Exploration of Multiconformers to Extract Information About Structural Deformation Undergone by a Protein Target: Illustration on the Bcl-xL Target
Abstract
1. Introduction
2. Results & Discussion
2.1. Structural Variability of Bcl-xL: Identification of Conserved and Variable Regions
2.1.1. Structural Conserved Positions and Weak Variable Positions
2.1.2. Structural Variable Positions
2.2. Amino Acid Substitutions in Bcl-xL Chains Induce Limited Backbone Deformations
2.3. Ligand Binding Induces Localized and Allosteric Structural Variability
2.4. Structural Variability of the Bcl-xL Binding Pocket
2.5. Ligand-Dependent Conformations in the Binding Pocket
3. Materials and Methods
3.1. Bcl-xL Structure
3.2. Dataset Presentation
3.3. Co-Crystallized Ligand Classification
3.4. Structural Variability Quantification Using the SA-conf Tool
- The structurally conserved positions ( = 1). For these positions, all structures share the same local conformation.
- The weakly variable positions (). For these positions, one structural letter predominates. These positions exhibit rare local conformational changes across structures.
- The structurally variable positions (). Multiple structural letters are observed at these positions, indicating distinct conformations across chains.
3.5. Quantification of Residue Flexibility
3.6. Binding Site Estimation
3.7. Protein–Ligand Network
3.7.1. Step 1: Clustering of Ligand Atoms Based on Their 3D Proximity
- Intra-cluster variability: Clusters with the lowest internal variance were favored.
- Ligand atom uniqueness per protein: Each cluster should ideally contain only one ligand atom per Bcl-xL structure, to preserve one-to-one mapping at the complex level.
3.7.2. Step 2: Identification of Potential Protein–Ligand Contacts
3.7.3. Step 3: Network Construction
3.7.4. Step 4: Network Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| Bcl-xL | B-cell lymphoma-extra large |
