Matrix-Assisted Laser Desorption/Ionization Imaging Mass Spectrometry
Abstract
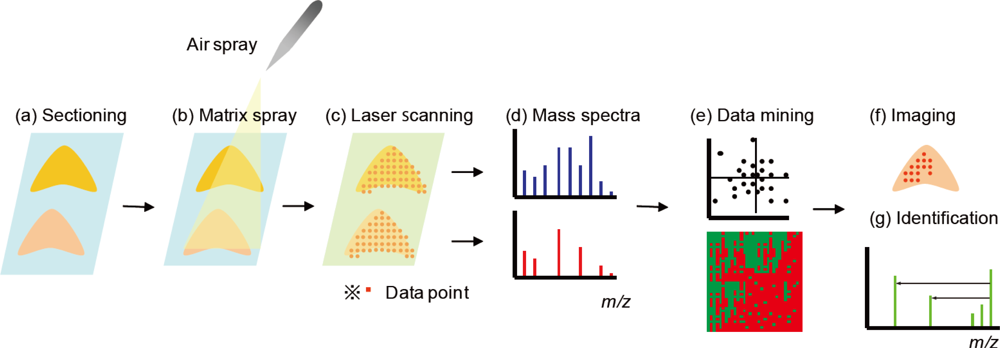
:1. Overview
2. Methodology
2.1. Biological Sample Preparation
2.1.1. Fixation and Embedding
2.1.2. Sectioning
2.1.3. Washing
2.2. Matrix Application
2.2.1. Choice of Matrix
2.2.2. Matrix Application Methods
2.3. Measurement and Data Analysis
2.3.1. Instruments
2.3.2. Measurement
2.3.3. Data Analysis
3. Applications of IMS
3.1. IMS for Proteins and Peptides
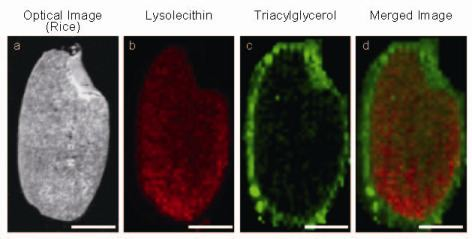
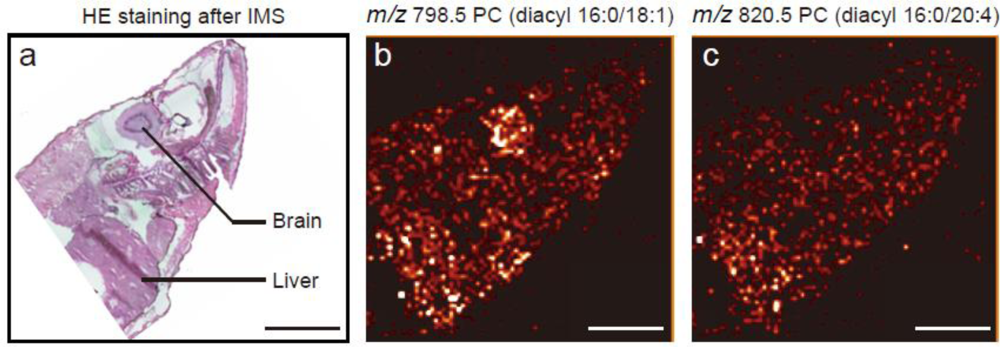
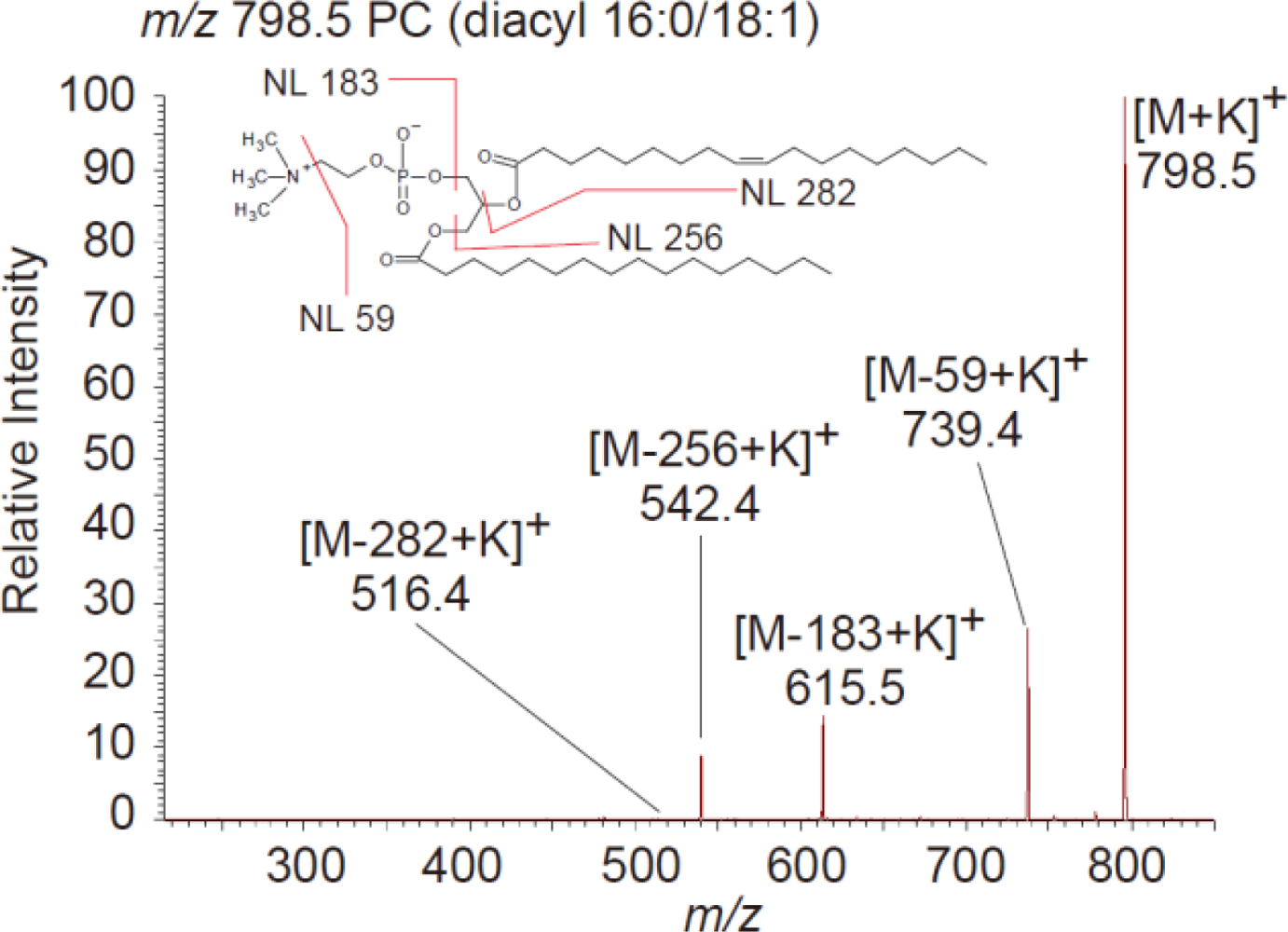
3.2. IMS for Lipids
3.3. IMS for Endogenous Metabolites
4. Future Perspectives
Acknowledgments
References
- Benninghoven, A; Sichtermann, WK. Detection, Identification and Structural Investigation of Biologically Important Compounds by Secondary Ion Mass Spectrometry. Anal. Chem 1978, 50, 1180–1184. [Google Scholar]
- Takats, Z; Wiseman, JM; Gologan, B; Cooks, RG. Mass Spectrometry Sampling under Ambient Conditions with Desorption Electrospray Ionization. Science 2004, 306, 471–473. [Google Scholar]
- Caprioli, RM; Farmer, TB; Gile, J. Molecular Imaging of Biological Samples: Localization of Peptides and Proteins Using Maldi-Tof Ms. Anal. Chem 1997, 69, 4751–4760. [Google Scholar]
- Karas, M; Hillenkamp, F. Laser Desorption Ionization of Proteins with Molecular Masses Exceeding 10,000 Daltons. Anal. Chem 1988, 60, 2299–2301. [Google Scholar]
- Tanaka, K; Waki, H; Ido, Y; Akita, S; Yoshida, Y; Yoshida, T. Protein and Polymer Analyses up to M/Z 100000 by Laser Ionization Time-of Flight Mass Spectrometry. Rapid Commun. Mass Spectrom 1988, 2, 151–153. [Google Scholar]
- Svatos, A. Mass Spectrometric Imaging of Small Molecules. Trends Biotechnol 2010, 28, 425–434. [Google Scholar]
- Yates, JR, 3rd. Mass Spectrometry and the Age of the Proteome. J. Mass Spectrom 1998, 33, 1–19. [Google Scholar]
- Yao, I; Sugiura, Y; Matsumoto, M; Setou, M. In Situ Proteomics with Imaging Mass Spectrometry and Principal Component Analysis in the Scrapper-Knockout Mouse Brain. Proteomics 2008, 8, 3692–3701. [Google Scholar]
- Groseclose, MR; Andersson, M; Hardesty, WM; Caprioli, RM. Identification of Proteins Directly from Tissue: In Situ Tryptic Digestions Coupled with Imaging Mass Spectrometry. J. Mass Spectrom 2007, 42, 254–262. [Google Scholar]
- Dani, FR; Francese, S; Mastrobuoni, G; Felicioli, A; Caputo, B; Simard, F; Pieraccini, G; Moneti, G; Coluzzi, M; della Torre, A; et al. Exploring Proteins in Anopheles Gambiae Male and Female Antennae through Maldi Mass Spectrometry Profiling. PLoS One 2008, 3, e2822. [Google Scholar]
- Murphy, RC; Hankin, JA; Barkley, RM. Imaging of Lipid Species by Maldi Mass Spectrometry. J Lipid Res 2009, 50(Suppl), S317–S322. [Google Scholar]
- Chan, K; Lanthier, P; Liu, X; Sandhu, JK; Stanimirovic, D; Li, J. Maldi Mass Spectrometry Imaging of Gangliosides in Mouse Brain Using Ionic Liquid Matrix. Anal. Chim. Acta 2009, 639, 57–61. [Google Scholar]
