Reticulate Evolution and Marine Organisms: The Final Frontier?
Abstract
:1. Introduction
2. Horizontal Transfer
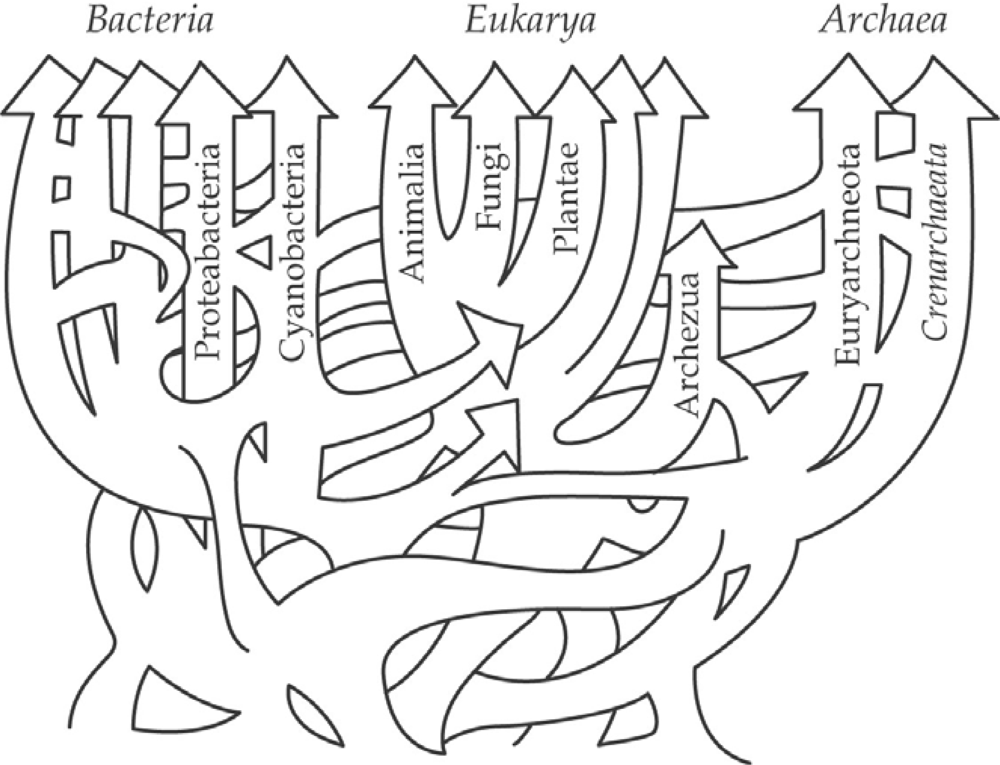
2.1. Horizontal Transfer and Adaptation in Marine Archaea, Bacteria and Cyanobacteria
2.2. Horizontal Transfer and the Evolution of the Marine Protist, Micromonas
2.3. Horizontal Transfer of Transposable Elements among Marine Invertebrates
3. Introgressive Hybridization and Hybrid Speciation
3.1. Introgressive Hybridization in Marine Angiosperms
3.2. Introgressive Hybridization and Hybrid Speciation in Corals
3.3. Introgressive Hybridization in Protists–The Diatom Genus Pseudo-nitzschia
3.4. Introgressive Hybridization in Crustacea
3.4.1. Genus Tetraclita (Acorn Barnacles)
3.4.2. Genus Mysis (Opossum Shrimp)
3.5. Introgressive Hybridization between Hydrothermal Vent Mussels of the Genus, Bathymodiolus
3.6. Introgressive Hybridization in Echinodermata
3.6.1. Sea Urchin Species
3.6.2. Genus Asterias (Sea Stars)
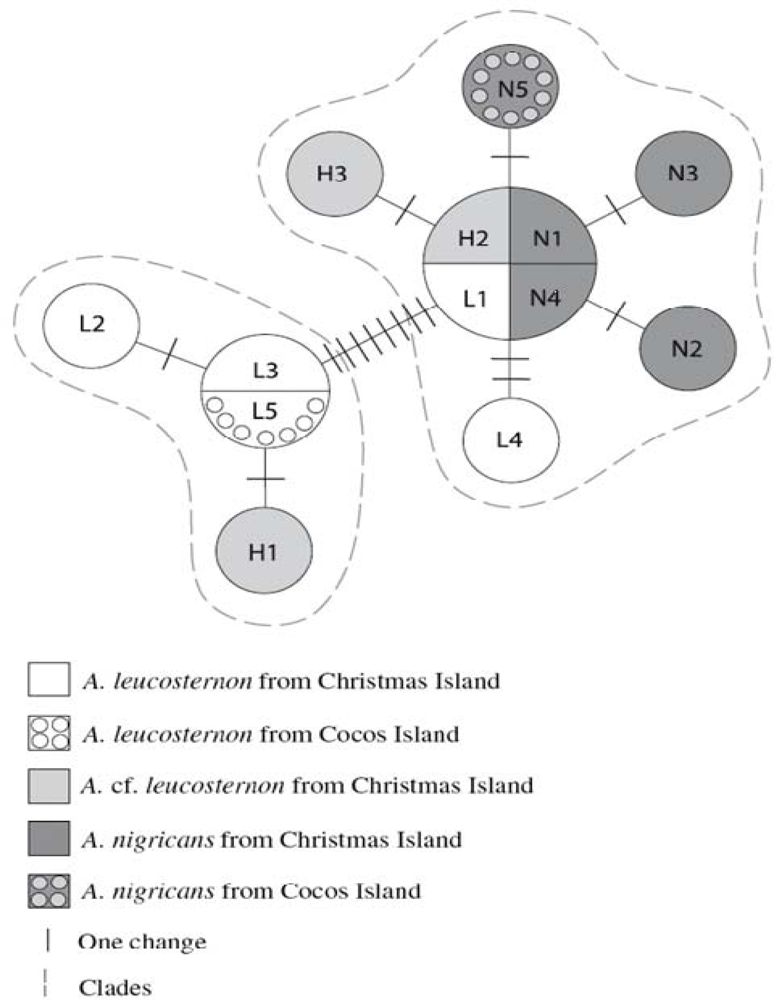
3.7. Introgressive Hybridization and Hybrid Speciation in Coral Reef Fishes Belonging to the Genera, Plectropomus and Acanthurus
3.8. Introgressive Hybridization in Marine Turtles
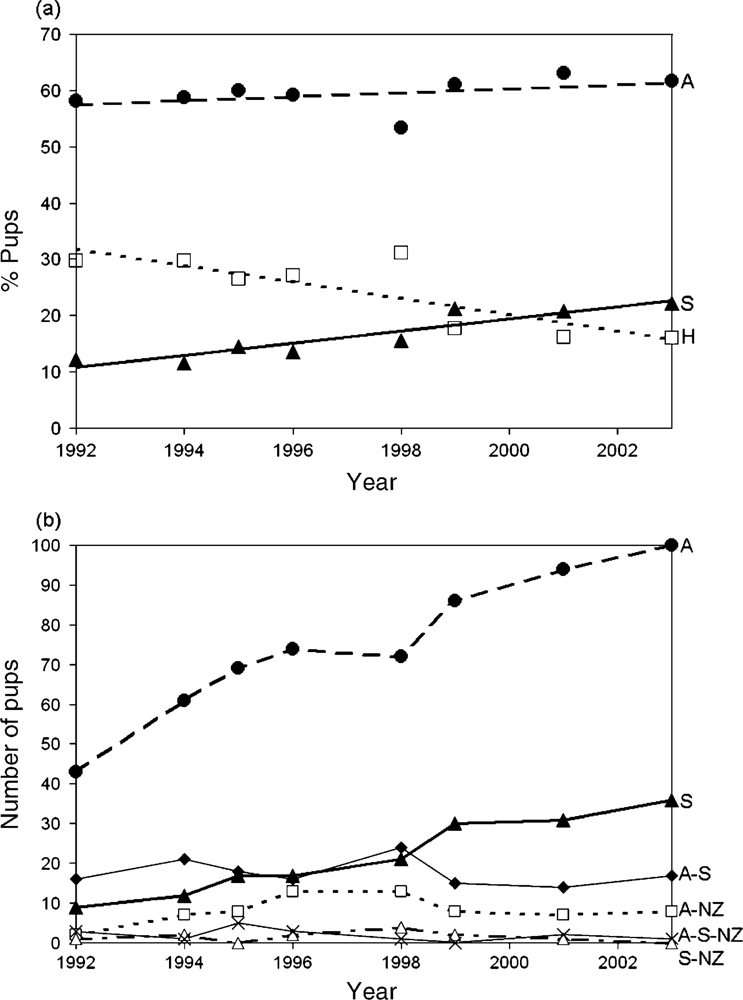
3.9. Introgressive Hybridization in Fur Seals
4. Conclusions and Future Directions
Acknowledgments
References
- Anderson, E. Introgressive Hybridization; John Wiley and Sons, Inc: New York, NY, USA, 1949. [Google Scholar]
- Anderson, E; Stebbins, GL, Jr. Hybridization as an evolutionary stimulus. Evolution 1954, 8, 378–388. [Google Scholar]
- Grant, V. Plant Speciation; Columbia University Press: New York, NY, USA, 1981. [Google Scholar]
- Dowling, TE; DeMarais, BD. Evolutionary significance of introgressive hybridization in cyprinid fishes. Nature 1993, 362, 444–446. [Google Scholar]
- Arnold, ML. Natural Hybridization and Evolution; Oxford University Press: Oxford, UK, 1997. [Google Scholar]
- Arnold, ML. Evolution through Genetic Exchange; Oxford University Press: Oxford, UK, 2006. [Google Scholar]
- Arnold, ML. Reticulate Evolution and Humans: Origins and Ecology; Oxford University Press: Oxford, UK, 2008. [Google Scholar]
- Doolittle, WF. Phylogenetic classification and the universal tree. Science 1999, 284, 2124–2128. [Google Scholar]
- Ochman, H; Lerat, E; Daubin, V. Examining bacterial species under the specter of gene transfer and exchange. Proc. Natl. Acad. Sci. USA 2005, 102, 6595–6599. [Google Scholar]
- Arnold, ML; Meyer, A. Natural hybridization in primates: One evolutionary mechanism. Zoology 2006, 109, 261–276. [Google Scholar]
- Lawrence, JG; Ochman, H. Molecular archaeology of the Escherichia coli genome. Proc. Natl. Acad. Sci. USA 1998, 95, 9413–9417. [Google Scholar]
- Dambroski, HR; Linn, C, Jr; Berlocher, SH; Forbes, AA; Roelofs, W; Feder, JL. The genetic basis for fruit odor discrimination in Rhagoletis flies and its significance for sympatric host shifts. Evolution 2005, 59, 1953–1964. [Google Scholar]
- Vana, G; Westover, KM. Origin of the 1918 Spanish influenza virus: A comparative genomic analysis. Mol. Phylogenet. Evol 2008, 47, 1100–1110. [Google Scholar]
- Hubbs, CL. Hybridization between fish species in nature. Syst. Zool 1955, 4, 1–20. [Google Scholar]
- Roques, S; Sévigny, J-M; Bernatchez, L. Evidence for broadscale introgressive hybridization between two redfish (genus Sebastes) in the North-west Atlantic: A rare marine example. Mol. Ecol 2001, 10, 149–165. [Google Scholar]
- Grant, PR; Grant, BR. Hybridization of bird species. Science 1992, 256, 193–197. [Google Scholar]
- Gardner, JPA. Hybridization in the sea. Adv. Mar. Biol 1997, 31, 1–78. [Google Scholar]
- Veron, JEN. Corals in Space and Time The Biogeography and Evolution of the Scleractinia; University of New South Wales Press: Sydney, Australia, 1995. [Google Scholar]
- Willis, BL; van Oppen, MJH; Miller, DJ; Vollmer, SV; Ayre, DJ. The role of hybridization in the evolution of reef corals. Ann. Rev. Ecol. Evol. Syst 2006, 37, 489–517. [Google Scholar]
- Ausich, WI; Meyer, DL. Hybrid crinoids in the fossil record (Early Mississippian, Phylum Echinodermata). Paleobiology 1994, 20, 362–367. [Google Scholar]
- Budd, AF; Pandolfi, JM. Overlapping species boundaries and hybridization within the Montastraea “annularis” reef coral complex in the Pleistocene of the Bahama Islands. Paleobiology 2004, 30, 396–425. [Google Scholar]
- Arnold, ML; Larson, EJ. Evolution’s new look. Wilson Quart 2004, 60–72. [Google Scholar]
