Comparative Study of Ni(II) Complexes with Dithiocarbazate- and Thiosemicarbazone-Based Ligands: Synthesis, Crystal Structures, and Anticancer Activity
Abstract
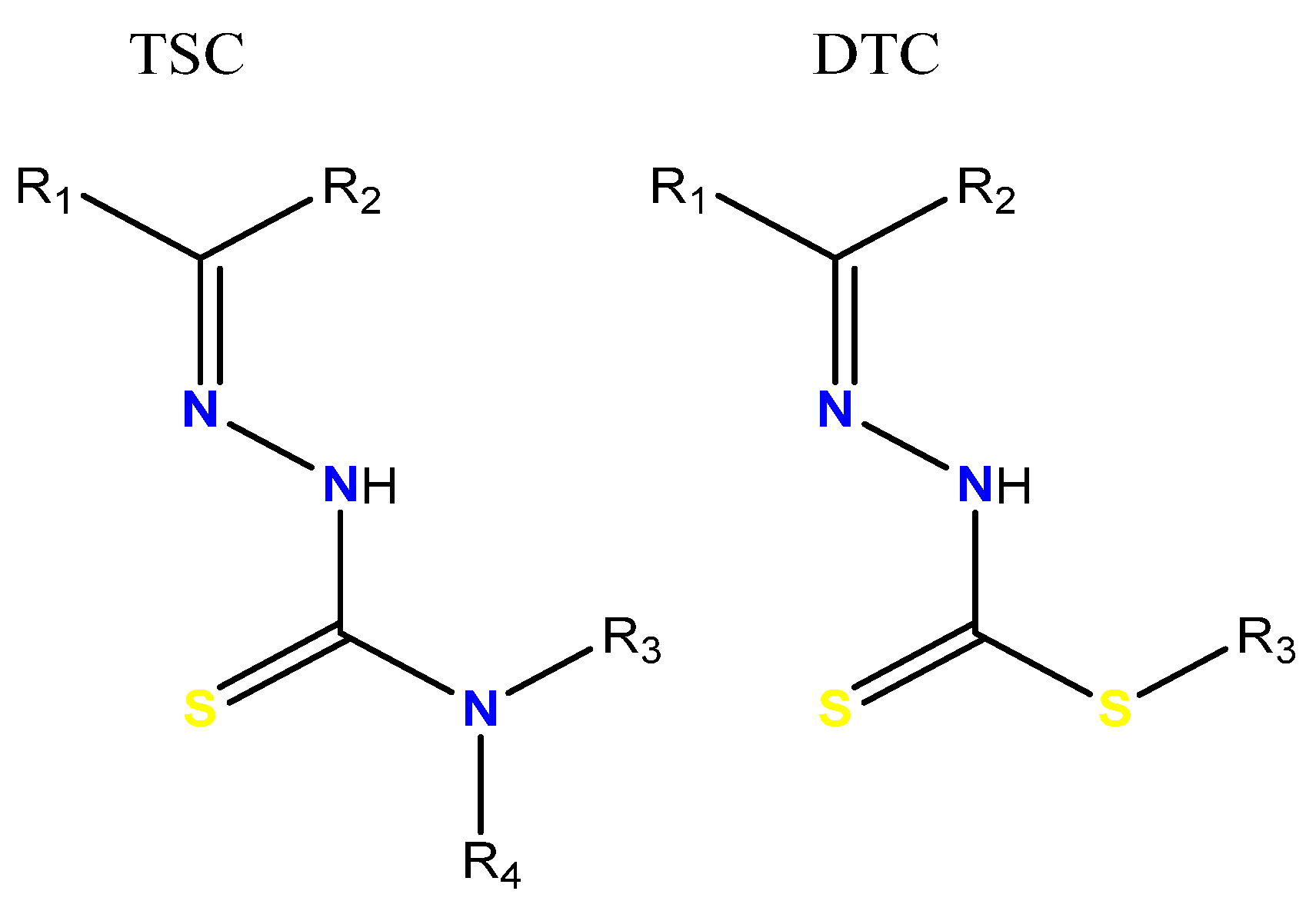
1. Introduction
2. Results and Discussion
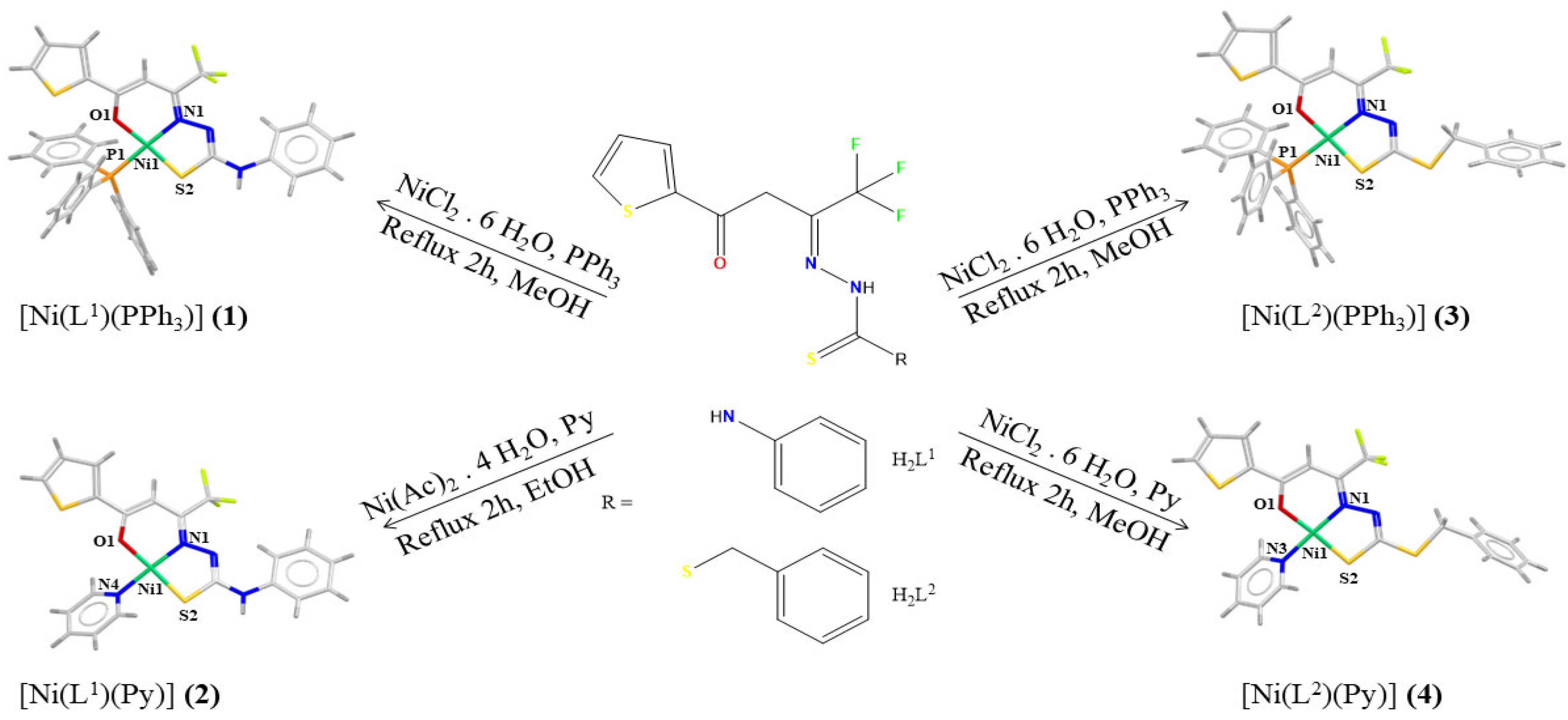
2.1. Structural Analyses
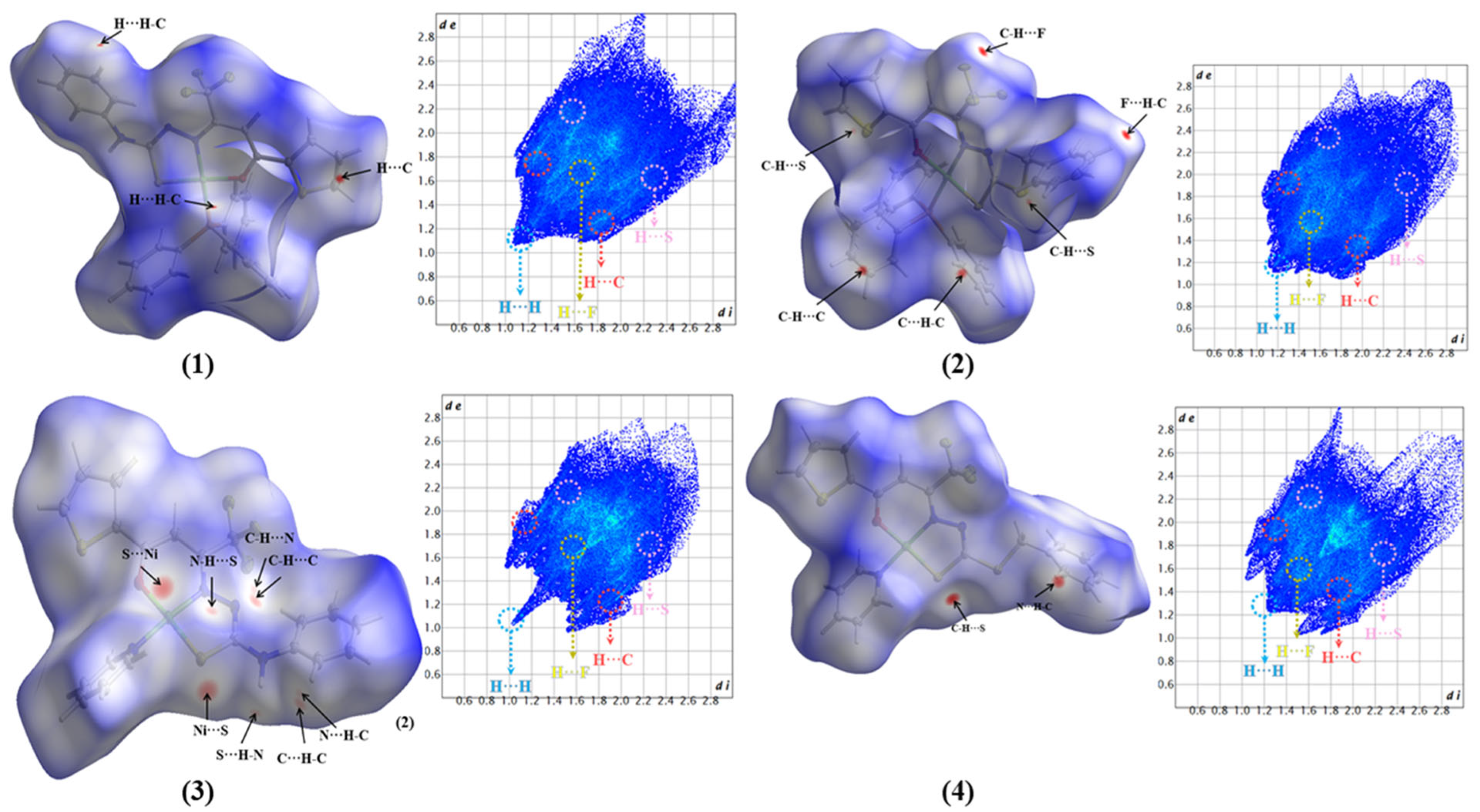
2.2. Hirshfeld Surface
2.3. Infrared Spectra
2.4. Electronic Spectra
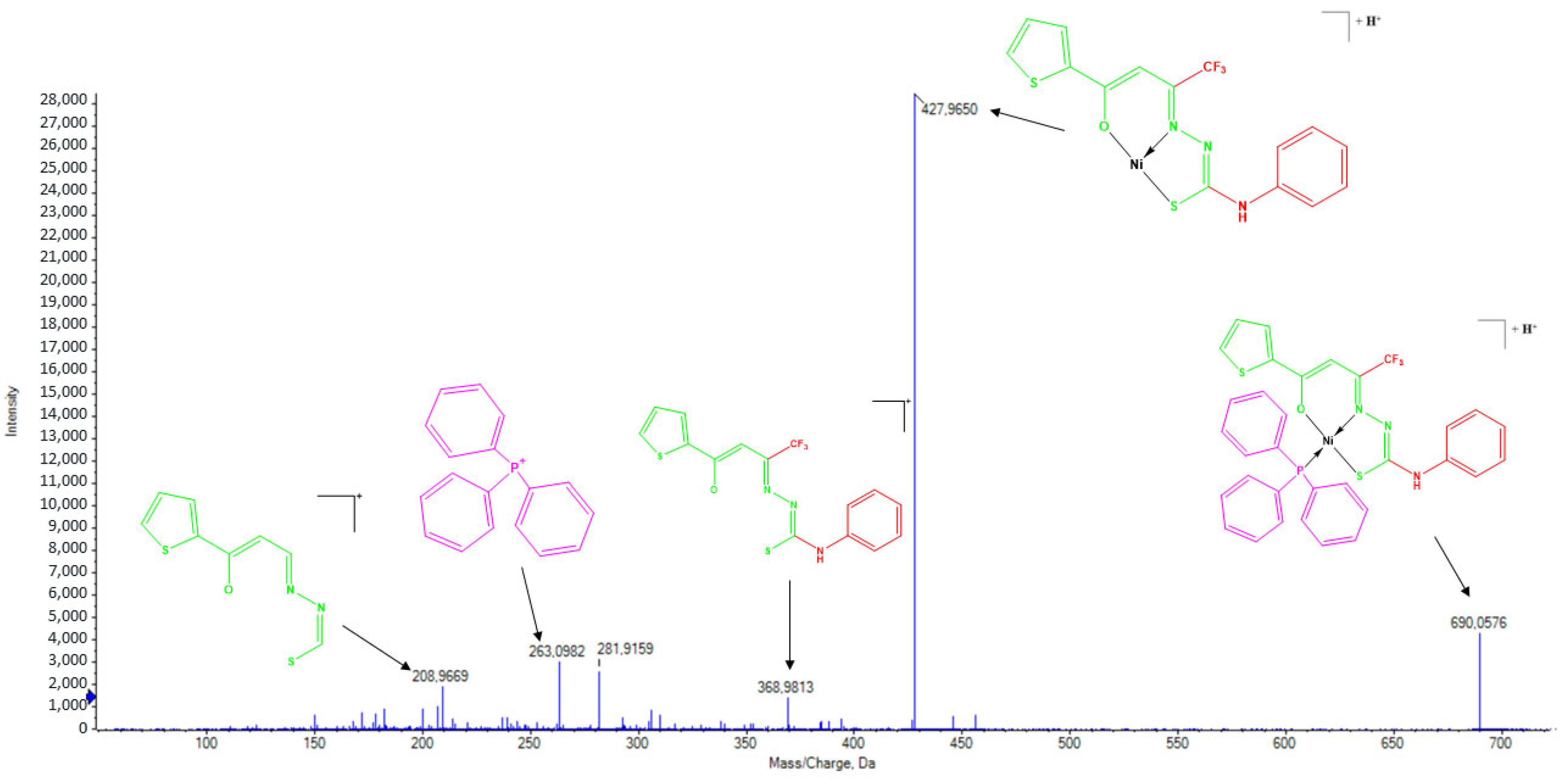
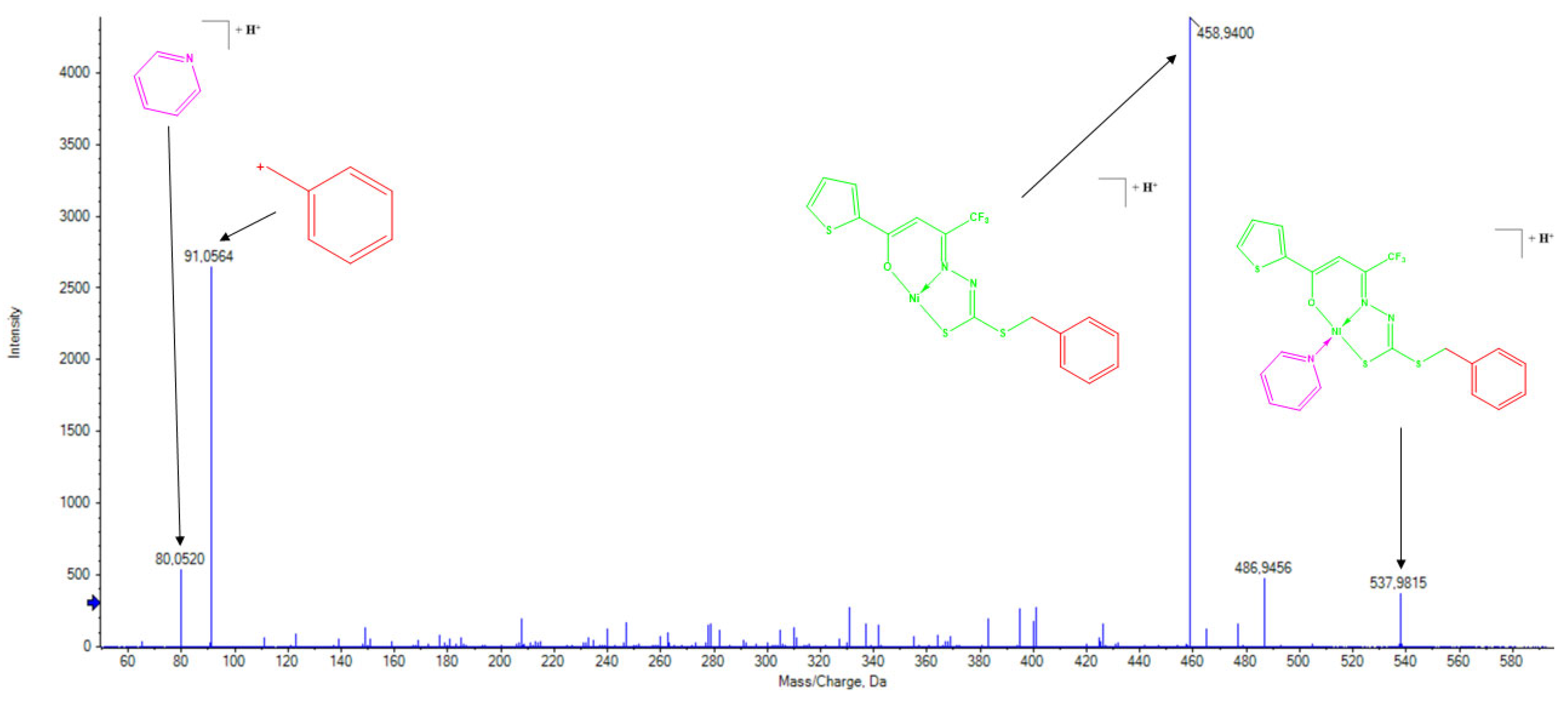
2.5. Mass Spectrometry
2.6. 1H, 19F, and 31P NMR Spectra
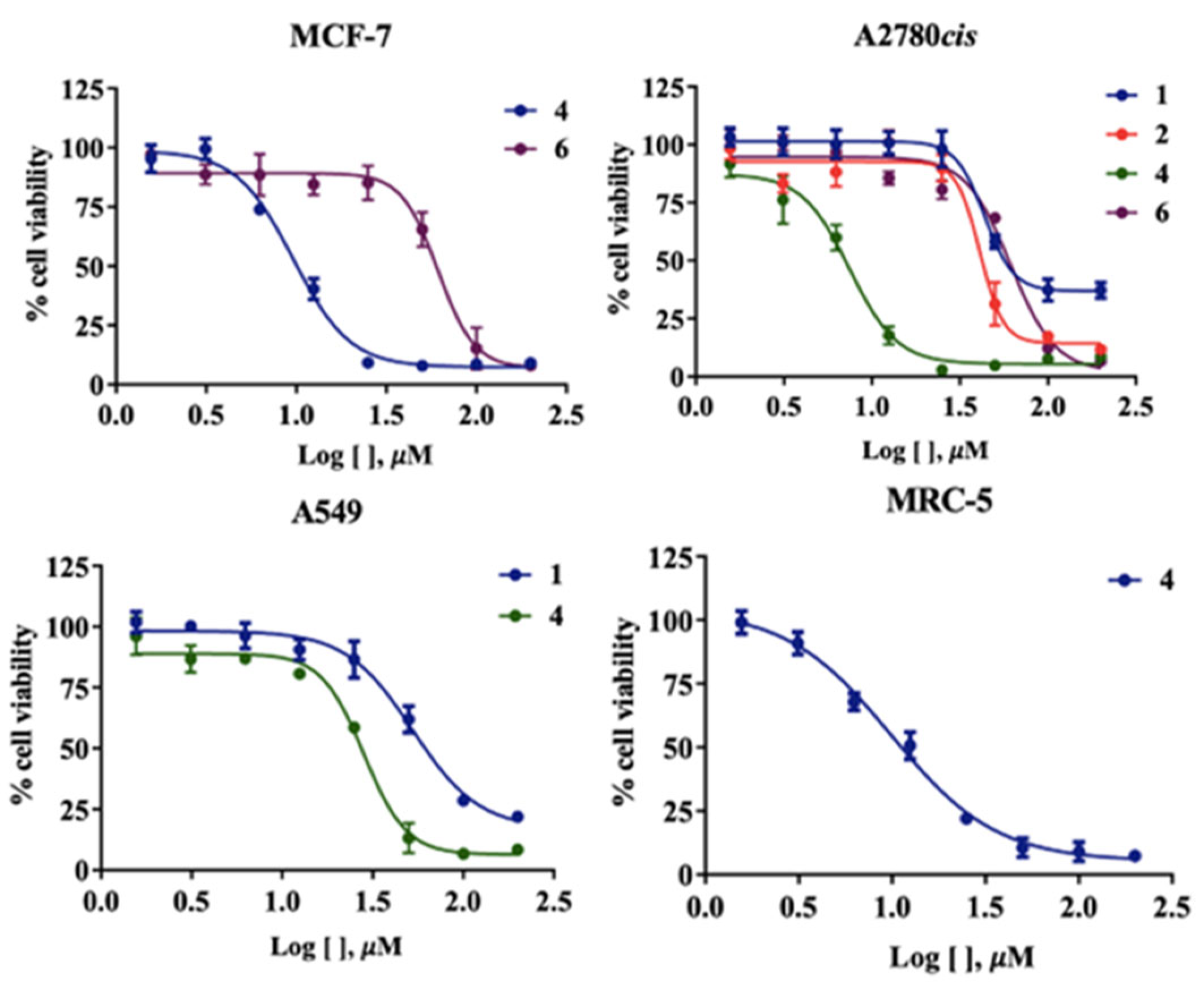
2.7. Biological Activity Analysis
3. Materials and Methods
3.1. Materials, Methods, and Instruments
3.2. Synthesis of Thiosemicarbazone Ligand (H2L1)
3.3. Synthesis of Dithiocarbazate Ligand (H2L2)
3.4. Synthesis of [Ni(L1)(PPh3)] (1)
3.5. Synthesis of [Ni(L1)(Py)] (2)
3.6. Synthesis of [Ni(L2)(PPh3)] (3)
3.7. Synthesis of [Ni(L2)(Py)] (4)
3.8. Crystal Structure Determination
3.9. Hirshfeld Surface
3.10. Anticancer Activity
3.10.1. Cell Culture
3.10.2. Cell Viability Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Edwards, I.R.; Aronson, J.K. Adverse Drug Reactions: Definitions, Diagnosis, and Management. Lancet 2000, 356, 1255–1259. [Google Scholar] [CrossRef]
- Due, A. What Are Side Effects? Eur. J. Philos. Sci. 2023, 13, 16. [Google Scholar] [CrossRef] [PubMed]
- Martinez, A.; Hemmert, C.; Loup, C.; Barré, G.; Meunier, B. Synthesis of New Macrocyclic Chiral Manganese(III) Schiff Bases as Catalysts for Asymmetric Epoxidation. J. Org. Chem. 2006, 71, 1449–1457. [Google Scholar] [CrossRef] [PubMed]
- Berhanu, A.L.; Gaurav; Mohiuddin, I.; Malik, A.K.; Aulakh, J.S.; Kumar, V.; Kim, K.-H. A Review of the Applications of Schiff Bases as Optical Chemical Sensors. TrAC Trends Anal. Chem. 2019, 116, 74–91. [Google Scholar] [CrossRef]
- Catalano, A.; Sinicropi, M.S.; Iacopetta, D.; Ceramella, J.; Mariconda, A.; Rosano, C.; Scali, E.; Saturnino, C.; Longo, P. A Review on the Advancements in the Field of Metal Complexes with Schiff Bases as Antiproliferative Agents. Appl. Sci. 2021, 11, 6027. [Google Scholar] [CrossRef]
- Ceramella, J.; Iacopetta, D.; Catalano, A.; Cirillo, F.; Lappano, R.; Sinicropi, M.S. A Review on the Antimicrobial Activity of Schiff Bases: Data Collection and Recent Studies. Antibiotics 2022, 11, 191. [Google Scholar] [CrossRef]
- Ejidike, I.P.; Ajibade, P.A. Transition Metal Complexes of Symmetrical and Asymmetrical Schiff Bases as Antibacterial, Antifungal, Antioxidant, and Anticancer Agents: Progress and Prospects. Rev. Inorg. Chem. 2015, 35, 191–224. [Google Scholar] [CrossRef]
