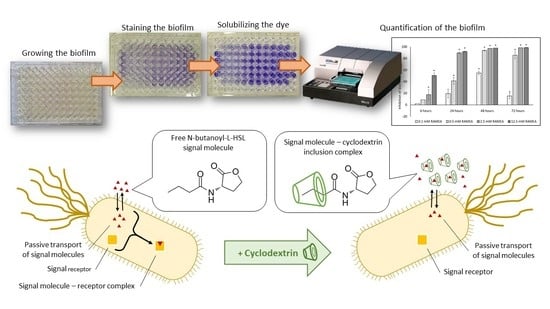
Effect of Cyclodextrins on the Biofilm Formation Capacity of Pseudomonas aeruginosa PAO1
Abstract
:1. Introduction
2. Results
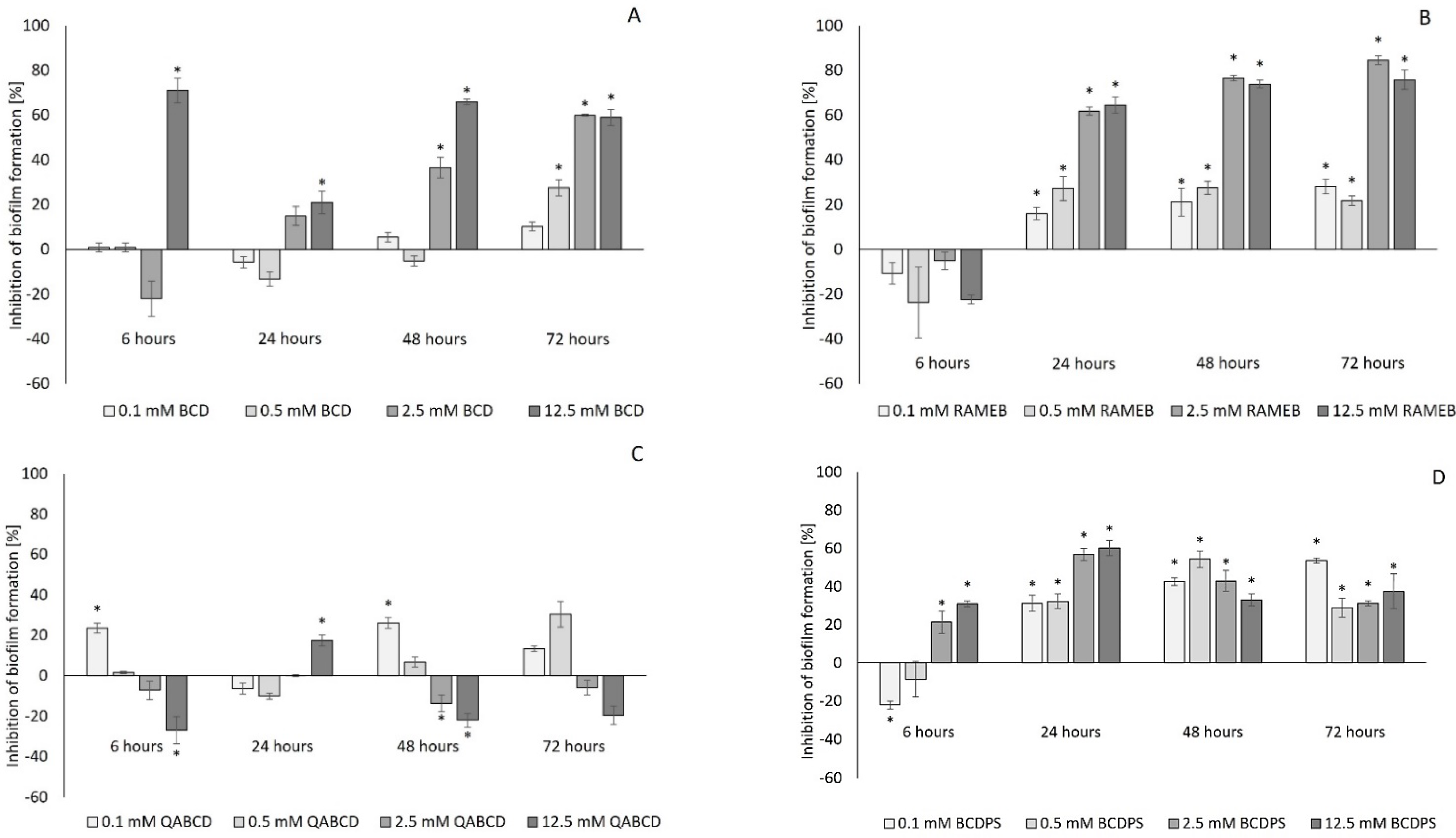
2.1. Effect of Cyclodextrins on Biofilm Formation of Pseudomonas aeruginosa
2.2. Efficiency of the Cyclodextrin-Mediated Effects on Biofilm Formation Based on Their Effective Concentration Values
3. Discussion
3.1. Evaluation of the High-Throughput Microtiter Plate Assay Applied for Quantification of Biofilm Formation
3.2. Quorum Quenching Effect of Cyclodextrins
3.3. Cytotoxicity of Cyclodextrins
3.4. Future Research Directions
4. Materials and Methods
4.1. Bacterial Strain and Culture Conditions
4.2. Tested Cyclodextrins
4.3. Biofilm Formation Assay—Examination of the QQ Effects of Cyclodextrins
4.4. Optical Density Assay—Examination of Population Growth
4.5. Resazurin Reduction Method (RRM)—Examination of Cell Viability
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Lewis, K. Riddle of biofilm resistance. Antimicrob. Agents Chemother. 2001, 45, 999–1007. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jimenez, P.N.; Koch, G.; Thompson, J.A.; Xavier, K.B.; Cool, R.H.; Quax, W.J. The multiple signaling systems regulating virulence in Pseudomonas aeruginosa. Microbiol. Mol. Biol. Rev. 2012, 76, 46–65. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, J.; Zhang, L. The hierarchy quorum sensing network in Pseudomonas aeruginosa. Protein Cell 2015, 6, 26–41. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ikeda, T.; Inoue, Y.; Suehiro, A.; Ikeshoji, H.; Ishida, T.; Takiguchi, N.; Kuroda, A.; Kato, J.; Ohtake, H. The effects of cyclodextrins on autoinducer activities of quorum sensing in Pseudomonas aeruginosa. J. Incl. Phenom. Macrocycl. Chem. 2002, 44, 381–382. [Google Scholar] [CrossRef]
- Flemming, H.C.; Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 2010, 8, 623–633. [Google Scholar] [CrossRef]
- Ikuma, K.; Decho, A.W.; Lau, B.L.T. The extracellular bastions of bacteria—A biofilm way of life. Nat. Ed. Knowl. 2013, 4, 2. Available online: https://www.nature.com/scitable/knowledge/library/the-extracellular-bastions-of-bacteria-nbsp-a-100450088/ (accessed on 31 March 2022).
- Sun, F.; Qu, F.; Ling, Y.; Mao, P.; Xia, P.; Chen, H.; Zhou, D. Biofilm-associated infections: Antibiotic resistance and novel therapeutic strategies. Future Microbiol. 2013, 8, 877–886. [Google Scholar] [CrossRef]
- Hozalski, R.M.; Goel, S.; Bouwer, E.J. TOC removal in biological filters. J. Am. Water Work. Assoc. 1995, 87, 40–54. [Google Scholar] [CrossRef]
- Zhu, I.X.; Getting, T.; Bruce, D. Review of biologically active filters in drinking water applications. J. Am. Water Work. Assoc. 2010, 102, 67–77. [Google Scholar] [CrossRef]
- Beech, I.B. Corrosion of technical materials in the presence of biofilms—Current understanding and state-of-the art methods of study. Int. Biodeterior. Biodegrad. 2004, 53, 177–183. [Google Scholar] [CrossRef]
- Bjarnsholt, T.; Tolker-Nielsen, T.; Hoiby, N.; Givskov, M. Interference of Pseudomonas aeruginosa signalling and biofilm formation for infection control. Expert Rev. Mol. Med. 2010, 12, 1–18. [Google Scholar] [CrossRef]