| BH | Bcl-2 homology |
| Bnorm | Normalized B-factor |
| C | Carbon |
| HMM-SA | Hidden Markov Model-Structural Alphabet |
| MSLA | Multiple Sequence Letter Alignment |
| MSA | Multiple Sequence Alignment |
| PDB | Protein Data Bank |
| PR2 | HIV-2 Protease |
| RMSD | Root Mean Square Deviation |
| SL | Structural letters |
| X-ray | Crystallographic |
| 3D | Three-dimensional |
Appendix A
Appendix A.1. Composition of the Bcl-xL Data Set
| PDB | Ligand | PDB | Ligand | PDB | Ligand | PDB | Ligand |
|---|---|---|---|---|---|---|---|
| 1MAZ_A | Apo | 3R85_D | P | 4TUH_F | SM | 6UVF_G | SM |
| 1R2D_A | Apo | 3SP7_A | SM | 4TUH_G | SM | 6UVF_H | SM |
| 1R2E_A | Apo | 3SPF_A | SM | 4TUH_H | SM | 6UVF_I | SM |
| 1R2G_A | Apo | 3ZK6_A | SM | 4Z9V_A | P | 6UVF_J | SM |
| 1R2H_A | Apo | 3ZLN_A | SM | 4Z9V_B | P | 6UVF_K | SM |
| 1R2I_A | Apo | 3ZLO_A | SM | 5B1Z_A | P | 6UVF_L | SM |
| 2B48_A | Apo | 3ZLR_A | SM | 5B1Z_B | P | 6UVG_A | SM |
| 2P1L_A | P | 3ZLR_B | SM | 5C3G_A | P | 6UVG_B | SM |
| 2P1L_C | P | 4A1U_A | pP | 5FMJ_A | P | 6UVG_C | SM |
| 2P1L_E | P | 4A1W_A | pP | 5FMK_A | P | 6UVG_D | SM |
| 2P1L_G | P | 4A1W_B | pP | 5VX3_A | P | 6UVG_E | SM |
| 2YJ1_A | pP | 4A1W_C | pP | 5VX3_C | P | 6UVG_F | SM |
| 2YJ1_C | pP | 4A1W_D | pP | 5VX3_E | P | 6UVG_G | SM |
| 2YQ6_A | P | 4BPK_A | pP | 5VX3_G | P | 6UVG_H | SM |
| 2YQ7_A | P | 4BPK_B | pP | 6DCN_A | P | 6UVG_I | SM |
| 2YXJ_A | SM | 4C52_A | SM | 6DCN_B | P | 6UVG_J | SM |
| 2YXJ_B | SM | 4C52_B | SM | 6DCO_A | P | 6UVG_K | SM |
| 3CVA_X | Apo | 4C5D_A | SM | 6DCO_B | P | 6UVG_L | SM |
| 3FDL_A | P | 4C5D_B | SM | 6RNU_A | SM | 6UVH_A | SM |
| 3FDM_A | pP | 4CIN_A | P | 6RNU_B | SM | 6UVH_B | SM |
| 3FDM_B | pP | 4CIN_B | P | 6UVC_A | SM | 6UVH_C | SM |
| 3FDM_C | pP | 4EHR_A | SM | 6UVC_B | SM | 6UVH_D | SM |
| 3INQ_A | SM | 4HNJ_B | Apo | 6UVD_A | SM | 6VWC_A | SM |