- Hu, W.; Zhou, W.; Xia, C.; Wen, X. Synthesis and Anticancer Activity of Thiosemicarbazones. Bioorg. Med. Chem. Lett. 2006, 16, 2213–2218. [Google Scholar] [CrossRef]
- Heffeter, P.; Pape, V.F.S.; Enyedy, É.A.; Keppler, B.K.; Szakacs, G.; Kowol, C.R. Anticancer Thiosemicarbazones: Chemical Properties, Interaction with Iron Metabolism, and Resistance Development. Antioxid. Redox Signal. 2019, 30, 1062–1082. [Google Scholar] [CrossRef]
- Singh, N.K.; Kumbhar, A.A.; Pokharel, Y.R.; Yadav, P.N. Anticancer Potency of Copper(II) Complexes of Thiosemicarbazones. J. Inorg. Biochem. 2020, 210, 111134. [Google Scholar] [CrossRef]
- Chang, E.L.; Simmers, C.; Knight, D.A. Cobalt Complexes as Antiviral and Antibacterial Agents. Pharmaceuticals 2010, 3, 1711–1728. [Google Scholar] [CrossRef]
- de Andrade Danin Barbosa, G.; Palermo de Aguiar, A. Synthesis of 1,3,4-Thiadiazole Derivatives and Microbiological Activities: A Review. Rev. Virtual Química 2019, 11, 806–848. [Google Scholar] [CrossRef]
- Gou, Y.; Chen, M.; Li, S.; Deng, J.; Li, J.; Fang, G.; Yang, F.; Huang, G. Dithiocarbazate-Copper Complexes for Bioimaging and Treatment of Pancreatic Cancer. J. Med. Chem. 2021, 64, 5485–5499. [Google Scholar] [CrossRef]
- Gou, Y.; Jia, X.; Hou, L.X.; Deng, J.G.; Huang, G.J.; Jiang, H.W.; Yang, F. Dithiocarbazate–Fe III, –Co III, –Ni II, and –Zn II Complexes: Design, Synthesis, Structure, and Anticancer Evaluation. J. Med. Chem. 2022, 65, 6677–6689. [Google Scholar] [CrossRef]
- Leite, T.S.O.; Magalhães, I.O.M.; Machado, D.F.S.; Martins, J.B.L.; Gatto, C.C. Experimental and Theoretical Investigation of New Mono and Binuclear Cu(II) and Zn(II) Complexes with Dithiocarbazate Ligand. Struct. Chem. 2025, 1–20. [Google Scholar] [CrossRef]
- Nanjundan, N.; Selvakumar, P.; Narayanasamy, R.; Haque, R.A.; Velmurugan, K.; Nandhakumar, R.; Silambarasan, T.; Dhandapani, R. Synthesis, Structure, DNA/BSA Interaction and in Vitro Cytotoxic Activity of Nickel(II) Complexes Derived from S-Allyldithiocarbazate. J. Photochem. Photobiol. B Biol. 2014, 141, 176–185. [Google Scholar] [CrossRef] [PubMed]
- Cavalcante, C.d.Q.O.; da Mota, T.H.A.; de Oliveira, D.M.; Nascimento, É.C.M.; Martins, J.B.L.; Pittella-Silva, F.; Gatto, C.C. Dithiocarbazate Ligands and Their Ni(II) Complexes with Potential Biological Activity: Structural, Antitumor and Molecular Docking Study. Front. Mol. Biosci. 2023, 10, 6677–6689. [Google Scholar] [CrossRef]
- Gatto, C.C.; Cavalcante, C.d.Q.O.; Lima, F.C.; Nascimento, É.C.M.; Martins, J.B.L.; Santana, B.L.O.; Gualberto, A.C.M.; Pittella-Silva, F. Structural Design, Anticancer Evaluation, and Molecular Docking of Newly Synthesized Ni(II) Complexes with ONS-Donor Dithiocarbazate Ligands. Molecules 2024, 29, 2759. [Google Scholar] [CrossRef]