- Rasamiravaka, T.; Labtani, Q.; Duez, P.; El Jaziri, M. The formation of biofilms by Pseudomonas aeruginosa: A review of the natural and synthetic compounds interfering with control mechanisms. Biomed Res. Int. 2015, 2015, 759348. [Google Scholar] [CrossRef] [Green Version]
- Ding, X.; Yin, B.; Qian, L.; Zeng, Z.; Yang, Z.; Li, H.; Lu, Y.; Zhou, S. Screening for novel quorum-sensing inhibitors to interfere with the formation of Pseudomonas aeruginosa biofilm. J. Med. Microbiol. 2011, 60, 1827–1834. [Google Scholar] [CrossRef]
- Zhou, L.; Zhang, Y.; Ge, Y.; Zhu, X.; Pan, J. Regulatory mechanisms and promising applications of quorum sensing-inhibiting agents in control of bacterial biofilm formation. Front. Microbiol. 2020, 11, 589640. [Google Scholar] [CrossRef]
- Nealson, K.H.; Platt, T.; Hastings, J.W. Cellular control of the synthesis and activity of the bacterial luminescent system. J. Bacteriol. 1970, 104, 313–322. [Google Scholar] [CrossRef] [Green Version]
- Greenberg, E.P. Quorum sensing in gram-negative bacteria. ASM News 1997, 63, 371–377. [Google Scholar]
- Davies, D.G.; Parsek, M.R.; Pearson, J.P.; Iglewski, B.H.; Costerton, J.W.; Greenberg, E.P. The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science 1998, 280, 295–298. [Google Scholar] [CrossRef] [Green Version]
- Williams, P. Quorum sensing, communication and crosskingdom signalling in the bacterial world. Microbiology 2007, 153, 3923–3938. [Google Scholar] [CrossRef] [Green Version]
- Pan, J.; Ren, D. Quorum sensing inhibitors: A patent overview. Expert Opin. Ther. Pat. 2009, 19, 1581–1601. [Google Scholar] [CrossRef]
- Boyer, M.; Wisniewski-Dye, F. Cell-cell signalling in bacteria: Not simply a matter of quorum. FEMS Microbiol. Ecol. 2009, 70, 1–19. [Google Scholar] [CrossRef]
- Waters, C.M.; Bassler, B.L. Quorum sensing: Cell-to-cell communication in bacteria. Annu. Rev. Cell Dev. Biol. 2005, 21, 319–346. [Google Scholar] [CrossRef] [Green Version]
- Reuter, K.; Steinbach, A.; Helms, V. Interfering with bacterial quorum sensing. Perspect. Med. Chem. 2016, 8, PMC-S13209. [Google Scholar] [CrossRef] [Green Version]
- Sperandio, V.; Torres, A.G.; Kaper, J.B. Quorum sensing Escherichia coli regulators B and C (QseBC): A novel two-component regulatory system involved in the regulation of flagella and motility by quorum sensing in E. coli. Mol. Microbiol. 2002, 43, 809–821. [Google Scholar] [CrossRef]
- Kim, C.S.; Gatsios, A.; Cuesta, S.; Lam, Y.C.; Wei, Z.; Chen, H.; Russell, R.M.; Shine, E.E.; Wang, R.; Wyche, T.P.; et al. Characterization of autoinducer-3 structure and biosynthesis in E. coli. ACS Cent. Sci. 2020, 6, 197–206. [Google Scholar] [CrossRef] [Green Version]
- Lee, K.; Yoon, S.S. Pseudomonas aeruginosa biofilm, a programmed bacterial life for fitness. J. Microbiol. Biotechnol. 2017, 27, 1053–1064. [Google Scholar] [CrossRef] [Green Version]
- García-Reyes, S.; Soberón-Chávez, G.; Cocotl-Yanez, M. The third quorum-sensing system of Pseudomonas aeruginosa: Pseudomonas quinolone signal and the enigmatic PqsE protein. J. Med. Microbiol. 2019, 69, 25–34. [Google Scholar] [CrossRef]
- Pesci, E.C.; Pearson, J.P.; Seed, P.C.; Iglewski, B.H. Regulation of las and rhl quorum sensing in Pseudomonas aeruginosa. J. Bacteriol. 1997, 179, 3127–3132. [Google Scholar] [CrossRef] [Green Version]
