoport, G.; Msadek, T. CtsR, a novel regulator of stress and heat shock response, controls clp and molecular chaperone gene expression in gram-positive bacteria. Mol. Microbiol 1999, 31, 117–131. [Google Scholar]
- Derré, I.; Rapoport, G.; Msadek, T. The CtsR regulator of stress response is active as a dimer and specifically degraded in vivo at 37 °C. Mol. Microbiol 2000, 38, 335–347. [Google Scholar]
- Elsholz, A.K.W.; Michalik, S.; Zühlke, D.; Hecker, M.; Gerth, U. CtsR, the Gram-positive master regulator of protein quality control, feels the heat. EMBO J 2010, 29, 3621–3629. [Google Scholar]
- Elsholz, A.K.W.; Gerth, U.; Hecker, M. Regulation of CtsR activity in low GC, Gram+ bacteria. Adv. Microb. Physiol 2010, 57, 119–144. [Google Scholar]
- Spano, G.; Massa, S. Environmental stress response in wine lactic acid bacteria: Beyond Bacillus subtilis. Crit. Rev. Microbiol 2006, 32, 77–86. [Google Scholar]
- Guarner, F.; Malagelada, J.-R. Gut flora in health and disease. Lancet 2003, 361, 512–519. [Google Scholar]
- Molin, G. Probiotics in foods not containing milk or milk constituents, with special reference to Lactobacillus plantarum 299v. Am. J. Clin. Nutr 2001, 73, 380–385. [Google Scholar]
- De Vries, M.C.; Vaughan, E.E.; Kleerebezem, M.; de Vos, W.M. Lactobacillus plantarum— Survival, functional and potential probiotic properties in the human intestinal tract. Int. Dairy J 2006, 16, 1018–1028. [Google Scholar]
- Fiocco, D.; Collins, M.; Muscariello, L.; Hols, P.; Kleerebezem, M.; Msadek, T.; Spano, G. The Lactobacillus plantarum ftsH gene is a novel member of the CtsR stress response regulon. J. Bacteriol 2009, 191, 1688–1694. [Google Scholar]
- Fiocco, D.; Capozzi, V.; Collins, M.; Gallone, A.; Hols, P.; Guzzo, J.; Weidmann, S.; Rieu, A.; Msadek, T.; Spano, G. Characterization of the CtsR stress response regulon in Lactobacillus plantarum. J. Bacteriol 2010, 192, 896–900. [Google Scholar]
- Cohen, D.P.A.; Renes, J.; Bouwman, F.G.; Zoetendal, E.G.; Mariman, E.; de Vos, W.M.; Vaughan, E.E. Proteomic analysis of log to stationary growth phase Lactobacillus plantarum cells and a 2-DE database. Proteomics 2006, 6, 6485–6493. [Google Scholar]
- Zhu, L.; Hu, W.; Liu, D.; Tian, W.; Yu, G.; Liu, X.; Wang, J.; Feng, E.; Zhang, X.; Chen, B.; et al. A reference proteomic database of Lactobacillus plantarum CMCC-P0002. PLoS One 2011, 6, e25596. [Google Scholar]
- De Angelis, M.; di Cagno, R.; Huet, C.; Crecchio, C.; Fox, P.F.; Gobbetti, M. Heat shock response in Lactobacillus plantarum. Appl. Environ. Microbiol 2004, 70, 1336–1346. [Google Scholar]
- Hamon, E.; Horvatovich, P.; Izquierdo, E.; Bringel, F.; Marchioni, E.; Aoudé-Werner, D.; Ennahar, S. Comparative proteomic analysis of Lactobacillus plantarum for the identification of key proteins in bile tolerance. BMC Microbiol 2011, 11, 63. [Google Scholar]
- Lee, K.; Rho, B.-S.; Pi, K.; Kim, H.-J.; Choi, Y.-J. Proteomic analysis of protein expression in Lactobacillus plantarum in response to alkaline stress. J. Biotechnol 2011, 153, 1–7. [Google Scholar]
