Resistance to β-lactams in Bacteria Isolated from Different Types of Portuguese Cheese
Abstract
:1. Introduction
2. Experimental Section
2.1. Sampling
2.2. Enumeration of ampicillin resistant bacteria
2.3. Detection of ampicillin resistant bacteria
2.4. Nitrocefin test and antimicrobial susceptibility tests
2.5. Bacterial identification
2.6. β-lactamase genes
3. Results and Discussion
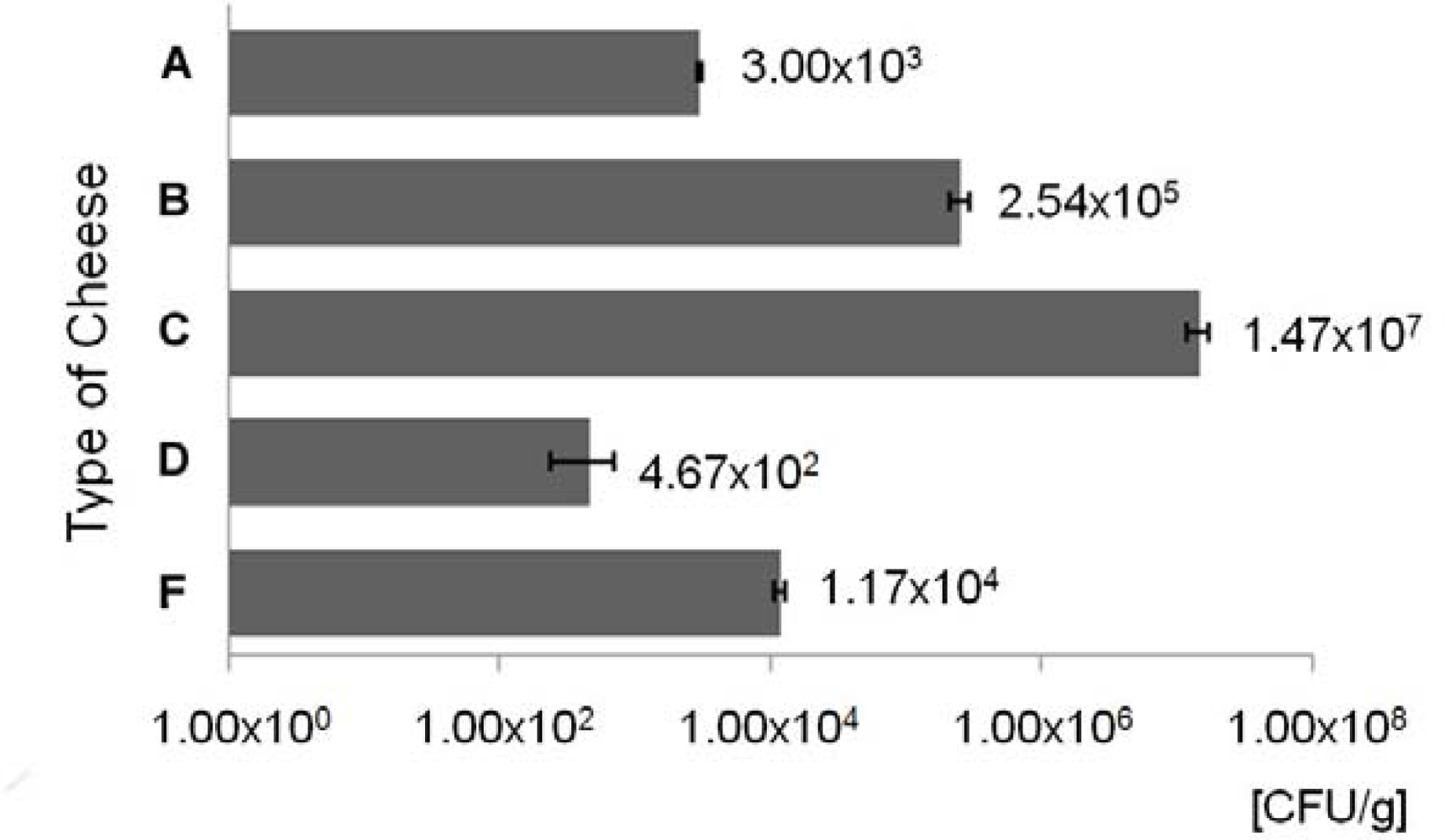
3.1. Enumeration of ampicillin resistant bacteria
3.2. Detection of β-lactam-resistant bacteria
3.3. Bacterial identification
3.4. Antimicrobial susceptibility tests
3.5. Molecular genetics of resistance genes
4. Conclusions
Acknowledgments
References and Notes
- Li, XZ; Mehrotra, M; Ghimire, S; Adewoye, L. β-lactam resistance and β-lactamases in bacteria of animal origin. Vet. Microbiol 2007, 121, 197–214. [Google Scholar]
- Silbergeld, EK; Graham, J; Price, LB. Industrial Food Animal Production, Antimicrobial Resistance and Human Health. Ann. Rev. Public Health 2008, 29, 151–169. [Google Scholar]
- Johnson, JR; Kuskowski, MA; Smith, K; O’Bryan, TT; Tatini, S. Antimicrobial-resistant and extra intestinal pathogenic Escherichia coli in retail foods. J. Infect. Dis 2005, 191, 1040–1049. [Google Scholar]
- Baylis, CL. Food Spoilage Microorganisms; Blackburn, CW, Ed.; CRC Press LLC: Cambridge, UK, 2006; Part 5, p. 635. [Google Scholar]
- Prudêncio, C. Multidrug resistance in yeast; PhD thesis, University of Minho: Braga, Portugal, 2000. [Google Scholar]
- Prudêncio, C; Sansonetty, F; Sousa, MJ; Côrte-Real, M; Leão, C. Rapid detection of efflux pumps and their relation with drug-resistance in yeast cells. Cytometry 2000, 39, 26–35. [Google Scholar]
- Walsh, C. Molecular mechanisms that confer antibacterial drug resistance. Rev. Nature 2000, 406, 775–781. [Google Scholar]