- Dubern, J.F.; Diggle, S.P. Quorum sensing by 2-alkyl-4-quinolones in Pseudomonas aeruginosa and other bacterial species. Mol. Biosyst. 2008, 4, 882–888. [Google Scholar] [CrossRef]
- Lee, J.; Wu, J.; Deng, Y.; Wang, J.; Wang, C.; Wang, J.; Chang, C.; Dong, Y.; Williams, P.; Zhang, L.H. A cell-cell communication signal integrates quorum sensing and stress response. Nat. Chem. Biol. 2013, 9, 339–343. [Google Scholar] [CrossRef]
- Pérez-Pérez, M.; Jorge, P.; Rodríguez, G.P.; Pereira, M.O.; Lourenço, A. Quorum sensing inhibition in Pseudomonas aeruginosa biofilms: New insights through network mining. Biofouling 2017, 33, 128–142. [Google Scholar] [CrossRef] [Green Version]
- Khan, F.; Pham, D.T.N.; Oloketuyi, S.F.; Kim, Y.M. Regulation and controlling the motility properties of Pseudomonas aeruginosa. Appl. Microbiol. Biotechnol. 2020, 104, 33–49. [Google Scholar] [CrossRef] [PubMed]
- Wade, D.S.; Calfee, M.W.; Rocha, E.R.; Ling, E.A.; Engstrom, E.; Coleman, J.P.; Pesci, E.C. Regulation of Pseudomonas quinolone signal synthesis in Pseudomonas aeruginosa. J. Bacteriol. 2005, 187, 4372–4380. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, Y.H.; Wang, L.H.; Xu, J.L.; Zhang, H.B.; Zhang, X.F.; Zhang, L.H. Quenching quorum-sensing- dependent bacterial infection by an N-acyl homoserine lactonase. Nature 2001, 411, 813–817. [Google Scholar] [CrossRef] [PubMed]
- Kalia, V.C.; Purohit, H.J. Quenching the quorum sensing system: Potential antibacterial drug targets. Crit. Rev. Microbiol. 2011, 37, 121–140. [Google Scholar] [CrossRef]
- Bhardwaj, A.K.; Vinothkumar, K.; Rajpara, N. Bacterial quorum sensing inhibitors: Attractive alternatives for control of infectious pathogens showing multiple drug resistance. Recent Pat. Antiinfect. Drug Discov. 2013, 8, 68–83. [Google Scholar] [CrossRef]
- Kalia, M.; Yadav, V.K.; Singh, P.K.; Sharma, D.; Narvi, S.S.; Agarwal, V. Exploring the impact of parthenolide as anti-quorum sensing and anti-biofilm agent against Pseudomonas aeruginosa. Life Sci. 2018, 199, 96–103. [Google Scholar] [CrossRef]
- Zhang, W.; Li, C. Exploiting quorum sensing interfering strategies in Gram-negative bacteria for the enhancement of environmental applications. Front. Microbiol. 2016, 6, 1535. [Google Scholar] [CrossRef]
- Lade, H.; Paul, D.; Kweon, J.H. Quorum quenching mediated approaches for control of membrane biofouling. Int. J. Biol. Sci. 2014, 10, 550–565. [Google Scholar] [CrossRef]
- Brackman, G.; Coenye, T. Quorum sensing inhibitors as anti-biofilm agents. Curr. Pharm. Des. 2015, 21, 5–11. [Google Scholar] [CrossRef]
- Kalia, V.C. Quorum sensing inhibitors: An overview. Biotechnol. Adv. 2013, 31, 224–245. [Google Scholar] [CrossRef]
- Abbas, H.A.; Shaldam, M.A.; Eldamasi, D. Curtailing quorum sensing in Pseudomonas aeruginosa by Sitagliptin. Curr. Microbiol. 2020, 77, 1051–1060. [Google Scholar] [CrossRef] [PubMed]
- El-Shaer, S.; Shaaban, M.; Barwa, R.; Hassan, R. Control of quorum sensing and virulence factors of Pseudomonas aeruginosa using phenylalanine arginyl beta-naphthylamide. J. Med. Microbiol. 2016, 65, 1194–1204. [Google Scholar] [CrossRef]