- Sugiura, Y; Shimma, S; Konishi, Y; Yamada, MK; Setou, M. Imaging Mass Spectrometry Technology and Application on Ganglioside Study; Visualization of Age-Dependent Accumulation of C20-Ganglioside Molecular Species in the Mouse Hippocampus. PLoS One 2008, 3, e3232. [Google Scholar]
- Goto-Inoue, N; Hayasaka, T; Sugiura, Y; Taki, T; Li, YT; Matsumoto, M; Setou, M. High-Sensitivity Analysis of Glycosphingolipids by Matrix-Assisted Laser Desorption/Ionization Quadrupole Ion Trap Time-of-Flight Imaging Mass Spectrometry on Transfer Membranes. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci 2008, 870, 74–83. [Google Scholar]
- Jackson, SN; Wang, HY; Woods, AS; Ugarov, M; Egan, T; Schultz, JA. Direct Tissue Analysis of Phospholipids in Rat Brain Using Maldi-Tofms and Maldi-Ion Mobility-Tofms. J. Am. Soc. Mass Spectrom 2005, 16, 133–138. [Google Scholar]
- Wang, HY; Jackson, SN; Woods, AS. Direct Maldi-Ms Analysis of Cardiolipin from Rat Organs Sections. J. Am. Soc. Mass Spectrom 2007, 18, 567–577. [Google Scholar]
- Jackson, SN; Woods, AS. Direct Profiling of Tissue Lipids by Maldi-Tofms. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci 2009, 877, 2822–2829. [Google Scholar]
- Goto-Inoue, N; Hayasaka, T; Taki, T; Gonzalez, TV; Setou, M. A New Lipidomics Approach by Thin-Layer Chromatography-Blot-Matrix-Assisted Laser Desorption/Ionization Imaging Mass Spectrometry for Analyzing Detailed Patterns of Phospholipid Molecular Species. J. Chromatogr. A 2009, 1216, 7096–7101. [Google Scholar]
- Sugiura, Y; Setou, M. Imaging Mass Spectrometry for Visualization of Drug and Endogenous Metabolite Distribution: Toward in Situ Pharmacometabolomes. J. Neuroimmun. Pharmacol 2010, 5, 31–43. [Google Scholar]
- Burrell, M; Earnshaw, C; Clench, M. Imaging Matrix Assisted Laser Desorption Ionization Mass Spectrometry: A Technique to Map Plant Metabolites within Tissues at High Spatial Resolution. J. Exp. Bot 2007, 58, 757–763. [Google Scholar]
- Goto-Inoue, N; Hayasaka, T; Zaima, N; Setou, M. The Specific Localization of Seminolipid Molecular Species on Mouse Testis During Testicular Maturation Revealed by Imaging Mass Spectrometry. Glycobiology 2009, 19, 950–957. [Google Scholar]
- Ageta, H; Asai, S; Sugiura, Y; Goto-Inoue, N; Zaima, N; Setou, M. Layer-Specific Sulfatide Localization in Rat Hippocampus Middle Molecular Layer Is Revealed by Nanoparticle-Assisted Laser Desorption/Ionization Imaging Mass Spectrometry. Med. Mol. Morphol 2009, 42, 16–23. [Google Scholar]
- Zaima, N; Hayasaka, T; Goto-Inoue, N; Setou, M. Imaging of Metabolites by Maldi Mass Spectrometry. J. Oleo Sci 2009, 58, 415–419. [Google Scholar]
- Stoeckli, M; Farmer, TB; Caprioli, RM. Automated Mass Spectrometry Imaging with a Matrix-Assisted Laser Desorption Ionization Time-of-Flight Instrument. J. Am. Soc. Mass Spectrom 1999, 10, 67–71. [Google Scholar]
- Stoeckli, M; Chaurand, P; Hallahan, DE; Caprioli, RM. Imaging Mass Spectrometry: A New Technology for the Analysis of Protein Expression in Mammalian Tissues. Nat. Med 2001, 7, 493–496. [Google Scholar]