- Abd El-Lateef, H.M.; Khalaf, M.M.; Gouda, M.; Amer, A.A.; Abdelhamid, A.A.; Abdou, A. Synthesis, Characterization, and Therapeutic Potential of Novel Schiff Base Metal Complexes: Nickel(II) and Copper(II) Complexes Based 2-[(2-Chloroquinolin-3-yl)Methylidene]Aminobenzenethiol (CQAT). Appl. Organomet. Chem. 2025, 39, e7843. [Google Scholar] [CrossRef]
- Özbağcı, D.İ.; Erdağı, S.İ.; Aydın, İ.; Aydın, R.; Arı, F.; Zorlu, Y. Crystal Structures and Biological Profiles of Novel Tridentate Schiff Bases Nickel (II) Complexes. Appl. Organomet. Chem. 2025, 39, e7860. [Google Scholar] [CrossRef]
- Tewari, B.B. Metal Complexes in Biology and Medicine (Part- Mercury(II) / Nickel(II) /Lead(II)–Isoleucine). In Novel Aspects on Chemistry and Biochemistry; Mohamadpour, D.F., Ed.; B P International (a part of SCIENCEDOMAIN International): Kolkata, India, 2023; Volume 3, pp. 111–127. ISBN 9788119315451. [Google Scholar]
- Kumar, B.; Devi, J.; Dubey, A.; Tufail, A.; Antil, N. Biological and Computational Investigation of Transition Metal(II) Complexes of 2-Phenoxyaniline-Based Ligands. Future Med. Chem. 2023, 15, 1919–1942. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Choudhary, M. An In Silico Design of Antivirus Nickel (II) Complexes as Therapeutic Drug Candidates. Indian J. Chem. 2023, 62, 586–599. [Google Scholar] [CrossRef]
- Ma, B.N.; Baecker, D.; Descher, H.; Brandstaetter, P.; Hermann, M.; Kircher, B.; Gust, R. Synthesis and Biological Evaluation of Salophen Nickel(II) and Cobalt(III) Complexes as Potential Anticancer Compounds. Arch. Pharm. 2023, 356, 2200655. [Google Scholar] [CrossRef]
- Abula, A.; Xu, Z.; Zhu, Z.; Peng, C.; Chen, Z.; Zhu, W.; Aisa, H.A. Substitution Effect of the Trifluoromethyl Group on the Bioactivity in Medicinal Chemistry: Statistical Analysis and Energy Calculations. J. Chem. Inf. Model. 2020, 60, 6242–6250. [Google Scholar] [CrossRef]
- Novás, M.; Matos, M.J. The Role of Trifluoromethyl and Trifluoromethoxy Groups in Medicinal Chemistry: Implications for Drug Design. Molecules 2025, 30, 3009. [Google Scholar] [CrossRef]
- Almeida, C.M.; Nascimento, É.C.M.; Martins, J.B.L.; da Mota, T.H.A.; de Oliveira, D.M.; Gatto, C.C. Crystal Design, Antitumor Activity and Molecular Docking of Novel Palladium(II) and Gold(III) Complexes with a Thiosemicarbazone Ligand. Int. J. Mol. Sci. 2023, 24, 11442. [Google Scholar] [CrossRef]