- Izquierdo, E.; Horvatovich, P.; Marchioni, E.; Aoude-Werner, D.; Sanz, Y.; Ennahar, S. 2-DE and MS analysis of key proteins in the adhesion of Lactobacillus plantarum, a first step toward early selection of probiotics based on bacterial biomarkers. Electrophoresis 2009, 30, 949–956. [Google Scholar]
- O’Donoghue, P.; Luthey-Schulten, Z. On the evolution of structure in aminoacyl-tRNA synthetases. Microbiol. Mol. Biol. Rev 2003, 67, 550–573. [Google Scholar]
- Varmanen, P.; Vogensen, F.K.; Hammer, K.; Palva, A.; Ingmer, H. ClpE from Lactococcus lactis promotes repression of CtsR-dependent gene expression. J. Bacteriol 2003, 185, 5117–5124. [Google Scholar]
- Miethke, M.; Hecker, M.; Gerth, U. Involvement of Bacillus subtilis ClpE in CtsR degradation and protein quality control. J. Bacteriol 2006, 188, 4610–4619. [Google Scholar]
- Elsholz, A.K.W.; Hempel, K.; Pöther, D.; Becher, D.; Hecker, M.; Gerth, U. CtsR inactivation during thiol specific stress in low GC, Gram+ bacteria. Mol. Microbiol 2011, 79, 772–785. [Google Scholar]
- Ingmer, H.; Vogensen, F.K.; Hammer, K.; Kilstrup, M. Disruption and analysis of the clpB, clpC, and clpE genes in Lactococcus lactis: ClpE, a new Clp family in Gram-positive bacteria. J. Bacteriol 1999, 181, 2075–2083. [Google Scholar]
- Helmann, J.D.; Wu, M.F.; Kobel, P.A.; Gamo, F.J.; Wilson, M.; Morshedi, M.M.; Navre, M.; Paddon, C. Global transcriptional response of Bacillus subtilis to heat shock. J. Bacteriol 2001, 183, 7318–7328. [Google Scholar]
- Krüger, E.; Völker, U.; Hecker, M. Stress induction of clpC in Bacillus subtilis and its involvement in stress tolerance. J. Bacteriol 1994, 176, 3360–3367. [Google Scholar]
- Chastanet, A.; Fert, J.; Msadek, T. Comparative genomics reveal novel heat shock regulatory mechanisms in Staphylococcus aureus and other Gram-positive bacteria. Mol. Microbiol 2003, 47, 1061–1073. [Google Scholar]
- Derré, I.; Rapoport, G.; Devine, K.; Rose, M.; Msadek, T. ClpE, a novel type of HSP100 ATPase, is part of the CtsR heat shock regulon of Bacillus subtilis. Mol. Microbiol 1999, 32, 581–593. [Google Scholar]
- Varmanen, P.; Ingmer, H.; Vogensen, F.K. ctsR of Lactococcus lactis encodes a negative regulator of clp gene expression. Microbiology 2000, 146, 1447–1455. [Google Scholar]
- Spano, G.; Capozzi, V.; Vernile, A.; Massa, S. Cloning, molecular characterization and expression analysis of two small heat shock genes isolated from wine Lactobacillus plantarum. J. Appl. Microbiol 2004, 97, 774–782. [Google Scholar]
- Kleerebezem, M.; Boekhorst, J.; van Kranenburg, R.; Molenaar, D.; Kuipers, O.P.; Leer, R.; Tarchini, R.; Peters, S.A.; Sandbrink, H.M.; Fiers, M.W.; et al. Complete genome sequence of Lactobacillus plantarum WCFS1. Proc. Natl. Acad. Sci. USA 2003, 100, 1990–1995. [Google Scholar]
- Mohedano, M.L.; Overweg, K.; de la Fuente, A.; Reuter, M.; Altabe, S.; Mulholland, F.; de Mendoza, D.; López, P.; Wells, J.M. Evidence that the essential response regulator YycF in Streptococcus pneumoniae modulates expression of fatty acid biosynthesis genes and alters membrane composition. J. Bacteriol 2005, 187, 2357–2367. [Google Scholar]