- Todd, PJ; Schaaff, TG; Chaurand, P; Caprioli, RM. Organic Ion Imaging of Biological Tissue with Secondary Ion Mass Spectrometry and Matrix-Assisted Laser Desorption/Ionization. J. Mass Spectrom 2001, 36, 355–369. [Google Scholar]
- Hayasaka, T; Goto-Inoue, N; Sugiura, Y; Zaima, N; Nakanishi, H; Ohishi, K; Nakanishi, S; Naito, T; Taguchi, R; Setou, M. Matrix-Assisted Laser Desorption/Ionization Quadrupole Ion Trap Time-of-Flight (Maldi-Qit-Tof)-Based Imaging Mass Spectrometry Reveals a Layered Distribution of Phospholipid Molecular Species in the Mouse Retina. Rapid Commun. Mass Spectrom 2008, 22, 3415–3426. [Google Scholar]
- Shimma, S; Sugiura, Y; Hayasaka, T; Hoshikawa, Y; Noda, T; Setou, M. Maldi-Based Imaging Mass Spectrometry Revealed Abnormal Distribution of Phospholipids in Colon Cancer Liver Metastasis. J. Chromatogr. B Analyt. Technol. Biomed. Life Sci 2007, 855, 98–103. [Google Scholar]
- Kimura, Y; Tsutsumi, K; Sugiura, Y; Setou, M. Medical Molecular Morphology with Imaging Mass Spectrometry. Med. Mol. Morphol 2009, 42, 133–137. [Google Scholar]
- Chughtai, K; Heeren, RM. Mass Spectrometric Imaging for Biomedical Tissue Analysis. Chem. Rev 2010, 110, 3237–3277. [Google Scholar]
- Mange, A; Chaurand, P; Perrochia, H; Roger, P; Caprioli, RM; Solassol, J. Liquid Chromatography-Tandem and Maldi Imaging Mass Spectrometry Analyses of Rcl2/Cs100-Fixed, Paraffin-Embedded Tissues: Proteomics Evaluation of an Alternate Fixative for Biomarker Discovery. J. Proteome. Res 2009, 8, 5619–5628. [Google Scholar]
- Meistermann, H; Norris, JL; Aerni, HR; Cornett, DS; Friedlein, A; Erskine, AR; Augustin, A; De Vera Mudry, MC; Ruepp, S; Suter, L; et al. Biomarker Discovery by Imaging Mass Spectrometry: Transthyretin Is a Biomarker for Gentamicin-Induced Nephrotoxicity in Rat. Mol. Cell Proteomics 2006, 5, 1876–1886. [Google Scholar]
- Lemaire, R; Menguellet, SA; Stauber, J; Marchaudon, V; Lucot, JP; Collinet, P; Farine, MO; Vinatier, D; Day, R; Ducoroy, P; et al. Specific Maldi Imaging and Profiling for Biomarker Hunting and Validation: Fragment of the 11s Proteasome Activator Complex, Reg Alpha Fragment, Is a New Potential Ovary Cancer Biomarker. J. Proteome Res 2007, 6, 4127–4134. [Google Scholar]
- Jehl, B; Bauer, R; Dorge, A; Rick, R. The Use of Propane/Isopentane Mixtures for Rapid Freezing of Biological Specimens. J. Microsc 1981, 123, 307–309. [Google Scholar]
- Schwartz, SA; Reyzer, ML; Caprioli, RM. Direct Tissue Analysis Using Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry: Practical Aspects of Sample Preparation. J. Mass Spectrom 2003, 38, 699–708. [Google Scholar]
- Zaima, N; Goto-Inoue, N; Setou, M. Application of Imaging Mass Spectrometry for Analysis of Rice Oryza Sativa. Rapid Commun. Mass Spectrom 2010, 24, 2723–2927. [Google Scholar]
- Stoeckli, M; Staab, D; Schweitzer, A. Compound and Metabolite Distribution Measured by Maldi Mass Spectrometric Imaging in Whole-Body Tissue Sections. Int. J. Mass Spectrom 2006, 260, 195–202. [Google Scholar]