- Doolittle, WF; Bapteste, E. Pattern pluralism and the Tree of Life hypothesis. Proc. Natl. Acad. Sci. USA 2007, 104, 2043–2049. [Google Scholar]
- DeLong, EF. Archaea in coastal marine environments. Proc. Natl. Acad. Sci. USA 1992, 89, 5685–5689. [Google Scholar]
- Frigaard, N-U; Martinez, A; Mincer, TJ; DeLong, EF. Proteorhodopsin lateral gene transfer between marine planktonic Bacteria and Archaea. Nature 2006, 439, 847–850. [Google Scholar]
- Béjà, O; Aravind, L; Koonin, EV; Suzuki, MT; Hadd, A; Nguyen, LP; Jovanovich, SB; Gates, CM; Feldman, RA; Spudich, JL; Spudich, EN; DeLong, EF. Bacterial rhodopsin: Evidence for a new type of phototrophy in the sea. Science 2000, 289, 1902–1906. [Google Scholar]
- Béjà, O; Spudich, EN; Spudich, JL; Leclerc, M; DeLong, EF. Proteorhodopsin phototrophy in the ocean. Nature 2001, 411, 786–789. [Google Scholar]
- Shi, T; Falkowski, PG. Genome evolution in cyanobacteria: The stable core and the variable shell. Proc. Natl. Acad. Sci. USA 2008, 105, 2510–2515. [Google Scholar]
- Swingley, WD; Chen, M; Cheung, PC; Conrad, AL; Dejesa, LC; Hao, JC; Honchak, BM; Karbach, LE; Kurdoglu, A; Lahiri, S; Mastrian, SD; Miyashita, H; Page, L; Ramakrishna, P; Satoh, S; Sattley, WM; Shimada, Y; Taylor, HL; Tomo, T; Tsuchiya, T; Wang, ZT; Raymond, J; Mimuro, M; Blankenship, RE; Touchman, JW. Niche adaptation and genome expansion in the chlorophyll d-producing cyanobacterium Acaryochloris marina. Proc. Natl. Acad. Sci. USA 2008, 105, 2005–2010. [Google Scholar]
- Worden, AZ; Lee, J-H; Mock, T; Rouzé, P; Simmons, MP; Aerts, AL; Allen, AE; Cuvelier, ML; Derelle, E; Everett, MV; Foulon, E; Grimwood, J; Gundlach, H; Henrissat, B; Napoli, C; McDonald, SM; Parker, MS; Rombauts, S; Salamov, A; von Dassow, P; Badger, JH; Coutinho, PM; Demir, E; Dubchak, I; Dubchak, C; Eikrem, W; Gready, JE; John, U; Lanier, W; Lindquist, EA; Lucas, S; Mayer, KFX; Moreau, H; Not, F; Otillar, R; Panaud, O; Pangilinan, J; Paulsen, I; Piegu, B; Poliakov, A; Robbens, S; Schmutz, J; Toulza, E; Wyss, T; Zelensky, A; Zhou, K; Armbrust, EV; Bhattacharya, D; Goodenough, UW; van de Peer, Y; Grigoriev, IV. Green evolution and dynamic adaptations revealed by genomes of the marine picoeukaryotes Micromonas. Science 2009, 324, 268–272. [Google Scholar]
- Kidwell, MG. Lateral transfer in natural populations of eukaryotes. Ann. Rev. Genet 1993, 27, 235–256. [Google Scholar]
- Casse, N; Bui, QT; Nicolas, V; Renault, S; Bigot, Y; Laulier, M. Species sympatry and horizontal transfers of Mariner transposons in marine crustacean genomes. Mol. Phylogenet. Evol 2006, 40, 609–619. [Google Scholar]
- Reusch, TBH. Microsatellites reveal high population connectivity in eelgrass (Zostera marina) in two contrasting coastal areas. Limnol. Oceanogr 2002, 47, 78–85. [Google Scholar]
- Coyer, JA; Miller, KA; Engle, JM; Veldsink, J; Cabello-Pasini, A; Stam, WT; Olsen, JL. Eelgrass meadows in the California Channel Islands and adjacent coast reveal a mosaic of two species, evidence for introgression and variable clonality. Ann. Bot 2008, 101, 73–87. [Google Scholar]
- Hatta, M; Fukami, H; Wang, W; Omori, M; Shimoike, K; Hayashibara, T; Ina, Y; Sugiyama, T. Reproductive and genetic evidence for a reticulate evolutionary history of mass-spawning corals. Mol. Biol. Evol 1999, 16, 1607–1613. [Google Scholar]
- Knowlton, N; Weil, E; Wright, LA; Guzman, HM. Sibling species in Montastraea annularis, coral bleaching, and the coral climate record. Science 1992, 255, 330–333. [Google Scholar]
- Levitan, DR; Fukami, H; Jara, J; Kline, D; McGovern, TM; McGhee, KE; Swanson, CA; Knowlton, N. Mechanisms of reproductive isolation among sympatric broadcast-spawning corals of the Montastraea annularis species complex. Evolution 2004, 58, 308–323. [Google Scholar]
- Fukami, H; Budd, AF; Levitan, DR; Jara, J; Kersanach, R; Knowlton, N. Geographic differences in species boundaries among members of the Montastraea annularis complex based on molecular and morphological markers. Evolution 2004, 58, 324–337. [Google Scholar]
- Huang, D; Meier, R; Todd, PA; Chou, LM. More evidence for pervasive paraphyly in scleractinian corals: Systematic study of Southeast Asian Faviidae (Cnidaria; Scleractinia) based on molecular and morphological data. Mol. Phylogenet. Evol 2009, 50, 102–116. [Google Scholar]
- McFadden, CS; Hutchinson, MB. Molecular evidence for the hybrid origin of species in the soft coral genus Alcyonium (Cnidaria: Anthozoa: Octocorallia). Mol. Ecol 2004, 13, 1495–1505. [Google Scholar]