- Almeida, C.M.; Marcon, P.H.S.; Nascimento, É.C.M.; Martins, J.B.L.; Chagas, M.A.S.; Fujimori, M.; De Marchi, P.G.F.; França, E.L.; Honorio-França, A.C.; Gatto, C.C. Organometallic Gold (III) and Platinum (II) Complexes with Thiosemicarbazone: Structural Behavior, Anticancer Activity, and Molecular Docking. Appl. Organomet. Chem. 2022, 36, e6761. [Google Scholar] [CrossRef]
- Okuniewski, A.; Rosiak, D.; Chojnacki, J.; Becker, B. Coordination Polymers and Molecular Structures among Complexes of Mercury(II) Halides with Selected 1-Benzoylthioureas. Polyhedron 2015, 90, 47–57. [Google Scholar] [CrossRef]
- Lima, F.C.; Só, Y.A.O.; Gargano, R.; de Oliveira, D.M.; Gatto, C.C. Structural, Theoretical and Biological Activity of Mono and Binuclear Nickel(II) Complexes with Symmetrical and Asymmetrical 4,6-Diacetylresorcinol-Dithiocarbazate Ligands. J. Inorg. Biochem. 2021, 224, 111559. [Google Scholar] [CrossRef] [PubMed]
- Datta, S.; Seth, D.K.; Gangopadhyay, S.; Karmakar, P.; Bhattacharya, S. Nickel Complexes of Some Thiosemicarbazones: Synthesis, Structure, Catalytic Properties and Cytotoxicity Studies. Inorganica Chim. Acta 2012, 392, 118–130. [Google Scholar] [CrossRef]
- De Sousa, G.F.; Marques, D.R.; Resck, I.S.; Sabino, J.R. Synthesis and Crystal Structures of the Five-Coordinate Diorganotin(IV) Complex [Ph2Sn(L)]·DMF and Its 5-Hydroxypyrazoline Ligand (H2L). J. Chem. Crystallogr. 2011, 41, 528–532. [Google Scholar] [CrossRef]
- Ali, M.A.; Mirza, A.H.; Butcher, R.J.; Chowdhury, A.K. Synthesis, Spectroscopy, and X-Ray Crystal Structures of Copper(II) Complexes of the Tridentate ONS Ligand Formed by Condensation of 4,4,4-Trifluoro-1-(2-Thienyl)-2,4-Butanedione with S-Benzyldithiocarbazate. Transit. Met. Chem. 2011, 36, 471–479. [Google Scholar] [CrossRef]
- McKinnon, J.J.; Spackman, M.A.; Mitchell, A.S. Novel Tools for Visualizing and Exploring Intermolecular Interactions in Molecular Crystals. Struct. Sci. 2004, 60, 627–668. [Google Scholar] [CrossRef]
- Spackman, P.R.; Turner, M.J.; McKinnon, J.J.; Wolff, S.K.; Grimwood, D.J.; Jayatilaka, D.; Spackman, M.A. CrystalExplorer: A Program for Hirshfeld Surface Analysis, Visualization and Quantitative Analysis of Molecular Crystals. J. Appl. Crystallogr. 2021, 54, 1006–1011. [Google Scholar] [CrossRef]
- Lakshmanan, P.; Gayathri, E.; Thirumaran, S.; Ciattini, S. Synthesis, Crystal Structure, DFT and Hirshfeld Surface Analysis of Ni(II) Complexes: Precursor for Nickel Sulfide Nanoparticles. J. Mol. Struct. 2023, 1273, 134306. [Google Scholar] [CrossRef]
