Pharmacogenetics of OATP Transporters Reveals That SLCO1B1 c.388A>G Variant Is Determinant of Increased Atorvastatin Response
Abstract
:Aims
Material and Methods
Results
Conclusion
1. Introduction
2. Results and Discussion
2.1. Characteristics of the Hypercholesterolemic Individuals
2.2. SLCO1B1 and SLCO2B1 Polymorphisms
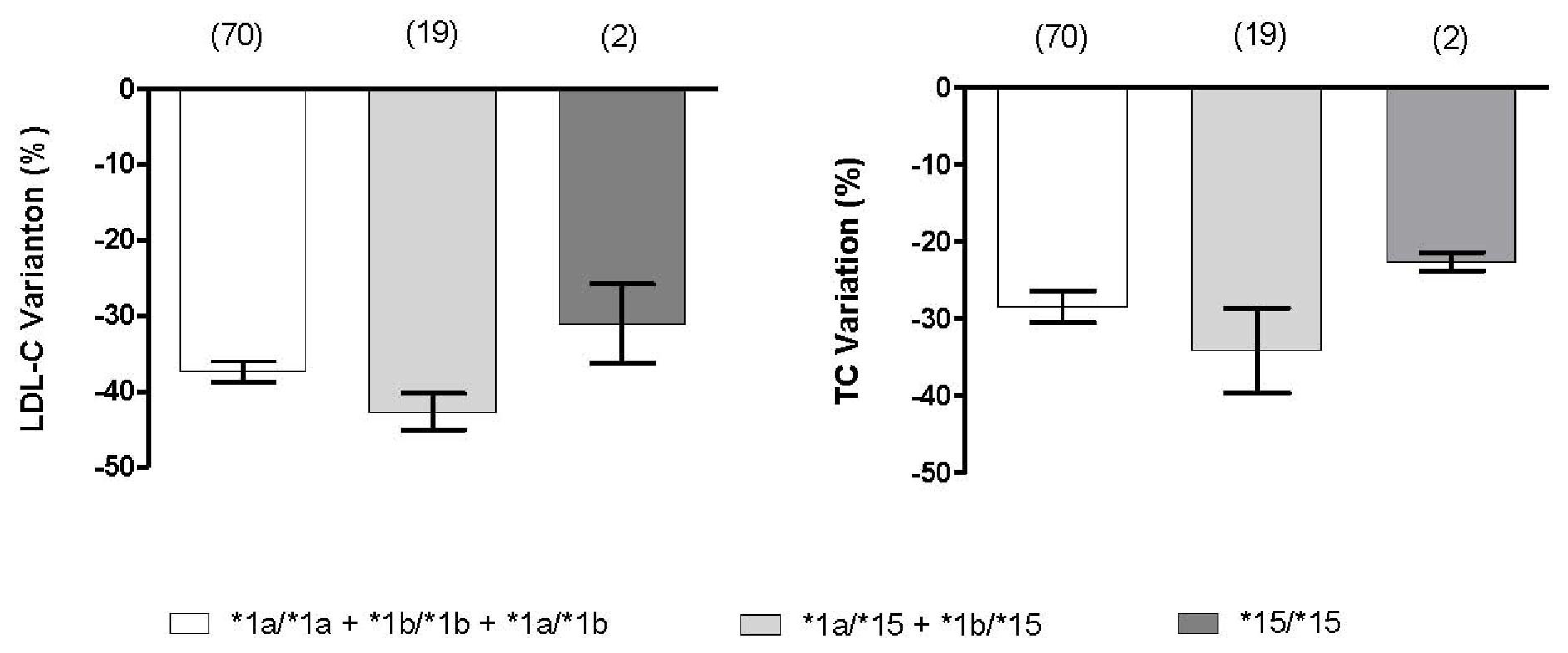
2.3. Effect of SLCO1B1 and SLCO2B1 Polymorphisms on Atorvastatin Response
3. Material and Methods
3.1. Subjects and Study Protocol
3.2. Biochemical Profile and SLCO Variants Genotyping
3.3. Ancestry Informative Markers (AIMs)
3.4. Statistical Analysis
4. Conclusions
Supplementary Information
ijms-12-05815-s001.pdfAcknowledgments
- DisclosureThis work was partially supported by Life Technologies, Sao Paulo, SP, Brazil. The sponsor of this study had no role in the study design, data collection, data analysis, data interpretation or writing of the report. S.G. Purim was salaried personnel of Life Technologies. The authors have no other relevant affiliation or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials in the manuscript apart from those disclosed.