| 3INQ_B | SM | 4PPI_A | Apo | 6UVD_B | SM | 6VWC_B | SM |
| 3IO8_A | P | 4QVE_A | P | 6UVE_A | SM | 6YLI_A | P |
| 3IO8_C | P | 4QVF_A | P | 6UVE_B | SM | 7CA4_A | Apo |
| 3PL7_A | P | 4QVX_A | SM | 6UVE_C | SM | 7JGV_A | SM |
| 3PL7_B | P | 4QVX_B | SM | 6UVF_A | SM | 7JGV_B | SM |
| 3QKD_A | SM | 4TUH_A | SM | 6UVF_B | SM | 7JGW_A | SM |
| 3QKD_B | SM | 4TUH_B | SM | 6UVF_C | SM | 7LH7_A | SM |
| 3R85_A | P | 4TUH_C | SM | 6UVF_D | SM | 7LH7_B | SM |
| 3R85_B | P | 4TUH_D | SM | 6UVF_E | SM | ||
| 3R85_C | P | 4TUH_E | SM | 6UVF_F | SM |
Appendix A.2. Long Range Impact of Mutations on the Bcl-xL Backbone Conformation
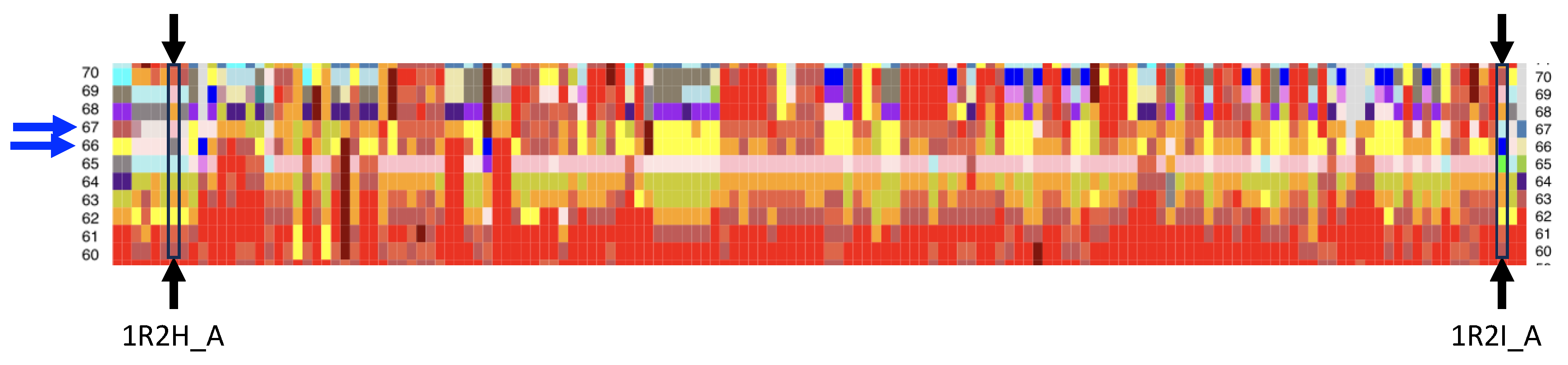
Appendix A.3. Local Conformations of the Pocket Extracted from the 130 Bcl-xL Chains
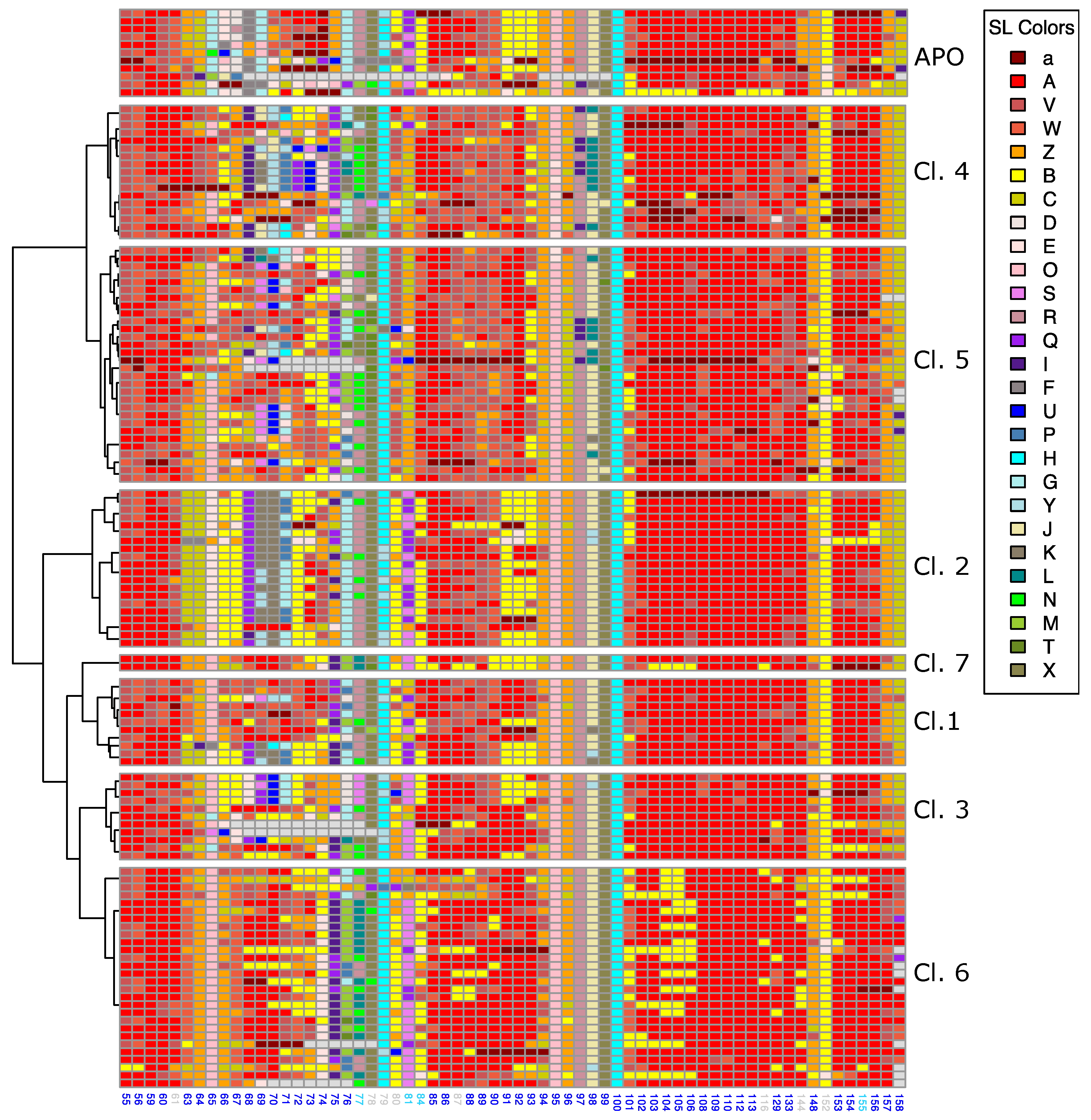
Appendix A.4. Multiple Sequence Alignment of the 60 X-Ray Structures Computed by SA-conf
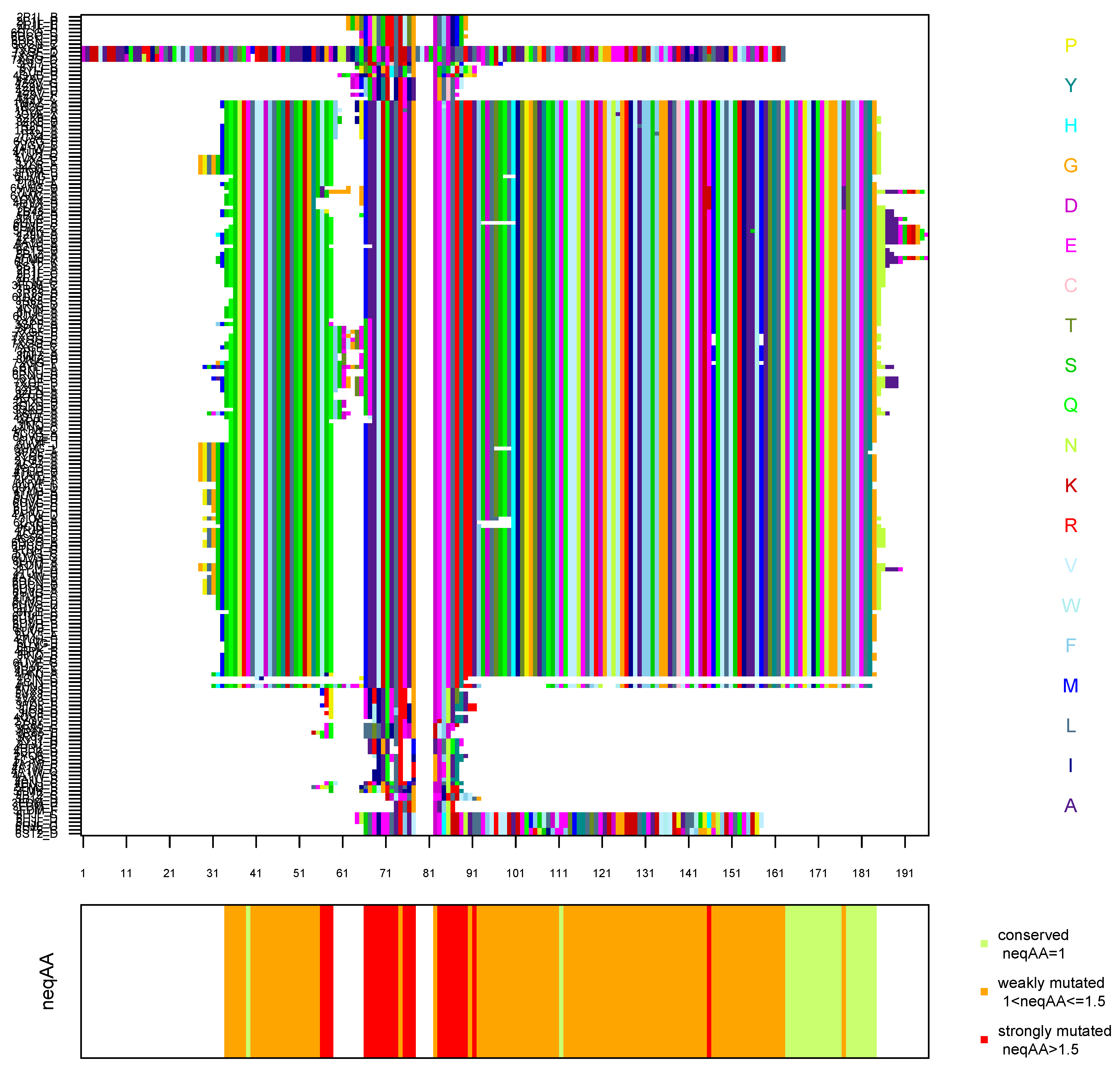
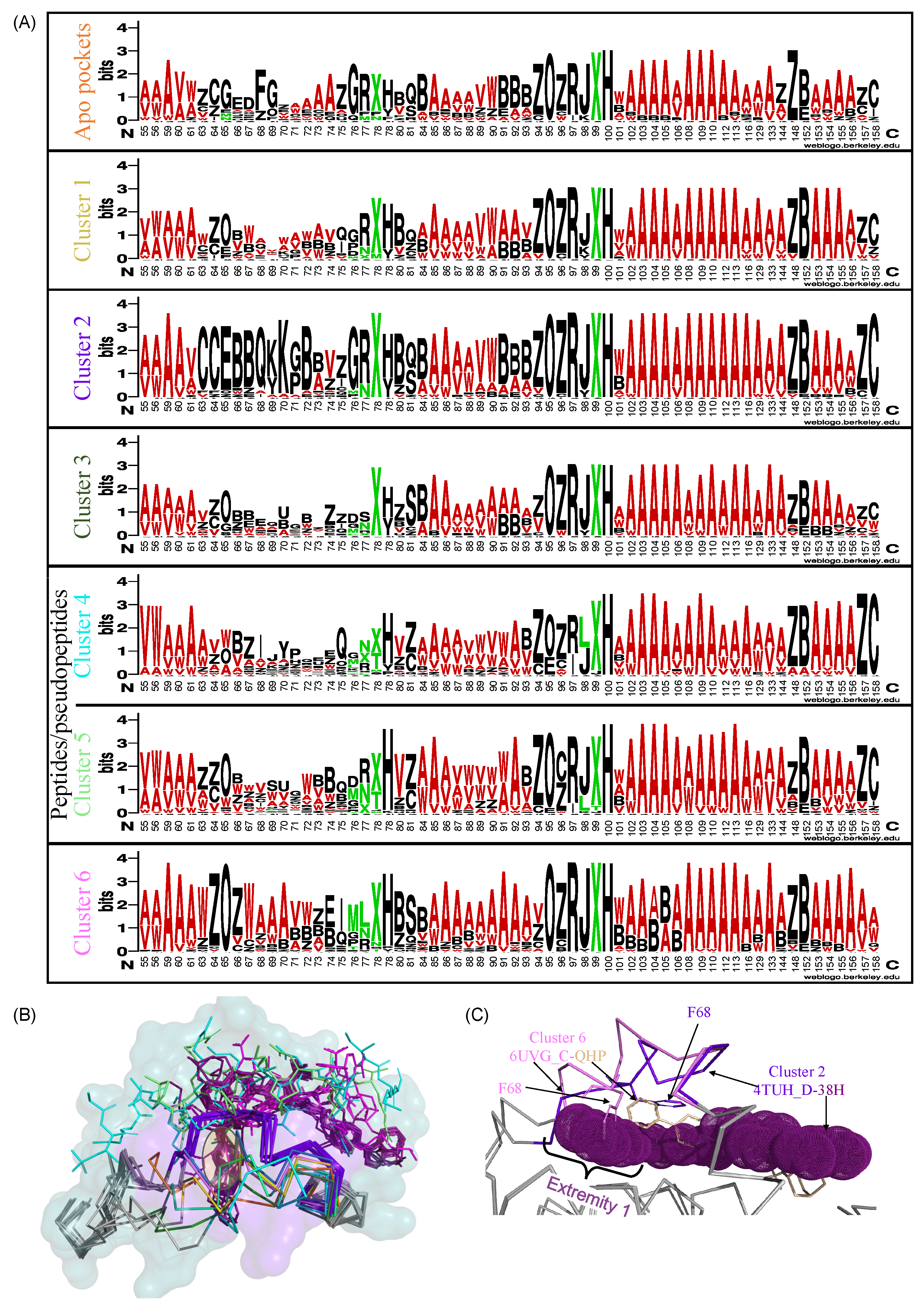
References
- Berman, H.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.; Weissig, H.; Shindyalov, I.N.; Bourne, P. The Protein Data Bank. Nucl. Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Murray, C.W.; Baxter, C.A.; Frenkel, A.D. The sensitivity of the results of molecular docking to induced fit effects: Application to thrombin, thermolysin and neuraminidase. J. Comput.-Aided Mol. Des. 1999, 13, 547–562. [Google Scholar] [CrossRef] [PubMed]
- Davis, A.M.; Teague, S.J. Hydrogen bonding, hydrophobic interactions, and failure of the rigid receptor hypothesis. Angew. Chem. Int. Ed. Engl. 1999, 38, 736–749. [Google Scholar] [CrossRef]
- Najmanovich, R.; Kuttner, J.; Sobolev, V.; Edelman, M. Side chain flexibility in proteins upon ligand binding. Proteins Struct. Funct. Bioinform. 2000, 39, 261–268. [Google Scholar] [CrossRef]