- Hussain Dar, S.; Hasan, N.; Rana, M.; Fatima, A.; Noorul Sabah Andrabi, S.; Ahmedi, S.; Manzoor, N.; Javed, S.; Rahisuddin. Synthesis and Spectral Studies of Ni(II) Complexes Involving Functionalized Dithiocarbamates and Triphenylphosphine: X-Ray Crystal Structure, Thermal Stability, Hirshfeld Surface Analysis, DFT and Biological Evaluation. Inorganica Chim. Acta 2023, 545, 121271. [Google Scholar] [CrossRef]
- Almeida, C.M.; de Carvalho, J.G.M.; Fujimori, M.; França, E.L.; Honorio-França, A.C.; Parreira, R.L.T.; Orenha, R.P.; Gatto, C.C. Structural Investigation of Group 10 Metal Complexes with Thiosemicarbazone: Crystal Structure, Mass Spectrometry, Hirshfeld Surface and in Vitro Antitumor Activity. Struct. Chem. 2020, 31, 2093–2103. [Google Scholar] [CrossRef]
- Lima, D.J.B.; Almeida, R.G.; Jardim, G.A.M.; Barbosa, B.P.A.; Santos, A.C.C.; Valença, W.O.; Scheide, M.R.; Gatto, C.C.; de Carvalho, G.G.C.; Costa, P.M.S.; et al. It Takes Two to Tango: Synthesis of Cytotoxic Quinones Containing Two Redox Active Centers with Potential Antitumor Activity. RSC Med. Chem. 2021, 12, 1709–1721. [Google Scholar] [CrossRef]
- Leovac, V.M.; Češljević, V.I.; Nešić, S.M. Transition Metal Complexes with the Thiosemicarbazide-based Ligands. XII. Complexes of Nickel(II) with Benzoylacetone S-methylisothiosemicarbazone and N(1)-2-butylidene-4-oxo-4-phenyl-N(4)-salicylidene-S-methylisothiosemicarbazide. Zeitschrift für Anorg. und Allg. Chemie 1991, 592, 217–223. [Google Scholar] [CrossRef]
- Almeida, C.M.; Nascimento, G.P.; Magalhães, K.G.; Iglesias, B.A.; Gatto, C.C. Crystal Structures, DNA-Binding Ability and Influence on Cellular Viability of Gold(I) Complexes of Thiosemicarbazones. J. Coord. Chem. 2018, 71, 502–519. [Google Scholar] [CrossRef]
- Cavalcante, C.D.Q.O.; Arcanjo, D.D.S.; Silva, G.G.D.; Oliveira, D.M.D.; Gatto, C.C. Solution and Solid Behavior of Mono and Binuclear Zinc(Ii) and Nickel(Ii) Complexes with Dithiocarbazates: X-Ray Analysis, Mass Spectrometry and Cytotoxicity against Cancer Cell Lines. New J. Chem. 2019, 43, 11209–11221. [Google Scholar] [CrossRef]
- Lima, F.C.; Silva, T.S.; Martins, C.H.G.; Gatto, C.C. Synthesis, Crystal Structures and Antimicrobial Activity of Dimeric Copper(II) Complexes with 2-Hydroxyphenyl-Ethylidene-Dithiocarbazates. Inorganica Chim. Acta 2018, 483, 464–472. [Google Scholar] [CrossRef]
- Haas, A.; Lieb, M.; Schelvis, M. Trifluoromethyl-Substituted Heteroolefins (Schiffs Bases): Syntheses and Chemical Properties. J. Fluor. Chem. 1997, 83, 133–143. [Google Scholar] [CrossRef]
- Colmenares, L.U.; Liu, R.S.H. 1H NMR and 19F NMR Chemical Shifts of Fluorinated Retinals, Schiff Bases and Protonated Schiff Bases. Magn. Reson. Chem. 1992, 30, 490–496. [Google Scholar] [CrossRef]