- Chen, R; Hui, L; Sturm, RM; Li, L. Three Dimensional Mapping of Neuropeptides and Lipids in Crustacean Brain by Mass Spectral Imaging. J. Am. Soc. Mass Spectrom 2009, 20, 1068–1077. [Google Scholar]
- Rohner, TC; Staab, D; Stoeckli, M. Maldi Mass Spectrometric Imaging of Biological Tissue Sections. Mech. Ageing Dev 2005, 126, 177–185. [Google Scholar]
- Djidja, MC; Francese, S; Loadman, PM; Sutton, CW; Scriven, P; Claude, E; Snel, MF; Franck, J; Salzet, M; Clench, MR. Detergent Addition to Tryptic Digests and Ion Mobility Separation Prior to Ms/Ms Improves Peptide Yield and Protein Identification for in Situ Proteomic Investigation of Frozen and Formalin-Fixed Paraffin-Embedded Adenocarcinoma Tissue Sections. Proteomics 2009, 9, 2750–2763. [Google Scholar]
- Sugiura, Y; Shimma, S; Setou, M. Thin Sectioning Improves the Peak Intensity and Signal-to-Noise Ratio in Direct Tissue Mass Spectrometry. J. Mass Spectrom. Soc. Jpn 2006, 54, 4. [Google Scholar]
- Goodwin, RJ; Pennington, SR; Pitt, AR. Protein and Peptides in Pictures: Imaging with Maldi Mass Spectrometry. Proteomics 2008, 8, 3785–3800. [Google Scholar]
- Kawamoto, T. Use of a New Adhesive Film for the Preparation of Multi-Purpose Fresh-Frozen Sections from Hard Tissues, Whole-Animals, Insects and Plants. Arch. Histol. Cytol 2003, 66, 123–143. [Google Scholar]
- Lemaire, R; Wisztorski, M; Desmons, A; Tabet, JC; Day, R; Salzet, M; Fournier, I. Maldi-Ms Direct Tissue Analysis of Proteins: Improving Signal Sensitivity Using Organic Treatments. Anal. Chem 2006, 78, 7145–7153. [Google Scholar]
- Aerni, HR; Cornett, DS; Caprioli, RM. Automated Acoustic Matrix Deposition for Maldi Sample Preparation. Anal. Chem 2006, 78, 827–834. [Google Scholar]
- Andersson, M; Groseclose, MR; Deutch, AY; Caprioli, RM. Imaging Mass Spectrometry of Proteins and Peptides: 3d Volume Reconstruction. Nature Methods 2008, 5, 101–108. [Google Scholar]
- Kaletas, BK; van der Wiel, IM; Stauber, J; Guzel, C; Kros, JM; Luider, TM; Heeren, RM. Sample Preparation Issues for Tissue Imaging by Imaging Ms. Proteomics 2009, 9, 2622–2633. [Google Scholar]
- McLean, JA; Stumpo, KA; Russell, DH. Size-Selected (2–10 nm) Gold Nanoparticles for Matrix Assisted Laser Desorption Ionization of Peptides. J. Am. Chem. Soc 2005, 127, 5304–5305. [Google Scholar]
- Su, CL; Tseng, WL. Gold Nanoparticles as Assisted Matrix for Determining Neutral Small Carbohydrates through Laser Desorption/Ionization Time-of-Flight Mass Spectrometry. Anal. Chem 2007, 79, 1626–1633. [Google Scholar]
- Hayasaka, T; Goto-Inoue, N; Zaima, N; Shrivas, K; Kashiwagi, Y; Yamamoto, M; Nakamoto, M; Setou, M. Imaging Mass Spectrometry with Silver Nanoparticles Reveals the Distribution of Fatty Acids in Mouse Retinal Sections. J. Am. Soc. Mass Spectrom 2010, 21, 1446–1454. [Google Scholar]
- Moritake, S; Taira, S; Sugiura, Y; Setou, M; Ichiyanagi, Y. Magnetic Nanoparticle-Based Mass Spectrometry for the Detection of Biomolecules in Cultured Cells. J. Nanosci. Nanotechnol 2009, 9, 169–176. [Google Scholar]