- Verkhivker, G.M.; Bouzida, D.; Glhaar, D.K.; Rejto, P.A.; Arthurs, S.; Colson, A.B.; Freer, S.T.; Larson, V.; Luty, B.A.; Marrone, T.; et al. Deciphering common failures in molecular docking of ligand–protein complexes. J. Comput.-Aided Mol. Des. 2000, 14, 731–751. [Google Scholar] [CrossRef]
- Halperin, I.; Ma, B.; Wolfson, H.; Nussinov, R. Principles of docking: An overview of search algorithms and a guide to scoring functions. Proteins Struct. Funct. Bioinform. 2002, 47, 409–443. [Google Scholar] [CrossRef]
- Brooijmans, N.; Kuntz, I.D. Molecular recognition and docking algorithms. Annu. Rev. Biophys. Biomol. Struct. 2003, 32, 335–373. [Google Scholar] [CrossRef]
- Teague, S.J. Implications of protein flexibility for drug discovery. Nat. Rev. Drug Discov. 2003, 2, 527–534. [Google Scholar] [CrossRef]
- Gutteridge, A.; Thornton, J. Conformational changes observed in enzyme crystal structures upon substrate binding. J. Mol. Biol. 2005, 346, 21–28. [Google Scholar] [CrossRef]
- Brylinski, M.; Skolnick, J. What is the relationship between the global structures of apo and holo proteins? Proteins 2008, 70, 363–377. [Google Scholar] [CrossRef]
- Clark, J.J.; Benson, M.L.; Smith, R.D.; Carlson, H.A. Inherent versus induced protein flexibility: Comparisons within and between apo and holo structures. PLoS Comput. Biol. 2019, 15, e1006705. [Google Scholar] [CrossRef] [PubMed]
- Amemiya, T.; Koike, R.; Fuchigami, S.; Ikeguchi, M.; Kidera, A. Classification and annotation of the relationship between protein structural change and ligand binding. J. Mol. Biol. 2011, 408, 568–584. [Google Scholar] [CrossRef] [PubMed]
- Regad, L.; Chéron, J.; Triki, D.; Senac, C.; Flatters, D.; Camproux, A. Exploring the potential of a structural alphabet-based tool for mining multiple target conformations and target flexibility insight. PLoS ONE 2017, 12, e0182972. [Google Scholar] [CrossRef]
