NotI Microarrays: Novel Epigenetic Markers for Early Detection and Prognosis of High Grade Serous Ovarian Cancer
Abstract
:1. Introduction
2. Results
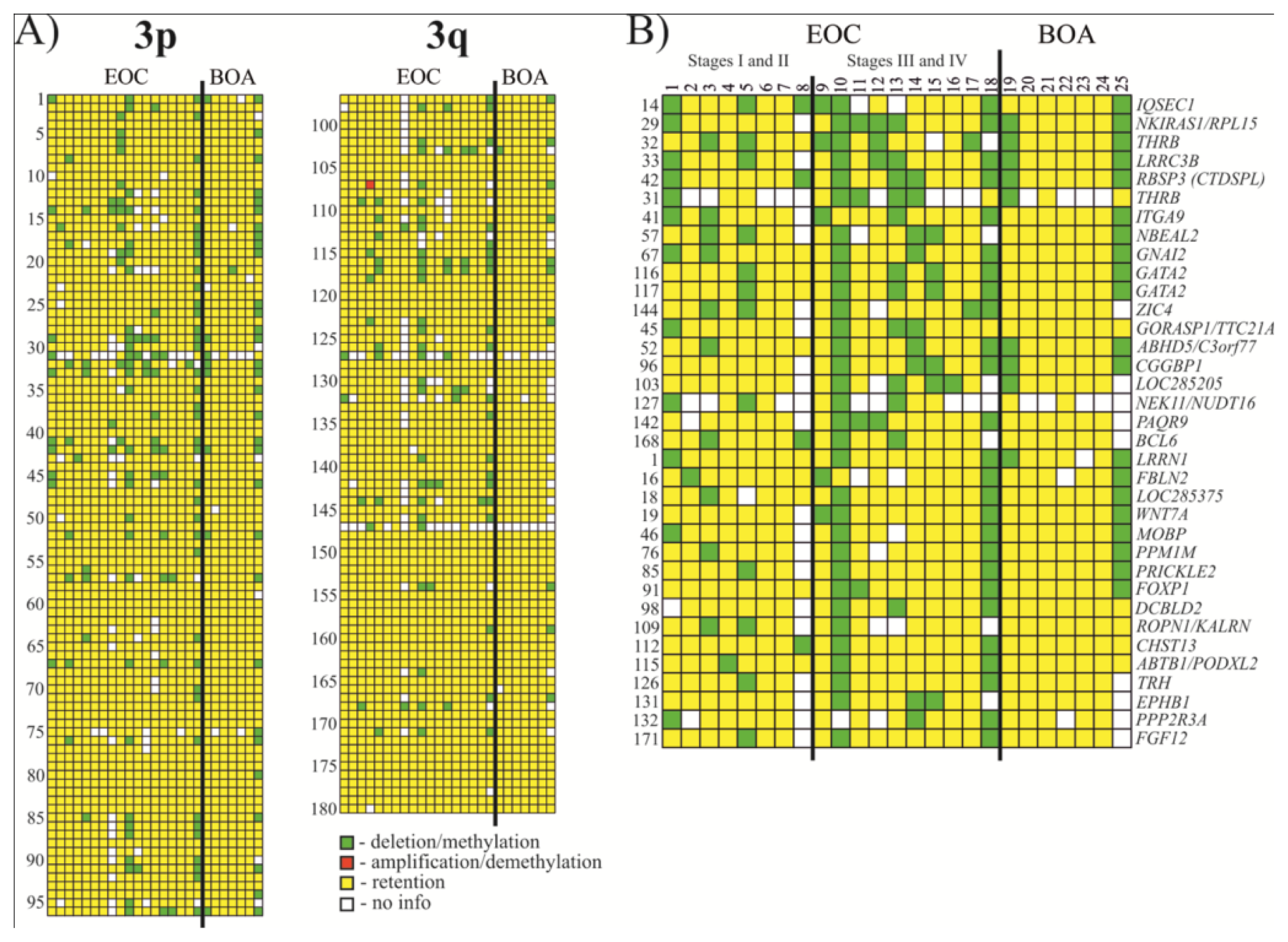
2.1. Analysis of Methylation Frequency Using NotI Microarrays
2.2. Selection of Genes/Biomarkers for Detection and Discrimination EOC with Different Histological Characteristics
3. Discussion
- EPHB1 for example participates in the Ephrin-EphR Signaling Pathway. In this pathway members of RAS family, MAPK, oncogene NCK1 and other genes important for cell developmental processes are included. Ephrin receptors make up the largest subgroup of the receptor tyrosine kinase (RTK) family. The protein encoded by this gene is a receptor for ephrin-B family members.
- NKIRAS1 is a potent regulator of NF-κ-B activity by preventing the degradation of NF-κ-B inhibitor beta (NFKBIB) by most signals, explaining why NFKBIB is more resistant to degradation.
- IQSEC1 links epidermal growth factor receptor signaling to ARF6 activation to induce breast cancer invasion (see Table 4).
- RBSP3 (CTDSPL) is a potential activator of the RB1 gene pathway [81].
4. Materials and Methods
4.1. Tissue Specimens
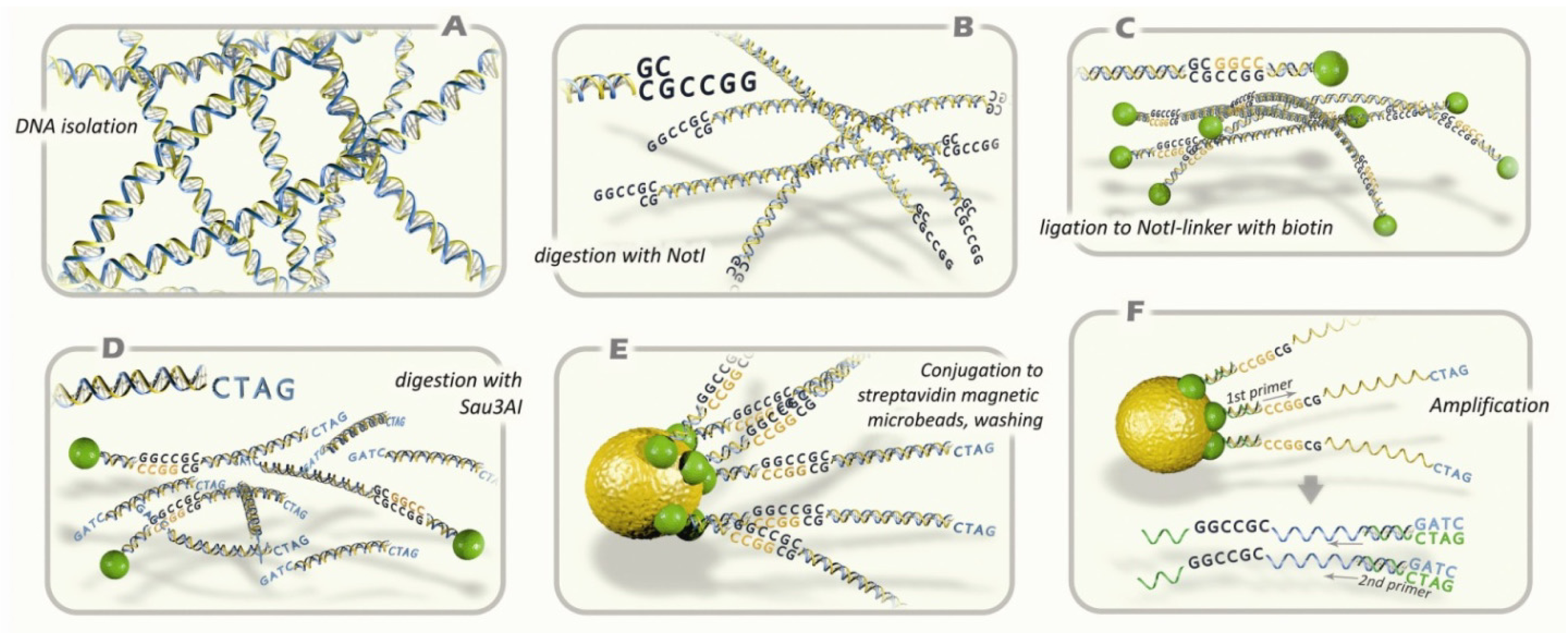
4.2. NotI-Microarrays
4.3. NotI Probes and Hybridization
4.4. Statistical Analysis
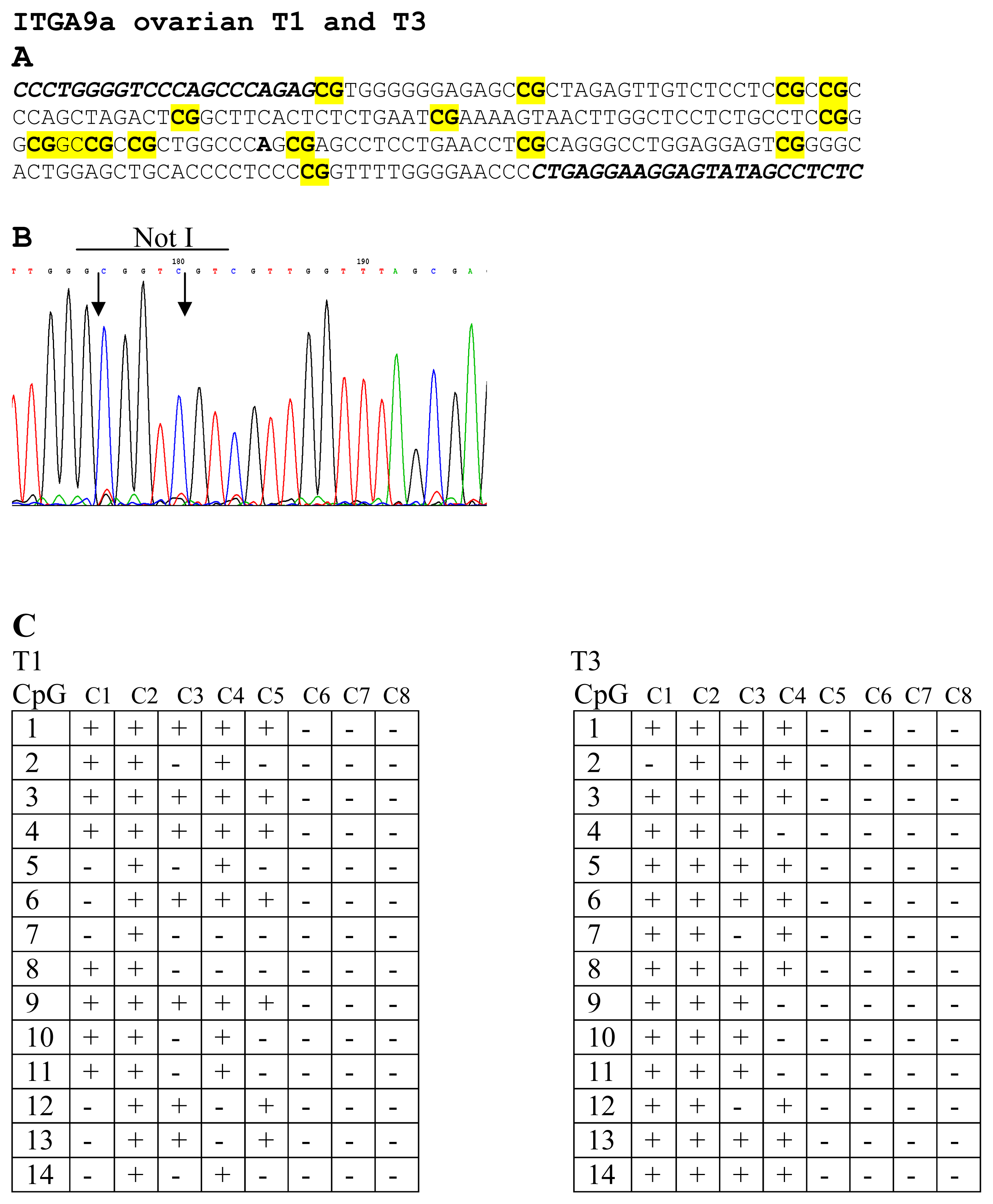
4.5. PCR, Cloning, Bisulfite Sequencing
5. Conclusions
Acknowledgements
- Conflict of interestThe authors declare no conflict of interest.
References
- Hunn, J.; Rodriguez, G.C. Ovarian cancer: Etiology, risk factors, and epidemiology. Clin. Obstet. Gynecol 2012, 55, 3–23. [Google Scholar]
- Chen, H.; Hardy, T.M.; Tollefsbol, T.O. Epigenomics of ovarian cancer and its chemoprevention. Front. Genet 2011, 2, 67. [Google Scholar]
- Yarbro, J.W.; Page, D.L.; Fielding, L.P.; Partridge, E.E.; Murphy, G.P. American Joint Committee on Cancer prognostic factors consensus conference. Cancer 1999, 86, 2436–2446. [Google Scholar]
- Jemal, A.; Siegel, R.; Xu, J.; Ward, E. Cancer statistics, 2010. CA Cancer J. Clin 2010, 60, 277–300. [Google Scholar]
- Landen, C.N., Jr; Birrer, M.J.; Sood, A.K. Early events in the pathogenesis of epithelial ovarian cancer. J. Clin. Oncol 2008, 26, 995–1005. [Google Scholar]
- Montavon, C.; Gloss, B.S.; Warton, K.; Barton, C.A.; Statham, A.L.; Scurry, J.P.; Tabor, B.; Nguyen, T.V.; Qu, W.; Samimi, G.; et al. Prognostic and diagnostic significance of DNA methylation patterns in high grade serous ovarian cancer. Gynecol. Oncol 2012, 124, 582–588. [Google Scholar]
- Barton, C.A.; Hacker, N.F.; Clark, S.J.; O’Brien, P.M. DNA methylation changes in ovarian cancer: Implications for early diagnosis, prognosis and treatment. Gynecol. Oncol 2008, 109, 129–139. [Google Scholar]
- Jones, P.A.; Baylin, S.B. The fundamental role of epigenetic events in cancer. Nat. Rev. Genet 2002, 3, 415–428. [Google Scholar]
- Ibanez de Caceres, I.; Battagli, C.; Esteller, M.; Herman, J.G.; Dulaimi, E.; Edelson, M.I.; Bergman, C.; Ehya, H.; Eisenberg, B.L.; Cairns, P. Tumor cell-specific BRCA1 and RASSF1A hypermethylation in serum, plasma, and peritoneal fluid from ovarian cancer patients. Cancer Res 2004, 64, 6476–6481. [Google Scholar]
- Kagan, J.; Srivastava, S.; Barker, P.E.; Belinsky, S.A.; Cairns, P. Towards Clinical Application of Methylated DNA Sequences as Cancer Biomarkers: A Joint NCI’s EDRN and NIST Workshop on Standards, Methods, Assays, Reagents and Tools. Cancer Res 2007, 67, 4545–4549. [Google Scholar]
- Laird, P.W. The power and the promise of DNA methylation markers. Nat. Rev. Cancer 2003, 3, 253–266. [Google Scholar]
- Wei, S.H.; Chen, C.M.; Strathdee, G.; Harnsomburana, J.; Shyu, C.R.; Rahmatpanah, F.; Shi, H.; Ng, S.W.; Yan, P.S.; Nephew, K.P.; et al. Methylation microarray analysis of late-stage ovarian carcinomas distinguishes progression-free survival in patients and identifies candidate epigenetic markers. Clin. Cancer Res 2002, 8, 2246–2252. [Google Scholar]
- Wei, S.H.; Balch, C.; Paik, H.H.; Kim, Y.S.; Baldwin, R.L.; Liyanarachchi, S.; Li, L.; Wang, Z.; Wan, J.C.; Davuluri, R.V.; et al. Prognostic DNA methylation biomarkers in ovarian cancer. Clin. Cancer Res 2006, 12, 2788–2794. [Google Scholar]
- Zabarovsky, E.R.; Klein, G.; Winberg, G. Lambda SK diphasmids: phage lambda vectors for genomic, jumping, linking and cDNA libraries. Gene 1993, 127, 1–14. [Google Scholar]
- Zabarovsky, E.R.; Boldog, F.; Erlandsson, R.; Kashuba, V.I.; Allikmets, R.L.; Marcsek, Z.; Kisselev, L.L.; Stanbridge, E.; Klein, G.; Sumegi, J.; et al. New strategy for mapping the human genome based on a novel procedure for construction of jumping libraries. Genomics 1991, 11, 1030–1039. [Google Scholar]
- Zabarovsky, E.R.; Boldog, F.; Thompson, T.; Scanlon, D.; Winberg, G.; Marcsek, Z.; Erlandsson, R.; Stanbridge, E.J.; Klein, G.; Sumegi, J. Construction of a human chromosome 3 specific NotI linking library using a novel cloning procedure. Nucleic Acids Res 1990, 18, 6319–6324. [Google Scholar]
- Allikmets, R.L.; Kashuba, V.I.; Pettersson, B.; Gizatullin, R.; Lebedeva, T.; Kholodnyuk, I.D.; Bannikov, V.M.; Petrov, N.; Zakharyev, V.M.; Winberg, G.; et al. NotI linking clones as a tool for joining physical and genetic maps of the human genome. Genomics 1994, 19, 303–309. [Google Scholar]