- Combosch, DJ; Guzman, HM; Schuhmacher, H; Vollmer, SV. Interspecific hybridization and restricted trans-Pacific gene flow in the tropical eastern Pacific. Pocillopora. Mol. Ecol 2008, 17, 1304–1312. [Google Scholar]
- Glynn, PW; Gassman, NJ; Eakin, CM; Cortes, J; Smith, DB; Guzman, HM. Reef coral reproduction in the eastern Pacific: Costa Rica, Panama, and Galapagos Islands (Ecuador) I. Pocilloporiidae. Mar. Biol 1991, 109, 355–368. [Google Scholar]
- Bates, SS. Domoic-Acid-producing diatoms: Another genus added! J. Phycol 2000, 36, 978–985. [Google Scholar]
- Hasle, GR. Are most domoic acid-producing species of the diatom genus Pseudo-nitzschia cosmopolites? Harmful Algae 2002, 1, 137–146. [Google Scholar]
- D’Alelio, D; Amato, A; Kooistra, WHCF; Procaccini, G; Casotti, R; Montresor, M. Internal transcribed spacer polymorphism in Pseudo-nitzschia multistriata (Bacillariophyceae) in the Gulf of Naples: Recent divergence or intraspecific hybridization? Protist 2009, 160, 9–20. [Google Scholar]
- Casteleyn, G; Adams, NG; Vanormelingen, P; Debeer, A-E; Sabbe, K; Vyverman, W. Natural hybrids in the marine diatom Pseudo-nitzschia pungens (Bacillariophyceae): Genetic and morphological evidence. Protist 2009, 160, 343–354. [Google Scholar]
- Tsang, LM; Chan, BKK; Ma, KY; Chu, KH. Genetic differentiation, hybridization and adaptive divergence in two subspecies of the acorn barnacle Tetraclita japonica in the northwestern Pacific. Mol. Ecol 2008, 17, 4151–4163. [Google Scholar]
- Audzijonyte, A; Damgaard, J; Varvio, S-L; Vainio, JK; Väinölä, R. Phylogeny of Mysis (Crustacea, Mysida): History of continental invasions inferred from molecular and morphological data. Cladistics 2005, 21, 575–596. [Google Scholar]
- Vainio, JK; Väinölä, R. Refugial races and postglacial colonization history of the freshwater amphipod Gammarus lacustris in northern Europe. Biol. J. Linn. Soc 2003, 79, 523–542. [Google Scholar]
- Audzijonyte, A; Väinölä, R. Mysis nordenskioldi n. sp. (Crustacaea, Mysida), a circumpolar coastal mysid separated from the NE Pacific M. litoralis (Banner, 1948). Polar Biol 2007, 30, 1137–1157. [Google Scholar]
- O’Mullan, GD; Maas, PAY; Lutz, RA; Vrijenhoek, RC. A hybrid zone between hydrothermal vent mussels (Bivalvia: Mytilidae) from the Mid-Atlantic Ridge. Mol. Ecol 2001, 10, 2819–2831. [Google Scholar]
- Won, Y; Hallam, SJ; O’Mullan, GD; Vrijenhoek, RC. Cytonuclear disequilibrium in a hybrid zone involving deep-sea hydrothermal vent mussels of the genus Bathymodiolus. Mol. Ecol 2003, 12, 3185–3190. [Google Scholar]
- Levitan, DR. The relationship between conspecific fertilization success and reproductive isolation among three congeneric sea urchins. Evolution 2002, 56, 1599–1609. [Google Scholar]
- Harper, FM; Addison, JA; Hart, MW. Introgression versus immigration in hybridizing high-dispersal echinoderms. Evolution 2007, 61, 2410–2418. [Google Scholar]
- Addison, JA; Hart, MW. Colonization, dispersal, and hybridization influence phylogeography of north Atlantic sea urchins (Strongylocentrotus droebachiensis). Evolution 2005, 59, 532–543. [Google Scholar]
- Lessios, HA; Pearse, JS. Hybridization and introgression between Indo-Pacific species of Diadema. Mar. Biol 1996, 126, 715–723. [Google Scholar]
- Zigler, KS; Lessios, HA. Speciation on the coasts of the New World: Phylogeography and the evolution of bindin in the sea urchin genus Lytechinus. Evolution 2004, 58, 1225–1241. [Google Scholar]
- Harper, FM; Hart, MW. Morphological and phylogenetic evidence for hybridization and introgression in a sea star secondary contact zone. Invert. Biol 2007, 126, 373–384. [Google Scholar]
- Scheibling, RE; Lauzon-Guay, J-S. Feeding aggregations of sea stars (Asterias spp. and Henricia sanguinolenta) associated with sea urchin (Strongylocentrotus droebachiensis) grazing fronts in Nova Scotia. Mar. Biol 2007, 151, 1175–1183. [Google Scholar]
- van Herwerden, L; Choat, JH; Dudgeon, CL; Carlos, G; Newman, SJ; Frisch, A; van Oppen, M. Contrasting patterns of genetic structure in two species of the coral trout Plectropomus (Serranidae) from east and west Australia: Introgressive hybridisation or ancestral polymorphisms. Mol. Phylogenet. Evol 2006, 41, 420–435. [Google Scholar]
- Marie, AD; van Herwerden, L; Choat, JH; Hobbs, J-PA. Hybridization of reef fishes at the Indo-Pacific biogeographic barrier: A case study. Coral Reefs 2007, 26, 841–850. [Google Scholar]
- Karl, SA; Bowen, BW; Avise, JC. Hybridization among the ancient mariners: Characterization of marine turtle hybrids with molecular genetic assays. J. Hered 1995, 86, 262–268. [Google Scholar]