- Kotra, LP; Samama, J; Mobashery, S. Bacterial Resistance to Antimicrobials; Lewis, K, Salyers, AA, Taber, HW, Wax, RG, Eds.; Marcel Decker: New York, NY, USA, 2000; pp. 123–160. [Google Scholar]
- Samaha-Kfoury, JN; Araj, GF. Recent developments in β-lactamases and extended-spectrum β-lactamases. BMJ 2003, 327, 1209–1213. [Google Scholar]
- Sanders, CC; Bradford, PA; Ehrhardt, AF; Bush, K; Young, KD; Henderson, TA; Sanders, WE, Jr. Penicillin-binding proteins and induction of AmpC beta-lactamase. Antimicrob. Agents Chemother 1997, 36, 2013–2015. [Google Scholar]
- Alvarez, M; Tran, JH; Chow, N; Jacoby, GA. Epidemiology of conjugative plasmid-mediated AmpC beta-lactamases in the United States. Antimicrob. Agents Chemother 2004, 48, 533–537. [Google Scholar]
- Philippon, A; Arlet, G; Jacoby, GA. Plasmid-determined AmpC-type β-lactamases. Antimicrob. Agents Chemother 2002, 46, 1–11. [Google Scholar]
- Mesa, RJ; Blanc, V; Blanch, AR; Cortés, P; González, JJ; Lavilla, S; Miró, E; Muniesa, M; Saco, M; Tórtola, MT; Mirelis, B; Coll, P; Llagostera, M; Prats, G; Navarro, F. Extended-spectrum β-lactamase-producing Enterobacteriaceae in different environments (humans, food, animal farms and sewage). J. Antimicrob. Chemother 2006, 58, 211–215. [Google Scholar]
- Bradford, PA. Extended-spectrum β-lactamases in the 21st century: Characterization, epidemiology, and detection of this important resistance threat. Clin. Microbiol. Rev 2001, 14, 933–951. [Google Scholar]
- Jorgensen, JH. NCCLS; Methods for dilution antimicrobial susceptibility test for bacteria that grow aerobically, 6th ed; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2003; pp. M7–A6. [Google Scholar]
- NCCLS; Performance standard for antimicrobial susceptibility testing; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2005; Supplement pp. M100–S15.