- Wang, J.Y.; Zabarovsky, E.R.; Talmadge, C.; Berglund, P.; Chan, K.W.; Pokrovskaya, E.S.; Kashuba, V.I.; Zhen, D.K.; Boldog, F.; Zabarovskaya, V.I.; et al. Somatic cell hybrid panel and NotI linking clones for physical mapping of human chromosome 3. Genomics 1994, 20, 105–113. [Google Scholar]
- Zabarovsky, E.R.; Allikmets, R.; Kholodnyuk, I.; Zabarovska, V.I.; Paulsson, N.; Bannikov, V.M.; Kashuba, V.I.; Dean, M.; Kisselev, L.L.; Klein, G. Construction of representative NotI linking libraries specific for the total human genome and for human chromosome 3. Genomics 1994, 20, 312–316. [Google Scholar]
- Kutsenko, A.S.; Gizatullin, R.Z.; Al-Amin, A.N.; Wang, F.; Kvasha, S.M.; Podowski, R.M.; Matushkin, Y.G.; Gyanchandani, A.; Muravenko, O.V.; Levitsky, V.G.; et al. NotI flanking sequences: A tool for gene discovery and verification of the human genome. Nucleic Acids Res 2002, 30, 3163–3170. [Google Scholar]
- Zabarovsky, E.R.; Gizatullin, R.; Podowski, R.M.; Zabarovska, V.V.; Xie, L.; Muravenko, O.V.; Kozyrev, S.; Petrenko, L.; Skobeleva, N.; Li, J.; et al. NotI clones in the analysis of the human genome. Nucleic Acids Res 2000, 28, 1635–1639. [Google Scholar]
- Li, J.; Protopopov, A.; Wang, F.; Senchenko, V.; Petushkov, V.; Vorontsova, O.; Petrenko, L.; Zabarovska, V.; Muravenko, O.; Braga, E.; et al. NotI subtraction and NotI-specific microarrays to detect copy number and methylation changes in whole genomes. Proc. Natl. Acad. Sci. USA 2002, 99, 10724–10729. [Google Scholar]
- Lind, G.E.; Skotheim, R.I.; Fraga, M.F.; Abeler, V.M.; Esteller, M.; Lothe, R.A. Novel epigenetically deregulated genes in testicular cancer include homeobox genes and SCGB3A1 (HIN-1). J. Pathol 2006, 210, 441–449. [Google Scholar]
- Singh, U.; Roswall, P.; Uhrbom, L.; Westermark, B. CGGBP1 regulates cell cycle in cancer cells. BMC Mol. Biol 2011, 12, 28. [Google Scholar]
- Cody, N.A.; Shen, Z.; Ripeau, J.S.; Provencher, D.M.; Mes-Masson, A.M.; Chevrette, M.; Tonin, P.N. Characterization of the 3p12.3-pcen region associated with tumor suppression in a novel ovarian cancer cell line model genetically modified by chromosome 3 fragment transfer. Mol. Carcinog 2009, 48, 1077–1092. [Google Scholar]
- Dong, Y.; Mao-Ying, Q.L.; Chen, J.W.; Yang, C.J.; Wang, Y.Q.; Tan, Z.M. Involvement of EphB1 receptor/ephrinB1 ligand in bone cancer pain. Neurosci. Lett 2011, 496, 163–167. [Google Scholar]
- Liu, S.; Liu, W.T.; Liu, Y.P.; Dong, H.L.; Henkemeyer, M.; Xiong, L.Z.; Song, X.J. Blocking EphB1 receptor forward signaling in spinal cord relieves bone cancer pain and rescues analgesic effect of morphine treatment in rodents. Cancer Res 2011, 71, 4392–4402. [Google Scholar]
- Kuang, S.Q.; Bai, H.; Fang, Z.H.; Lopez, G.; Yang, H.; Tong, W.; Wang, Z.Z.; Garcia-Manero, G. Aberrant DNA methylation and epigenetic inactivation of Eph receptor tyrosine kinases and ephrin ligands in acute lymphoblastic leukemia. Blood 2010, 115, 2412–2419. [Google Scholar]
- Sheng, Z.; Wang, J.; Dong, Y.; Ma, H.; Zhou, H.; Sugimura, H.; Lu, G.; Zhou, X. EphB1 is underexpressed in poorly differentiated colorectal cancers. Pathobiology 2008, 75, 274–280. [Google Scholar]
- Wang, J.D.; Dong, Y.C.; Sheng, Z.; Ma, H.H.; Li, G.L.; Wang, X.L.; Lu, G.M.; Sugimura, H.; Jin, J.; Zhou, X.J. Loss of expression of EphB1 protein in gastric carcinoma associated with invasion and metastasis. Oncology 2007, 73, 238–245. [Google Scholar]
- Berardi, A.C.; Marsilio, S.; Rofani, C.; Salvucci, O.; Altavista, P.; Perla, F.M.; Diomedi-Camassei, F.; Uccini, S.; Kokai, G.; Landuzzi, L.; et al. Up-regulation of EphB and ephrin-B expression in rhabdomyosarcoma. Anticancer Res 2008, 28, 763–769. [Google Scholar]
- Huynh-Do, U.; Vindis, C.; Liu, H.; Cerretti, D.P.; McGrew, J.T.; Enriquez, M.; Chen, J.; Daniel, T.O. Ephrin-B1 transduces signals to activate integrin-mediated migration, attachment and angiogenesis. J. Cell Sci 2002, 115, 3073–3081. [Google Scholar]
- Pejovic, T.; Pande, N.T.; Mori, M.; Mhawech-Fauceglia, P.; Harrington, C.; Mongoue-Tchokote, S.; Dim, D.; Andrews, C.; Beck, A.; Tarumi, Y.; et al. Expression profiling of the ovarian surface kinome reveals candidate genes for early neoplastic changes. Translational Oncology 2009, 2, 341–349. [Google Scholar]
- Chen, Z.; Xiao, Y.; Zhang, J.; Li, J.; Liu, Y.; Zhao, Y.; Ma, C.; Luo, J.; Qiu, Y.; Huang, G.; Korteweg, C.; Gu, J. Transcription factors E2A, FOXO1 and FOXP1 regulate recombination activating gene expression in cancer cells. PLoS One 2011, 6, e20475. [Google Scholar]
- Goatly, A.; Bacon, C.M.; Nakamura, S.; Ye, H.; Kim, I.; Brown, P.J.; Ruskone-Fourmestraux, A.; Cervera, P.; Streubel, B.; Banham, A.H.; Du, M.Q. FOXP1 abnormalities in lymphoma: translocation breakpoint mapping reveals insights into deregulated transcriptional control. Mod. Pathol 2008, 21, 902–911. [Google Scholar]
- Streubel, B.; Vinatzer, U.; Lamprecht, A.; Raderer, M.; Chott, A. T(3;14)(p14.1;q32) involving IGH and FOXP1 is a novel recurrent chromosomal aberration in MALT lymphoma. Leukemia 2005, 19, 652–658. [Google Scholar]
- Fenton, J.A.; Schuuring, E.; Barrans, S.L.; Banham, A.H.; Rollinson, S.J.; Morgan, G.J.; Jack, A.S.; van Krieken, J.H.; Kluin, P.M. t(3;14)(p14;q32) results in aberrant expression of FOXP1 in a case of diffuse large B-cell lymphoma. Genes Chromosomes Cancer 2006, 45, 164–168. [Google Scholar]