- Camproux, A.; Gautier, R.; Tufféry, P. A hidden Markov model derived structural alphabet for proteins. J. Mol. Biol. 2004, 339, 591–605. [Google Scholar] [CrossRef] [PubMed]
- Triki, D.; Fartek, S.; Visseaux, B.; Descamps, D.; Camproux, A.; Regad, L. Characterizing the structural variability of HIV-2 protease upon the binding of diverse ligands using a structural alphabet approach. J. Biomol. Struct. Dyn. 2019, 37, 4658–4670. [Google Scholar] [CrossRef]
- Reed, J. Bcl-2 family proteins. Oncogene 1998, 17, 3225–3236. [Google Scholar] [CrossRef]
- Certo, M.; Del Gaizo Moore, V.; Nishino, M.; Wei, G.; Korsmeyer, S.; Armstrong, S.; Letai, A. Mitochondria primed by death signals determine cellular addiction to antiapoptotic BCL-2 family members. Cancer Cell. 2006, 9, 351–365. [Google Scholar] [CrossRef]
- Feng, Y.; Zhang, L.; Hu, T.; Shen, X.; Ding, J.; Chen, K.; Jiang, H.; Liu, D. A conserved hydrophobic core at Bcl-xL mediates its structural stability and binding affinity with BH3-domain peptide of pro-apoptotic protein. Arch. Biochem. Biophys. 2009, 484, 46–54. [Google Scholar] [CrossRef]
- Manion, M.; O’Neill, J.; Giedt, C.; Kim, K.; Zhang, K.; Hockenbery, D. Bcl-XL mutations suppress cellular sensitivity to antimycin A. J. Biol. Chem. 2004, 279, 2159–2165. [Google Scholar] [CrossRef]
- Wakui, N.; Yoshino, R.; Yasuo, N.; Ohue, M.; Sekijima, M. Exploring the selectivity of inhibitor complexes with Bcl-2 and Bcl-XL: A molecular dynamics simulation approach. J. Mol. Graph. Model. 2018, 79, 166–174. [Google Scholar] [CrossRef]
- Salam, A.; Nayek, U.; Sunil, D. Homology Modeling and Docking Studies of Bcl-2 and Bcl-xL with Small Molecule Inhibitors: Identification and Functional Studies. Curr. Top. Med. Chem. 2018, 18, 2633–2663. [Google Scholar] [CrossRef]
- Maity, A.; Majumdar, S.; Dastidar, S. Flexibility enables to discriminate between ligands: Lessons from structural ensembles of Bcl-xl and Mcl-1. Comput. Biol. Chem. 2018, 77, 17–27. [Google Scholar] [CrossRef]
