Extraction, Purification, and Elucidation of Six Ginkgol Homologs from Ginkgo biloba Sarcotesta and Evaluation of Their Anticancer Activities
Abstract
1. Introduction
2. Results and Discussion
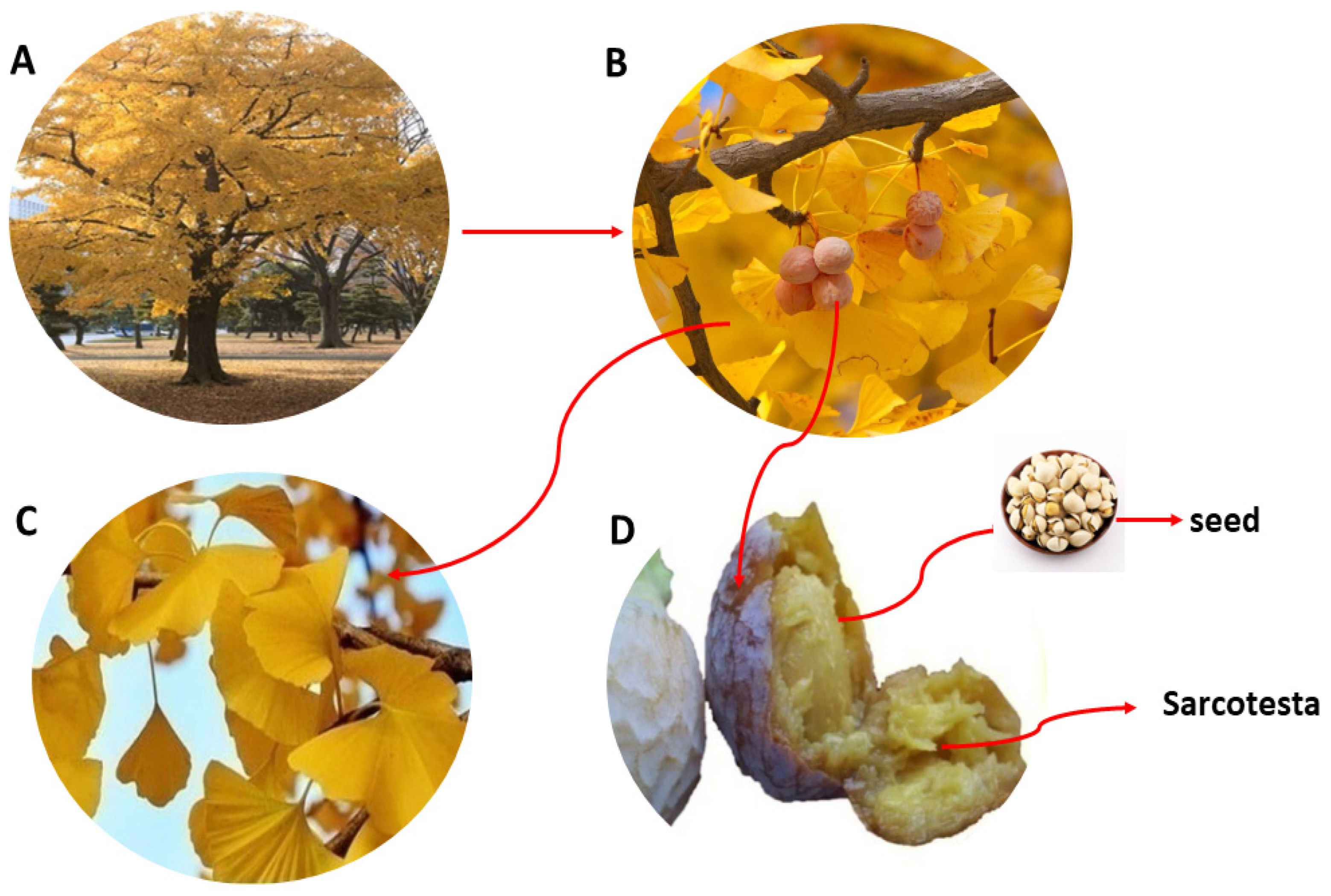
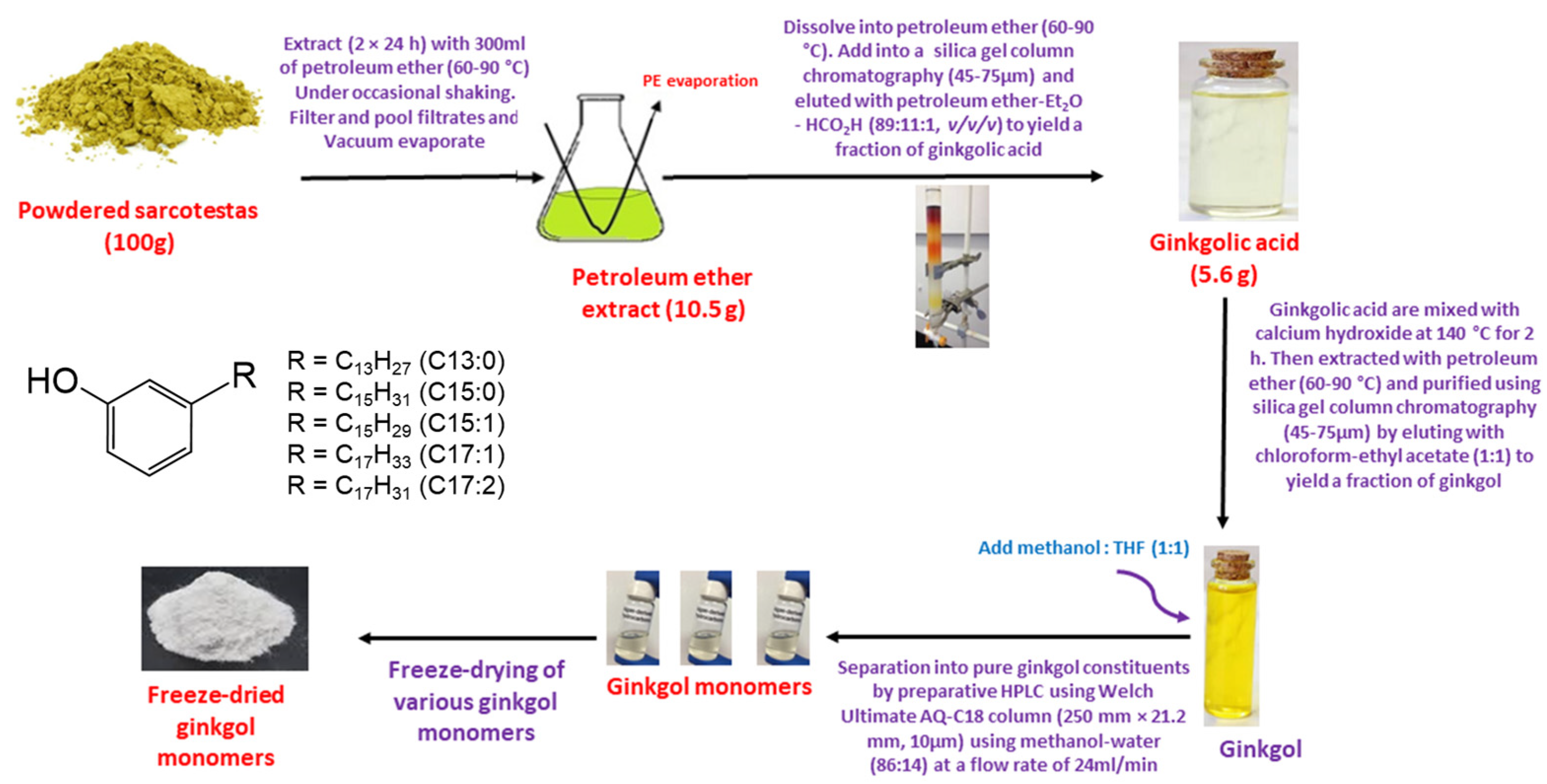
2.1. Preparation of Ginkgol
2.2. Identification of Ginkgols
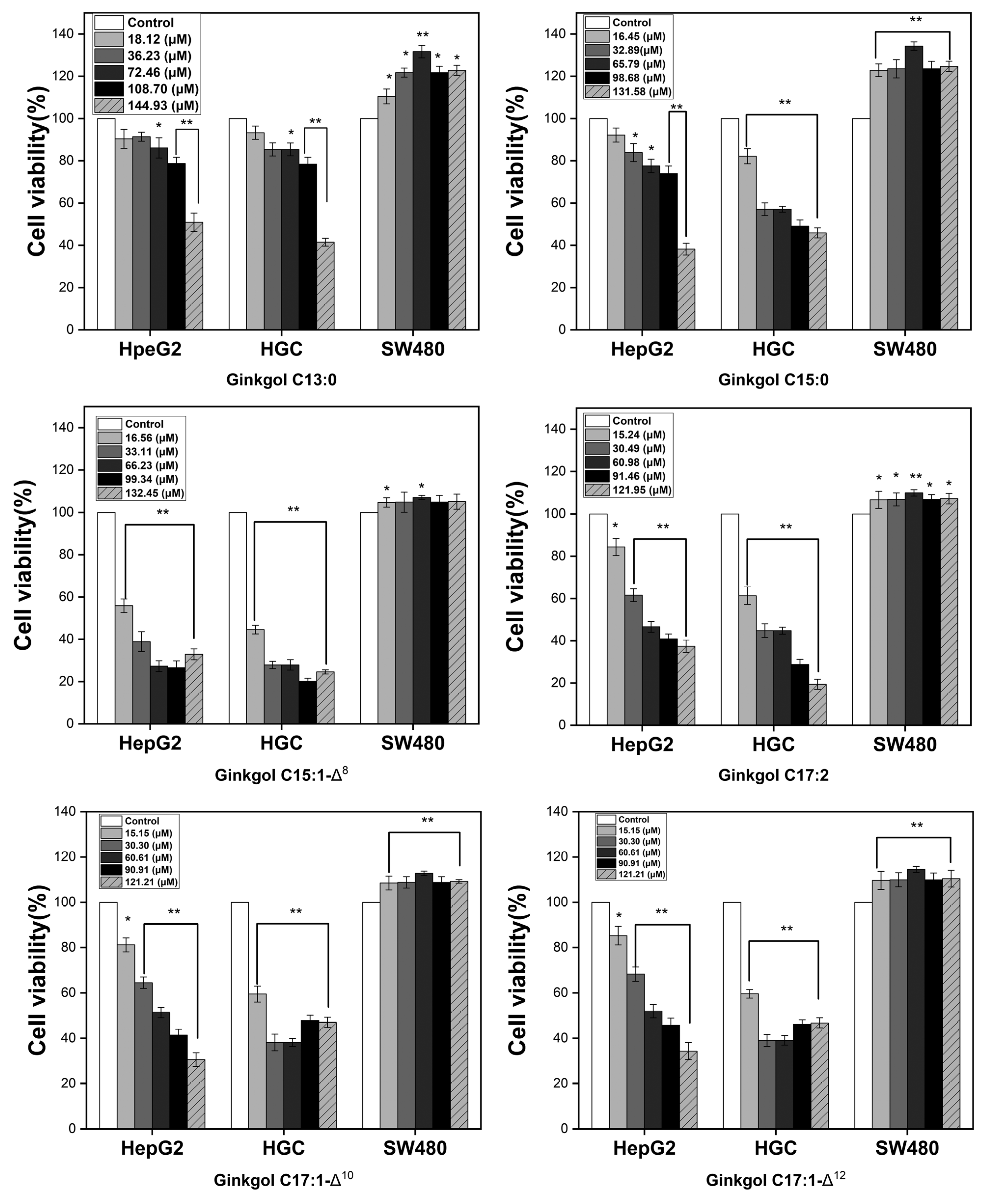
2.3. Anticancer Activity of Ginkgol Monomers
3. Materials and Methods
3.1. Materials and Chemicals
3.2. Extraction and Decarboxylation of Gingkolic Acids
3.3. Isolation and Purification of Ginkgol Monomers
3.4. Structural Characterization of Ginkgol Monomers
3.4.1. Ultraviolet (UV), Fourier Transform Infrared (FTIR), and NMR Spectra
3.4.2. GC-MS Analysis
3.4.3. Assessment of Ginkgols’ Olefinic Chains
3.5. Anticancer Activity Assay
3.6. Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| GB | Ginkgo biloba |
| GA | Ginkgolic acid |
| GBE | Ginko biloba extract |
| HPLC | High-performance liquid chromatography |
| RP-HPLC | Reversed-phase high-performance liquid chromatography |
| UV | Ultraviolet |
| FTIR | Fourier transform infrared |
| GC-MS | Gas chromatograph mass spectrometer |
| 1H-NMR | Proton nuclear magnetic resonance |
| 13C-NMR | Carbon-13 nuclear magnetic resonance |
| IC50 | Half-maximal inhibition concentration |
| MTT | 3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2-H-tetrazolium bromide |
| HepG2 | Human hepatocellular carcinomas |
| TIC | Total ion chromatogram |
| MEP | Methyl esters product |
| SW480 | Colon cancer |
| HGC | Human gastric cancer |
| DMSO | Dimethyl sulfoxide |
| DMEM | Dulbecco’s modified eagle medium |
References
- Boateng, I.D. A critical review of ginkgolic acids in Ginkgo biloba leaf extract (EGb): Toxicity and technologies to remove ginkgolic acids and their promising bioactivities. Food Funct. 2022, 13, 9226–9242. [Google Scholar] [CrossRef] [PubMed]