- Holt, JG. Bergey’s Manual of Determinative Bacteriology; Lippincott Williams & Wilkins: Philadelphia, USA, 1994. [Google Scholar]
- Hugh, R; Leifson, E. The taxonomic significance of fermentative versus oxidative metabolism of carbohydrates by various Gram negative bacteria. J. Bacteriol 1953, 66, 24–26. [Google Scholar]
- Monstein, HJ; Östholm-Balkhed, Å; Nilsson, NV; Nilsson, M; Dornbusch, K; Nilsson, LE. Multiplex PCR amplification assay for the detection of blaSHV, blaTEM and blaCTX-M genes in Enterobacteriaceae. Apmis 2007, 115, 1400–1408. [Google Scholar]
- Pitcher, DG; Saunders, NA; Owen, RJ. Rapid extraction of bacterial genomic DNA with guanidium thiocyanate. Lett. Appl. Microbiol 1989, 8, 151–156. [Google Scholar]
- Commission Regulation (EC 1441/2007). Official J. Eur. Commun 2007, L322, 12–29.
- Morales, P; Fernández-Garcia, E; Nuñez, M. Caseinolysis in cheese by Enterobacteriaceae strains of dairy origin. Lett. Appl. Microbiol 2003, 37, 410–414. [Google Scholar]
- Ferech, M; Coenen, S; Malhotra-Kumar, S; Dvorakova, K; Hendrickx, E; Suetens, C; Goossens, H. esac project group. European Surveillance of Antimicrobial Consumption (ESAC): Outpatient quinolone use in Europe. J. Antimicrob. Chemother 2006, 58, 423–427. [Google Scholar]
- Ferech, M; Coenen, S; Dvorakova, K; Hendrickx, E; Suetens, C; Goossens, H. esac project group. European Surveillance of Antimicrobial Consumption (ESAC): Outpatient penicillin use in Europe. J. Antimicrob. Chemother 2006, 58, 408–412. [Google Scholar]
- Coenen, S; Ferech, M; Dvorakova, K; Hendrickx, E; Suetens, C; Goossens, H. esac project group. European Surveillance of Antimicrobial Consumption (ESAC): outpatient cephalosporin use in Europe. J. Antimicrob. Chemother 2006, 58, 413–417. [Google Scholar]
- Emborg, HD; Vigre, H; Jensen, VF; Vieira, ARP; Baggesen, DL; Aarestrup, FM. Tetracycline consumption and occurrence of tetracycline resistance in Salmonella Typhimurium phage types from danish pigs. Microb. Drug Res 2007, 13, 289–294. [Google Scholar]
- Nawaz, M; Khan, AA; Khan, S; Sung, K; Steele, R. Isolation and characterization of tetracycline-resistant Citrobacter spp. from catfish. Food Microbiol 2008, 25, 85–91. [Google Scholar]
- Schwarz, S; Roberts, MC; Werckenthin, C; Pang, Y; Lange, C. Tetracycline resistance in Staphylococcus spp. from domestic animals. Vet. Microbiol 1998, 63, 217–227. [Google Scholar]
- Stine, OC; Johnson, JA; Keefer-Norris, A; Perry, KL; Tigno, J; Qaiyumi, S; Stine, MS; Morris, JG, Jr. Widespread distribution of tetracycline resistance genes in a confined animal feeding facility. Int. J. Antimicrob. Agents 2007, 29, 348–352. [Google Scholar]
- Nguyen, TV; Le, PV; Le, CH; Weintraub, A. Antibiotic resistance in diarrheagenic Escherichia coli and Shigella strains isolated from children in Hanoi, Vietnam. Antimicrob. Agents Chemother 2005, 49, 816–819. [Google Scholar]