- Bass, AL; Good, DA; Bjorndal, KA; Richardson, JI; Hillis, Z-M; Horrocks, JA; Bowen, BW. Testing models of female reproductive migratory behaviour and population structure in the Caribbean hawksbill turtle, Eretmochelys imbricata, with mtDNA sequences. Mol. Ecol 1996, 5, 321–328. [Google Scholar]
- Lara-Ruiz, P; Lopez, GG; Santos, FR; Soares, LS. Extensive hybridization in hawksbill turtles (Eretmochelys imbricata) nesting in Brazil revealed by mtDNA analyses. Conserv. Genet 2006, 7, 773–781. [Google Scholar]
- Lancaster, ML; Gemmel, NJ; Negro, S; Goldsworthy, S; Sunnucks, P. Ménage à trois on Macquarie Island: Hybridization among three species of fur seal (Arctocephalus spp.) following historical population extinction. Mol. Ecol 2006, 15, 3681–3692. [Google Scholar]
- Kingston, JJ; Gwilliam, J. Hybridization between two sympatrically breeding species of fur seal at Iles Crozet revealed by genetic analysis. Conserv. Genet 2007, 8, 1133–1145. [Google Scholar]
- Lancaster, ML; Bradshaw, CJA; Goldsworthy, S; Sunnucks, P. Lower reproductive success in hybrid fur seal males indicates fitness costs to hybridization. Mol. Ecol 2007, 16, 3187–3197. [Google Scholar]
- Lancaster, ML; Goldsworthy, S; Sunnucks, P. Multiple mating strategies explain unexpected genetic mixing of New Zealand fur seals with two congenerics in a recently recolonized population. Mol. Ecol 2007, 16, 5267–5276. [Google Scholar]
- Dufresne, A; Salanoubat, M; Partensky, F; Artiguenave, F; Axmann, IM; Barbe, V; Duprat, S; Galperin, MY; Koonin, EV; Le Gall, F; Makarova, KS; Ostrowski, M; Oztas, S; Robert, C; Rogozin, IB; Scanlan, DJ; de Marsac, NT; Weissenbach, J; Wincker, P; Wolf, YI; Hess, WR. Genome sequence of the cyanobacterium Prochlorococcus marinus SS120, a nearly minimal oxyphototrophic genome. Proc. Natl. Acad. Sci. USA 2003, 100, 10020–10025. [Google Scholar]
- Palenik, B; Brahamsha, B; Larimer, FW; Land, M; Hauser, L; Chain, P; Lamerdin, J; Regala, W; Allen, EE; McCarren, J; Paulsen, I; Dufresne, A; Partensky, F; Webb, EA; Waterbury, J. The genome of a motile marine Synechococcus. Nature 2003, 424, 1037–1042. [Google Scholar]
- Rocap, G; Larimer, FW; Lamerdin, J; Malfatti, S; Chain, P; Ahlgren, NA; Arellano, A; Coleman, M; Hauser, L; Hess, WR; Johnson, ZI; Land, M; Lindell, D; Post, AF; Regala, W; Shah, M; Shaw, SL; Steglich, C; Sullivan, MB; Ting, CS; Tolonen, A; Webb, EA; Zinser, ER; Chisholm, SW. Genome divergence in two Prochlorococcus ecotypes reflects oceanic niche differentiation. Nature 2003, 424, 1042–1047. [Google Scholar]
- Lindell, D; Sullivan, MB; Johnson, ZI; Tolonen, AC; Rohwer, F; Chisholm, SW. Transfer of photosynthesis genes to and from Prochlorococcus viruses. Proc. Natl. Acad. Sci. USA 2004, 101, 11013–11018. [Google Scholar]
- Millard, A; Clokie, MRJ; Shub, DA; Mann, NH. Genetic organization of the psbAD region in phages infecting marine Synechococcus strains. Proc. Natl. Acad. Sci. USA 2004, 101, 11007–11012. [Google Scholar]
- Filée, J; Tétart, F; Suttle, CA; Krisch, HM. Marine T4-type bacteriophages, a ubiquitous component of the dark matter of the biosphere. Proc. Natl. Acad. Sci. USA 2005, 102, 12471–12476. [Google Scholar]
- Daehler, CC; Strong, DR. Hybridization between introduced smooth cordgrass (Spartina alterniflora; Poaceae) and native California cordgrass (S. foliosa) in San Francisco Bay, California, USA. Amer. J. Bot 1997, 84, 607–611. [Google Scholar]
- Anttila, CK; King, RA; Ferris, C; Ayres, DR; Strong, DR. Reciprocal hybrid formation of Spartina in San Francisco Bay. Mol. Ecol 2000, 9, 765–770. [Google Scholar]
- Baumel, A; Ainouche, ML; Bayer, RJ; Ainouche, AK; Misset, MT. Molecular phylogeny of hybridizing species from the genus Spartina Schreb. (Poaceae). Mol. Phylogenet. Evol 2002, 22, 303–314. [Google Scholar]
- Baumel, A; Ainouche, ML; Misset, MT; Gourret, J-P; Bayer, RJ. Genetic evidence for hybridization between the native Spartina maritima and the introduced Spartina alterniflora (Poaceae) in south-west France: Spartina x neyrautii re-examined. Plant Syst. Evol 2003, 237, 87–97. [Google Scholar]
- Ayres, DR; Grotkopp, E; Zaremba, K; Sloop, CM; Blum, MJ; Bailey, JP; Anttila, CK; Strong, DR. Hybridization between invasive Spartina densiflora (Poaceae) and native S. foliosa in San Francisco Bay, California, USA. Amer. J. Bot 2008, 95, 713–719. [Google Scholar]