- Boateng, I.D. A critical review of current technologies used to reduce ginkgotoxin, ginkgotoxin-5′-glucoside, ginkgolic acid, allergic glycoprotein, and cyanide in Ginkgo biloba L. seed. Food Chem. 2022, 382, 132408. [Google Scholar] [CrossRef] [PubMed]
- Boateng, I.D.; Yang, X.-M. Ginkgo biloba L. seed; A comprehensive review of bioactives, toxicants, and processing effects. Ind. Crops Prod. 2022, 176, 114281. [Google Scholar] [CrossRef]
- Liu, L.; Wang, Y.; Zhang, J.; Wang, S. Advances in the chemical constituents and chemical analysis of Ginkgo biloba leaf, extract, and phytopharmaceuticals. J. Pharm. Biomed. Anal. 2021, 193, 113704. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.G.; Lu, X.; Gao, W.; Li, P.; Yang, H. Structure, synthesis, biosynthesis, and activity of the characteristic compounds from Ginkgo biloba L. Nat. Prod. Rep. 2022, 39, 474–511. [Google Scholar] [CrossRef] [PubMed]
- Berg, K.; Braun, C.; Krug, I.; Schrenk, D. Evaluation of the cytotoxic and mutagenic potential of three ginkgolic acids. Toxicology 2015, 327, 47–52. [Google Scholar] [CrossRef]
- Ryckewaert, L.; Sacconnay, L.; Carrupt, P.-A.; Nurisso, A.; Simões-Pires, C. Non-specific SIRT inhibition as a mechanism for the cytotoxicity of ginkgolic acids and urushiols. Toxicol. Lett. 2014, 229, 374–380. [Google Scholar] [CrossRef]
- Chen, H.Y.; Cheng, K.C.; Hsu, R.J.; Hsieh, C.W.; Wang, H.T.; Ting, Y.W. Enzymatic degradation of ginkgolic acid by laccase immobilized on novel electrospun nanofiber mat. J. Sci. Food Agric. 2020, 100, 2705–2712. [Google Scholar] [CrossRef]
- Qi, L.W.S.; Yan, J.; Fuliang, C.; Guibin, W.; Lginguo, Z. Extraction and biodegradation of ginkgolic acids from Ginkgo biloba sarcotestae. Front. Agric. Sci. Eng. 2017, 4, 465–472. [Google Scholar]