- Goettsch, W; van Pelt, W; Nagelkerke, N; Hendrix, MG; Buiting, AG; Petit, PL; Sabbe, LJ; van Griethuysen, AJ; de Neeling, AJ. Increasing resistance to fluoroquinolones in Escherichia coli from urinary tract infections in the Netherlands. J. Antimicrob. Chemother 2000, 46, 223–228. [Google Scholar]
- Fernandes, R; Vieira, M; Ferraz, R; Prudêncio, C. Bloodstream infections caused by multidrug-resistant Enterobacteriaceae: report from two Portuguese hospitals. J. Hosp. Infect 2008, 70, 93–95. [Google Scholar]
- Livermore, DM; Canton, R; Gniadkowski, M; Nordmann, P; Rossolini, GM; Arlet, G; Ayala, J; Coque, TM; Kern-Zdanowicz, I; Luzzaro, F; Poirel, L; Woodford, N. CTX-M: changing the face of ESBLs in Europe. J. Antimicrob. Chemother 2007, 59, 165–174. [Google Scholar]
- Bhatta, DR; Bangtrakulnonth, A; Tishyadhigama, P; Saroj, SD; Bandekar, JR; Hendriksen, RS; Kapadnis, BP. Serotyping, PCR, phage-typing and antibiotic sensitivity testing of Salmonella serovars isolated from urban drinking water supply systems of Nepal. Lett. Appl. Microbiol 2007, 44, 588–594. [Google Scholar]
- Lachmayr, KL; Kerkhof, LJ; Dirienzo, AG; Cavanaugh, CM; Ford, TE. Quantifying nonspecific TEM beta-lactamase (blaTEM) genes in a wastewater stream. Appl. Environ. Microbiol 2009, 75, 203–211. [Google Scholar]
- Sunde, M; Tharaldsen, H; Slettemeås, JS; Norström, M; Carattoli, A; Bjorland, J. Escherichia coli of animal origin in Norway contains a blaTEM-20-carrying plasmid closely related to blaTEM-20 and blaTEM-52 plasmids from other European countries. J. Antimicrob. Chemother 2009, 63, 215–216. [Google Scholar]
- Kim, SH; Wei, CI; An, H. Molecular characterization of multidrug-resistant Proteus mirabilis isolates from retail meat products. J. Food Prot 2005, 68, 1408–1413. [Google Scholar]
- Bibbal, D; Dupouy, V; Ferré, JP; Toutain, PL; Fayet, O; Prère, MF; Bousquet-Mélou, A. Impact of three ampicillin dosage regimens on selection of ampicillin resistance in Enterobacteriaceae and excretion of blaTEM genes in swine feces. Appl. Environ. Microbiol 2007, 73, 4785–4790. [Google Scholar]
- Cloeckaert, A; Praud, K; Doublet, B; Bertini, A; Carattoli, A; Butaye, P; Imberechts, H; Bertrand, S; Collard, JM; Arlet, G; Weill, FX. Dissemination of an extended-spectrum-beta-lactamase blaTEM-52 gene-carrying IncI1 plasmid in various Salmonella enterica serovars isolated from poultry and humans in Belgium and France between 2001 and 2005. Antimicrob. Agents Chemother 2007, 51, 1872–1875. [Google Scholar]
- Demanèche, S; Sanguin, H; Poté, J; Navarro, E; Bernillon, D; Mavingui, P; Wildi, W; Vogel, TM; Simonet, P. Antibiotic-resistant soil bacteria in transgenic plant fields. Proc. Natl. Acad. Sci. USA 2008, 105, 3957–3962. [Google Scholar]
- Partridge, SR; Hall, RM. Evolution of transposons containing blaTEM genes. Antimicrob. Agents Chemother 2005, 49, 1267–1268. [Google Scholar]