- Banham, A.H.; Beasley, N.; Campo, E.; Fernandez, P.L.; Fidler, C.; Gatter, K.; Jones, M.; Mason, D.Y.; Prime, J.E.; Trougouboff, P.; et al. The FOXP1 winged helix transcription factor is a novel candidate tumor suppressor gene on chromosome 3p. Cancer Res 2001, 61, 8820–8829. [Google Scholar]
- Toma, M.I.; Grosser, M.; Herr, A.; Aust, D.E.; Meye, A.; Hoefling, C.; Fuessel, S.; Wuttig, D.; Wirth, M.P.; Baretton, G.B. Loss of heterozygosity and copy number abnormality in clear cell renal cell carcinoma discovered by high-density affymetrix 10K single nucleotide polymorphism mapping array. Neoplasia 2008, 10, 634–642. [Google Scholar]
- Wlodarska, I.; Veyt, E.; de Paepe, P.; Vandenberghe, P.; Nooijen, P.; Theate, I.; Michaux, L.; Sagaert, X.; Marynen, P.; Hagemeijer, A.; de Wolf-Peeters, C. FOXP1, a gene highly expressed in a subset of diffuse large B-cell lymphoma, is recurrently targeted by genomic aberrations. Leukemia 2005, 19, 1299–1305. [Google Scholar]
- Giatromanolaki, A.; Koukourakis, M.I.; Sivridis, E.; Gatter, K.C.; Harris, A.L.; Banham, A.H. Loss of expression and nuclear/cytoplasmic localization of the FOXP1 forkhead transcription factor are common events in early endometrial cancer: Relationship with estrogen receptors and HIF-1alpha expression. Mod. Pathol 2006, 19, 9–16. [Google Scholar]
- Brown, P.J.; Ashe, S.L.; Leich, E.; Burek, C.; Barrans, S.; Fenton, J.A.; Jack, A.S.; Pulford, K.; Rosenwald, A.; Banham, A.H. Potentially oncogenic B-cell activation-induced smaller isoforms of FOXP1 are highly expressed in the activated B cell-like subtype of DLBCL. Blood 2008, 111, 2816–2824. [Google Scholar]
- Green, M.R.; Gandhi, M.K.; Courtney, M.J.; Marlton, P.; Griffiths, L. Relative abundance of full-length and truncated FOXP1 isoforms is associated with differential NFκB activity in Follicular Lymphoma. Leuk. Res 2009, 33, 1699–1702. [Google Scholar]
- Taylor, B.S.; Schultz, N.; Hieronymus, H.; Gopalan, A.; Xiao, Y.; Carver, B.S.; Arora, V.K.; Kaushik, P.; Cerami, E.; Reva, B.; et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 2010, 18, 11–22. [Google Scholar]
- Kim, Y.S.; Hwan, J.D.; Bae, S.; Bae, D.H.; Shick, W.A. Identification of differentially expressed genes using an annealing control primer system in stage III serous ovarian carcinoma. BMC Cancer 2010, 10, 576. [Google Scholar]
- Winn, R.A.; van Scoyk, M.; Hammond, M.; Rodriguez, K.; Crossno, J.T., Jr; Heasley, L.E.; Nemenoff, R.A. Antitumorigenic effect of Wnt 7a and Fzd 9 in non-small cell lung cancer cells is mediated through ERK-5-dependent activation of peroxisome proliferator-activated receptor gamma. J. Biol. Chem 2006, 281, 26943–26950. [Google Scholar]
- Bren-Mattison, Y.; van Putten, V.; Chan, D.; Winn, R.; Geraci, M.W.; Nemenoff, R.A. Peroxisome proliferator-activated receptor-gamma (PPAR(gamma)) inhibits tumorigenesis by reversing the undifferentiated phenotype of metastatic non-small-cell lung cancer cells (NSCLC). Oncogene 2005, 24, 1412–1422. [Google Scholar]
- Leung, W.K.; Bai, A.H.; Chan, V.Y.; Yu, J.; Chan, M.W.; To, K.F.; Wu, J.R.; Chan, K.K.; Fu, Y.G.; Chan, F.K.; Sung, J.J. Effect of peroxisome proliferator activated receptor gamma ligands on growth and gene expression profiles of gastric cancer cells. Gut 2004, 53, 331–338. [Google Scholar]
- Tennis, M.A.; Vanscoyk, M.M.; Wilson, L.A.; Kelley, N.; Winn, R.A. Methylation of Wnt7a is modulated by DNMT1 and cigarette smoke condensate in non-small cell lung cancer. PLoS One 2012, 7, e32921. [Google Scholar]
- Yoshioka, S.; King, M.L.; Ran, S.; Okuda, H.; Maclean, J.A., 2nd; McAsey, M.E.; Sugino, N.; Brard, L.; Watabe, K.; Hayashi, K. WNT7A Regulates Tumor Growth and Progression in Ovarian Cancer through the WNT/beta-Catenin Pathway. Mol. Cancer Res 2012, 10, 496–482. [Google Scholar]
- Calvo, R.; West, J.; Franklin, W.; Erickson, P.; Bemis, L.; Li, E.; Helfrich, B.; Bunn, P.; Roche, J.; Brambilla, E.; et al. Altered HOX and WNT7A expression in human lung cancer. Proc. Natl. Acad. Sci. USA 2000, 97, 12776–12781. [Google Scholar]
- Winn, R.A.; Marek, L.; Han, S.Y.; Rodriguez, K.; Rodriguez, N.; Hammond, M.; van Scoyk, M.; Acosta, H.; Mirus, J.; Barry, N.; et al. Restoration of Wnt-7a expression reverses non-small cell lung cancer cellular transformation through frizzled-9-mediated growth inhibition and promotion of cell differentiation. J. Biol. Chem 2005, 280, 19625–19634. [Google Scholar]
- Li, S.; Chiang, T.C.; Davis, G.R.; Williams, R.M.; Wilson, V.P.; McLachlan, J.A. Decreased expression of Wnt7a mRNA is inversely associated with the expression of estrogen receptor-alpha in human uterine leiomyoma. J. Clin. Endocrinol. Metabol 2001, 86, 454–457. [Google Scholar]
- Fluge, O.; Bruland, O.; Akslen, L.A.; Lillehaug, J.R.; Varhaug, J.E. Gene expression in poorly differentiated papillary thyroid carcinomas. Thyroid 2006, 16, 161–175. [Google Scholar]
- Merritt, M.A.; Parsons, P.G.; Newton, T.R.; Martyn, A.C.; Webb, P.M.; Green, A.C.; Papadimos, D.J.; Boyle, G.M. Expression profiling identifies genes involved in neoplastic transformation of serous ovarian cancer. BMC Cancer 2009, 9, 378. [Google Scholar]
- Zhang, X.L.; Peng, C.J.; Peng, J.; Jiang, L.Y.; Ning, X.M.; Zheng, J.H. Prognostic role of Wnt7a expression in ovarian carcinoma patients. Neoplasma 2010, 57, 545–551. [Google Scholar]
- Prado, S.M.D.; Villaamil, V.M.; Gallego, G.A.; Calvo, M.B.; Cedrun, J.L.L.; Soliva, S.S.; Ayerbes, M.V.; Campelo, R.G.; Aparicio, L.M.A. Expression of Wnt gene family and frizzled receptors in head and neck squamous cell carcinomas. Virchows. Arch 2009, 455, 67–75. [Google Scholar]