- Liu, Y.; Xin, H.; Zhang, Y.; Che, F.; Shen, N.; Cui, Y. Leaves, seeds and exocarp of Ginkgo biloba L. (Ginkgoaceae): A comprehensive review of traditionalues, phytochemistry, pharmacology, resource utilization and toxicity. J. Ethnopharmacol. 2022, 298, 115645. [Google Scholar] [CrossRef]
- Yang, X.-M.; Wang, Y.-F.; Li, Y.-Y.; Ma, H.-L. Thermal stability of ginkgolic acids from Ginkgo biloba and the effects of ginkgol C17:1 on the apoptosis and migration of SMMC7721 cells. Fitoterapia 2014, 98, 66–76. [Google Scholar] [CrossRef]
- Fang, Y.-Y.; Yang, X.-M.; Li, Y.-Y.; Feng, C.-L. Spectroscopic studies on the interaction of bovine serum albumin with Ginkgol C15:1 from Ginkgo biloba L. J. Lumin. 2015, 162, 203–211. [Google Scholar] [CrossRef]
- Li, Y.Y.; Liu, J.; Liu, Y.L.; Yang, X.M.; Huang, B.Z.; Chen, M. Inhibitory effect of ginkgol C17:1 on the biological behavior of tumor cells. Oncol. Lett. 2017, 13, 1873–1879. [Google Scholar] [CrossRef]
- Lepoittevin, J.P.; Benezra, C.; Asakawa, Y. Allergic contact dermatitis to Ginkgo biloba L.: Relationship with urushiol. Arch. Dermatol. Res. 1989, 281, 227–230. [Google Scholar] [CrossRef]
- van Beek, T.A. Chemical analysis of Ginkgo biloba leaves and extracts. J. Chromatogr. A 2002, 967, 21–55. [Google Scholar] [CrossRef]
- van Beek, T.A.; Wintermans, M.S. Preparative isolation and dual column high-performance liquid chromatography of ginkgolic acids from Ginkgo biloba. J. Chromatogr. A. 2001, 930, 109–117. [Google Scholar] [CrossRef]
- Stefanuto, P.-H.; Smolinska, A.; Focant, J.-F. Advanced chemometric and data handling tools for GC×GC-TOF-MS: Application of chemometrics and related advanced data handling in chemical separations. TrAC Trends Anal. Chem. 2021, 139, 116251. [Google Scholar] [CrossRef]
- Boateng, I.D.; Yang, X.-M. Thermal and non-thermal processing affect Maillard reaction products, flavor, and phytochemical profiles of Ginkgo biloba seed. Food Biosci. 2021, 41, 101044. [Google Scholar] [CrossRef]