- Okano, C.; Arai, M.; Nasuno, E.; Iimura, K.; Morohoshi, T.; Ikeda, T.; Kato, N. β-Cyclodextrin interaction with N-hexanoyl homoserine lactone as quorum sensing signal produced in Gram-negative bacteria. Trans. Mater. Res. Soc. Jpn. 2012, 37, 315–318. [Google Scholar] [CrossRef] [Green Version]
- Okano, C.; Arai, M.; Nasuno, E.; Iimura, K.; Kato, N. Cyclodextrin-immobilized microspheres for uptake of the quorum-sensing signaling molecule N-acylhomoserine lactone. J. Appl. Polym. Sci. 2016, 133, 43198. [Google Scholar] [CrossRef]
- Bender, M.L.; Komiyama, M. Cyclodextrin Chemistry, 1st ed.; Springer: Berlin/Heidelberg, Germany, 1978; pp. 1–96. [Google Scholar]
- Szejtli, J. Introduction and general overview of cyclodextrin chemistry. Chem. Rev. 1998, 98, 1743–1753. [Google Scholar] [CrossRef]
- Simoes, S.M.N.; Rey-Rico, A.; Concheiro, A.; Alvarez-Lorenzo, C. Supramolecular cyclodextrin-based drug nanocarriers. Chem. Commun. 2015, 51, 6275–6289. [Google Scholar] [CrossRef]
- Muankaew, C.; Loftsson, T. Cyclodextrin-based formulations: A non-invasive platform for targeted drug delivery. Basic Clin. Pharmacol. Toxicol. 2018, 122, 46–55. [Google Scholar] [CrossRef] [Green Version]
- Szente, L.; Szejtli, J. Cyclodextrins as food ingredients. Trends Food Sci. Technol. 2004, 15, 137–142. [Google Scholar] [CrossRef]
- Molnár, M.; Gruiz, K.; Fenyvesi, E. Biodegradation-based remediation. Overview and case studies. In Engineering Tools for Environmental Risk Management: 4. Risk Reduction Technologies and Case Studies; Gruiz, K., Meggyes, T., Fenyvesi, E., Eds.; CRC Press: London, UK, 2019; Volume 4, pp. 243–275. [Google Scholar]
- Molnár, M.; Fenyvesi, É.; Berkl, Z.; Németh, I.; Fekete-Kertész, I.; Márton, R.; Vaszita, E.; Varga, E.; Ujj, D.; Szente, L. Cyclodextrin-mediated quorum quenching in the Aliivibrio fischeri bioluminescence model system—Modulation of bacterial communication. Int. J. Pharm. 2021, 594, 120–150. [Google Scholar] [CrossRef]
- Szeman, J.; Szente, L.; Szabó, T.; Szejtli, J. Highly soluble beta-cyclodextrin derivatives, a comparative study. In Proceedings of the Fourth International Symposium on Cyclodextrins, Munich, Germany, 20–22 April 1988; Huber, O., Szejtli, J., Eds.; Kluwer: Dordrecht, The Netherland, 2004; Volume 1, pp. 393–398. [Google Scholar]
- Kato, N.; Morohoshi, T.; Nozawa, T.; Matsumoto, H.; Ikeda, T. Control of Gram-negative bacterial quorum sensing with cyclodextrin immobilized cellulose ether gel. J. Incl. Phenom. Macrocycl. Chem. 2006, 56, 55–59. [Google Scholar] [CrossRef]
- Kato, N.; Tanaka, T.; Nakagawa, S.; Morohoshi, T.; Hiratani, K.; Ikeda, T. Control of virulence factor expression in opportunistic pathogens using cyclodextrin immobilized gel. J. Incl. Phenom. Macrocycl. Chem. 2007, 57, 419–423. [Google Scholar] [CrossRef]
- Morohoshi, T.; Tokita, K.; Ito, S.; Saito, Y.; Maeda, S.; Kato, N.; Ikeda, T. Inhibition of quorum sensing in Gram-negative bacteria by alkylamine-modified cyclodextrins. J. Biosci. Bioeng. 2013, 116, 175–179. [Google Scholar] [CrossRef] [PubMed]
- Vandeputte, O.M.; Kiendrebeogo, M.; Rajaonson, S.; Diallo, B.; Mol, A.; El Jaziri, M.; Baucher, M. Identification of catechin as one of the flavonoids from Combretum albiflorum bark extract that reduces the production of quorum-sensing-controlled virulence factors in Pseudomonas aeruginosa PAO1. Appl. Environ. Microbiol. 2010, 76, 243–253. [Google Scholar] [CrossRef] [Green Version]