- Lee, E.; Czabotar, P.; Smith, B.; Deshayes, K.; Zobel, K.; Colman, P.M.; Fairlie, W. Crystal structure of abt-737 complexed with bcl-xl: Implications for selectivity of antagonists of the bcl-2 family. Cell Death Differ. 2007, 14, 1711–1713. [Google Scholar] [CrossRef] [PubMed]
- Thébault, S.; Agez, M.; Chi, X.; Stojko, J.; Cura, V.; Telerman, S.B.; Maillet, L.; Gautier, F.; Billas-Massobrio, I.; Birck, C.; et al. TCTP contains a BH3-like domain, which instead of inhibiting, activates Bcl-xL. Sci. Rep. 2016, 6, 19725. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.Y.; Wang, S. Analysis of Flexibility and Hotspots in Bcl-xL and Mcl-1 Proteins for the Design of Selective Small-Molecule Inhibitors. ACS Med. Chem. Lett. 2012, 3, 308–312. [Google Scholar] [CrossRef] [PubMed]
- Aguirre, C.; Ten Brink, T.; Walker, O.; Guillière, F.; Davesne, D.; Krimm, I. BcL-xL conformational changes upon fragment binding revealed by NMR. PLoS ONE 2013, 8, e64400. [Google Scholar] [CrossRef]
- Crooks, G.; Hon, G.; Chandonia, J.; Brenner, S. WebLogo: A sequence logo generator. Genome Res. 2004, 14, 1188–1190. [Google Scholar] [CrossRef]
- Sattler, M.; Liang, H.; Nettesheim, J.; Meadows, R.; Yoon, H.; Chung, S.; Thompson, J.; Nathanson, S.; Wickramasinghe, S.; Korsmeyer, S.; et al. Structure of Bcl-xL–Bak peptide complex: Recognition between regulators of apoptosis. Science 1997, 275, 983–986. [Google Scholar] [CrossRef]
- Oberstein, A.; Jeffrey, P.; Shi, Y. Crystal structure of the Bcl-XL-Beclin 1 peptide complex: Beclin 1 is a novel BH3-only protein. J. Biol. Chem. 2007, 282, 13123–13132. [Google Scholar] [CrossRef]
- Feng, W.; Huang, S.; Wu, H.; Zhang, M. Molecular basis of Bcl-xL’s target recognition versatility revealed by the structure of Bcl-xL in complex with the BH3 domain of Beclin-1. J. Mol. Biol. 2007, 372, 223–235. [Google Scholar] [CrossRef]