- Li, S.-C.; Yang, X.-M.; Ma, H.-L.; Yan, J.-K.; Guo, D.-Z. Purification, characterization and antitumor activity of polysaccharides extracted from Phellinus igniarius mycelia. Carbohydr. Polym. 2015, 133, 24–30. [Google Scholar] [CrossRef]
- Irie, J.; Murata, M.; Homma, S. Glycerol-3-phosphate dehydrogenase inhibitors, anacardic acids, from Ginkgo biloba. Biosci. Biotechnol. Biochem. 1996, 60, 240–243. [Google Scholar] [CrossRef]
- Itokawa, H.; Totsuka, N.; Nakahara, K.; Takeya, K.; Lepoittevin, J.-P.; Asakawa, Y. Antitumor Principles from Ginkgo biloba L. Chem. Pharm. Bull. 1987, 35, 3016–3020. [Google Scholar] [CrossRef]
- Oguadinma, P.; Bilodeau, F.; LaPlante, S.R. NMR strategies to support medicinal chemistry workflows for primary structure determination. Bioorg. Med. Chem. Lett. 2017, 27, 242–247. [Google Scholar] [CrossRef]
- Gellerman, J.L.; Schlenk, H. Methods for isolation and determination of anacardic acids. Anal. Chem. 1968, 40, 739–743. [Google Scholar] [CrossRef]
- Rossi, R.; Carpita, A.; Quirici, M.G.; Veracini, C.A. Insect pheromone components: Use of 13C NMR spectroscopy for assigning the configuration of C=C double bonds of monoenic or dienic pheromone components and for quantitative determination of Z/E mixtures. Tetrahedron 1982, 38, 639–644. [Google Scholar] [CrossRef]
- Sun, Y.; Tang, C.L.; Wu, X.W.; Pan, Z.F.; Wang, L.L. Characterization of alkylphenol components in Ginkgo biloba sarcotesta by thermochemolysis-gas chromatography/mass spectrometry in the presence of trimethylsulfonium hydroxide. Chromatographia 2012, 75, 387–395. [Google Scholar] [CrossRef]
- Qiao, L.; Zheng, J.; Jin, X.; Wei, G.; Wang, G.; Sun, X.; Li, X. Ginkgolic acid inhibits the invasiveness of colon cancer cells through AMPK activation. Oncol. Lett. 2017, 14, 5831–5838. [Google Scholar] [CrossRef]