- Hsu, Y.A.; Lin, H.J.; Sheu, J.J.; Shieh, F.K.; Chen, S.Y.; Lai, C.H.; Tsai, F.J.; Wan, L.; Chen, B.H. A novel interaction between interferon-inducible protein p56 and ribosomal protein L15 in gastric cancer cells. DNA Cell Biol 2011, 30, 671–679. [Google Scholar]
- Gerashchenko, G.V.; Bogatyrova, O.O.; Rudenko, E.E.; Kondratov, A.G.; Gordiyuk, V.V.; Zgonnyk, Y.M.; Vozianov, O.F.; Pavlova, T.V.; Zabarovsky, E.R.; Rynditch, A.V.; et al. Genetic and epigenetic changes of NKIRAS1 gene in human renal cell carcinomas. Exp. Oncol 2010, 32, 71–75. [Google Scholar]
- Lin, H.; Wang, Y.; Zhang, X.; Liu, B.; Zhang, W.; Cheng, J. Prognostic significance of kappaB-Ras1 expression in gliomas. Med. Oncol 2011, 29, 1272–1279. [Google Scholar]
- Wang, H.; Zhao, L.N.; Li, K.Z.; Ling, R.; Li, X.J.; Wang, L. Overexpression of ribosomal protein L15 is associated with cell proliferation in gastric cancer. BMC Cancer 2006, 6, 91. [Google Scholar]
- Dang, C.; Gottschling, M.; Manning, K.; O’Currain, E.; Schneider, S.; Sterry, W.; Stockfleth, E.; Nindl, I. Identification of dysregulated genes in cutaneous squamous cell carcinoma. Oncol. Rep 2006, 16, 513–519. [Google Scholar]
- Willett, R.T.; Greene, L.A. Gata2 is required for migration and differentiation of retinorecipient neurons in the superior colliculus. J. Neurosci 2011, 31, 4444–4455. [Google Scholar]
- Kashiwabara, Y.; Sasaki, S.; Matsushita, A.; Nagayama, K.; Ohba, K.; Iwaki, H.; Matsunaga, H.; Suzuki, S.; Misawa, H.; Ishizuka, K.; et al. Functions of PIT1 in GATA2-dependent transactivation of the thyrotropin beta promoter. J. Mol. Endocrinol 2009, 42, 225–237. [Google Scholar]
- Kandimalla, R.; van Tilborg, A.A.; Kompier, L.C.; Stumpel, D.J.; Stam, R.W.; Bangma, C.H.; Zwarthoff, E.C. Genome-wide Analysis of CpG Island Methylation in Bladder Cancer Identified TBX2, TBX3, GATA2, and ZIC4 as pTa-Specific Prognostic Markers. Eur. Urol 2012, 61, 1245–1256. [Google Scholar]
- Bodor, C.; Renneville, A.; Smith, M.; Charazac, A.; Iqbal, S.; Etancelin, P.; Cavenagh, J.; Barnett, M.J.; Kramarzova, K.; Krishnan, B.; et al. Germ-line GATA2 p.THR354MET mutation in familial myelodysplastic syndrome with acquired monosomy 7 and ASXL1 mutation demonstrating rapid onset and poor survival. Haematologica 2012, 97, 890–894. [Google Scholar]
- Hahn, C.N.; Chong, C.E.; Carmichael, C.L.; Wilkins, E.J.; Brautigan, P.J.; Li, X.C.; Babic, M.; Lin, M.; Carmagnac, A.; Lee, Y.K.; et al. Heritable GATA2 mutations associated with familial myelodysplastic syndrome and acute myeloid leukemia. Nat. Genet 2011, 43, 1012–1017. [Google Scholar]
- Acosta, D.; Suzuki, M.; Connolly, D.; Thompson, R.F.; Fazzari, M.J.; Greally, J.M.; Montagna, C. DNA methylation changes in murine breast adenocarcinomas allow the identification of candidate genes for human breast carcinogenesis. Mamm. Genome 2011, 22, 249–259. [Google Scholar]
- Wang, Y.; He, X.; Ngeow, J.; Eng, C. GATA2 negatively regulates PTEN by preventing nuclear translocation of androgen receptor and by androgen-independent suppression of PTEN transcription in breast cancer. Hum. Mol. Genet 2012, 21, 569–576. [Google Scholar]
- Jazdzewski, K.; Boguslawska, J.; Jendrzejewski, J.; Liyanarachchi, S.; Pachucki, J.; Wardyn, K.A.; Nauman, A.; de la Chapelle, A. Thyroid hormone receptor beta (THRB) is a major target gene for microRNAs deregulated in papillary thyroid carcinoma (PTC). J. Clin. Endocrinol. Metab 2011, 96, E546–E553. [Google Scholar]
- Master, A.; Wojcicka, A.; Piekielko-Witkowska, A.; Boguslawska, J.; Poplawski, P.; Tanski, Z.; Darras, V.M.; Williams, G.R.; Nauman, A. Untranslated regions of thyroid hormone receptor beta 1 mRNA are impaired in human clear cell renal cell carcinoma. Biochim. Biophys. Acta 2010, 1802, 995–1005. [Google Scholar]
- Ling, Y.; Xu, X.; Hao, J.; Ling, X.; Du, X.; Liu, X.; Zhao, X. Aberrant methylation of the THRB gene in tissue and plasma of breast cancer patients. Cancer Genet. Cytogenet 2010, 196, 140–145. [Google Scholar]
- Vasiljevic, N.; Wu, K.; Brentnall, A.R.; Kim, D.C.; Thorat, M.A.; Kudahetti, S.C.; Mao, X.; Xue, L.; Yu, Y.; Shaw, G.L.; et al. Absolute quantitation of DNA methylation of 28 candidate genes in prostate cancer using pyrosequencing. Dis. Markers 2011, 30, 151–161. [Google Scholar]
- Muller, I.; Urban, K.; Pantel, K.; Schwarzenbach, H. Comparison of genetic alterations detected in circulating microsatellite DNA in blood plasma samples of patients with prostate cancer and benign prostatic hyperplasia. Ann. N. Y. Acad. Sci 2006, 1075, 222–229. [Google Scholar]
- Schwarzenbach, H.; Chun, F.K.; Muller, I.; Seidel, C.; Urban, K.; Erbersdobler, A.; Huland, H.; Pantel, K.; Friedrich, M.G. Microsatellite analysis of allelic imbalance in tumour and blood from patients with prostate cancer. BJU Int 2008, 102, 253–258. [Google Scholar]
- Qiu, H.; Lotan, R.; Lippman, S.M.; Xu, X.C. Lack of correlation between expression of retinoic acid receptor-beta and loss of heterozygosity on chromosome band 3p24 in esophageal cancer. Genes Chromosomes Cancer 2000, 28, 196–202. [Google Scholar]
- Mannion, C.; Park, W.S.; Man, Y.G.; Zhuang, Z.; Albores-Saavedra, J.; Tavassoli, F.A. Endocrine tumors of the cervix: Morphologic assessment, expression of human papillomavirus, and evaluation for loss of heterozygosity on 1p,3p, 11q, and 17p. Cancer 1998, 83, 1391–1400. [Google Scholar]