- Petros, A.; Dinges, D.; Augeri, J.; Baumeister, B.; Betebenner, B.; Bures, W.; Elmore, D.; Hajduk, J.; Harlan, E.; Joseph, M.; et al. Solution structure of the antiapoptotic protein bcl-2. Proc. Natl. Acad. Sci. USA 2001, 98, 3012–3017. [Google Scholar] [CrossRef] [PubMed]
- Muchmore, S.; Sattler, M.; Liang, H.; Meadows, R.; Harlan, J.; Yoon, H.; Nettesheim, D.; Chang, B.; Thompson, C.; Wong, S.L.; et al. X-ray and NMR structure of human Bcl-xL, an inhibitor of programmed cell death. Nature 1996, 381, 335–341. [Google Scholar] [CrossRef] [PubMed]
- Aritomi, M.; Kunishima, N.; Inohara, N.; Ishibashi, Y.; Ohta, S.; Morikawa, K. Crystal structure of rat Bcl-xL. Implications for the function of the Bcl-2 protein family. J. Biol. Chem. 1997, 272, 27886–27892. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, J.; Manion, M.; Maguire, B.; Hockenbery, D. BCL-XL dimerization by three-dimensional domain swapping. J. Mol. Biol. 2006, 356, 367–381. [Google Scholar] [CrossRef]
- Follis, A.; Chipuk, J.; Fisher, J.; Yun, M.K.; Grace, C.; Nourse, A.; Baran, K.; Ou, L.; Min, L.; White, S.; et al. PUMA binding induces partial unfolding within BCL-xL to disrupt p53 binding and promote apoptosis. Nat. Chem. Biol. 2013, 9, 163–168. [Google Scholar] [CrossRef]
- Tanaka, Y.; Aikawa, K.; Nishida, G.; Homma, M.; Sogabe, S.; Igaki, S.; Hayano, Y.; Sameshima, T.; Miyahisa, I.; Kawamoto, T.; et al. Discovery of potent Mcl-1/Bcl-xL dual inhibitors by using a hybridization strategy based on structural analysis of target proteins. J. Med. Chem. 2013, 56, 9635–9645. [Google Scholar] [CrossRef]
- Rajan, S.; Choi, M.; Baek, K.; Yoon, H. Bh3 induced conformational changes in Bcl-Xl revealed by crystal structure and comparative analysis. Proteins 2015, 83, 1262–1272. [Google Scholar] [CrossRef]
- RDKit: Open-Source Cheminformatics. Available online: https://www.rdkit.org (accessed on 4 October 2023).