- Coyer, JA; Peters, AF; Hoarau, G; Stam, WT; Olsen, JL. Hybridization of the marine seaweeds, Fucus serratus and Fucus evanescens (Heterokontophyta: Phaeophyceae) in a 100-year-old zone of secondary contact. Proc. R. Soc. Lond. B 2002, 269, 1829–1834. [Google Scholar]
- Wallace, AL; Klein, AS; Mathieson, AC. Determining the affinities of salt marsh fucoids using microsatellite markers: Evidence of hybridization and introgression between two species of Fucus (Phaeophyta) in a marine estuary. J. Phycol 2004, 40, 1013–1027. [Google Scholar]
- Coyer, JA; Hoarau, G; Oudot-Le Secq, M-P; Stam, WT; Olsen, JL. A mtDNA-based phylogeny of the brown algal genus Fucus (Heterokontophyta: Phaeophyta). Mol. Phylogenet. Evol 2006, 39, 209–222. [Google Scholar]
- Coyer, JA; Hoarau, G; Pearson, GA; Serrão, EA; Stam, WT; Olsen, JL. Convergent adaptation to a marginal habitat by homoploid hybrids and polyploidy ecads in the seaweed genus Fucus. Biol. Let 2006, 2, 405–408. [Google Scholar]
- Coyer, JA; Hoarau, G; Stam, WT; Olsen, JL. Hybridization and introgression in a mixed population of the intertidal seaweeds Fucus evanescens and F. serratus. J. Evol. Biol 2007, 20, 2322–2333. [Google Scholar]
- Bert, TM. Speciation in western Atlantic stone crabs (genus Menippe): The role of geological processes and climatic events in the formation and distribution of species. Mar. Biol 1986, 93, 157–170. [Google Scholar]
- Bert, TM; Harrison, RG. Hybridization in western Atlantic stone crabs (genus Menippe): Evolutionary history and ecological context influence species interactions. Evolution 1988, 42, 528–544. [Google Scholar]
- Bert, TM; McCarthy, KJ; Cruz-Lopez, H; Bogdanowicz, S. Character discriminatory power, character-set congruence, and the classification of individuals from hybrid zones: An example using stone crabs (Menippe). Evolution 1996, 50, 655–671. [Google Scholar]
- Muths, D; Davoult, D; Gentil, F; Jollivet, D. Incomplete cryptic speciation between intertidal and subtidal morphs of Acrocnida brachiata (Echinodermata: Ophiuroidea) in the northeast Atlantic. Mol. Ecol 2006, 15, 3303–3318. [Google Scholar]
- Wallace, CC; Willis, BL. Systematics of the coral genus Acropora: Implications of new biological findings for species concepts. Ann. Rev. Ecol. Syst 1994, 25, 237–262. [Google Scholar]
- van Oppen, MJH; McDonald, BJ; Willis, B; Miller, DJ. The evolutionary history of the coral genus Acropora (Scleractinia, Cnidaria) based on a mitochondrial and a nuclear marker: Reticulation, incomplete lineage sorting, or morphological convergence? Mol. Biol. Evol 2001, 18, 1315–1329. [Google Scholar]
- Márquez, LM; van Oppen, MJH; Willis, BL; Reyes, A; Miller, DJ. The highly cross-fertile coral species, Acropora hyacinthus and Acropora cytherea, constitute statistically distinguishable lineages. Mol. Ecol 2002, 11, 1339–1349. [Google Scholar]
- van Oppen, MJH; Willis, BL; van Rheede, T; Miller, DJ. Spawning times, reproductive compatibilities and genetic structuring in the Acropora aspera group: Evidence for natural hybridization and semi-permeable species boundaries in corals. Mol. Ecol 2002, 11, 1363–1376. [Google Scholar]
- Vollmer, SV; Palumbi, SR. Hybridization and the evolution of reef coral diversity. Science 2002, 296, 2023–2025. [Google Scholar]
- Vollmer, SV; Palumbi, SR. Restricted gene flow in the Caribbean staghorn coral Acropora cervicornis: Implications for the recovery of endangered reefs. J. Hered 2007, 98, 40–50. [Google Scholar]
- Rolán-Alvarez, E; Johannesson, K; Erlandsson, J. The maintenance of a cline in the marine snail Littorina saxtilis: The role of home site advantage and hybrid fitness. Evolution 1997, 51, 1838–1847. [Google Scholar]
- Cruz, R; Vilas, C; Mosquera, J; García, C. The close relationship between estimated divergent selection and observed differentiation supports the selective origin of a marine snail hybrid zone. J. Evol. Biol 2004, 17, 1221–1229. [Google Scholar]
- Bert, TM; Hesselman, DM; Arnold, WS; Moore, WS; Cruz-Lopez, H; Marelli, DC. High frequency of gonadal neoplasia in a hard clam (Mercenaria spp.) hybrid zone. Mar. Biol 1993, 117, 97–104. [Google Scholar]
- Bert, TM; Arnold, WS. An empirical test of predictions of two competing models for the maintenance and fate of hybrid zones: Both models are supported in a hard-clam hybrid zone. Evolution 1995, 49, 276–289. [Google Scholar]
- Strelkov, P; Nikula, R; Väinölä, R. Macoma balthica in the White and Barents Seas: Properties of a widespread marine hybrid swarm (Mollusca: Bivalvia). Mol. Ecol 2007, 16, 4110–4127. [Google Scholar]