- Gokulnath, G.; Manikandan, R.; Anitha, P.; Umarani, C. Synthesis, characterization and antibacterial studies of CO(II), NI(II) and CU(II) complexes containing triphenylphosphine and schiff base ligand based on salicylaldehyde. Rasayan J. Chem. 2022, 14, 2692–2697. [Google Scholar] [CrossRef]
- Muthu Tamizh, M.; Mereiter, K.; Kirchner, K.; Ramachandra Bhat, B.; Karvembu, R. Synthesis, Crystal Structures and Spectral Studies of Square Planar Nickel(II) Complexes Containing an ONS Donor Schiff Base and Triphenylphosphine. Polyhedron 2009, 28, 2157–2164. [Google Scholar] [CrossRef]
- Mosmann, T. Rapid Colorimetric Assay for Cellular Growth and Survival: Application to Proliferation and Cytotoxicity Assays. J. Immunol. Methods 1983, 65, 55–63. [Google Scholar] [CrossRef]
- Jiang, C.; Shen, C.; Ni, M.; Huang, L.; Hu, H.; Dai, Q.; Zhao, H.; Zhu, Z. Molecular Mechanisms of Cisplatin Resistance in Ovarian Cancer. Genes Dis. 2024, 11, 101063. [Google Scholar] [CrossRef] [PubMed]
- West, D.X.; Ives, J.S.; Bain, G.A.; Liberta, A.E.; Valdés-Martínez, J.; Ebert, K.H.; Hernández-Ortega, S. Copper(II) and Nickel(II) Complexes of 2,3-Butanedione Bis(N(3)-Substituted Thiosemicarbazones). Polyhedron 1997, 16, 1895–1905. [Google Scholar] [CrossRef]
- Akbar Ali, M.; Tarafdar, M.T.H. Metal Complexes of Sulphur and Nitrogen-Containing Ligands: Complexes of s-Benzyldithiocarbazate and a Schiff Base Formed by Its Condensation with Pyridine-2-Carboxaldehyde. J. Inorg. Nucl. Chem. 1977, 39, 1785–1791. [Google Scholar] [CrossRef]
- SMART and SAINT. Area Detector Control Integration Software; Bruker AXS: Karlsruhe, Germany, 1999. [Google Scholar]
- Sheldrick, G.M. SADABS, Program for Empirical Absorption Correction of Area Detector Data; University of Göttingen: Göttingen, Germany, 1997. [Google Scholar]
- Sheldrick, G.M. A Short History of SHELX. Acta Crystallogr. Sect. A 2008, 64, 112–122. [Google Scholar] [CrossRef]
- Sheldrick, G.M. Crystal Structure Refinement with SHELXL. Acta Crystallogr. Sect. C 2015, 71, 3–8. [Google Scholar] [CrossRef] [PubMed]
- Dolomanov, O.V.; Bourhis, L.J.; Gildea, R.J.; Howard, J.A.K.; Puschmann, H. OLEX2: A Complete Structure Solution, Refinement and Analysis Program. J. Appl. Crystallogr. 2009, 42, 339–341. [Google Scholar] [CrossRef]
- MacRae, C.F.; Sovago, I.; Cottrell, S.J.; Galek, P.T.A.; McCabe, P.; Pidcock, E.; Platings, M.; Shields, G.P.; Stevens, J.S.; Towler, M.; et al. Mercury 4.0: From Visualization to Analysis, Design and Prediction. J. Appl. Crystallogr. 2020, 53, 226–235. [Google Scholar] [CrossRef] [PubMed]


| Bond Lengths (Å) | ||||
|---|---|---|---|---|
| (1) | (2) | (3) | (4) | |
| C5–O1 | 1.295(4) | 1.280(11) | 1.297(4) | 1.299(7) |