- Larkin, M.; Blackshields, G.; Brown, N.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. ClustalW and ClustalX version 2. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Regad, L.; Guyon, F.; Maupetit, J.; Tufféry, P.; Camproux, A.C. A Hidden Markov Model applied to the protein 3D structure analysis. Comput. Stat. Data Anal. 2008, 52, 3198–3207. [Google Scholar] [CrossRef]
- Drenth, J. Principles of Protein X-ray Crystallography; Springer: New York, NY, USA, 1994; pp. 189–198. [Google Scholar]
- Csàrdi, G.; Nepusz, T. The igraph software package for complex network research. InterJournal Complex Syst. 2006, 1695, 1–9. [Google Scholar]




| Aligned Position | PDB Residue | Position Category | Substitution | SL in Substituted Sequences | SL Distribution |
|---|---|---|---|---|---|
| 6 | - | / | M → H | 7XGG_F: N | / |
| 29 | 24 | Structurally variable | W → A | 6VWC_A: N | E: 1 F: 3 G: 16 I: 1 K: 1 |
| 6VWC_B: NA | M: 1 N: 1 P: 118 | ||||
| 7LH7_A: X | X: 1 NA: 6 | ||||
| 31 | 26 | Structurally variable | Q → G | 7LH7_A: U | A: 1 B: 17 D: 2 E: 10 H: 92 O: 1 P: 3 U: 1 X: 1 Y: 6 Z: 2 NA: 13 |
| 55 | 92 | Structurally variable | E → L | 1R2E_A: A | a: 2 A: 69 B: 1 V: 71 W: 6 |
| 60 | 97 | Structurally variable | F → W | 1R2G_A: V | a: 2 A: 100 B: 2 V: 20 W: 25 |
| 100 | 137 | Structurally conserved | W → A | 3CVA_X: H | H: 149 |
| 105 | 142 | Structurally variable | A → L | 1R2H_A: A | a: 8 A: 122 B: 18 V: 1 |
| 109 | 146 | Weakly structurally variable | F → L | 1R2I_A: A | a: 4 A: 136 V: 8 W: 1 |
| 121 | 158 | Weakly structurally variable | E → K | 4QVX_A: D | A: 1 D: 142 H: 1 V: 1 W: 4 |
| 4QVX_B: D | |||||
| 6VWC_A: D | |||||
| 6VWC_B: D | |||||
| 7LH7_A: D | |||||
| 7LH7_B: D | |||||
| 131 | 168 | Structurally variable | A → S | 5FMK_A: A | a: 1 A: 128 B: 3 V: 14 W: 3 |
| 152 | 189 | Weakly structurally variable | D → A | 4QVX_A: B | a: 2 B: 133 E: 14 |
| 4QVX_B: B | |||||
| 6VWC_B: B | |||||
| 6VWC_A: B | |||||
| 7LH7_A: B | |||||
| 7LH7_B: B |
| Aligned Position Type | Average Number of Links | Average Conservation of Link | Average Size of Ligand Clusters |
|---|---|---|---|
| Rigid and structurally variable positions | 9.83 (8.8) | 0.26 (0.05) | 66.64 (9.58) |
| Flexible and structurally variable positions | 8.18 (8.41) | 0.16 (0.03) | 54.64 (11.75) |
| Structurally conserved positions | 6.75 (4.92) | 0.22 (0.03) | 62.79 (4.89) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Baillif, M.; Tempez, E.; Badel, A.; Regad, L. Exploration of Multiconformers to Extract Information About Structural Deformation Undergone by a Protein Target: Illustration on the Bcl-xL Target. Molecules 2025, 30, 3355. https://doi.org/10.3390/molecules30163355
Baillif M, Tempez E, Badel A, Regad L. Exploration of Multiconformers to Extract Information About Structural Deformation Undergone by a Protein Target: Illustration on the Bcl-xL Target. Molecules. 2025; 30(16):3355. https://doi.org/10.3390/molecules30163355
Chicago/Turabian StyleBaillif, Marine, Eliott Tempez, Anne Badel, and Leslie Regad. 2025. "Exploration of Multiconformers to Extract Information About Structural Deformation Undergone by a Protein Target: Illustration on the Bcl-xL Target" Molecules 30, no. 16: 3355. https://doi.org/10.3390/molecules30163355
APA StyleBaillif, M., Tempez, E., Badel, A., & Regad, L. (2025). Exploration of Multiconformers to Extract Information About Structural Deformation Undergone by a Protein Target: Illustration on the Bcl-xL Target. Molecules, 30(16), 3355. https://doi.org/10.3390/molecules30163355