- Chatterjee, M.; D’Morris, S.; Paul, V.; Warrier, S.; Vasudevan, A.K.; Vanuopadath, M.; Nair, S.S.; Paul-Prasanth, B.; Mohan, C.G.; Biswas, R. Mechanistic understanding of phenyllactic acid mediated inhibition of quorum sensing and biofilm development in Pseudomonas aeruginosa. Appl. Microbiol. Biotechnol. 2017, 101, 8223–8236. [Google Scholar] [CrossRef]
- Hernando-Amado, S.; Alcalde-Rico, M.; Gil-Gil, T.; Valverde, J.R.; Martínez, J.L. Naringenin inhibition of the Pseudomonas aeruginosa quorum sensing response is based on its time-dependent competition with N-(3-oxo-dodecanoyl)-L-homoserine lactone for LasR binding. Front. Mol. Biosci. 2020, 7, 25. [Google Scholar] [CrossRef]
- Ishida, T.; Ikeda, T.; Takiguchi, N.; Kuroda, A.; Kato, J.; Ohtake, H. Inhibition of quorum sensing in Pseudomonas aeruginosa by N-acyl-cyclopentylamides. Appl. Environ. Microbiol. 2007, 73, 3183–3188. [Google Scholar] [CrossRef] [Green Version]
- Sahner, J.H.; Empting, M.; Kamal, A.; Weidel, E.; Groh, M.; Börger, C.; Hartmann, R.W. Exploring the chemical space of ureidothiophene-2-carboxylicacids as inhibitors of the quorum sensing enzyme PqsD from Pseudomonas aeruginosa. Eur. J. Med. Chem. 2015, 96, 14–21. [Google Scholar] [CrossRef]
- Barnaby, R.; Koeppen, K.; Stanton, B.A. Cyclodextrins reduce the ability of Pseudomonas aeruginosa outer-membrane vesicles to reduce CFTR Cl_ secretion. Am. J. Physiol. Lung Cell. Mol. Physiol. 2019, 316, 206–215. [Google Scholar] [CrossRef] [Green Version]
- Ziegler, E.W.; Brown, A.B.; Nesnas, N.; Chouinard, C.D.; Mehta, A.K.; Palmer, A.G. β-Cyclodextrin encapsulation of synthetic AHLs: Drug delivery implications and quorum-quenching exploits. Chem. Biochem. 2021, 22, 1292–1301. [Google Scholar] [CrossRef]
- Präbst, K.; Engelhardt, H.; Ringgeler, S.; Hübner, H. Basic Colorimetric Proliferation Assays: MTT, WST, and Resazurin. In Cell Viability Assays. Methods in Molecular Biology; Gilbert, D., Friedrich, O., Eds.; Humana Press: New York, NY, USA, 2017; Volume 1601, pp. 1–17. [Google Scholar]
- O’Toole, G.A. Microtiter dish biofilm formation assay. J. Vis. Exp. 2011, 47, e2437. [Google Scholar] [CrossRef]
- Palomino, J.C.; Martin, A.; Camacho, M.; Guerra, H.; Swings, J.; Portaels, F. Resazurin microtiter assay plate: Simple and inexpensive method for detection of drug resistance in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 2002, 46, 2720–2722. [Google Scholar] [CrossRef] [Green Version]
- Ikeda, T.; Morohoshi, T.; Kato, N.; Inoyama, M.; Nakazawa, S.; Hiratani, K.; Ishida, T.; Kato, J.; Ohtake, H. Quorum sensing control using synthetic autoinducer analogues and cyclodextrins in Gram-negative bacteria. In Proceedings of the 10th Asia Pacific Confederation of Chemical Engineering, Kitakyushu, Japan, 17–21 October 2004; Volume 390, pp. 1–5. [Google Scholar]
- Poulson, B.G.; Alsulami, Q.A.; Sharfalddin, A.; El Agammy, E.F.; Mouffouk, F.; Emwas, A.H.; Jaremko, L.; Jaremko, M. Cyclodextrins: Structural, chemical, and physical properties, and applications. Polysaccharides 2022, 3, 1–31. [Google Scholar] [CrossRef]