| C7–N1 | 1.324(4) | 1.312(10) | 1.321(5) | 1.325(7) |
| N1–N2 | 1.392(3) | 1.408(10) | 1.396(4) | 1.406(5) |
| N2–C9 | 1.293(4) | 1.309(11) | 1.291(5) | 1.292(7) |
| C9–S2 | 1.732(3) | 1.745(9) | 1.732(4) | 1.735(6) |
| Ni1–O1 | 1.841(2) | 1.857(6) | 1.849(3) | 1.835(4) |
| Ni1–N1 | 1.898(3) | 1.863(7) | 1.874(3) | 1.859(4) |
| Ni1–S2 | 2.112(10) | 2.143(3) | 2.137(12) | 2.128(16) |
| Ni1–PPh3 | 2.204(10) | – | 2.210(12) | – |
| Ni–Py | – | 1.919(8) | – | 1.903(5) |
| Bond Angles (°) | ||||
| (1) | (2) | (3) | (4) | |
| N1–Ni1–S2 | 87.28(8) | 86.7(2) | 87.46(10) | 88.04(14) |
| O1–Ni1–N1 | 95.98(10) | 95.6(3) | 93.99(13) | 95.82(19) |
| O1–Ni–S2 | 175.94(8) | 177.6(19) | 174.23(10) | 175.39(14) |
| O1–Ni1–PPh3 | 85.87(7) | – | 86.50(9) | – |
| S2–Ni1–PPh3 | 90.85(4) | – | 92.24(5) | – |
| N1–Ni1–PPh3 | 178.12(8) | – | 178.07(10) | – |
| O1–Ni1–Py | – | 86.7(3) | – | 86.11(18) |
| S2–Ni1–Py | – | 91.0(2) | – | 90.00(13) |
| N1–Ni1–Py | – | 174.6(3) | – | 177.96(19) |
| H2L1 | H2L2 | (1) | (2) | (3) | (4) | |
|---|---|---|---|---|---|---|
| ν(C=S) | 1307 | 1306 | – | – | – | – |
| ν(C–S) | 728 | 732 | 746 | 746 | 746 | 759 |
| ν(N–N) | 1038 | 1064 | 1097 | 1066 | 1095 | – |
| ν(C=O) | 1651 | 1637 | – | – | – | – |
| ν(C=N) | 1597 | – | 1544 | 1544 | 1544 | 1544 |
| ν(N–H) | 3333/3304 | 3195 | 3401 | 3343 | – | – |
| δ(Py) | – | – | – | 690 | – | 694 |
| ν(Ni–PPh3) | – | – | 692 | – | 692 | – |
| π–π* | Log ε | n–π* | Log ε | LMCT | Log ε | ||
|---|---|---|---|---|---|---|---|
| H2L1 | MeOH | 284 | 4.09 | 415 | 3.59 | – | – |
| DMF | 283 | 4.26 | 431 | – | – | – | |
| H2L2 | MeOH | 262 and 295 | 4.03 and 4.10 | 445 | – | – | – |
| DMF | 294 | 4.19 | 451 | – | – | – | |
| (1) | MeOH | 230 and 273 | 4.13 and 4.14 | 441 | 3.88 | 461 and 490 | 3.87 and 3.79 |
| DMF | – | – | 445 | 4.36 | 465 and 498 | 4.34 and 4.25 | |
| (2) | MeOH | 257 | 4.24 | 336 | 3.89 | 462 and 493 | 3.87 and 3.70 |
| DMF | 279 | 4.24 | 446 | 4.15 | 468 and 497 | 4.13 and 4.04 | |
| (3) | MeOH | 254 and 275 | 4.18 and 4.16 | – | – | 449 | 4.08 |
| DMF | – | – | 455 | 4.29 | 488 | 4.10 | |
| (4) | MeOH | 257 | 4.18 | 301 | 3.97 | 454 | 4.10 |
| DMF | – | – | 455 | 4.35 | 488 | 4.16 |
| Compound | 1H (ppm) | 19F (ppm) | 31P (ppm) |
|---|---|---|---|
| H2L1 | 11.72 (s, 1H, N–H), 10.19 (s, 1H, N–H), 8.12–8.10 (m, 1H, –CH), 8.10–8.08 (m, 1H, –CH), 7.53 (d, J = 7.20 Hz, 1H, Ar), 7.39 (dd, J = 7.40 and 7.20 Hz, 1H, Ar), 7.33 (dd, J = 3.89 and 3.74 Hz, 1H, Ar), 7.25 (t, J = 7.40 Hz, 1H, Ar), 4.67 (s, 2H, CH2) | −67.57 (s, CF3) | −6.50 (s, PPh3-free) |