References
- Ieri, I; Higuchi, S; Sugiyama, Y. Genetic polymorphisms of uptake (OATP1B1, 1B3) and efflux (MRP2, BCRP) transporters: Implications for inter-individual differences in the pharmacokinetics and pharmacodynamics of statins and other clinically relevant drugs. Expert Opin Drug Metab Toxicol 2009, 5, 703–729. [Google Scholar]
- Nawrocki, JW; Weiss, SR; Davidson, MH; Sprecher, DL; Schwartz, SL; Lupien, PJ; Jones, PH; Haber, HE; Black, DM. Reduction of LDL cholesterol by 25% to 60% in patients with primary hypercholesterolemia by atorvastatin, a new HMG-CoA reductase inhibitor. Arterioscler Thromb Vasc Biol 1995, 15, 678–682. [Google Scholar]
- Vaughan, CJ; Gotto, AM, Jr. Update on statins: 2003. Circulation 2004, 110, 886–892. [Google Scholar]
- Shitara, Y; Sugiyama, Y. Pharmacokinetic and pharmacodynamic alterations of 3-hydroxy-3-methylglutaryl coenzyme A (HMG-CoA) reductase inhibitors: Drug-drug interactions and interindividual differences in transporter and metabolic enzyme functions. Pharmacol Ther 2006, 112, 71–105. [Google Scholar]
- Rodrigues, AC. Efflux and uptake transporters as determinants of statin response. Expert Opin Drug Metab Toxicol 2010, 6, 621–632. [Google Scholar]
- Nishizato, Y; Ieiri, I; Suzuki, H; Kimura, M; Kawabata, K; Hirota, T; Takane, H; Irie, S; Kusuhara, H; Urasaki, Y; Urae, A; Higuchi, S; Otsubo, K; Sugiyama, Y. Polymorphisms of OATP-C (SLC21A6) and OAT3 (SLC22A8) genes: Consequences for pravastatin pharmacokinetics. Clin Pharmacol Ther 2003, 73, 554–565. [Google Scholar]
- Niemi, M; Schaeffeler, E; Lang, T; Fromm, MF; Neuvonen, M; Kyrklund, C; Backman, JT; Kerb, R; Schwab, M; Neuvonen, PJ; Eichelbaum, M; Kivistö, KT. High plasma pravastatin concentrations are associated with single nucleotide polymorphisms and haplotypes of organic transporting polypeptide-C (OATP-C, SLCO1B1). Pharmacogenetics 2004, 14, 429–440. [Google Scholar]
- Chung, JY; Cho, JY; Yu, KS; Kim, JR; Oh, DS; Jung, HR; Lim, KS; Moon, KH; Shin, SG; Jang, IJ. Effect of OATP1B1 (SLCO1B1) variants on the pharmacokinetics of pitavastatin in healthy volunteers. Clin Pharmacol Ther 2005, 78, 342–350. [Google Scholar]
- Niemi, M; Pasanen, MK; Neuvonen, PJ. SLCO1B1 polymorphisms and sex affect the pharmacokinetics of pravastatin but not fluvastatin. Clin Pharmacol Ther 2006, 80, 356–366. [Google Scholar]
- Pasanen, MK; Fredrikson, H; Neuvonen, PJ; Niemi, M. Different effects of SLCO1B1 polymorphisms on the pharmacokinetics of atorvastatin and rosuvastatin. Clin Pharmacol Ther 2007, 82, 726–733. [Google Scholar]
- Ieiri, I; Suwannakul, S; Maeda, K; Uchimaru, H; Hashimoto, K; Kimura, M; Fujino, H; Hirano, M; Kusuhara, H; Irie, S; Higuchi, S; Sugiyama, Y. SLCO1B1 (OATP1B1, an uptake transporter) and ABCG2 (BCRP, an efflux transporter) variant alleles and pharmacokinetics of pitavastatin in healthy volunteers. Clin Pharmacol Ther 2007, 82, 541–547. [Google Scholar]
- Choi, JH; Lee, MG; Cho, JY; Lee, JE; Kim, KH; Park, K. Influence of OATP1B1 genotype on the pharmacokinetics of rosuvastatin in Koreans. Clin Pharmacol Ther 2008, 83, 251–257. [Google Scholar]