- Puskás, I.; Szemjonov, A.; Fenyvesi, É.; Milo, M.; Szente, L. Aspects of determining the molecular weight of cyclodextrin polymers and oligomers by static light scattering. Carbohydr. Polym. 2013, 94, 124–128. [Google Scholar] [CrossRef]
- Coleman, A.W.; Nicolis, I.; Keller, N.; Dalbiez, J.P. Aggregation of cyclodextrins: An explanation of the abnormal solubility of β-cyclodextrin. J. Incl. Phenom. Mol. Recognit. Chem. 1992, 13, 139–143. [Google Scholar] [CrossRef]
- Loftsson, T.; Saokham, P.; Sá Couto, A.R. Self-association of cyclodextrins and cyclodextrin complexes in aqueous solutions. Int. J. Pharm. 2019, 560, 228–234. [Google Scholar] [CrossRef]
- González-Gaitano, G.; Rodríguez, P.; Isasi, J.R.; Fuentes, M.; Tardajos, G.; Sanchez, M. The aggregation of cyclodextrins as studied by photon correlation spectroscopy. J. Incl. Phenom. 2002, 44, 101–105. [Google Scholar] [CrossRef]
- Miller, K.P.; Wang, L.; Chen, Y.P.; Pellechia, P.J.; Benicewicz, B.C.; Decho, A.W. Engineering nanoparticles to silence bacterial communication. Front. Microbiol. 2015, 6, 189. [Google Scholar] [CrossRef] [Green Version]
- Takayama, Y.; Kato, N. Inhibition of quorum sensing in opportunistic pathogen, Serratia marcescens, using cyclodextrin-immobilized, multiple parallel gel filaments fabricated with dynamic flow of polymer blend solution. Mater. Sci. Eng. C 2020, 107, 110331. [Google Scholar] [CrossRef]
- Zare, M.; Amin, M.M.; Nikaeen, M.; Bina, B.; Pourzamani, H.; Fatehizadeh, A.; Taheri, E. Resazurin reduction assay, a useful tool for assessment of heavy metal toxicity in acidic conditions. Environ. Monit. Assess. 2015, 187, 276. [Google Scholar] [CrossRef]
- Csepregi, R.; Lemli, B.; Kunsági-Máté, S.; Szente, L.; Kőszegi, T.; Németi, B.; Poór, M. Complex formation of resorufin and resazurin with Β-cyclodextrins: Can cyclodextrins interfere with a resazurin cell viability assay? Molecules 2018, 23, 382. [Google Scholar] [CrossRef] [Green Version]
- Stella, V.J.; He, Q. Cyclodextrins. Toxicol. Pathol. 2008, 36, 30–42. [Google Scholar] [CrossRef] [PubMed]
- Kiss, T.; Fenyvesi, F.; Bácskay, I.; Váradi, J.; Fenyvesi, E.; Iványi, R.; Szente, L.; Tósaki, A.; Vecsernyés, M. Evaluation of the cytotoxicity of beta-cyclodextrin derivatives: Evidence for the role of cholesterol extraction. Eur. J. Pharm. Sci. 2010, 40, 376–380. [Google Scholar] [CrossRef] [PubMed]
- Aachmann, F.L.; Aune, T.E. Use of cyclodextrin and its derivatives for increased transformation efficiency of competent bacterial cells. Appl. Microbiol. Biotechnol. 2009, 83, 589–596. [Google Scholar] [CrossRef] [PubMed]





| Source of Variation | Df 1 | MS 2 | F 3 | p 4 |
|---|---|---|---|---|
| ACD | ||||
| ACD treatment | 4 | 1.14 | 138.57 | 0.000 |
| Time | 3 | 2.17 | 459.61 | 0.000 |
| Time × ACD treatment | 12 | 0.16 | 33.41 | 0.000 |
| RAMEA | ||||
| RAMEA treatment | 4 | 1.26 | 175.74 | 0.000 |
| Time | 3 | 1.19 | 217.22 | 0.000 |
| Time × RAMEA treatment | 12 | 0.15 | 26.95 | 0.000 |
| QAACD | ||||
| QAACD treatment | 4 | 0.09 | 12.68 | 0.000 |
| Time | 3 | 3.92 | 500.30 | 0.000 |
| Time × QAACD treatment | 12 | 0.08 | 10.51 | 0.000 |
| ACDPS | ||||
| ACDPS treatment | 4 | 1.68 | 461.42 | 0.000 |
| Time | 3 | 1.23 | 444.38 | 0.000 |
| Time × ACDPS treatment | 12 | 0.26 | 94.76 | 0.000 |
| Source of Variation | Df 1 | MS 2 | F 3 | p 4 |
|---|---|---|---|---|