| H2L2 | 13.20 (s, 1H, N–H), 8.10 (m, J = 4.92 Hz, 1H, –CH), 8.07 (m, J = 3.80 Hz, 1H, –CH), 7.42 (d, J = 7.50 Hz, 1H, Ar), 7.36–7.27 (m, 4H, Ar), 4.65 (s, 2H, –CH2), 4.46 (s, 2H, CH2) | −68.40 (s, CF3) | −6.50 (s, PPh3-free) |
| (1) | 9.40 (s, 1H, N–H), 7.78 (t, 6H, PPh3), 7.64–7.51 (m, 13H, Ar and PPh3), 7.20 (t, 2H, Ar), 6.92–6.89 (m, 1H, –CH), 6.87 (t, J = 7.35 Hz, 1H, Ar), 6.26 (s, 1H, –CH) | −63.63 (s, CF3) | 22.92 (s, PPh3) |
| (2) | 9.40 (s, 1H, N–H), 9.06 (s, 2H, –CH Py), 8.04 (s, 1H, –CH Py), 7.72–7.51 (m, 6H, Ar and Py), 7.21 (t, J = 7.31 Hz, 2H, Ar), 7.14–7.07 (m, 1H, –CH), 6.88 (t, J = 7.31 Hz, 1H, Ar), 6.17 (s, 1H, –CH) | −63.13 (s, CF3) | – |
| (3) | 7.80–7.65 (m, 6H, PPh3), 7.65–7.42 (m, 13H, Ar and PPh3), 7.34–7.29 (m, 4H, Ar), 7.29–7.22 (m, 1H, Ar), 4.33 (s, 2H, –CH2) | −62.04 (s, CF3) | 22.30 (s, PPh3) |
| (4) | 10.39 (s, 2H, Py), 8.16–7.79 (m, 13H, Ar and Py), 7.46–7.37 (m, 1H, Ar), 7.37–7.26 (m, 4H, Ar), 7.26–7.15 (m, 1H, Ar), 4.10 (s, 2H, –CH2) | −62.33 (s, CF3) | – |
| Compounds | IC50 (µM) | |||
|---|---|---|---|---|
| MCF-7 | A2780cis | A549 | MRC-5 | |
| H2L1 | >100 | (42.78 ± 0.66) | (51.38 ± 4.37) | >100 |
| H2L2 | >100 | (78.93 ± 1.96) | >100 | >100 |
| (1) | >100 | >100 | >100 | >100 |
| (2) | (8.48 ± 1.37) | (7.20 ± 0.73) | (27.18 ± 2.91) | (9.89 ± 0.17) |
| (3) | >100 | >100 | >100 | >100 |
| (4) | (56.54 ± 1.52) | (65.92 ± 3.52) | >100 | >100 |
| Cisplatin | (13.98 ± 2.02) | (37.03 ± 5.11) | (11.54 ± 1.19) | (29.09 ± 0.78) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pessoa, G.S.; Viana, M.P.; Oliveira, K.M.; Gatto, C.C. Comparative Study of Ni(II) Complexes with Dithiocarbazate- and Thiosemicarbazone-Based Ligands: Synthesis, Crystal Structures, and Anticancer Activity. Molecules 2025, 30, 3516. https://doi.org/10.3390/molecules30173516
Pessoa GS, Viana MP, Oliveira KM, Gatto CC. Comparative Study of Ni(II) Complexes with Dithiocarbazate- and Thiosemicarbazone-Based Ligands: Synthesis, Crystal Structures, and Anticancer Activity. Molecules. 2025; 30(17):3516. https://doi.org/10.3390/molecules30173516
Chicago/Turabian StylePessoa, Gabriel S., Mariana P. Viana, Katia M. Oliveira, and Claudia C. Gatto. 2025. "Comparative Study of Ni(II) Complexes with Dithiocarbazate- and Thiosemicarbazone-Based Ligands: Synthesis, Crystal Structures, and Anticancer Activity" Molecules 30, no. 17: 3516. https://doi.org/10.3390/molecules30173516
APA StylePessoa, G. S., Viana, M. P., Oliveira, K. M., & Gatto, C. C. (2025). Comparative Study of Ni(II) Complexes with Dithiocarbazate- and Thiosemicarbazone-Based Ligands: Synthesis, Crystal Structures, and Anticancer Activity. Molecules, 30(17), 3516. https://doi.org/10.3390/molecules30173516