- Sugiura, Y; Setou, M. Matrix-Assisted Laser Desorption/Ionization and Nanoparticle-Based Imaging Mass Spectrometry for Small Metabolites: A Practical Protocol. Methods Mol. Biol 2010, 656, 173–195. [Google Scholar]
- Taira, S; Sugiura, Y; Moritake, S; Shimma, S; Ichiyanagi, Y; Setou, M. Nanoparticle-Assisted Laser Desorption/Ionization Based Mass Imaging with Cellular Resolution. Anal. Chem 2008, 80, 4761–4766. [Google Scholar]
- Goto-Inoue, N; Hayasaka, T; Zaima, N; Kashiwagi, Y; Yamamoto, M; Nakamoto, M; Setou, M. The Detection of Glycosphingolipids in Brain Tissue Sections by Imaging Mass Spectrometry Using Gold Nanoparticlesthe Detection of Glycosphingolipids in Brain Tissue Sections by Imaging Mass Spectrometry Using Gold Nanoparticles. J Am Soc Mass Spectrom 2010, in press.. [Google Scholar]
- Zaima, N; Matsuyama, Y; Setou, M. Principal Component Analysis of Direct Matrix-Assisted Laser Desorption/Ionization Mass Spectrometric Data Related to Metabolites of Fatty Liver. J. Oleo Sci 2009, 58, 267–273. [Google Scholar]
- Morita, Y; Ikegami, K; Goto-Inoue, N; Hayasaka, T; Zaima, N; Tanaka, H; Uehara, T; Setoguchi, T; Sakaguchi, T; Igarashi, H; et al. Imaging Mass Spectrometry of Gastric Carcinoma in Formalin-Fixed Paraffin-Embedded Tissue Microarray. Cancer Sci 2010, 101, 267–273. [Google Scholar]
- Cornett, DS; Reyzer, ML; Chaurand, P; Caprioli, RM. Maldi Imaging Mass Spectrometry: Molecular Snapshots of Biochemical Systems. Nature Methods 2007, 4, 828–833. [Google Scholar]
- Hankin, JA; Barkley, RM; Murphy, RC. Sublimation as a Method of Matrix Application for Mass Spectrometric Imaging. J. Am. Soc. Mass Spectrom 2007, 18, 1646–1652. [Google Scholar]
- Dekker, LJ; van Kampen, JJ; Reedijk, ML; Burgers, PC; Gruters, RA; Osterhaus, AD; Luider, TM. A Mass Spectrometry Based Imaging Method Developed for the Intracellular Detection of Hiv Protease Inhibitors. Rapid Commun. Mass Spectrom 2009, 23, 1183–1188. [Google Scholar]
- Wiseman, JM; Ifa, DR; Song, Q; Cooks, RG. Tissue Imaging at Atmospheric Pressure Using Desorption Electrospray Ionization (Desi) Mass Spectrometry. Angew. Chem. Int. Ed. Engl 2006, 45, 7188–7192. [Google Scholar]
- Landgraf, RR; Conaway, MCP; Garrett, TJ; Stacpoole, PW; Yost, RA. Imaging of Lipids in Spinal Cord Using Intermediate Pressure Matrix-Assisted Laser Desorption-Linear Ion Trap/Orbitrap Ms. Anal. Chem 2009, 81, 8488–8495. [Google Scholar]
- Hopfgartner, G; Varesio, E; Stoeckli, M. Matrix-Assisted Laser Desorption/Ionization Mass Spectrometric Imaging of Complete Rat Sections Using a Triple Quadrupole Linear Ion Trap. Rapid Commun. Mass Spectrom 2009, 23, 733–736. [Google Scholar]
- Taban, IM; Altelaar, AFM; van der Burgt, YEM; McDonnell, LA; Heeren, RMA; Fuchser, J; Baykut, G. Imaging of Peptides in the Rat Brain Using Maldi-Fticr Mass Spectrometry. J. Am. Soc. Mass Spectrom 2007, 18, 145–151. [Google Scholar]