- Rowley, H.; Jones, A.; Spandidos, D.; Field, J. Definition of a tumor suppressor gene locus on the short arm of chromosome 3 in squamous cell carcinoma of the head and neck by means of microsatellite markers. Arch. Otolaryngol. Head Neck Surg 1996, 122, 497–501. [Google Scholar]
- Buchhagen, D.L. Frequent involvement of chromosome 3p alterations in lung carcinogenesis: allelotypes of 215 established cell lines at six chromosome 3p loci. J. Cell. Biochem. Suppl 1996, 24, 198–209. [Google Scholar]
- Guigon, C.J.; Kim, D.W.; Willingham, M.C.; Cheng, S.Y. Mutation of thyroid hormone receptor-beta in mice predisposes to the development of mammary tumors. Oncogene 2011, 30, 3381–3390. [Google Scholar]
- Kashuba, V.I.; Li, J.; Wang, F.; Senchenko, V.N.; Protopopov, A.; Malyukova, A.; Kutsenko, A.S.; Kadyrova, E.; Zabarovska, V.I.; Muravenko, O.V.; et al. RBSP3 (HYA22) is a tumor suppressor gene implicated in major epithelial malignancies. Proc. Natl. Acad. Sci. USA 2004, 101, 4906–4911. [Google Scholar]
- Anedchenko, E.A.; Dmitriev, A.A.; Krasnov, G.S.; Kondrat’eva, T.T.; Kopantsev, E.P.; Vinogradova, T.V.; Zinov’eva, M.V.; Zborovskaia, I.B.; Polotskii, B.E.; Sakharova, O.V.; et al. Down-regulation of RBSP3/CTDSPL, NPRL2/G21, RASSF1A, ITGA9, HYAL1 and HYAL2 genes in non-small cell lung cancer. Mol. Biol (Mosk) 2008, 42, 965–976. [Google Scholar]
- Anedchenko, E.A.; Kiseleva, N.P.; Dmitriev, A.A.; Kiselev, F.L.; Zabarovskii, E.R.; Senchenko, V.N. Tumor suppressor gene RBSP3 in cervical carcinoma: Copy number and transcriptional level. Mol. Biol (Mosk) 2007, 41, 86–95. [Google Scholar]
- Pavlova, T.V.; Kashuba, V.I.; Muravenko, O.V.; Yenamandra, S.P.; Ivanova, T.A.; Zabarovskaia, V.I.; Rakhmanaliev, E.R.; Petrenko, L.A.; Pronina, I.V.; Loginov, V.I.; et al. Technology of analysis of epigenetic and structural changes of epithelial tumors genome with NotI-microarrays by the example of human chromosome. Mol. Biol (Mosk) 2009, 43, 339–347. [Google Scholar]
- Senchenko, V.N.; Anedchenko, E.A.; Kondratieva, T.T.; Krasnov, G.S.; Dmitriev, A.A.; Zabarovska, V.I.; Pavlova, T.V.; Kashuba, V.I.; Lerman, M.I.; Zabarovsky, E.R. Simultaneous down-regulation of tumor suppressor genes RBSP3/CTDSPL, NPRL2/G21 and RASSF1A in primary non-small cell lung cancer. BMC Cancer 2010, 10, 75. [Google Scholar]
- Sinha, S.; Singh, R.K.; Alam, N.; Roy, A.; Roychoudhury, S.; Panda, C.K. Frequent alterations of hMLH1 and RBSP3/HYA22 at chromosomal 3p22.3 region in early and late-onset breast carcinoma: Clinical and prognostic significance. Cancer Sci 2008, 99, 1984–1991. [Google Scholar]
- Ghosh, A.; Ghosh, S.; Maiti, G.P.; Sabbir, M.G.; Zabarovsky, E.R.; Roy, A.; Roychoudhury, S.; Panda, C.K. Frequent alterations of the candidate genes hMLH1, ITGA9 and RBSP3 in early dysplastic lesions of head and neck: Clinical and prognostic significance. Cancer Sci 2010, 101, 1511–1520. [Google Scholar]
- Mitra, S.; Mazumder Indra, D.; Bhattacharya, N.; Singh, R.K.; Basu, P.S.; Mondal, R.K.; Roy, A.; Zabarovsky, E.R.; Roychoudhury, S.; Panda, C.K. RBSP3 is frequently altered in premalignant cervical lesions: Clinical and prognostic significance. Genes Chromosomes Cancer 2010, 49, 155–170. [Google Scholar]
- Kashuba, V.I.; Pavlova, T.V.; Grigorieva, E.V.; Kutsenko, A.; Yenamandra, S.P.; Li, J.; Wang, F.; Protopopov, A.I.; Zabarovska, V.I.; Senchenko, V.; et al. High mutability of the tumor suppressor genes RASSF1 and RBSP3 (CTDSPL) in cancer. PLoS One 2009, 4, e5231. [Google Scholar]
- Zheng, Y.S.; Zhang, H.; Zhang, X.J.; Feng, D.D.; Luo, X.Q.; Zeng, C.W.; Lin, K.Y.; Zhou, H.; Qu, L.H.; Zhang, P.; et al. MiR-100 regulates cell differentiation and survival by targeting RBSP3, a phosphatase-like tumor suppressor in acute myeloid leukemia. Oncogene 2012, 31, 80–92. [Google Scholar]
- Shoubridge, C.; Walikonis, R.S.; Gecz, J.; Harvey, R.J. Subtle functional defects in the Arf-specific guanine nucleotide exchange factor IQSEC2 cause non-syndromic X-linked intellectual disability. Small GTPases 2010, 1, 98–103. [Google Scholar]
- Someya, A.; Moss, J.; Nagaoka, I. The guanine nucleotide exchange protein for ADP-ribosylation factor 6, ARF-GEP100/BRAG2, regulates phagocytosis of monocytic phagocytes in an ARF6-dependent process. J. Biol. Chem 2010, 285, 30698–30707. [Google Scholar]
- Morishige, M.; Hashimoto, S.; Ogawa, E.; Toda, Y.; Kotani, H.; Hirose, M.; Wei, S.; Hashimoto, A.; Yamada, A.; Yano, H.; et al. GEP100 links epidermal growth factor receptor signalling to Arf6 activation to induce breast cancer invasion. Nat. Cell. Biol 2008, 10, 85–92. [Google Scholar]
- Sabe, H.; Hashimoto, S.; Morishige, M.; Ogawa, E.; Hashimoto, A.; Nam, J.M.; Miura, K.; Yano, H.; Onodera, Y. The EGFR-GEP100-Arf6-AMAP1 signaling pathway specific to breast cancer invasion and metastasis. Traffic 2009, 10, 982–993. [Google Scholar]
- Gunay-Aygun, M.; Falik-Zaccai, T.C.; Vilboux, T.; Zivony-Elboum, Y.; Gumruk, F.; Cetin, M.; Khayat, M.; Boerkoel, C.F.; Kfir, N.; Huang, Y.; et al. NBEAL2 is mutated in gray platelet syndrome and is required for biogenesis of platelet alpha-granules. Nat. Genet 2011, 43, 732–734. [Google Scholar]
- SABiosciences Gene Network Central Home Page. Available online: http://www.sabiosciences.com/genenetwork/genenetworkcentral.php accessed on 20 June 2012.