- Ji, S.L.; You, S.C.; Park, E.J.; Kim, J.; Oh, W.K.; Lee, H.S.; Ahn, J.S. Phospholipase Cγ1 Inhibitory Principles from the Sarcotestas of Ginkgo biloba. J. Nat. Prod. 1998, 61, 867–871. [Google Scholar]
- Kim, M.-G.; Shin, H.-S.; Kim, J.-H. Exploring structure-activity relationships for the in vitro cytotoxicity of alkylphenols (APs) toward HeLa cell. Mol. Cell. Toxicol. 2009, 5, 14–22. [Google Scholar]
- Li, Y.Y.; Zhang, X.C.; Yang, X.M.; Liu, J.; Li, L.J.; Ma, W.B.; Chen, M. Differential effects of ginkgol C17:1 on cisplatin-induced cytotoxicity: Protecting human normal L02 hepatocytes versus sensitizing human hepatoma HepG2 cells. Oncol. Lett. 2019, 17, 3181–3190. [Google Scholar] [CrossRef]
- Liu, J.; Li, Y.Y.; Yang, X.M.; Dong, Y.; Wu, J.; Chen, M. Effects of ginkgol C17:1 on cisplatin-induced autophagy and apoptosis in HepG2 cells. Oncol. Lett. 2018, 15, 1021–1029. [Google Scholar] [CrossRef]
- Boulikas, T.; Vougiouka, M. Cisplatin and platinum drugs at the molecular level. (Review). Oncol. Rep. 2003, 10, 1663–1682. [Google Scholar] [CrossRef] [PubMed]
- Paramashivappa, R.; Kumar, P.P.; Vithayathil, P.J.; Rao, A.S. Novel method for isolation of major phenolic constituents from cashew (Anacardium occidentale L.) nut shell liquid. J. Agric. Food Chem. 2001, 49, 2548–2551. [Google Scholar] [CrossRef] [PubMed]

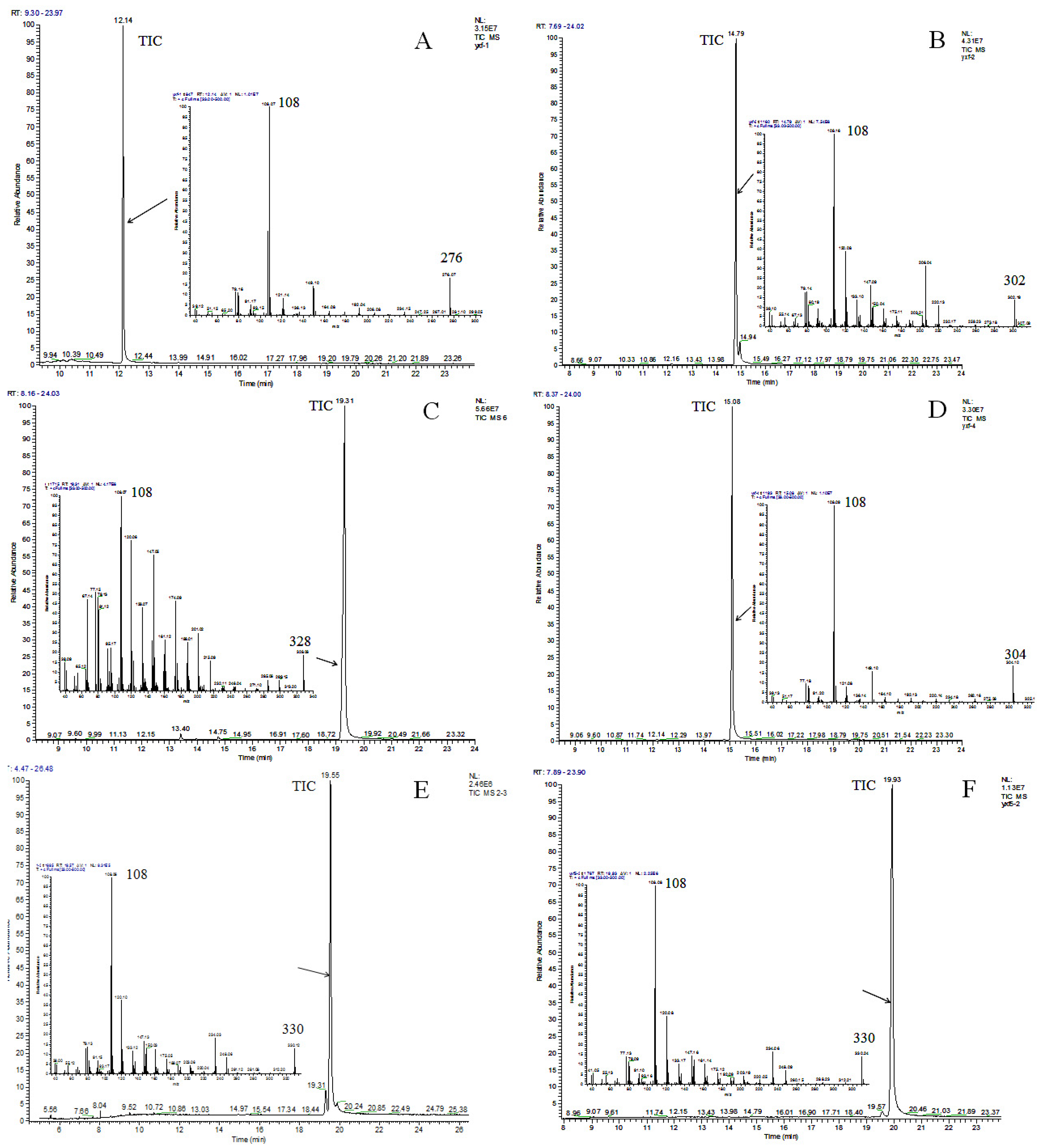
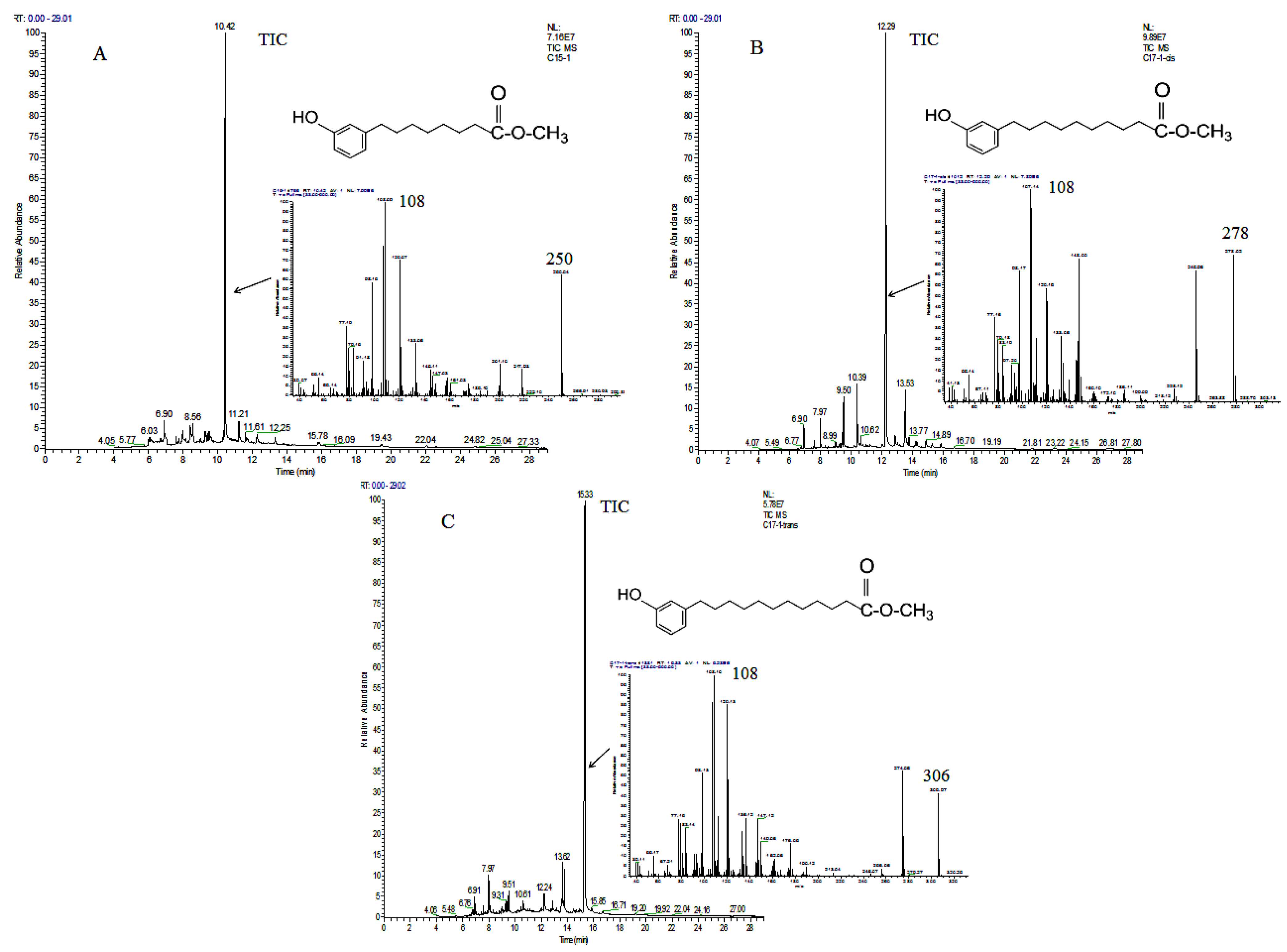

| Peak | Retention Time (min) | Compound | m/z | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| G-1 | 12.14 | C13:0 | 276 (M+) | 175 | 147 | 133 | 108 (100%) | 91 | 77 | |
| G-2 | 14.79 | C15:1-Δ8 | 302 (M+) | 206 | 175 | 147 | 133 | 108 (100%) | 91 | 77 |
| G-3 | 19.31 | C17:2 | 328 (M+) | 232 | 175 | 147 | 133 | 108 (100%) | 91 | 77 |
| G-4 | 15.08 | C15:0 | 304 (M+) | 175 | 147 | 133 | 108 (100%) | 91 | 77 | |