- Link, E; Parish, S; Armitage, J; Bowman, L; Heath, S; Matsuda, F; Gut, I; Lathrop, M; Collins, R. SLCO1B1 variants and statin-induced myopathy—A genomewide study. N Engl J Med 2008, 359, 789–799. [Google Scholar]
- Mwinyi, J; Johne, A; Bauer, S; Roots, I; Gerloff, T. Evidence for inverse effects of OATP-C (SLC21A6) *5 and *1b haplotypes on pravastatin kinetics. Clin Pharmacol Ther 2004, 75, 415–421. [Google Scholar]
- Takane, H; Miyata, M; Burioka, N; Shigemasa, C; Shimizu, E; Otsubo, K; Ieiri, I. Pharmacogenetic determinants of variability in lipid-lowering response to pravastatin therapy. J Hum Gene 2006, 51, 822–826. [Google Scholar]
- Couvert, P; Giral, P; Dejager, S; Gu, J; Huby, T; Chapman, MJ; Bruckert, E; Carrié, A. Association between a frequent allele of the gene encoding OATP1B1 and enhanced LDL-lowering response to fluvastatin therapy. Pharmacogenomics 2008, 9, 1217–1227. [Google Scholar]
- Knauer, MJ; Urquhart, BL; Meyer zu Schwabedissen, HE; Schwarz, UI; Lemke, CJ; Leake, BF; Kim, RB; Tirona, RG. Human skeletal muscle drug transporters determine local exposure and toxicity of statins. Circ Res 2010, 106, 297–306. [Google Scholar]
- Rebecchi, IM; Rodrigues, AC; Arazi, SS; Genvigir, FD; Willrich, MA; Hirata, MH; Soares, SA; Bertolami, MC; Faludi, AA; Bernik, MM; Dorea, EL; Dagli, ML; Avanzo, JL; Hirata, RD. ABCB1 and ABCC1 expression in peripheral mononuclear cells is influenced by gene polymorphisms and atorvastatin treatment. Biochem Pharmacol 2009, 77, 66–75. [Google Scholar]
- Pasanen, MK; Neuvonen, PJ; Niemi, M. Global analysis of genetic variation in SLCO1B1. Pharmacogenomics 2008, 9, 19–33. [Google Scholar]
- Mwinyi, J; Köpke, K; Schaefer, M; Roots, I; Gerloff, T. Comparison of SLCO1B1 sequence variability among German, Turkish, and African populations. Eur J Clin Pharmacol 2008, 64, 257–266. [Google Scholar]
- Xu, LY; He, YJ; Zhang, W; Deng, S; Li, Q; Zhang, WX; Liu, ZQ; Wang, D; Huang, YF; Zhou, HH; Sun, ZQ. Organic anion transporting polypeptide-1B1 haplotypes in Chinese patients. Acta Pharmacol Sin 2007, 28, 1693–1697. [Google Scholar]
- Pena, SD; Bastos-Rodrigues, L; Pimenta, JR; Bydlowski, SP. DNA tests probe the genomic ancestry of Brazilians. Braz J Med Biol Res 2009, 42, 870–876. [Google Scholar]
- Suarez-Kurtz, G; Perini, JA; Bastos-Rodrigues, L; Pena, SD; Struchiner, C. Impact of population admixture on the distribution of the CYP3A5*3 polymorphism. Pharmacogenomics 2007, 8, 1299–1306. [Google Scholar]
- Estrela, RC; Ribeiro, FS; Carvalho, RS; Gregório, SP; Dias-Neto, E; Struchiner, CJ; Suarez-Kurtz, G. Distribution of ABCB1 polymorphisms among Brazilians: Impact of population admixture. Pharmacogenomics 2008, 9, 267–276. [Google Scholar]
- Suarez-Kurtz, G; Amorim, A; Damasceno, A; Hutz, MH; de Moraes, MO; Ojopi, EB; Pena, SD; Perini, JA; Prata, MJ; Ribeiro-dos-Santos, A; Romano-Silva, MA; Teixeira, D; Struchiner, CJ. VKORC1 polymorphisms in Brazilians: Comparison with the Portuguese and Portuguese-speaking Africans and pharmacogenetic implications. Pharmacogenomics 2010, 11, 1257–1267. [Google Scholar]