- Riginos, C; Cunningham, CW. Hybridization in postglacial marine habitats. Mol. Ecol 2007, 16, 3971–3972. [Google Scholar]
- Hare, MP; Avise, JC. Molecular genetic analysis of a stepped multilocus cline in the American oyster (Crassostrea virginica). Evolution 1996, 50, 2305–2315. [Google Scholar]
- Murray, MC; Hare, MP. A genomic scan for divergent selection in a secondary contact zone between Atlantic and Gulf of Mexico oysters, Crassostrea virginica. Mol. Ecol 2006, 15, 4229–4242. [Google Scholar]
- Skibinski, DOF; Ahmad, M; Beardmore, JA. Genetic evidence for naturally occurring hybrids between Mytilus edulis and Mytilus galloprovincialis. Evolution 1978, 32, 354–364. [Google Scholar]
- Rawson, PD; Hilbish, TJ. Asymmetric introgression of mitochondrial DNA among European populations of blue mussels (Mytilus spp.). Evolution 1998, 52, 100–108. [Google Scholar]
- Daguin, C; Bonhomme, F; Borsa, P. The zone of sympatry and hybridization of Mytilus edulis and M. galloprovincialis, as described by intron length polymorphism at locus mac-1. Heredity 2001, 86, 342–354. [Google Scholar]
- Riginos, C; Sukhdeo, K; Cunningham, CW. Evidence for selection at multiple allozyme loci across a mussel hybrid zone. Mol. Biol. Evol 2002, 19, 347–351. [Google Scholar]
- Bierne, N; Borsa, P; Daguin, C; Jollivet, D; Viard, F; Bonhomme, F; David, P. Introgression patterns in the mosaic hybrid zone between Mytilus edulis and M. galloprovincialis. Mol. Ecol 2003, 12, 447–461. [Google Scholar]
- Gilg, MR; Hilbish, TJ. Patterns of larval dispersal and their effect on the maintenance of a blue mussel hybrid zone in southwestern England. Evolution 2003, 57, 1061–1077. [Google Scholar]
- Riginos, C; Hickerson, MJ; Henzler, CM; Cunningham, CW. Differential patterns of male and female mtDNA exchange across the Atlantic Ocean in the blue mussel, Mytilus edulis. Evolution 2004, 58, 2438–2451. [Google Scholar]
- Toro, J; Innes, DJ; Thompson, RJ. Genetic variation among life-history stages of mussels in a Mytilus edulis—M. trossulus hybrid zone. Mar. Biol 2004, 145, 713–725. [Google Scholar]
- Riginos, C; Cunningham, CW. Local adaptation and species segregation in two mussel (Mytilus edulis x Mytilus trossulus) hybrid zones. Mol. Ecol 2005, 14, 381–400. [Google Scholar]
- Borsa, P; Daguin, C; Bierne, N. Genomic reticulation indicates mixed ancestry in southern-hemisphere Mytilus spp. mussels. Biol. J. Linn. Soc 2007, 92, 747–754. [Google Scholar]
- Coghlan, B; Gosling, E. Genetic structure of hybrid mussel populations in the west of Ireland: Two hypotheses revisited. Mar. Biol 2007, 150, 841–852. [Google Scholar]
- Shields, JL; Barnes, P; Heath, DD. Growth and survival differences among native, introduced and hybrid blue mussels (Mytilus spp.): Genotype, environments and interaction effects. Mar. Biol 2008, 154, 919–928. [Google Scholar]
- Nielsen, EE; Nielsen, PH; Meldrup, D; Hansen, MM. Genetic population structure of turbot (Scophthalmus maximus L.) supports the presence of multiple hybrid zones for marine fishes in the transition zone between the Baltic Sea and the North Sea. Mol. Ecol 2004, 13, 585–595. [Google Scholar]
- Bekkevold, D; Andre, C; Dahlgren, TG; Clausen, LAW; Torstensen, E; Mosegaard, H; Carvalho, GR; Christensen, TB; Norlinder, E; Ruzzante, DE. Environmental correlates of population differentiation in Atlantic herring. Evolution 2005, 59, 2656–2668. [Google Scholar]
- Nielsen, EE; Hansen, MM; Ruzzante, DE; Meldrup, D; Grønkjær, P. Evidence of a hybrid-zone in Atlantic cod (Gadus morhua) in the Baltic and the Danish Belt Sea revealed by individual admixture analysis. Mol. Ecol 2003, 12, 1497–1508. [Google Scholar]
- Nielsen, EE; Grønkjær, P; Meldrup, D; Paulsen, H. Retention of juveniles within a hybrid zone between North Sea and Baltic Sea Atlantic cod (Gadus morhua). Can. J. Fish. Aquat. Sci 2005, 62, 2219–2225. [Google Scholar]
- Hemmer-Hansen, J; Nielsen, EEG; Grønkjær, P; Loeschcke, V. Evolutionary mechanisms shaping the genetic population structure of marine fishes; lessons from the European flounder (Platichthys flesus L.). Mol. Ecol 2007, 16, 3104–3118. [Google Scholar]
- Chow, S; Kishino, H. Phylogenetic relationships between tuna species of the genus Thunnus (Scombridae: Teleostei): Inconsistent implications from morphology, nuclear and mitochondrial genomes. J. Mol. Evol 1995, 41, 741–748. [Google Scholar]
- Alvarado Bremer, JR; Naseri, I; Ely, B. Orthodox and unorthodox phylogenetic relationships among tunas revealed by the nucleotide sequence analysis of the mitochondrial DNA control region. J. Fish Biol 1997, 50, 540–554. [Google Scholar]