- García-Quintáns, N.; Repizo, G.; Martín, M.; Magni, C.; López, P. Activation of diacetyl/acetoin pathway in Lactococcus lactis subsp. lactis biovar diacetylactis CRL264 by acidic growth. Appl. Environ. Microbiol 2008, 74, 1988–1996. [Google Scholar]
- Shevchenko, A.; Tomas, H.; Havlis, J.; Olsen, J.V.; Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc 2007, 1, 2856–2860. [Google Scholar]

| SSP | Identified Protein | Mascot Score | Sequence Coverage (%) | Peptides (n) | MW (kDa)/pI | Induction Factor | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| Theoretical | Observed | Lp ΔctsR/Lp WCFS1 | Stress/Control | |||||||
| Control | Stress | Lp WCFS1 | Lp ΔctsR | |||||||
| 1 | cold shock protein CspC | 122 | 68 | 3 | 7.30/4.57 | 11.21/4.40 | 0.63 ± 0.06 | 1.15 ± 0.32 | 0.64 ± 0.05 | 1.16 ± 0.33 |
| 2 | small heat shock protein Hsp1 | 340 | 54 | 8 | 15.99/4.53 | 14.32/4.50 | n.d. | 1.11 ± 0.24 | On/Off | On/Off |
| 3 | small heat shock protein Hsp3 | 224 | 51 | 6 | 16.66/5.00 | 15.33/4.79 | 0.56 ± 0.09 | 1.56 ± 0.18 | 4.49 ± 1.09 | 9.36 ± 1.10 |
| 4 | small heat shock protein Hsp3 | 255 | 51 | 6 | 16.66/5.00 | 16.93/4.76 | Off/On | 1.72 ± 0.14 | 12.72 ± 3.68 | On/Off |
| 5 | small heat shock protein Hsp3 | 212 | 51 | 5 | 16.66/5.00 | 15.45/4.99 | 0.72 ± 0.44 | 0.83 ± 0.11 | 27.23 ± 1.21 | 31.29 ± 4.31 |
| ∑ small heat shock protein Hsp3 | 0.75 ± 0.25 | 1.02 ± 0.12 | 14.85 ± 1.38 | 19.97 ± 2.35 | ||||||
| 6 | GroES co-chaperonin | 291 | 50 | 4 | 10.28/4.95 | 11.39/5.02 | 0.64 ± 0.09 | 0.90 ± 0.14 | 4.03 ± 0.26 | 5.70 ± 0.89 |
| 7 | GroEL chaperonin | 319 | 22 | 9 | 57.40/4.69 | 56.60/4.67 | 0.68 ± 0.10 | 0.96 ± 0.15 | 2.47 ± 0.17 | 3.51 ± 0.53 |
| 8 | molecular chaperone DnaK | 481 | 41 | 14 | 66.69/4.68 | 67.33/4.67 | 0.62 ± 0.07 | 0.94 ± 0.14 | 1.25 ± 0.06 | 1.87 ± 0.28 |
| 9 | Seryl-tRNA synthetase SerRS2 | 282 | 13 | 4 | 48.22/5.16 | 48.41/5.13 | Off/On | Off/On | 0.69 ± 0.20 | n.d. |
| 10 | Seryl-tRNA synthetase SerRS2 | 327 | 19 | 6 | 48.22/5.16 | 48.41/5.17 | On/Off | On/Off | n.d. | 0.67 ± 0.49 |
| 11 | ATP-dependent Clp protease proteolytic subunit ClpP | 129 | 42 | 5 | 21.51/4.87 | 32.90/5.24 | 1.75 ± 0.19 | 1.14 ± 0.20 | 3.20 ± 0.39 | 2.10 ± 0.36 |
| 12 | ATP-dependent Clp protease proteolytic subunit ClpP | 277 | 55 | 8 | 21.51/4.87 | 21.46/4.76 | 4.77 ± 0.68 | 1.99 ± 0.39 | 3.90 ± 0.60 | 1.63 ± 0.32 |
| 13 | ATP-dependent Clp protease proteolytic subunit ClpP | 154 | 37 | 5 | 21.51/4.87 | 21.46/4.88 | On/Off | 1.89 ± 0.10 | On/Off | 1.75 ± 0.09 |
| * ∑ ATP-dependent Clp protease proteolytic subunit ClpP | 5.85 ± 0.84 | 1.97 ± 0.30 | 4.91 ± 0.60 | 1.65 ± 0.25 | ||||||
| 14 | ATP-dependent Clp protease, ATP-binding subunit ClpE | 492 | 27 | 14 | 81.90/5.28 | 77.09/5.15 | 3.41 ± 0.14 | 1.79 ± 0.17 | 2.88 ± 0.20 | 1.52 ± 0.14 |
| 15 | ATP-dependent Clp protease, ATP-binding subunit ClpE | 135 | 15 | 10 | 81.90/5.28 | 77.09/5.19 | 2.70 ± 0.13 | 1.49 ± 0.13 | 2.12 ± 0.11 | 1.17 ± 0.10 |