- Niemi, M; Neuvonen, PJ; Hofmann, U; Backman, JT; Schwab, M; Lütjohann, D; von Bergmann, K; Eichelbaum, M; Kivistö, KT. Acute effects of pravastatin on cholesterol synthesis are associated with SLCO1B1 (encoding OATP1B1) haplotype *17. Pharmacogenet Genomics 2005, 15, 303–309. [Google Scholar]
- Igel, M; Arnold, KA; Niemi, M; Hofmann, U; Schwab, M; Lütjohann, D; von Bergmann, K; Eichelbaum, M; Kivistö, KT. Impact of the SLCO1B1 polymorphism on the pharmacokinetics and lipid-lowering efficacy of multiple-dose pravastatin. Clin Pharmacol Ther 2006, 79, 419–426. [Google Scholar]
- Gerloff, T; Schaefer, M; Mwinyi, J; Johne, A; Sudhop, T; Lütjohann, D; Roots, I; von Bergmann, K. Influence of the SLCO1B1*1b and *5 haplotypes on pravastatin’s cholesterol lowering capabilities and basal sterol serum levels. Naunyn Schmiedebergs Arch Pharmacol 2006, 373, 45–50. [Google Scholar]
- Hedman, M; Antikainen, M; Holmberg, C; Neuvonen, M; Eichelbaum, M; Kivistö, KT; Neuvonen, PJ; Niemi, M. Pharmacokinetics and response to pravastatin in paediatric patients with familial hypercholesterolaemia and in paediatric cardiac transplant recipients in relation to polymorphisms of the SLCO1B1 and ABCB1 genes. Br J Clin Pharmacol 2006, 61, 706–715. [Google Scholar]
- Zhang, W; Chen, BL; Ozdemir, V; He, YJ; Zhou, G; Peng, DD; Deng, S; Xie, QY; Xie, W; Xu, LY; Wang, LC; Fan, L; Wang, A; Zhou, HH. SLCO1B1 521>T functional genetic polymorphism and lipid-lowering efficacy of multiple-dose pravastatin, in Chinese coronary heart disease patients. Br J Clin Pharmacol 2007, 64, 346–352. [Google Scholar]
- Thompson, JF; Man, M; Johnson, KJ; Wood, LS; Lira, ME; Lloyd, DB; Banerjee, P; Milos, PM; Myrand, SP; Paulauskis, J; Milad, MA; Sasiela, WJ. An association study of 43 SNPs in 16 candidate genes with atorvastatin response. Pharmacogenomics J 2005, 5, 352–358. [Google Scholar]
- Wen, J; Xiong, Y. OATP1B1 388A>G polymorphism and pharmacokinetics of pitavastatin in Chinese healthy volunteers. J Clin Pharm Ther 2010, 35, 99–104. [Google Scholar]
- Iwai, M; Suzuki, H; Ieiri, I; Otsubo, K; Sugiyama, Y. Functional analysis of single nucleotide polymorphisms of hepatic organic anion transporter OATP1B1 (OATP-C). Pharmacogenetics 2004, 14, 749–757. [Google Scholar]
- Kameyama, Y; Yamashita, K; Kobayashi, K; Hosokawa, M; Chiba, K. Functional characterization of SLCO1B1 (OATP-C) variants, SLCO1B1*5, SLCO1B1*15 and SLCO1B1*15+C1007G, by using transient expression systems of HeLa and HEK293 cells. Pharmacogenet Genomics 2005, 15, 513–522. [Google Scholar]
- Lee, YJ; Lee, MG; Lim, LA; Jang, SB; Chung, JY. Effects of SLCO1B1 and ABCB1 genotypes on the pharmacokinetics of atorvastatin and 2-hydroxyatorvastatin in healthy Korean subjects. Int J Clin Pharmacol Ther 2010, 48, 36–45. [Google Scholar]
- National Cholesterol Education Program (NCEP). Third Report of the Expert Panel on Detection, Evaluation, and Treatment of High Blood Cholesterol in Adults (Adult Treatment Panel III), Final report. Circulation 2002, 106, 3143–3421.