| G-5 | 19.55 | C17:1-Δ10 | 330 (M+) | 234 | 175 | 147 | 133 | 108 (100%) | 91 | 77 |
| G-6 | 19.93 | C17:1-Δ12 | 330 (M+) | 234 | 175 | 147 | 133 | 108 (100%) | 91 | 77 |
| Ginkgol | Spectrum | Aryl |
| C13:0 | 1H a | 6.67 (m, 2H, H2′, H4′), 6.76 (d, 1H, H6′), 7.15 (t, 1H, H5′) |
| 13C a | 112.5 (C6′), 115.3 (C2′), 120.7 (C4′), 129.3 (C5′), 144.9 (C3′), 155.5 (C1′) | |
| UV-(MeOH) | λmax 275 nm | |
| FTIR (cm−1) | 3353 (νO-H), 1593 (νC=C), 1458 (νC=C), 1265 (νC-O), 1154 (δ=C-H), 778 (δ=C-H), 695 (δC-O-H) | |
| C15:1-Δ8 | 1H | 6.66 (m, 2H, H2′, H4′), 6.77 (d, 1H, H6′), 7.15 (t, 1H, H5′) |
| 13C | 112.5 (C6′), 115.3 (C2′), 120.7 (C4′), 129.3 (C5′), 144.9 (C3′), 155.5 (C1′) | |
| UV-(MeOH) | λmax 275 nm | |
| FTIR (cm−1) | 3360 (νO-H), 1593 (νC=C), 1457 (νC=C), 1265 (νC-O), 1154 (δ=C-H), 778 (δ=C-H), 694 (δC-O-H) | |
| C17:2 | 1H | 6.66 (m, 2H, H2′, H4′), 6.76 (d, 1H, H6′), 7.15 (t, 1H, H5′) |
| 13C | 112.3 (C6′), 115.3 (C2′), 121.0, 129.4 (C5′), 144.9 (C3′), 155.5 (C1′) | |
| UV-(MeOH) | λmax 275 nm | |
| FTIR (cm−1) | 3338 (νO-H), 1581 (νC=C), 1456 (νC=C), 1406 (δO-H), 1265 (νC-O), 1154 (δ=C-H), 778 (δ=C-H), 694 (δC-O-H) | |
| C15:0 | 1H | 6.68 (m, 2H, H2′, H4′), 6.77 (d, 1H, H6′), 7.15 (t, 1H, H5′) |
| 13C | 112.5 (C6′), 115.3 (C2′), 120.8 (C4′), 129.3 (C5′), 144.9 (C3′), 155.5 (C1′) | |
| UV-(MeOH) | λmax 275 nm | |
| FTIR (cm−1) | 3328 (νO-H), 1593 (νC=C), 1458 (νC=C), 1265 (νC-O), 1154 (δ=C-H), 777 (δ=C-H), 695 (δC-O-H) | |
| C17:1-Δ10 | 1H | 6.65 (m, 2H, H2′, H4′), 6.76 (d, 1H, H6′), 7.15 (t, 1H, H5′) |
| 13C | 112.4 (C6′), 115.3 (C2′), 120.9 (C4′), 129.3 (C5′), 144.9 (C3′), 155.4 (C1′) | |
| UV-(MeOH) | λmax 275 nm | |
| FTIR (cm−1) | 3307 (νO-H), 1596 (νC=C), 1456 (νC=C), 1266 (νC-O), 1153 (δ=C-H), 738 (δ=C-H), 697 (δC-O-H) | |
| C17:1-Δ12 | 1H | 6.65 (m, 2H, H2′, H4′), 6.76 (d, 1H, H6′), 7.15 (t, 1H, H5′) |
| 13C | 112.5 (C6′), 115.3 (C2′), 120.9 (C4′), 129.3 (C5′), 145 (C3′), 155.4 (C1′) | |