| 16 | ATP-dependent Clp protease, ATP-binding subunit ClpE | 529 | 20 | 10 | 81.90/5.28 | 77.09/5.22 | 4.12 ± 0.89 | 1.99 ± 0.28 | 2.52 ± 0.83 | 1.22 ± 0.17 |
| 17 | ATP-dependent Clp protease, ATP-binding subunit ClpE | 344 | 24 | 14 | 81.90/5.28 | 82.95/5.17 | 7.34 ± 2.26 | 2.15 ± 0.68 | 5.26 ± 0.86 | 1.54 ± 0.49 |
| 18 | ATP-dependent Clp protease, ATP-binding subunit ClpE | 450 | 23 | 13 | 81.90/5.28 | 82.95/5.20 | 12.08 ± 0.63 | 2.24 ± 0.03 | 8.66 ± 0.82 | 1.60 ± 0.02 |
| 19 | ATP-dependent Clp protease, ATP-binding subunit ClpE | 253 | 15 | 8 | 81.90/5.28 | 82.95/5.24 | 9.74 ± 1.20 | 1.68 ± 0.40 | 9.39 ± 1.03 | 1.62 ± 0.39 |
| 20 | ATP-dependent Clp protease, ATP-binding subunit ClpE | 509 | 20 | 10 | 81.90/5.28 | 82.95/5.28 | On/Off | 2.32 ± 0.63 | On/Off | 1.17 ± 0.32 |
| ∑ ATP-dependent Clp protease, ATP-binding subunit ClpE | 4.64 ± 0.45 | 1.66 ± 0.25 | 3.50 ± 0.25 | 1.26 ± 0.19 | ||||||
| 21 | ATP-dependent Clp protease, ATP-binding subunit ClpB | 279 | 25 | 17 | 96.51/5.19 | 95.63/5.12 | On/Off | 2.10 ± 0.08 | On/Off | 3.82 ± 0.14 |
| 22 | ATP-dependent Clp protease, ATP-binding subunit ClpB | 228 | 25 | 16 | 96.51/5.19 | 95.63/5.16 | 7.12 ± 1.42 | 1.82 ± 0.19 | 12.13 ± 4.92 | 3.10 ± 0.32 |
| 23 | ATP-dependent Clp protease, ATP-binding subunit ClpB | 378 | 35 | 22 | 96.51/5.19 | 95.63/5.19 | On/Off | 1.83 ± 0.22 | On/Off | 2.86 ± 0.35 |
| ∑ ATP-dependent Clp protease, ATP-binding subunit ClpB | 10.74 ± 2.19 | 1.83 ± 0.29 | 17.23 ± 5.77 | 2.93 ± 0.77 | ||||||
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Russo, P.; De la Luz Mohedano, M.; Capozzi, V.; De Palencia, P.F.; López, P.; Spano, G.; Fiocco, D. Comparative Proteomic Analysis of Lactobacillus plantarum WCFS1 and ΔctsR Mutant Strains Under Physiological and Heat Stress Conditions. Int. J. Mol. Sci. 2012, 13, 10680-10696. https://doi.org/10.3390/ijms130910680
Russo P, De la Luz Mohedano M, Capozzi V, De Palencia PF, López P, Spano G, Fiocco D. Comparative Proteomic Analysis of Lactobacillus plantarum WCFS1 and ΔctsR Mutant Strains Under Physiological and Heat Stress Conditions. International Journal of Molecular Sciences. 2012; 13(9):10680-10696. https://doi.org/10.3390/ijms130910680
Chicago/Turabian StyleRusso, Pasquale, María De la Luz Mohedano, Vittorio Capozzi, Pilar Fernández De Palencia, Paloma López, Giuseppe Spano, and Daniela Fiocco. 2012. "Comparative Proteomic Analysis of Lactobacillus plantarum WCFS1 and ΔctsR Mutant Strains Under Physiological and Heat Stress Conditions" International Journal of Molecular Sciences 13, no. 9: 10680-10696. https://doi.org/10.3390/ijms130910680
APA StyleRusso, P., De la Luz Mohedano, M., Capozzi, V., De Palencia, P. F., López, P., Spano, G., & Fiocco, D. (2012). Comparative Proteomic Analysis of Lactobacillus plantarum WCFS1 and ΔctsR Mutant Strains Under Physiological and Heat Stress Conditions. International Journal of Molecular Sciences, 13(9), 10680-10696. https://doi.org/10.3390/ijms130910680