- Salazar, LA; Hirata, MH; Cavalli, SA; Machado, MO; Hirata, RD. Optimized procedure for DNA isolation from fresh and cryopreserved clotted human blood useful in clinical molecular testing. Clin Chem 1998, 44, 1748–1750. [Google Scholar]
- Phillips, C; Salas, A; Sánchez, JJ; Fondevila, M; Gómez-Tato, A; Alvarez-Dios, J; Calaza, M; de Cal, MC; Ballard, D; Lareu, MV; Carracedo, A. SNPforID Consortium. Inferring ancestral origin using a single multiplex assay of ancestry-informative marker SNPs. Forensic Sci Int Genet 2007, 1, 273–280. [Google Scholar]
- Tirona, RG; Leake, BF; Merino, G; Kim, RB. Polymorphisms in OATP-C: Identification of multiple allelic variants associated with altered transport activity among European- and African-Americans. J Biol Chem 2001, 276, 35669–35675. [Google Scholar]
| Variables | Basal | Atorvastatin * | Change (%) | P |
|---|---|---|---|---|
| TC (mg/dL) | 281 ± 38 | 198 ± 30 | −28.9 ± 9.5 | <0.001 |
| LDL-C (mg/dL) | 193 ± 55 | 118 ± 27 | −38.3 ± 12.4 | <0.001 |
| HDL-C (mg/dL) | 56 ± 14 | 54 ± 13 | −2.5 ± 10.5 | <0.002 |
| TG (mg/dL) | 160 ± 66 | 132 ± 52 | −26.9 ± 52.5 | <0.001 |
| CK (U/L) | 102 ± 80 | 104 ± 88 | 4.9 ± 36.5 | 0.606 |
| ALT (U/L) | 22 ± 10 | 25 ± 15 | 23.0 ± 63.2 | <0.001 |
| ApoAI (g/L) | 130 ± 25 | 136 ± 27 | 4.9 ± 15.4 | 0.013 |
| ApoB (g/L) | 140 ± 22 | 102 ± 22 | −28 ± 46.1 | <0.001 |
| SNP | Basal | Atorvastatin | Change (%) | |||
|---|---|---|---|---|---|---|
| TC | LDL-C | TC | LDL-C | TC | LDL-C | |
| SLCO1B1 | ||||||
| c.521T>C | ||||||
| TT (108) | 281 ± 37 | 192 ± 34 | 199 ± 29 | 118 ± 26 | 28.7 ± 9.1 | 38.1 ± 12.4 |
| TC + CC (28) | 282 ± 35 | 193 ± 31 | 192 ± 32 | 114 ± 28 | 31.8 ± 9.3 | 40.9 ± 11.6 |
| P | 0.890 | 0.942 | 0.253 | 0.442 | 0.171 | 0.433 |
| c.388A>G | ||||||
| GG (49) | 279 ± 32 | 193 ± 36 | 193 ± 29 | 111 ± 25 | 30.6 ± 9.8 | 41.3 ± 12.4 |
| AA + AG (82) | 280 ± 40 | 191 ± 28 | 200 ± 31 | 121 ± 27 | 28.0 ± 9.2 | 36.6 ± 12.1 |
| P | 0.550 | 0.527 | 0.162 | 0.077 | 0.123 | 0.034 |
| c.463C>A | ||||||
| CC (95) | 283 ± 38 | 196 ± 35 | 199 ± 32 | 120 ± 27 | 29.4 ± 9.3 | 38.4 ± 11.5 |
| CA + AA (41) | 271 ± 31 | 184 ± 27 | 194 ± 27 | 113 ± 26 | 27.9 ± 9.9 | 38.0 ± 14.4 |
| P | 0.070 | 0.072 | 0.374 | 0.198 | 0.414 | 0.871 |
| SLCO2B1 | ||||||
| −71T>C | ||||||