| UV-(MeOH) | λmax 275 nm | |
| FTIR (cm−1) | 3306 (νO-H), 1594 (νC=C), 1458 (νC=C), 1266 (νC-O), 1154 (δ=C-H), 738 (δ=C-H), 697 (δC-O-H) | |
| Ginkgol | Spectrum | Side Chain |
| C13:0 | 1H a | 0.9 (t, 3H, H13′), 1.29 (m, 20H, H3′–12′), 1.60 (m, 2H, H2′), 2.56 (t, 2H, H1′) |
| 13C a | 14.1 (C13′), 22.7 (C12′), 29.3, 29.5, 29.7, 30.9, 31.3 (C2′), 31.9 (C11′), 35.8 (C1′) | |
| FTIR (cm−1) | 2925 (νC-H), 2855 (νC-H) | |
| C15:1-Δ8 | 1H | 0.90 (t, 3H, H15′),1.31 (m, 16H, H3′–6′, H11′–14′), 1.61 (m, 2H, H2′), 2.02 (m, 4H, H7′, H10′), 2.57 (t, 2H, H1′), 5.35 (m, 2H, CH=CH, H8′, H9′) |
| 13C | 14.1 (C15′), 22.7 (C14′), 27.2 (C8′), 27.2 (C9′), 29.0, 29.2, 29.3, 29.4, 29.7, 31.3 (C2′), 31.8 (C13′), 35.8 (C1′), 129.6 (C9′), 130.0 (C8′) | |
| FTIR (cm−1) | 2926 (νC-H), 2855 (νC-H) | |
| C17:2 | 1H | 0.90 (t, 3H, H17′), 1.32 (m, 14H, H3′–7′, H15′–16′), 1.60 (m, 2H, H2′), 2.06 (m, 4H, H8′, H14′), 2.56 (t, 2H, H1′), 2.78 (t, 2H, =CH-CH2-CH=, H14′), 5.37 (m, 4H, CH=CH, H9′–10′, H12′–13′) |
| 13C | 14.1 (C17′), 22.6 (C16′), 25.7, 27.2, 29.2, 29.3 (2C), 29.4, 29.7, 29.7, 31.3 (C2′), 31.6 (C15′), 35.8 (C1′), 128.0, 128.0, 130.1, 130.2 | |
| FTIR (cm−1) | 2925 (νC-H), 2855 (νC-H) | |
| C15:0 | 1H | 0.91 (t, 3H, H15′), 1.30 (m, 24H, H3′–14′), 1.61 (m, 2H, H2′), 2.57 (t, 2H, H1′) |
| 13C | 14.1 (C15′), 22.7 (C14′), 29.3, 29.3, 29.5, 29.6, 29.7, 30.9, 31.3, 31.9 (C13′), 35.8 (C1′) | |
| FTIR (cm−1) | 2925 (νC-H), 2855 (νC-H) | |
| C17:1-Δ10 | 1H | 0.89 (t, 3H, H17′), 1.29 (m, 20H, H3′–8′,H13′–16′), 1.60 (m, 2H, H2′), 2.02 (m, 4H, H8′, H9′), 2.56 (t, 2H, H1′), 5.36 (m, 2H,CH=CH, H10′, CH11′) |
| 13C | 14.1 (C17′), 22.7 (C16′), 27.2 (C9′), 27.2 (C12′), 29.0, 29.3, 29.5, 29.5, 29.7, 29.7, 29.8, 31.3 (C2′), 31.8 (C15′), 35.8 (C1′), 129.9 (C10′), 129.9 (C11′) | |
| FTIR (cm−1) | 2925 (νC-H), 2855 (νC-H) | |