- Simeoni, D; Rizzotti, L; Cocconcelli, P; Gazzola, S; Dellaglio, F; Torriani, S. Antibiotic resistance genes and identification of staphylococci collected from the production chain of swine meat commodities. Food Microbiol 2008, 25, 196–201. [Google Scholar]
- Angulo, FJ; Nargund, VN; Chiller, TC. Evidence of an association between use of antimicrobial agents in food animals and antimicrobial resistance among bacteria isolated from humans and the human health consequences of such resistance. J. Vet. Med. B 2004, 51, 374–379. [Google Scholar]
- Phillips, I; Casewell, M; Cox, T; De Groot, B; Friis, C; Jones, R; Nightingale, C; Preston, R; Waddell, J. Does the use of antibiotics in food animals pose a risk to human health? A critical review of published data. J. Antimicrob. Chemother 2004, 53, 28–52. [Google Scholar]
- Goossens, H; Ferech, M; Vander Stichele, R; Elseviers, M. ESAC Project Group. Outpatient antibiotic use in Europe and association with resistance: a cross-national database study. Lancet 2005, 365, 579–587. [Google Scholar]
| Type of cheese | Milk | Number of cheeses | |
|---|---|---|---|
| Origin | Thermal processing | ||
| A* | ewe | no | 2 |
| B | ewe | no | 13 |
| C | goat | no | 2 |
| D | goat | pasteurization | 1 |
| E | cow | pasteurization | 1 |
| F | mixture (cow, ewe and goat) | no | 1 |
| Family/Genus | Species | Number of isolates | % | Type of cheese | Recovery medium |
|---|---|---|---|---|---|
| Enterobacteriaceae | NI | 17 | 28.3 | A, B, C, D | VRBG, SS |
| Proteus vulgaris | 9 | 15.0 | B, C, F | VRBG, SS | |
| Escherichia coli | 6 | 10.0 | B, D, F | VRBG, SS | |
| Morganella morganii | 6 | 10 | B, C, F | VRBG, SS | |
| Enterobacter cloacae | 4 | 6.7 | A, B, F | VRBG | |
| Proteus mirabilis | 2 | 3.3 | B, F | VRBG | |
| Serratia rubidaea | 2 | 3.3 | A | SS | |
| Citrobacter freundii | 1 | 1.7 | B | VRBG | |
| Enterobacter sakazakii | 1 | 1.7 | B | VRBG | |
| Enterobacter alvei | 1 | 1.7 | B | VRBG | |
| Providencia retgeri | 1 | 1.7 | B | SS | |
| Shigella boydii | 1 | 1.7 | D | SS | |
| Pseudomonas spp. | NI | 5 | 8.3 | B, C | VRBG, SS |
| Aeromonas spp. | NI | 4 | 6.7 | B, C, D | VRBG, SS |
| All | 60 | 100 | A, B, C, D, F | VRBG, SS |
| Susceptible | Resistant | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Antimicrobial agent | Φ | MIC a) | % | Φ | MIC a) | n | % | ||
| (mm) | (μg/mL) | n | (n = 172) | (mm) | (μg/mL) | (n = 172) | |||
| β - lactams | AMC (30:10 μg) | ≥ 18 | ≤ 8/4 | 67 | 39.0 | ≤ 13 | ≥ 32/16 | 89 | 51.7 |
| FOX (30 μg) | ≥ 18 | ≤ 8 | 70 | 40.7 | ≤ 14 | ≥ 32 | 80 | 46.5 | |
| CTX (30 μg) | ≥ 23 | ≤ 8 | 115 | 66.9 | ≤ 14 | ≥ 32 | 23 | 13.4 | |
| CPO (30 μg) | ≥ 18 | ≤ 8 | 150 | 87.2 | ≤ 14 | ≥ 32 | 10 | 5.8 | |
| ATM (30 μg) | ≥ 22 | ≤ 8 | 142 | 82.6 | ≤ 15 | ≥ 32 | 10 | 5.8 | |
| CAZ (30 μg) | ≥ 18 | ≤ 8 | 158 | 91.9 | ≤ 14 | ≥ 32 | 9 | 5.2 | |