| TT (42) | 281 ± 43 | 194 ± 37 | 200 ± 34 | 120 ± 28 | 28.6 ± 8.6 | 37.6 ± 10.6 |
| TC + CC (94) | 282 ± 35 | 198 ± 29 | 198 ± 29 | 116 ± 27 | 29.4 ± 9.4 | 39.0 ± 12.9 |
| P | 0.891 | 0.794 | 0.598 | 0.324 | 0.463 | 0.394 |
| Variables | p-value | Odds Ratio | 95% CI |
|---|---|---|---|
| Basal LDL cholesterol (≥208 mg/dL) | 0.012 | 3.47 | 1.32–9.14 |
| Age (<60 years) | 0.077 | 0.45 | 0.19–1.09 |
| SLCO1B1 c.388G allele (dominant) | 0.012 | 3.23 | 1.30–8.04 |
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Rodrigues, A.C.; Perin, P.M.S.; Purim, S.G.; Silbiger, V.N.; Genvigir, F.D.V.; Willrich, M.A.V.; Arazi, S.S.; Luchessi, A.D.; Hirata, M.H.; Bernik, M.M.S.; et al. Pharmacogenetics of OATP Transporters Reveals That SLCO1B1 c.388A>G Variant Is Determinant of Increased Atorvastatin Response. Int. J. Mol. Sci. 2011, 12, 5815-5827. https://doi.org/10.3390/ijms12095815
Rodrigues AC, Perin PMS, Purim SG, Silbiger VN, Genvigir FDV, Willrich MAV, Arazi SS, Luchessi AD, Hirata MH, Bernik MMS, et al. Pharmacogenetics of OATP Transporters Reveals That SLCO1B1 c.388A>G Variant Is Determinant of Increased Atorvastatin Response. International Journal of Molecular Sciences. 2011; 12(9):5815-5827. https://doi.org/10.3390/ijms12095815
Chicago/Turabian StyleRodrigues, Alice C., Paula M. S. Perin, Sheila G. Purim, Vivian N. Silbiger, Fabiana D. V. Genvigir, Maria Alice V. Willrich, Simone S. Arazi, Andre D. Luchessi, Mario H. Hirata, Marcia M. S. Bernik, and et al. 2011. "Pharmacogenetics of OATP Transporters Reveals That SLCO1B1 c.388A>G Variant Is Determinant of Increased Atorvastatin Response" International Journal of Molecular Sciences 12, no. 9: 5815-5827. https://doi.org/10.3390/ijms12095815
APA StyleRodrigues, A. C., Perin, P. M. S., Purim, S. G., Silbiger, V. N., Genvigir, F. D. V., Willrich, M. A. V., Arazi, S. S., Luchessi, A. D., Hirata, M. H., Bernik, M. M. S., Dorea, E. L., Santos, C., Faludi, A. A., Bertolami, M. C., Salas, A., Freire, A., Lareu, M. V., Phillips, C., Porras-Hurtado, L., ... Hirata, R. D. C. (2011). Pharmacogenetics of OATP Transporters Reveals That SLCO1B1 c.388A>G Variant Is Determinant of Increased Atorvastatin Response. International Journal of Molecular Sciences, 12(9), 5815-5827. https://doi.org/10.3390/ijms12095815