- Tohyama, J.; Kato, M.; Kawasaki, S.; Harada, N.; Kawara, H.; Matsui, T.; Akasaka, N.; Ohashi, T.; Kobayashi, Y.; Matsumoto, N. Dandy-Walker malformation associated with heterozygous ZIC1 and ZIC4 deletion: Report of a new patient. Am. J. Med. Genet. A 2011, 155A, 130–133. [Google Scholar]
- Aruga, J.; Nozaki, Y.; Hatayama, M.; Odaka, Y.S.; Yokota, N. Expression of ZIC family genes in meningiomas and other brain tumors. BMC Cancer 2010, 10, 79. [Google Scholar]
- Pourebrahim, R.; Van Dam, K.; Bauters, M.; De Wever, I.; Sciot, R.; Cassiman, J.J.; Tejpar, S. ZIC1 gene expression is controlled by DNA and histone methylation in mesenchymal proliferations. FEBS Lett 2007, 581, 5122–5126. [Google Scholar]
- Bataller, L.; Wade, D.F.; Graus, F.; Stacey, H.D.; Rosenfeld, M.R.; Dalmau, J. Antibodies to Zic4 in paraneoplastic neurologic disorders and small-cell lung cancer. Neurology 2004, 62, 778–782. [Google Scholar]
- Sabater, L.; Bataller, L.; Suarez-Calvet, M.; Saiz, A.; Dalmau, J.; Graus, F. ZIC antibodies in paraneoplastic cerebellar degeneration and small cell lung cancer. J. Neuroimmunol 2008, 201–202, 163–165. [Google Scholar]
- GeneCards Home Page. Available online: http://genecards.org/ accessed on 20 June 2012.
- Dunwell, T.L.; Hesson, L.B.; Pavlova, T.; Zabarovska, V.; Kashuba, V.; Catchpoole, D.; Chiaramonte, R.; Brini, A.T.; Griffiths, M.; Maher, E.R.; et al. Epigenetic analysis of childhood acute lymphoblastic leukemia. Epigenetics 2009, 4, 185–193. [Google Scholar]
- Haraldson, K.; Kashuba, V.I.; Dmitriev, A.A.; Senchenko, V.N.; Kudryavtseva, A.V.; Pavlova, T.V.; Braga, E.A.; Pronina, I.V.; Kondratov, A.G.; Rynditch, A.V.; et al. LRRC3B gene is frequently epigenetically inactivated in several epithelial malignancies and inhibits cell growth and replication. Biochimie 2012, 94, 1151–1157. [Google Scholar]
- Law, E.W.; Cheung, A.K.; Kashuba, V.I.; Pavlova, T.V.; Zabarovsky, E.R.; Lung, H.L.; Cheng, Y.; Chua, D.; Lai-Wan Kwong, D.; Tsao, S.W.; et al. Anti-angiogenic and tumor-suppressive roles of candidate tumor-suppressor gene, Fibulin-2, in nasopharyngeal carcinoma. Oncogene 2012, 31, 728–738. [Google Scholar]
- Buys, S.S.; Partridge, E.; Greene, M.H.; Prorok, P.C.; Reding, D.; Riley, T.L.; Hartge, P.; Fagerstrom, R.M.; Ragard, L.R.; Chia, D.; et al. Ovarian cancer screening in the Prostate, Lung, Colorectal and Ovarian (PLCO) cancer screening trial: Findings from the initial screen of a randomized trial. Am. J. Obstet. Gynecol 2005, 193, 1630–1639. [Google Scholar]
- Balch, C.; Fang, F.; Matei, D.E.; Huang, T.H.; Nephew, K.P. Minireview: epigenetic changes in ovarian cancer. Endocrinology 2009, 150, 4003–4011. [Google Scholar]
- Baylin, S.B.; Jones, P.A. A decade of exploring the cancer epigenome—Biological and translational implications. Nat. Rev. Cancer 2011, 11, 726–734. [Google Scholar]
- Baylin, S.B.; Ohm, J.E. Epigenetic gene silencing in cancer—A mechanism for early oncogenic pathway addiction? Nat. Rev. Cancer 2006, 6, 107–116. [Google Scholar]
- Widschwendter, M.; Jiang, G.; Woods, C.; Muller, H.M.; Fiegl, H.; Goebel, G.; Marth, C.; Muller-Holzner, E.; Zeimet, A.G.; Laird, P.W.; et al. DNA hypomethylation and ovarian cancer biology. Cancer Res 2004, 64, 4472–4480. [Google Scholar]
- Collins, Y.; Dicioccio, R.; Keitz, B.; Lele, S.; Odunsi, K. Methylation of death-associated protein kinase in ovarian carcinomas. Int. J. Gynecol. Cancer 2006, 16 Suppl 1, 195–199. [Google Scholar]
- Gifford, G.; Paul, J.; Vasey, P.A.; Kaye, S.B.; Brown, R. The acquisition of hMLH1 methylation in plasma DNA after chemotherapy predicts poor survival for ovarian cancer patients. Clin. Cancer Res 2004, 10, 4420–4426. [Google Scholar]
- Liggett, T.E.; Melnikov, A.; Yi, Q.; Replogle, C.; Hu, W.; Rotmensch, J.; Kamat, A.; Sood, A.K.; Levenson, V. Distinctive DNA methylation patterns of cell-free plasma DNA in women with malignant ovarian tumors. Gynecol. Oncol 2011, 120, 113–120. [Google Scholar]
- Melnikov, A.; Scholtens, D.; Godwin, A.; Levenson, V. Differential methylation profile of ovarian cancer in tissues and plasma. J. Molecular Diagn 2009, 11, 60–65. [Google Scholar]
- Gloss, B.S.; Samimi, G. Epigenetic biomarkers in epithelial ovarian cancer. Cancer Lett 2012. [Google Scholar] [CrossRef]
- Kashuba, V.I.; Gizatullin, R.Z.; Protopopov, A.I.; Li, J.; Vorobieva, N.V.; Fedorova, L.; Zabarovska, V.I.; Muravenko, O.V.; Kost-Alimova, M.; Domninsky, D.A.; et al. Analysis of NotI linking clones isolated from human chromosome 3 specific libraries. Gene 1999, 239, 259–271. [Google Scholar]
- Dmitriev, A.A.; Kashuba, V.I.; Haraldson, K.; Senchenko, V.N.; Pavlova, T.V.; Kudryavtseva, A.V.; Anedchenko, E.A.; Krasnov, G.S.; Pronina, I.V.; Loginov, V.I.; et al. Genetic and epigenetic analysis of non-small cell lung cancer with NotI-microarrays. Epigenetics 2012, 7, 502–513. [Google Scholar]
- AnalystSoft BioStat home page. Available online: http://www.analystsoft.com/en/products/biostat/ accessed on 27 November 2009.