| ACD | ||||
| ACD treatment | 4 | 2.80 | 400.89 | 0.000 |
| Time | 3 | 0.93 | 59.09 | 0.000 |
| Time × ACD treatment | 12 | 0.33 | 21.03 | 0.000 |
| RAMEA | ||||
| RAMEA treatment | 4 | 3.74 | 471.10 | 0.000 |
| Time | 3 | 0.67 | 52.21 | 0.000 |
| Time × RAMEA treatment | 12 | 0.48 | 37.10 | 0.000 |
| QAACD | ||||
| QAACD treatment | 4 | 0.45 | 8.22 | 0.001 |
| Time | 3 | 2.97 | 46.82 | 0.000 |
| Time × QAACD treatment | 12 | 0.08 | 1.31 | 0.244 |
| ACDPS | ||||
| ACDPS treatment | 4 | 0.88 | 172.58 | 0.000 |
| Time | 3 | 1.22 | 107.87 | 0.000 |
| Time × ACDPS treatment | 12 | 0.21 | 18.16 | 0.000 |
| Degree of Inhibition [%] | BCD | RAMEB | QABCD | BCDPS |
|---|---|---|---|---|
| 12.5 mM concentration | 39 ± 2 (72 h) | 23 ± 3 (48 h) | 31 ± 3 (72 h) | 38 ± 1 (48 h), 44 ± 2 (72 h) |
| Source of Variation | Df 1 | MS 2 | F 3 | p 4 |
|---|---|---|---|---|
| BCD | ||||
| BCD treatment | 4 | 0.46 | 55.84 | 0.000 |
| Time | 3 | 3.63 | 395.76 | 0.000 |
| Time × BCD treatment | 12 | 0.13 | 14.69 | 0.000 |
| RAMEB | ||||
| RAMEB treatment | 4 | 1.71 | 304.23 | 0.000 |
| Time | 3 | 1.71 | 228.14 | 0.000 |
| Time × RAMEB treatment | 12 | 0.28 | 37.64 | 0.000 |
| QABCD | ||||
| QABCD treatment | 4 | 0.01 | 5.92 | 0.004 |
| Time | 3 | 0.86 | 671.35 | 0.000 |
| Time × QABCD treatment | 12 | 0.01 | 9.18 | 0.000 |
| BCDPS | ||||
| BCDPS treatment | 4 | 0.08 | 93.23 | 0.000 |
| Time | 3 | 0.30 | 330.56 | 0.000 |
| Time × BCDPS treatment | 12 | 0.02 | 20.27 | 0.000 |
| Source of Variation | Df 1 | MS 2 | F 3 | p 4 |
|---|---|---|---|---|
| BCD | ||||
| BCD treatment | 4 | 2.76 | 222.94 | 0.000 |
| Time | 3 | 2.16 | 134.13 | 0.000 |
| Time × BCD treatment | 12 | 0.36 | 22.39 | 0.000 |
| RAMEB | ||||
| RAMEB treatment | 4 | 4.29 | 51.89 | 0.000 |
| Time | 3 | 1.47 | 14.44 | 0.000 |
| Time × RAMEB treatment | 12 | 0.59 | 5.76 | 0.000 |
| QABCD | ||||
| QABCD treatment | 4 | 0.22 | 45.60 | 0.000 |
| Time | 3 | 0.25 | 44.79 | 0.000 |
| Time × QABCD treatment | 12 | 0.17 | 30.35 | 0.000 |
| BCDPS | ||||
| BCDPS treatment | 4 | 0.44 | 15.23 | 0.000 |
| Time | 3 | 0.76 | 24.48 | 0.000 |
| Time × BCDPS treatment | 12 | 0.32 | 10.31 | 0.000 |
| Effective Concentration Values—EC10 (22 °C) [mM] | ||||||||
|---|---|---|---|---|---|---|---|---|
| ACD | RAMEA | QAACD | ACDPS | BCD | RAMEB | QABCD | BCDPS | |
| 6 h | 0.01 | 0.29 | >12.50 | n.d. | 3.51 | n.d. | n.d. | 1.50 |
| 24 h | 1.49 | 0.36 | n.d. | 0.21 | 1.84 | 0.45 | n.d. | 0.02 |
| 48 h | 3.23 | 0.60 | n.d. | 0.77 | 1.25 | 0.29 | n.d. | n.d. |
| 72 h | 0.87 | 0.58 | 2.10 | 0.34 | 0.79 | 0.16 | n.d. | n.d. |
| Effective Concentration Values—EC10 (30 °C) [mM] | ||||||||
| ACD | RAMEA | QAACD | ACDPS | BCD | RAMEB | QABCD | BCDPS | |
| 6 h | 5.36 | 3.04 | 0.95 | 0.19 | >12.50 | 3.82 | n.d. | 0.91 |
| 24 h | 0.72 | 0.32 | 1.37 | 0.01 | 0.12 | 0.06 | n.d. | 3.56 |
| 48 h | 0.38 | 0.05 | 0.54 | 0.22 | 0.16 | 0.09 | n.d. | n.d. |
| 72 h | 0.29 | 0.13 | 0.70 | 0.07 | 0.65 | 0.29 | 0.22 | 0.35 |
| Effective Concentration Values—EC50 (22 °C) [mM] | ||||||||
|---|---|---|---|---|---|---|---|---|
| ACD | RAMEA | QAACD | ACDPS | BCD | RAMEB | QABCD | BCDPS | |
| 6 h | 0.03 | 0.62 | >12.50 | n.d. | 8.50 | n.d. | n.d. | >12.50 |
| 24 h | 3.91 | 0.84 | >12.50 | 0.21 | >12.50 | 1.20 | n.d. | 1.11 |