- Harada, T; Yuba-Kubo, A; Sugiura, Y; Zaima, N; Hayasaka, T; Goto-Inoue, N; Wakui, M; Suematsu, M; Takeshita, K; Ogawa, K; Yoshida, Y; Setou, M. Visualization of Volatile Substances in Different Organelles with an Atmospheric-Pressure Mass Microscope. Anal. Chem 2009, 81, 9153–9157. [Google Scholar]
- Setou, M; Shrivas, K; Sroyraya, M; Yang, H; Sugiura, Y; Moribe, J; Kondo, A; Tsutsumi, K; Kimura, Y; Kurabe, N; et al. Developments and Applications of Mass Microscopy. Med. Mol. Morphol 2010, 43, 1–5. [Google Scholar]
- Zhang, X; Leung, SM; Morris, CR; Shigenaga, MK. Evaluation of a Novel, Integrated Approach Using Functionalized Magnetic Beads, Bench-Top Maldi-Tof-Ms with Prestructured Sample Supports, and Pattern Recognition Software for Profiling Potential Biomarkers in Human Plasma. J. Biomol. Tech 2004, 15, 167–175. [Google Scholar]
- Deininger, SO; Ebert, MP; Futterer, A; Gerhard, M; Rocken, C. Maldi Imaging Combined with Hierarchical Clustering as a New Tool for the Interpretation of Complex Human Cancers. J. Proteome Res 2008, 7, 5230–5236. [Google Scholar]
- Stoeckli, M; Staab, D; Staufenbiel, M; Wiederhold, KH; Signor, L. Molecular Imaging of Amyloid Beta Peptides in Mouse Brain Sections Using Mass Spectrometry. Anal. Biochem 2002, 311, 33–39. [Google Scholar]
- Yao, I; Takagi, H; Ageta, H; Kahyo, T; Sato, S; Hatanaka, K; Fukuda, Y; Chiba, T; Morone, N; Yuasa, S; Inokuchi, K; Ohtsuka, T; MacGregor, GR; Tanaka, K; Setou, M. Scrapper-Dependent Ubiquitination of Active Zone Protein Rim1 Regulates Synaptic Vesicle Release. Cell 2007, 130, 943–957. [Google Scholar]
- Skold, K; Svensson, M; Nilsson, A; Zhang, X; Nydahl, K; Caprioli, RM; Svenningsson, P; Andren, PE. Decreased Striatal Levels of Pep-19 Following Mptp Lesion in the Mouse. J. Proteome Res 2006, 5, 262–269. [Google Scholar]
- Pierson, J; Norris, JL; Aerni, HR; Svenningsson, P; Caprioli, RM; Andren, PE. Molecular Profiling of Experimental Parkinson’s Disease: Direct Analysis of Peptides and Proteins on Brain Tissue Sections by Maldi Mass Spectrometry. J. Proteome Res 2004, 3, 289–295. [Google Scholar]
- Herring, KD; Oppenheimer, SR; Caprioli, RM. Direct Tissue Analysis by Matrix-Assisted Laser Desorption Ionization Mass Spectrometry: Application to Kidney Biology. Semin. Nephrol 2007, 27, 597–608. [Google Scholar]
- Xu, BJ; Shyr, Y; Liang, X; Ma, LJ; Donnert, EM; Roberts, JD; Zhang, X; Kon, V; Brown, NJ; Caprioli, RM; et al. Proteomic Patterns and Prediction of Glomerulosclerosis and Its Mechanisms. J. Am. Soc. Nephrol 2005, 16, 2967–2975. [Google Scholar]
- Chen, Y; Allegood, J; Liu, Y; Wang, E; Cachon-Gonzalez, B; Cox, TM; Merrill, AH, Jr; Sullards, MC. Imaging Maldi Mass Spectrometry Using an Oscillating Capillary Nebulizer Matrix Coating System and Its Application to Analysis of Lipids in Brain from a Mouse Model of Tay-Sachs/Sandhoff Disease. Anal. Chem 2008, 80, 2780–2788. [Google Scholar]
- Rubakhin, SS; Li, L; Moroz, TP; Sweedler, JV. Characterization of the Aplysia Californica Cerebral Ganglion F Cluster. J. Neurophysiol 1999, 81, 1251–1260. [Google Scholar]