- Altman, D.G.; Bland, J.M. Diagnostic tests. 1: Sensitivity and specificity. BMJ 1994, 308, 1552. [Google Scholar]
- Dreijerink, K.; Braga, E.; Kuzmin, I.; Geil, L.; Duh, F.M.; Angeloni, D.; Zbar, B.; Lerman, M.I.; Stanbridge, E.J.; Minna, J.D.; et al. The candidate tumor suppressor gene, RASSF1A, from human chromosome 3p21.3 is involved in kidney tumorigenesis. Proc. Natl. Acad. Sci. USA 2001, 98, 7504–7509. [Google Scholar]
| No. | NotI-site | Gene | Locus | Event frequency, (%) | |
|---|---|---|---|---|---|
| Met/Del | Met/Del without no info | ||||
| 1 | NR1-XM13C | IQSEC1 | 3p25.2 | 33 (6/18) | 38 (6/16) |
| 2 | NL1-CJ4R (C) | NKIRAS1/RPL15 | 3p24.2 | 33 (6/18) | 35 (6/17) |
| 3 | NL4-BB6R (C) | THRB | 3p24.2 | 33 (6/18) | 38 (6/16) |
| 4 | NL3-CA11RS | LRRC3B | 3p24 | 33 (6/18) | 35 (6/17) |
| 5 | NLJ-003RD | RBSP3(CTDSPL) | 3p21.3 | 33 (6/18) | 33 (6/18) |
| 6 | NR1-KA8R (C) | THRB | 3p24.2 | 28 (5/18) | 56 (5/9) |
| 7 | NL1A401R (D) | ITGA9 | 3p21.3 | 28 (5/18) | 29 (5/17) |
| 8 | NL3A006R (D) | NBEAL2 | 3p21.31 | 28 (5/18) | 33 (5/15) |
| 9 | NL3A001R (D) | GNAI2 | 3p21.31 | 28 (5/18) | 28 (5/18) |
| 10 | NL1-DE18R | GATA2 | 3q21.3 | 28 (5/18) | 28 (5/18) |
| 11 | NL4-BH3R (C) | GATA2 | 3q21.3 | 28 (5/18) | 28 (5/18) |
| 12 | NR1-PD1R | ZIC4 | 3q24 | 28 (5/18) | 31 (5/16) |
| 13 | NL3003R (U) | GORASP1/TTC21A | 3p22–p21.33 | 22 (4/18) | 24 (4/17) |
| 14 | NR1-AN24RS | ABHD5/C3orf77 | 3p21 | 22 (4/18) | 22 (4/18) |
| 15 | NR1-WE11RS | CGGBP1 | 3p12–p11.1 | 22 (4/18) | 24 (4/17) |
| 16 | NL3-CI2R (C) | LOC285205 | 3p13.12 | 22 (4/18) | 27 (4/15) |
| 17 | NR1-WD21R (C) | NEK11/NUDT16 | 3q22.1 | 22 (4/18) | 44 (4/9) |
| 18 | NR5-IO11R (C) | PAQR9 | 3q23 | 22 (4/18) | 25 (4/16) |
| 19 | NR1-AK24R | BCL6 | 3q27 | 22 (4/18) | 24 (4/17) |
| 20 | NL6-FJ5R (C) | LRRN1 | 3p26.2 | 17 (3/18) | 17 (3/18) |
| 21 | NR1-KJ5R (C) | FBLN2 | 3p25.1 | 17 (3/18) | 19 (3/16) |
| 22 | NR1-PL22R (C) | LOC285375 | 3p25.1 | 17 (3/18) | 18 (3/17) |
| 23 | NL4-BK12R (C) | WNT7A | 3p25 | 17 (3/18) | 18 (3/17) |
| 24 | NL1308R (D) | MOBP | 3p22.1 | 17 (3/18) | 19 (3/16) |
| 25 | NR1-NC7RS | PPM1M | 3p21.2 | 17 (3/18) | 19 (3/16) |
| 26 | NR1-NJ9R (C) | PRICKLE2 | 3p14.1 | 17 (3/18) | 18 (3/17) |
| 27 | NL1-BA6R | FOXP1 | 3p14.1 | 17 (3/18) | 17 (3/18) |
| 28 | NL6-F020R (C) | DCBLD2 | 3q12.1 | 17 (3/18) | 19 (3/16) |
| 29 | NL1-GK21R (C) | ROPN1/KALRN | 3q13.3 | 17 (3/18) | 21 (3/14) |
| 30 | NL1290R (D) | CHST13 | 3q21.3 | 17 (3/18) | 17 (3/18) |
| 31 | NL2A230R | ABTB1/PODXL2 | 3q21 | 17 (3/18) | 17 (3/18) |
| 32 | NL4-DJ11RS | TRH | 3q13.3–q21 | 17 (3/18) | 18 (3/17) |
| 33 | NL1A079R (D) | EPHB1 | 3q21–q23 | 17 (3/18) | 19 (3/16) |
| 34 | NL1-FK10R (C) | PPP2R3A | 3q22.1 | 17 (3/18) | 21 (3/14) |
| 35 | NR1-NH1R (C) | FGF12 | 3q28 | 17 (3/18) | 18 (3/17) |
| Gene | Methylation/Deletion frequency, % | p-Parameter * | |
|---|---|---|---|
| Stages I + II | Stages III + IV | ||
| LOC285205 | 0 (0/7) | 50 (4/8) | 0.08 |
| CGGBP1 | 0 (0/7) | 40 (4/10) | 0.10 |
| EPHB1 | 0 (0/7) | 33 (3/9) | 0.21 |
| FOXP1 | 0 (0/8) | 30 (3/10) | 0.22 |
| WNT7A | 0 (0/7) | 30 (3/10) | 0.23 |
| NKIRAS1/RPL15 | 14 (1/7) | 50 (5/10) | 0.30 |
| GATA2 | 13 (1/8) | 40 (4/10) | 0.31 |
| Use | Sets of markers |
|---|---|
| Early detection | NKIRAS1/RPL15, THRB, RBPS3 (CTDSPL), IQSEC1, NBEAL2, ZIC4, LOC285205, FOXP1 Sp = (94 ± 5)% Sn = (72 ± 11)% p < 0.01 |
| Discrimination of BOA and EOC | NKIRAS1/RPL15, THRB, RBPS3 (CTDSPL), IQSEC1, NBEAL2, ZIC4, LOC285205, FOXP1 Sp = (71 ± 17)% Sn = (72 ± 11)% p = 0.04 |
| Discrimination of Stages I + II and Stages III + IV | LOC285205, CGGBP1, EPHB1, NKIRAS1/RPL15 Sp = (88 ± 12)% Sn = (80 ± 13)% p < 0.01 |
| Gene symbol and location | Protein Function | Involvment in cancer |
|---|---|---|
| LOC285205 3q13.12 | This gene encodes uncharacterized protein with moderate expression level in ovary, low level in brain, bladder, skin, breast, and testis (according to the dbEST and SAGE). Rather high expression level of this gene is observed in ovarian normal tissue (*). | Only EST and SAGE data is available. According to this, expression level in ovary, testis and some types of brain tumors is expected to be decreased. |
| CGGBP1 3p12–p11.1 | Binds to unmethylated 5′-d(CGG)(n)-3′ trinucleotide repeats in the FMR1 (fragile X mental retardation gene) promoter and the ribosomal RNA gene clusters. Regulates FMR1 gene expression. Regulates gene expression during heat shock stress response. CGGBP1 is known to be a cell cycle regulatory midbody protein required for normal cytokinetic abscission in normal human fibroblasts (*). | Decreased mRNA level in testis cancer and various cell lines [23,24]. Microsatellite instability in ovarian cancer cell line [25]. The role of CGGBP1 in cell cycle involves multiple mechanisms: depletion of CGGBP1 mRNA observed in tumor cells leads to increase of the expression of cell cycle regulatory genes CDKN1A and GAS1; otherwise, a presence of CGGBP1 is required for the ability of cancer cells to progress cell cycle beyond G0/G1 [24]. |
| EPHB1 3q21–q23 | Encodes a member of attractive and repulsive axon-guidance molecules family (that includes SEMA5A, in addition); mediates numerous developmental processes, particularly in the nervous system. Receptor for members of the ephrin-B family. Binds to ephrin-B1, -B2 and -B3. Binding with the guidance cue ephrin-B2 at the optic chiasm midline redirect ventrotemporal (VT) retinal ganglion cells (RGCs) axons ipsilaterally. May be involved in cell-cell interactions in the nervous system (*) | Involvement in bone cancer pain [26,27]. Aberrant DNA methylation and epigenetic inactivation in acute lymphoblastic leukemia [28]. Underexpressed in poorly differentiated colorectal cancers [29]. Loss of expression in gastric carcinoma associated with invasion and metastasis [30]. Up-regulation in rhabdomyosarcoma [31]. Transduces signals to activate integrin-mediated migration, attachment and angiogenesis [32]. Expression level alterations in ovarian cancer [33] |
| FOXP1 3p14.1 | FOXP1 belongs to the family of Forkhead box proteins, which contain a common DNA-binding domain termed the forkhead box or winged helix domain. FOXP1 is involved in the negative regulation of tissue- and cell type-specific gene transcription. FOXO1 and FOXP1 also have regulatory function in recombination activating gene 1 (RAG) expression in cancer cells [34]. | FOXP1 has been reported to be associated with development of various types of tumors. Involved in chromosomal translocation in MALT lymphoma [35,36] and in large B-cell lymphoma [37]. Deletions, both mRNA and protein down-regulation in a wide range of tumors [38]. LOH and copy number alterations in kidney cancer [39]. Highly expressed in a subset of B-cell lymphoma [40]. Down-regulated in endometrial cancer [41]. High expression of tumor-specific smaller isoforms in B-cell lymphoma and Follicular lymphomas [42,43]. FOXP1 is located in the chromosomal region 3p14.1 reported to contain a number of TSGs [38,44]. FOXP1 is found to be significantly down-regulated in stage III serous ovarian carcinoma [45]. |
| WNT7A 3p25 | A member of the WNT gene family, which consists of structurally related genes that encode secreted signaling proteins. These proteins have been implicated in oncogenesis and in several developmental processes, including regulation of cell development and patterning during embryogenesis. WNT7A binds to the Fzd9 receptor and signals through ERK-5 to activate the tumor suppressor peroxisome proliferator-activated receptor γ (PPARγ) [46]. PAPRγ inhibits transformed cells growth and metastasis and promote epithelial differentiation and have demonstrated tumor prevention efficacy [47,48]. | Methylation of WNT7A promoter modulated with DNMT1 has been reported for non-small cell lung cancer [49]. It was shown that WNT7A regulates tumor growth and progression in ovarian cancer through the WNT/β-catenin pathway abnormally activated in ovarian cancer. Abundant WNT7A was found in the epithelium of serous ovarian carcinomas, but not detected in borderline and benign tumors, normal ovary, or endometrioid carcinomas [50]. Down-regulation in lung cancer [51,52], in uterine leiomyoma [53]. Overexpression in thyroid cancer [54], in ovarian cancer, associated with poor prognosis [55,56]. Differential expression (down-regulation), associated with poor prognosis in head and neck squamous cell carcinoma [57]. |