| C17:1-Δ12 | 1H | 0.91 (t, 3H, H17′), 1.31 (m, 20H, H3′–H10′, H15′–16′), 1.60 (m, 2HC2′), 2.02 (m, 4H, H11′, H14′), 2.56 (t, 2H, H1′), 5.36 (m, 2H, CH=CH, H12′, H13′) |
| 13C | 14.0 (C17′), 22.3 (C16′), 26.9 (C14′), 27.2 (C11′), 29.3, 29.5, 29.5, 29.6, 29.6, 29.6, 29.7, 29.8, 31.3 (C2′), 32.0 (C15′), 35.8 (C1′), 129.8 (C13′), 129.9 (C12′) | |
| FTIR (cm−1) | 2925 (νC-H), 2855 (νC-H) |
| Cell Lines | 24 h-IC50 (μM) | |||||
|---|---|---|---|---|---|---|
| C13:0 | C15:1-Δ8 | C17:2 | C15:0 | C17:1-Δ10 | C17:1-Δ12 | |
| HepG2 | 156.81 ± 3.01 Bd | 18.84 ± 2.58 Ba | 60.82 ± 2.90 Bb | 128.09 ± 2.60 Bc | 59.97 ± 3.01 Bb | 68.97 ± 3.00 Bb |
| HGC | 137.31 ± 3.22 Ad | 13.15 ± 2.91 Aa | 33.81 ± 2.74 Ab | 97.80 ± 3.22 Ac | 30.97 ± 1.03 Ab | 34.55 ± 1.45 Ab |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, F.; Boateng, I.D.; Yang, X.; Li, Y. Extraction, Purification, and Elucidation of Six Ginkgol Homologs from Ginkgo biloba Sarcotesta and Evaluation of Their Anticancer Activities. Molecules 2022, 27, 7777. https://doi.org/10.3390/molecules27227777
Li F, Boateng ID, Yang X, Li Y. Extraction, Purification, and Elucidation of Six Ginkgol Homologs from Ginkgo biloba Sarcotesta and Evaluation of Their Anticancer Activities. Molecules. 2022; 27(22):7777. https://doi.org/10.3390/molecules27227777
Chicago/Turabian StyleLi, Fengnan, Isaac Duah Boateng, Xiaoming Yang, and Yuanyuan Li. 2022. "Extraction, Purification, and Elucidation of Six Ginkgol Homologs from Ginkgo biloba Sarcotesta and Evaluation of Their Anticancer Activities" Molecules 27, no. 22: 7777. https://doi.org/10.3390/molecules27227777
APA StyleLi, F., Boateng, I. D., Yang, X., & Li, Y. (2022). Extraction, Purification, and Elucidation of Six Ginkgol Homologs from Ginkgo biloba Sarcotesta and Evaluation of Their Anticancer Activities. Molecules, 27(22), 7777. https://doi.org/10.3390/molecules27227777