- Alvarado Bremer, JR; Viñas, J; Mejuto, J; Ely, B; Pla, C. Comparative phylogeography of Atlantic bluefin tuna and swordfish: The combined effects of vicariance, secondary contact, introgression, and population expansion on the regional phylogenies of two highly migratory pelagic fishes. Mol. Phylogenet. Evol 2005, 36, 169–187. [Google Scholar]
- Avise, JC; Nelson, WS; Arnold, J; Koehn, RK; Williams, GC; Thorsteinsson, V. The evolutionary genetic status of Icelandic eels. Evolution 1990, 44, 1254–1262. [Google Scholar]
- Albert, V; Jónsson, B; Bernatchez, L. Natural hybrids in Atlantic eels (Anguilla anguilla, A. rostrata): Evidence for successful reproduction and fluctuating abundance in space and time. Mol. Ecol 2006, 15, 1903–1916. [Google Scholar]
- Burford, MO; Bernardi, G. Incipient speciation within a subgenus of rockfish (Sebastosomus) provides evidence of recent radiations within an ancient species flock. Mar. Biol 2008, 154, 701–717. [Google Scholar]
- Planes, S; Doherty, PJ. Genetic and color interactions at a contact zone of Acanthochromis polyacanthus: A marine fish lacking pelagic larvae. Evolution 1997, 51, 1232–1243. [Google Scholar]
- van Herwerden, L; Doherty, PJ. Contrasting genetic structure across two hybrid zones of a tropical reef fish, Acanthochromis polyacanthus (Bleeker 1855). J. Evol. Biol 2006, 19, 239–252. [Google Scholar]
- McMillan, WO; Weigt, LA; Palumbi, SR. Color pattern evolution, assortative mating, and genetic differentiation in brightly colored butterflyfishes (Chaetodontidae). Evolution 1999, 53, 247–260. [Google Scholar]
- Crespi, BJ; Fulton, MJ. Molecular systematics of Salmonidae: Combined nuclear data yields a robust phylogeny. Mol. Phylogenet. Evol 2004, 31, 658–679. [Google Scholar]
- Campos, JL; Posada, D; Morán, P. Introgression and genetic structure in northern Spanish Atlantic salmon (Salmo salar L.) populations according to mtDNA data. Conserv. Genet 2008, 9, 157–169. [Google Scholar]
| Taxon | Common Name | Type of Genetic Exchange | Data | Reference(s) |
|---|---|---|---|---|
| Multiple | Cyanobacteria | Horizontal transfer | Genome sequence analyses, Phylogenetic discordance | [28,29,69–73] |
| Multiple | Bacteriophage | Horizontal transfer | Genome sequence analyses, Phylogenetic discordance | [72–74] |
| Multiple | Bacteria | Horizontal transfer | Genome sequence analyses, Phylogenetic discordance | [25] |
| Multiple | Archaebacteria | Horizontal transfer | Genome sequence analyses, Phylogenetic discordance | [25] |
| Spartina (Multiple) | Cordgrass | Introgressive hybridization, Hybrid speciation | Population genetic surveys, Phylogenetic discordance | [75–79] |
| Zostera (Multiple) | Eelgrass | Introgressive hybridization | Population genetic surveys | [34] |
| Fucus (Multiple) | Seaweed | Introgressive hybridization, Hybrid speciation | Population genetic surveys, Phylogenetic discordance | [80–84] |
| Pseudo-nitzschia pungens | Diatom | Introgressive hybridization | Morphological analyses, Population genetic surveys | [44–46] |
| Bythograea thermydron | Hydrothermal crab | Horizontal transfer (transposable elements) | Phylogenetic discordance | [32] |
| Ventiella sulfuris | Hydrothermal amphipod | Horizontal transfer (transposable elements) | Phylogenetic discordance | [32] |
| Maia brachydactila | Sea shoe | Horizontal transfer (transposable elements) | Phylogenetic discordance | [32] |
| Cancer pagurus | Crab | Horizontal transfer (transposable elements) | Phylogenetic discordance | [32] |
| Menippe (Multiple) | Stone crab | Introgressive hybridization | Population genetic surveys | [85–87] |
| Mysis (Multiple) | Opossum shrimp | Introgressive hybridization | Phylogenetic discordance | [48,50] |
| Diadema (Multiple) | Sea urchins | Introgressive hybridization | Population genetic surveys | [56] |
| Lytechinus (Multiple) | Sea urchins | Introgressive hybridization | Population genetic surveys, Phylogenetic discordance | [57] |
| Strongylocentrotus (Multiple) | Sea urchins | Introgressive hybridization | Population genetic surveys, Phylogenetic discordance | [53–55] |
| Asterias (Multiple) | Sea stars | Introgressive hybridization | Morphological analyses, Population genetic surveys, Phylogenetic discordance | [54,58,59] |
| Acrocnida brachiata | Brittle-star | Introgressive hybridization | Population genetic surveys | [88] |
| Alcyonium hibernicum | Coral | Hybrid speciation | Population genetic surveys, Phylogenetic discordance | [40] |