| NKIRAS1/RPL15 3p24.2 | NKIRAS1: Atypical Ras-like protein that acts as a potent regulator of NF-κ-B activity by preventing the degradation of NF-κ-B inhibitor beta (NFKBIB). Both GTP- and GDP-bound forms block phosphorylation of NFKBIB (*) RPL15: A ribosomal protein that is a component of the large 60S subunit. The protein belongs to the L15E family of ribosomal proteins. Transcript variants utilizing alternative polyA signals exist. Interacts with IFIT1 [58]; up-regulation of both IFIT1 and RPL15 may lead to proliferative inhibition of gastric cancer cells [58]. | NKIRAS1: Chromosomal aberrations and subsequent down-regulation in kidney cancer; furthermore, high grade kidney tumors (III and IV stage) revealed lower NKIRAS1 mRNA level than low grade ones (stage I and II) [59]. Overexpression, associated with poor prognosis in gliomas [60]. RPL15: Overexpression in gastric cancer [61]. Differentially expressed in cutaneous squamous cell carcinoma [62]. |
| GATA2 3q21.3 | This gene encodes a member of the GATA family of zinc-finger transcription factors. GATA proteins bind the DNA sequence WGATAR and, along with other cofactors, drive expression of target genes important in development of a variety of tissues [63]. For example, the encoded protein plays an essential role in regulating transcription of genes involved in the development and proliferation of hematopoietic and endocrine cell lineages, e.g., activation of beta-thyrotropin (thyroid-stimulating hormone) expression [64]. | GATA2 along with ZIC4 was found to have methylated CpG islands in bladder cancer [65]. GATA2 mutations are associated with hereditary myelodysplastic syndrome and extreme risk of acute myelogenous leukemia development [66,67]. Considering murine model, GATA2 promoter methylation was found to be associated with development of breast cancer (BC); its down-regulation was seen for human BC [68]. However, GATA2 negatively regulates PTEN (phosphatase and tensin homolog deleted on chromosome 10) tumor suppressor by preventing nuclear translocation of androgen receptor and by androgen-independent suppression of PTEN transcription in breast cancer [69]. |
| THRB 3p24.2 | Encodes receptor of nuclear hormone receptor for triiodothyronine. The thyroid hormone receptors (TRs) are transcription factors that mediate the pleiotropic activities of the thyroid hormone, T3. TRs regulate cell proliferation, differentiation, and apoptosis [70]. These TRs are expressed in a tissue-dependent and developmentally regulated manner. Different hormone receptors, while having certain extent of redundancy, may mediate different functions of thyroid hormone. THRB acts as a tumor suppressor and disturbances of the THRB gene are frequent findings in cancer [71]. | In mouse models, a truncated THRB gene leads to thyroid cancer (TC); it can be down-regulated at least with seven miRNAs overexpressed in papillary TC [70]. THRB aberrant methylation can be found in tissue and plasma of BC patients [72]. THRB revealed a low frequency of methylation in prostate cancer samples [73], but high frequency of LOH in prostate [74,75], esophageal cancer [76], endocrine tumors of the cervix [77], head and neck cancer [78]; also small LOH frequencies were shown for NSCLC [79]. Mutation of this gene in mice predisposes to the development of mammary tumors [80]. Reduced THRB expression was shown for clear cell renal cell cancer samples which can be resulted from regulatory effects of THRB 5′ and 3′ UTRs on THRB protein translation [71]. |
| RBPS3 (CTDSPL) 3p21.3 | RBSP3/CTDSPL belongs to a gene family of small CTD phosphatases that preferentially catalyzes serine-5 dephosphorylation in the specific sequence of the RNA polymerase II (Pol II) large subunit and in other proteins. This leads to inactivation of Pol II and negative regulation of transcriptional activity. RBSP3 is thought also to activate RB1 (retinoblastoma 1) tumor suppressor precursor, that leads to cell cycle arrest at G1/S phases boundary. RBSP3 is TSG whose product is likely to be an important component of the Rb cycle regulation pathway. RBSP3 transcribes two isoforms with antitumor activity, which is more pronounced for the product of isoform B [81]. | RBSP3 showed a low expression level because of deletions and methylation in various epithelial tumors [82–86]; these aberrations were also found in early dysplastic lesions of head and neck [87], premalignant cervical lesions [88]. RBSP3 gene revealed high mutability rate in various primary tumors and cell lines [89]; tumor suppressor activity revealed for lung and renal cancer cell lines ACC-LC5 and KRC/Y, in vitro and in vivo [81]; transient protein expression resulting in a significant decrease of phosphorylated RB1 level, which may lead to cell cycle arrest between G1 and S phases [81]. Acute myeloid leukemia reveals specific overexpression of mir-100 targeting RBSP3, which promotes cell proliferation and blocks granulocyte/monocyte differentiation [90]. |
| IQSEC1 3p25.2 | The representative of guanine-exchange proteins binding to ADP-rybosylation factors. This protein preferentially works as a guanine nucleotide exchange protein for ARF6 (ADP-ribosylation factor), a member of a family of small GTPases, mediating internalization of beta-1 integrin [91]. Regulates phagocytosis of monocytic phagocytes [92]. | The EGFR-IQSEC1-ARF6-AMAP1 signaling pathway is essential for breast cancer (BC) invasion and metastasis. Overexpressed IQSEC1 is responsible for activation of ARF6 which leads to BC invasion and metastasis. IQSEC1, in turn, is activated by ligand-dependent epidermal growth factor receptor (EGFR) [93,94]. |
| NBEAL2 3p21.31 | Encodes a BEACH/ARM/WD40 domain protein. Mutations in this gene are leading to gray platelet syndrome (a rare congenital bleeding disorder caused by a reduction or absence of alpha-granules in blood platelets) [95]. NBEAL2 protein is predicted to interact with WDFY3 (WD repeat and FYVE domain containing 3), which itself interacts with CHS1, and with DLL1 and JAG1 [96], known to have roles in hematopoiesis. | The gene is located in close proximity to the LUCA and AP20 regions subject to frequent aberrations in various tumors, but there are no literature data concerning such NBEAL2 alterations. GeneNote, EST and SAGE analysis reveal omnipresent expression character of NBEAL and allow to expect its probable mRNA level decreases in thymus, brain (various type of tumors), liver, pancreas, prostate cancer and leukemia (*). |
| ZIC4 3q24 | Encodes a member of the ZIC family of C2H2-type zinc finger proteins. Members of this family plays important roles during development, and have been associated with X-linked visceral heterotaxy and holoprosencephaly type 5; heterozygous deletion of ZIC4 are associated with Dandy-Walker cerebellum malformation syndrome [97]. | ZIC4 along with GATA2 was found to have methylated CpG islands in bladder cancer; it was associated with high extent of progression and invasive character of bladder tumors [65]. Zic4 along with Zic1–5, other members of this family, was shown to suppress β-catenin-mediated transcriptional activation within the Wnt/β-catenin signaling pathway (in Xenopus laevis). ZIC1, ZIC2, and ZIC5 were found to be novel molecular markers for meningiomas whereas ZIC4 expression is highly selective for medulloblastomas [98]. Using NotI-microarrays, ZIC4 aberrant methylation/deletions were found for various types of tumors [84]. Consistent up-regulation of the neural transcription factors ZIC1 and ZIC4 was shown for desmoid tumors and other fibroproliferative disorders [99]. ZIC4 aberrations are associated with paraneoplastic neurologic disorders and small-cell lung cancer [100,101]. |
© 2012 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kashuba, V.; Dmitriev, A.A.; Krasnov, G.S.; Pavlova, T.; Ignatjev, I.; Gordiyuk, V.V.; Gerashchenko, A.V.; Braga, E.A.; Yenamandra, S.P.; Lerman, M.; et al. NotI Microarrays: Novel Epigenetic Markers for Early Detection and Prognosis of High Grade Serous Ovarian Cancer. Int. J. Mol. Sci. 2012, 13, 13352-13377. https://doi.org/10.3390/ijms131013352
Kashuba V, Dmitriev AA, Krasnov GS, Pavlova T, Ignatjev I, Gordiyuk VV, Gerashchenko AV, Braga EA, Yenamandra SP, Lerman M, et al. NotI Microarrays: Novel Epigenetic Markers for Early Detection and Prognosis of High Grade Serous Ovarian Cancer. International Journal of Molecular Sciences. 2012; 13(10):13352-13377. https://doi.org/10.3390/ijms131013352
Chicago/Turabian StyleKashuba, Vladimir, Alexey A. Dmitriev, George S. Krasnov, Tatiana Pavlova, Ilya Ignatjev, Vasily V. Gordiyuk, Anna V. Gerashchenko, Eleonora A. Braga, Surya P. Yenamandra, Michael Lerman, and et al. 2012. "NotI Microarrays: Novel Epigenetic Markers for Early Detection and Prognosis of High Grade Serous Ovarian Cancer" International Journal of Molecular Sciences 13, no. 10: 13352-13377. https://doi.org/10.3390/ijms131013352
APA StyleKashuba, V., Dmitriev, A. A., Krasnov, G. S., Pavlova, T., Ignatjev, I., Gordiyuk, V. V., Gerashchenko, A. V., Braga, E. A., Yenamandra, S. P., Lerman, M., Senchenko, V. N., & Zabarovsky, E. (2012). NotI Microarrays: Novel Epigenetic Markers for Early Detection and Prognosis of High Grade Serous Ovarian Cancer. International Journal of Molecular Sciences, 13(10), 13352-13377. https://doi.org/10.3390/ijms131013352