| 48 h | 7.54 | 1.29 | n.d. | 0.77 | 3.23 | 0.61 | n.d. | >12.50 |
| 72 h | 1.89 | 1.25 | 10.23 | 0.34 | 2.39 | 1.04 | n.d. | n.d. |
| Effective Concentration Values—EC50 (30 °C) [mM] | ||||||||
| ACD | RAMEA | QAACD | ACDPS | BCD | RAMEB | QABCD | BCDPS | |
| 6 h | >12.50 | >12.50 | >12.50 | 0.36 | >12.50 | >12.50 | n.d. | >12.50 |
| 24 h | 1.52 | 0.74 | >12.50 | >12.5 | 0.51 | 0.22 | n.d. | >12.50 |
| 48 h | 0.82 | 0.09 | >12.50 | 0.48 | 0.42 | 0.19 | n.d. | >12.50 |
| 72 h | 0.60 | 0.29 | 1.62 | 0.19 | 1.30 | 0.65 | 0.45 | 0.78 |
| Effective Concentration Values—EC90 (22 °C) [mM] | ||||||||
|---|---|---|---|---|---|---|---|---|
| ACD | RAMEA | QAACD | ACDPS | BCD | RAMEB | QABCD | BCDPS | |
| 6 h | 12.49 | >12.50 | >12.50 | n.d. | >12.50 | n.d. | n.d. | >12.50 |
| 24 h | >12.50 | >12.50 | >12.50 | >12.50 | >12.50 | >12.50 | n.d. | >12.50 |
| 48 h | >12.50 | >12.50 | n.d. | 1.50 | >12.50 | >12.50 | n.d. | >12.50 |
| 72 h | 7.99 | 3.71 | >12.50 | >12.50 | >12.50 | >12.50 | n.d. | n.d. |
| Effective Concentration Values—EC90 (30 °C) [mM] | ||||||||
| ACD | RAMEA | QAACD | ACDPS | BCD | RAMEB | QABCD | BCDPS | |
| 6 h | >12.50 | >12.50 | >12.50 | >12.50 | >12.50 | >12.50 | n.d. | >12.50 |
| 24 h | 4.73 | 2.49 | >12.50 | >12.50 | >12.50 | >12.50 | n.d. | >12.50 |
| 48 h | 1.94 | 0.19 | >12.50 | >12.50 | >12.50 | 0.65 | n.d. | >12.50 |
| 72 h | 1.31 | 0.66 | >12.50 | >12.50 | 3.11 | 1.69 | 5.73 | 2.51 |
| α-Cyclodextrins | A 1 | AMF 2 | MW 3 [g/mol] | WS 4 [g/L] | DS 5 |
|---|---|---|---|---|---|
| Native α-CD | ACD | C36H60O30 | 972 | 145 | - |
| Randomly methylated α-CD | RAMEA | C36H60-nO30 · (CH3)n | 1127 | >500 | 11 |
| Trimethyl-aminopropyl α-CD | QAACD | C48H80-nO40 · (C6H15ONCl)n | 1430 | >500 | 2.5–4 |
| α-CD polymer | ACDPS | - | 40,000 * | >500 | - |
| β-Cyclodextrins | A 1 | AMF 2 | MW 3 [g/mol] | WS 4 [g/L] | DS 5 |
|---|---|---|---|---|---|
| Native β-CD | BCD | C42H70O35 | 1135 | 18 | - |
| Randomly methylated β-CD | RAMEB | C42H70-nO35 · (CH3)n | 1303 | >500 | 12 |
| Trimethyl-aminopropyl β-CD | QABCD | C42H70-nO35 · (C6H15ONCl)n | 1665 | >500 | 3–4 |
| β-CD polymer | BCDPS | - | 87,000 * | >500 | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Berkl, Z.; Fekete-Kertész, I.; Buda, K.; Vaszita, E.; Fenyvesi, É.; Szente, L.; Molnár, M. Effect of Cyclodextrins on the Biofilm Formation Capacity of Pseudomonas aeruginosa PAO1. Molecules 2022, 27, 3603. https://doi.org/10.3390/molecules27113603
Berkl Z, Fekete-Kertész I, Buda K, Vaszita E, Fenyvesi É, Szente L, Molnár M. Effect of Cyclodextrins on the Biofilm Formation Capacity of Pseudomonas aeruginosa PAO1. Molecules. 2022; 27(11):3603. https://doi.org/10.3390/molecules27113603
Chicago/Turabian StyleBerkl, Zsófia, Ildikó Fekete-Kertész, Kata Buda, Emese Vaszita, Éva Fenyvesi, Lajos Szente, and Mónika Molnár. 2022. "Effect of Cyclodextrins on the Biofilm Formation Capacity of Pseudomonas aeruginosa PAO1" Molecules 27, no. 11: 3603. https://doi.org/10.3390/molecules27113603
APA StyleBerkl, Z., Fekete-Kertész, I., Buda, K., Vaszita, E., Fenyvesi, É., Szente, L., & Molnár, M. (2022). Effect of Cyclodextrins on the Biofilm Formation Capacity of Pseudomonas aeruginosa PAO1. Molecules, 27(11), 3603. https://doi.org/10.3390/molecules27113603