| Bellonella bocagei | Coral | Hybrid speciation | Population genetic surveys, Phylogenetic discordance | [40] |
| Pocillopora (Multiple) | Corals | Introgressive hybridization | Population genetic surveys, Phylogenetic discordance | [41] |
| Acropora (Multiple) | Corals | Introgressive hybridization, Hybrid speciation | Morphological analyses, Population genetic surveys, Phylogenetic discordance | [19,35,89–94] |
| Littorina saxtilis | Snail | Introgressive hybridization | Morphological analyses | [95,96] |
| Mercenaria (Multiple) | Clams | Introgressive hybridization | Population genetic surveys | [97,98] |
| Macoma balthica | Clam | Introgressive hybridization | Population genetic surveys | [99,100] |
| Crassostrea virginica | American oyster | Introgressive hybridization | Population genetic surveys | [101,102] |
| Mytilus (Multiple) | Mussels | Introgressive hybridization | Morphological analyses, Population genetic surveys, Phylogenetic discordance | [103–114] |
| Bathymodiolus (Multiple) | Hydrothermal vent mussels | Introgressive hybridization | Morphological analyses, Population genetic surveys | [51,52] |
| Scophthalmus maximus | Turbot | Introgressive hybridization | Population genetic surveys | [115] |
| Clupea harengus | Atlantic herring | Introgressive hybridization | Population genetic surveys | [116] |
| Gadus morhua | Atlantic cod | Introgressive hybridization | Population genetic surveys | [117,118] |
| Platichthys flesus | European flounder | Introgressive hybridization | Population genetic surveys | [119] |
| Pleuronectes platessa | Plaice | Introgressive hybridization | Population genetic surveys | [119] |
| Thunnus (Multiple) | Tuna and Albacore | Introgressive hybridization | Population genetic surveys, Phylogenetic discordance | [120–122] |
| Anguilla (Multiple) | Eels | Introgressive hybridization | Population genetic surveys | [123,124] |
| Sebastosomus (Multiple) | Rockfish | Introgressive hybridization | Population genetic surveys | [125] |
| Acanthochromis (Multiple) | Damselfish | Introgressive hybridization | Population genetic surveys | [126,127] |
| Plectropomus (Multiple) | Coral trout | Introgressive hybridization | Population genetic surveys, Phylogenetic discordance | [60] |
| Acanthurus (Multiple) | Surgeonfish | Introgressive hybridization, Hybrid speciation | Population genetic surveys, Phylogenetic discordance | [61] |
| Chaetodon (Multiple) | Butterflyfish | Introgressive hybridization | Population genetic surveys, Phylogenetic discordance | [128] |
| Sebastes (Multiple) | Redfish | Introgressive hybridization | Population genetic surveys | [15] |
| Salmonidae (Multiple) | Charr, Salmon | Introgressive hybridization | Population genetic surveys, Phylogenetic discordance | [129,130] |
| Caretta caretta | Loggerhead turtle | Introgressive hybridization | Population genetic surveys, Phylogenetic discordance | [62–64] |
| Lepidochelys (Multiple) | Kemp’s ridley and Olive ridley turtles | Introgressive hybridization | Population genetic surveys, Phylogenetic discordance | [62–64] |
| Eretmochelys imbricata | Hawksbill turtle | Introgressive hybridization | Population genetic surveys, Phylogenetic discordance | [62–64] |
| Chelonia mydas | Green turtle | Introgressive hybridization | Population genetic surveys | [62] |
| Arctocephalus (Multiple) | Fur seals | Introgressive hybridization | Morphological analyses, Population genetic surveys | [65–68] |
| Eretmocrinus (Fossils) | Crinoids | Introgressive hybridization | Morphological analyses | [20] |
| Montastraea (Fossils) | Corals | Introgressive hybridization | Morphological analyses | [21] |
© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Arnold, M.L.; Fogarty, N.D. Reticulate Evolution and Marine Organisms: The Final Frontier? Int. J. Mol. Sci. 2009, 10, 3836-3860. https://doi.org/10.3390/ijms10093836
Arnold ML, Fogarty ND. Reticulate Evolution and Marine Organisms: The Final Frontier? International Journal of Molecular Sciences. 2009; 10(9):3836-3860. https://doi.org/10.3390/ijms10093836
Chicago/Turabian StyleArnold, Michael L., and Nicole D. Fogarty. 2009. "Reticulate Evolution and Marine Organisms: The Final Frontier?" International Journal of Molecular Sciences 10, no. 9: 3836-3860. https://doi.org/10.3390/ijms10093836
APA StyleArnold, M. L., & Fogarty, N. D. (2009). Reticulate Evolution and Marine Organisms: The Final Frontier? International Journal of Molecular Sciences, 10(9), 3836-3860. https://doi.org/10.3390/ijms10093836