| MEM (10 μg) | ≥ 16 | ≤ 4 | 156 | 90.7 | ≤ 13 | ≥ 16 | 9 | 5.2 | |
| IPM (10 μg) | ≥ 16 | ≤ 4 | 164 | 95.3 | ≤ 13 | ≥ 16 | 2 | 1.2 | |
| Non β – lactams | TET (30 μg) | ≥ 19 | ≤ 4 | 56 | 32.6 | ≤ 14 | ≥ 16 | 80 | 46.5 |
| SXT (1:19 μg) | ≥ 16 | ≤ 2/38 | 103 | 59.9 | ≤ 10 | ≥ 8/152 | 66 | 38.4 | |
| CHL (30 μg) | ≥ 18 | ≤ 8 | 108 | 62.8 | ≤ 12 | ≥ 32 | 35 | 20.3 | |
| GEN (10 μg) | ≥ 15 | ≤ 4 | 169 | 98.3 | ≤ 12 | ≥ 8 | 3 | 1.7 | |
| CIP (5 μg) | ≥ 21 | ≤ 1 | 168 | 97.7 | ≤ 15 | ≥ 4 | 0 | 0.0 | |
| Resistance Phenotype | Number of isolates | % | |
|---|---|---|---|
| Multidrug resistant (MDR) phenotypes | AMP, SXT, TET | 8 | 4.65 |
| AMP, AMC, FOX, SXT, TET | 8 | 4.65 | |
| AMP, AMC, FOX, CHL, TET | 7 | 4.07 | |
| AMP, AMC, FOX, CHL, SXT, TET | 4 | 2.33 | |
| AMP, AMC, FOX, CHL, SXT | 4 | 2.33 | |
| AMP, AMC, CHL, TET | 2 | 1.16 | |
| AMP, CHL, SXT, TET | 2 | 1.16 | |
| AMP, AMC, SXT, TET | 2 | 1.16 | |
| AMP, AMC, CHL, SXT, TET | 2 | 1.16 | |
| AMP, AMC, FOX, CEPH3, CPO, CHL, SXT, TET | 2 | 1.16 | |
| AMP, CHL, TET | 2 | 1.16 | |
| AMP, FOX, CHL, TET | 2 | 1.16 | |
| Others with only one representative | 9 | 5,22 | |
| Sub-total | 54 | 31.40 | |
| Non-multidrug resistant (MDR) phenotypes | AMP | 38 | 22.09 |
| AMP, TET | 19 | 11.05 | |
| AMP, AMC, FOX, SXT | 8 | 4.65 | |
| AMP, AMC, FOX, TET | 7 | 4.07 | |
| AMP, AMC, FOX | 6 | 3.49 | |
| AMP, AMC, TET | 5 | 2.91 | |
| AMP, SXT | 5 | 2.91 | |
| AMP, AMC | 4 | 2.33 | |
| AMP, AMC, FOX, CEPH3, ATM | 4 | 2.33 | |
| AMP, AMC, FOX, CEPH3, CPO, ATM | 4 | 2.33 | |
| AMP, FOX, SXT | 2 | 1.16 | |
| AMP, AMC, FOX, CEPH3, CARB, SXT | 2 | 1.16 | |
| AMP, AMC, FOX, CEPH3, SXT | 2 | 1.16 | |
| AMP, AMC, FOX, CEPH3, CPO, CARB, SXT | 2 | 1.16 | |
| AMP, AMC, FOX, SXT | 2 | 1.16 | |
| Others with only one representative | 8 | 4.64 | |
| Sub-Total | 118 | 68.60 | |
| Total | 172 | 100 | |
© 2009 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Amador, P.; Fernandes, R.; Prudêncio, C.; Brito, L. Resistance to β-lactams in Bacteria Isolated from Different Types of Portuguese Cheese. Int. J. Mol. Sci. 2009, 10, 1538-1551. https://doi.org/10.3390/ijms10041538
Amador P, Fernandes R, Prudêncio C, Brito L. Resistance to β-lactams in Bacteria Isolated from Different Types of Portuguese Cheese. International Journal of Molecular Sciences. 2009; 10(4):1538-1551. https://doi.org/10.3390/ijms10041538
Chicago/Turabian StyleAmador, Paula, Ruben Fernandes, Cristina Prudêncio, and Luísa Brito. 2009. "Resistance to β-lactams in Bacteria Isolated from Different Types of Portuguese Cheese" International Journal of Molecular Sciences 10, no. 4: 1538-1551. https://doi.org/10.3390/ijms10041538
APA StyleAmador, P., Fernandes, R., Prudêncio, C., & Brito, L. (2009). Resistance to β-lactams in Bacteria Isolated from Different Types of Portuguese Cheese. International Journal of Molecular Sciences, 10(4), 1538-1551. https://doi.org/10.3390/ijms10041538