- Sugiura, Y; Konishi, Y; Zaima, N; Kajihara, S; Nakanishi, H; Taguchi, R; Setou, M. Visualization of the Cell-Selective Distribution of Pufa-Containing Phosphatidylcholines in Mouse Brain by Imaging Mass Spectrometry. J. Lipid Res 2009, 50, 1776–1788. [Google Scholar]
- Garrett, TJ; Prieto-Conaway, MC; Kovtoun, V; Bui, H; Izgarian, N; Stafford, G; Yost, RA. Imaging of Small Molecules in Tissue Sections with a New Intermediate-Pressure Maldi Linear Ion Trap Mass Spectrometer. Int. J. Mass Spectrom 2007, 260, 166–176. [Google Scholar]
- Koizumi, S; Yamamoto, S; Hayasaka, T; Konishi, Y; Yamaguchi-Okada, M; Goto-Inoue, N; Sugiura, Y; Setou, M; Namba, H. Imaging Mass Spectrometry Revealed the Production of Lyso-Phosphatidylcholine in the Injured Ischemic Rat Brain. Neuroscience 2010, 168, 219–225. [Google Scholar]
- Kobayashi, Y; Hayasaka, T; Setou, M; Itoh, H; Kanayama, N. Comparison of Phospholipid Molecular Species between Terminal and Stem Villi of Human Term Placenta by Imaging Mass Spectrometry. Placenta 2010, 31, 245–248. [Google Scholar]
- Goto-Inoue, N; Hayasaka, T; Setou, M. Imaging Mass Spectrometry of Glycolipids. Methods Enzymol 2010, 478, 287–301. [Google Scholar]
- Hayasaka, T; Goto-Inoue, N; Zaima, N; Kimura, Y; Setou, M. Organ-Specific Distributions of Lysophosphatidylcholine and Triacylglycerol in Mouse Embryo. Lipids 2009, 44, 837–848. [Google Scholar]
- Zaima, N; Goto-Inoue, N; Adachi, K; Setou, M. Selective Analysis of Lipids by Thin-Layer Chromatography Blot Matrix-Assisted Laser Desorption/Ionization Imaging Mass Spectrometry. J Oleo Sci 2010, in press.. [Google Scholar]
- Tanaka, H; Zaima, N; Yamamoto, N; Sagara, D; Suzuki, M; Nishiyama, M; Mano, Y; Sano, M; Hayasaka, H; Goto-Inoue, N; et al. Imaging Mass Spectrometry Reveals Unique Lipid Distribution in Primary Varicose Veins. Eur J Vasc Endovasc Surg 2010, in press.. [Google Scholar]
- Hirano, K; Ikeda, Y; Zaima, N; Sakata, Y; Matsumiya, G. Triglyceride Deposit Cardiomyovasculopathy. N. Engl. J. Med 2008, 359, 2396–2398. [Google Scholar]
- Benabdellah, F; Touboul, D; Brunelle, A; Laprevote, O. In Situ Primary Metabolites Localization on a Rat Brain Section by Chemical Mass Spectrometry Imaging. Anal. Chem 2009, 81, 5557–5560. [Google Scholar]
- Hattori, K; Kajimura, M; Hishiki, T; Nakanishi, T; Kubo, A; Nagahata, Y; Ohmura, M; Yachie-Kinoshita, A; Matsuura, T; Morikawa, T; et al. Paradoxical Atp Elevation in Ischemic Penumbra Revealed by Quantitative Imaging Mass Spectrometry. Antioxid Redox Signal 2010, 1157–1167. [Google Scholar]
- Goto-Inoue, N; Setou, M; Zaima, N. Visualization of Spatial Distribution of Gamma-Aminobutyric Acid in Eggplant (Solanum Melongena) by Matrix-Assisted Laser Desorption/Ionization Imaging Mass Spectrometry. Anal. Sci 2010, 26, 821–825. [Google Scholar]
- Taira, S; Ikeda, R; Yokota, N; Osaka, I; Sakamoto, M; Kato, M; Sahashi, Y. Mass Spectrometric Imaging of Ginsenosides Localization in Panax Ginseng Root. Am. J. Chin. Med 2010, 38, 485–493. [Google Scholar]
- McKay, K. New Techniques in the Pharmacokinetic Analysis of Cancer Drugs. Ii. The Ultratrace Determination of Platinum in Biological Samples by Inductively Coupled Plasma-Mass Spectrometry. Cancer Surv 1993, 17, 407–414. [Google Scholar]
- Yang, HJ; Ishizaki, I; Sanada, N; Zaima, N; Sugiura, Y; Yao, I; Ikegami, K; Setou, M. Detection of Characteristic Distributions of Phospholipid Head Groups and Fatty Acids on Neurite Surface by Time-of-Flight Secondary Ion Mass Spectrometry. Med. Mol. Morphol 2010, 43, 158–164. [Google Scholar]
- Slaveykova, VI; Guignard, C; Eybe, T; Migeon, HN; Hoffmann, L. Dynamic Nanosims Ion Imaging of Unicellular Freshwater Algae Exposed to Copper. Anal. Bioanal. Chem 2009, 393, 583–589. [Google Scholar]
- Finzi-Hart, JA; Pett-Ridge, J; Weber, PK; Popa, R; Fallon, SJ; Gunderson, T; Hutcheon, ID; Nealson, KH; Capone, DG. Fixation and Fate of C and N in the Cyanobacterium Trichodesmium Using Nanometer-Scale Secondary Ion Mass Spectrometry. Proc. Natl. Acad. Sci. USA 2009, 106, 9931–9931. [Google Scholar]
- Yang, HJ; Sugiura, Y; Ishizaki, I; Sanada, N; Ikegami, K; Zaima, N; Shrivas, K; Setou, M. Imaging of Lipids in Cultured Mammalian Neurons by Matrix Assisted Laser/Desorption Ionization and Secondary Ion Mass Spectrometry. Surf. Interf. Anal 2010, 42, 1606–1611. [Google Scholar]
- Wiseman, JM; Ifa, DR; Zhu, Y; Kissinger, CB; Manicke, NE; Kissinger, PT; Cooks, RG. Desorption Electrospray Ionization Mass Spectrometry: Imaging Drugs and Metabolites in Tissues. Proc. Natl. Acad. Sci. USA 2008, 105, 18120–18125. [Google Scholar]
- Ifa, DR; Manicke, NE; Dill, AL; Cooks, G. Latent Fingerprint Chemical Imaging by Mass Spectrometry. Science 2008, 321, 805–805. [Google Scholar]
- Becker, JS; Zoriy, M; Matusch, A; Wu, B; Salber, D; Palm, C; Becker, JS. Bioimaging of Metals by Laser Ablation Inductively Coupled Plasma Mass Spectrometry (La-Icp-Ms). Mass Spectrom. Rev 2010, 29, 156–175. [Google Scholar]
© 2010 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zaima, N.; Hayasaka, T.; Goto-Inoue, N.; Setou, M. Matrix-Assisted Laser Desorption/Ionization Imaging Mass Spectrometry. Int. J. Mol. Sci. 2010, 11, 5040-5055. https://doi.org/10.3390/ijms11125040
Zaima N, Hayasaka T, Goto-Inoue N, Setou M. Matrix-Assisted Laser Desorption/Ionization Imaging Mass Spectrometry. International Journal of Molecular Sciences. 2010; 11(12):5040-5055. https://doi.org/10.3390/ijms11125040
Chicago/Turabian StyleZaima, Nobuhiro, Takahiro Hayasaka, Naoko Goto-Inoue, and Mitsutoshi Setou. 2010. "Matrix-Assisted Laser Desorption/Ionization Imaging Mass Spectrometry" International Journal of Molecular Sciences 11, no. 12: 5040-5055. https://doi.org/10.3390/ijms11125040
APA StyleZaima, N., Hayasaka, T., Goto-Inoue, N., & Setou, M. (2010). Matrix-Assisted Laser Desorption/Ionization Imaging Mass Spectrometry. International Journal of Molecular Sciences, 11(12), 5040-5055. https://doi.org/10.3390/ijms11125040