Copper Binding and Oligomerization Studies of the Metal Resistance Determinant CrdA from Helicobacter pylori
Abstract
:1. Introduction
2. Materials and Methods
2.1. Molecular Cloning
2.2. Expression and Purification
2.3. Characterization in Solution
2.4. Ultraviolet-Visible (UV-VIS) Titration
2.5. Microscale Thermophoresis (MST)
3. Results and Discussion
3.1. Expression Level of HpCrdA
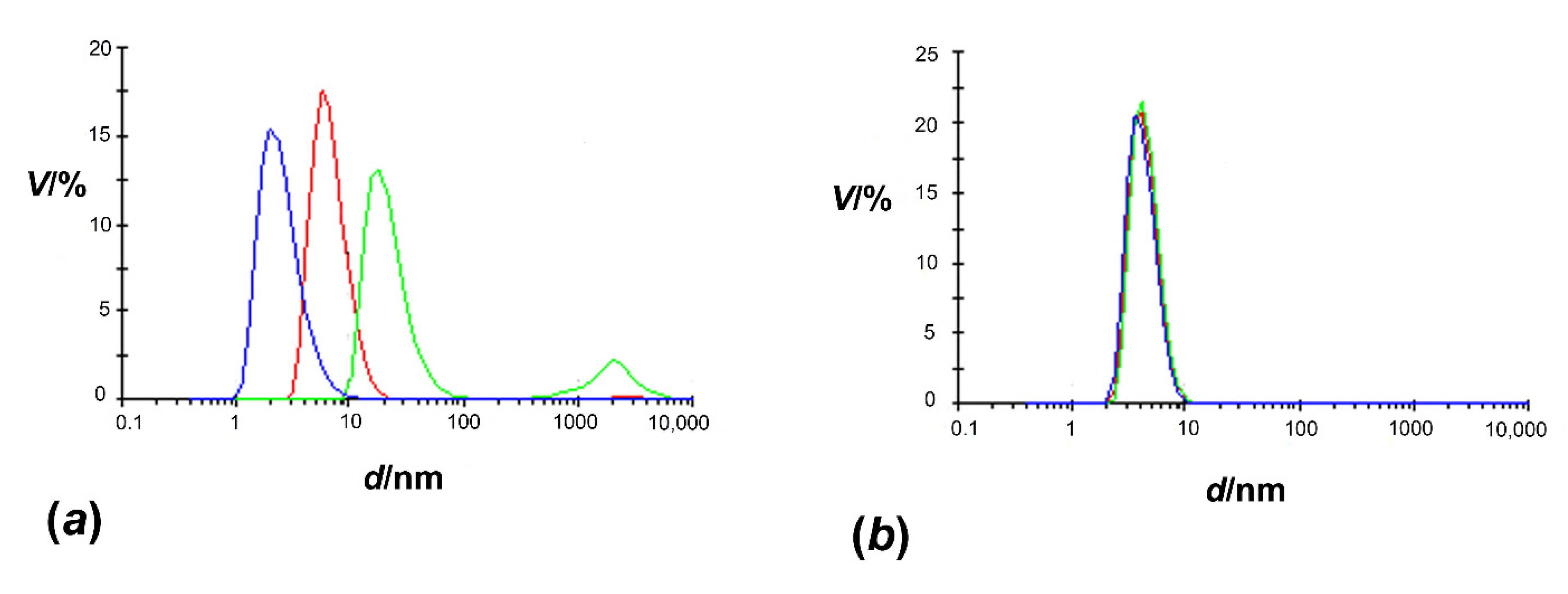
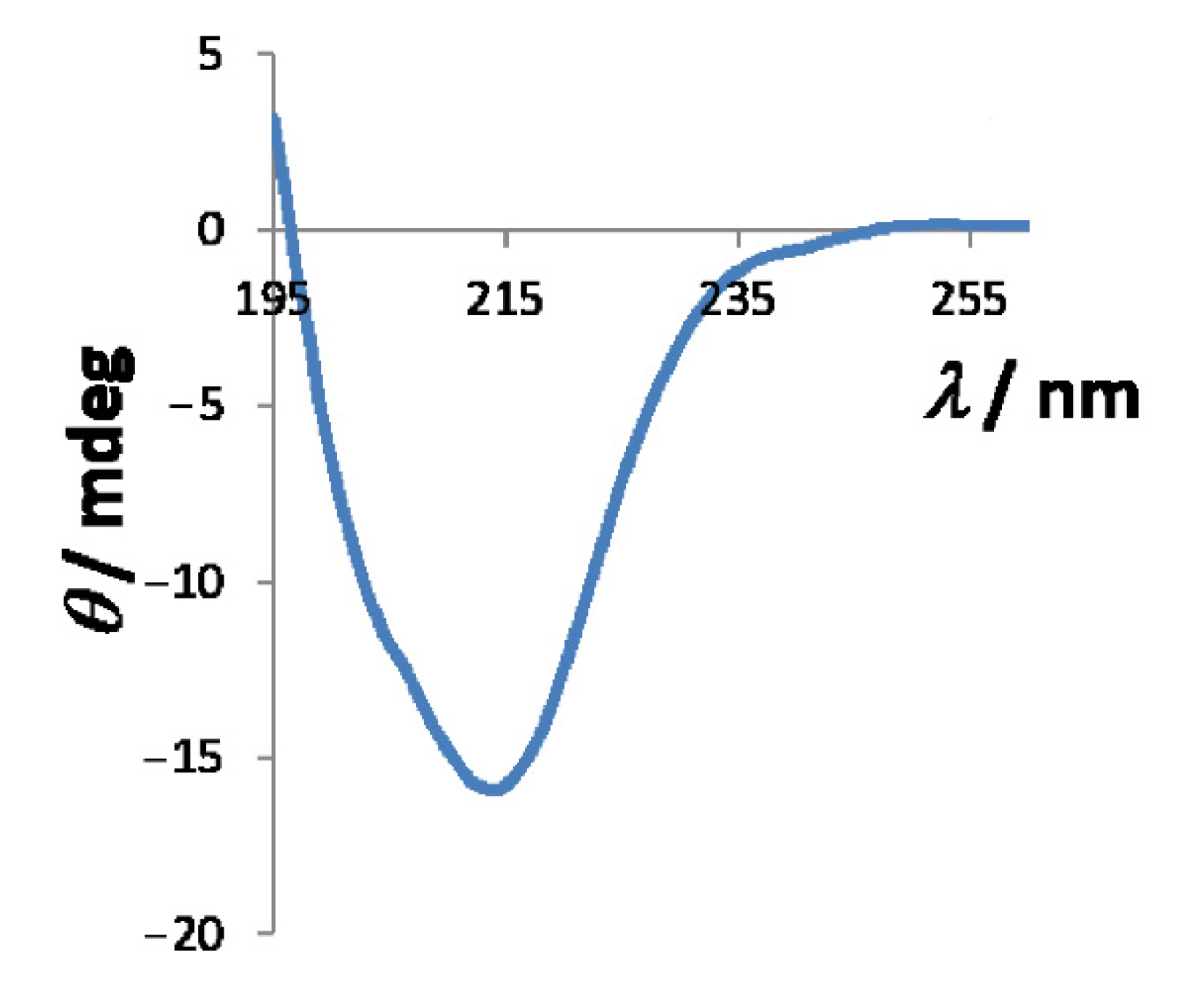
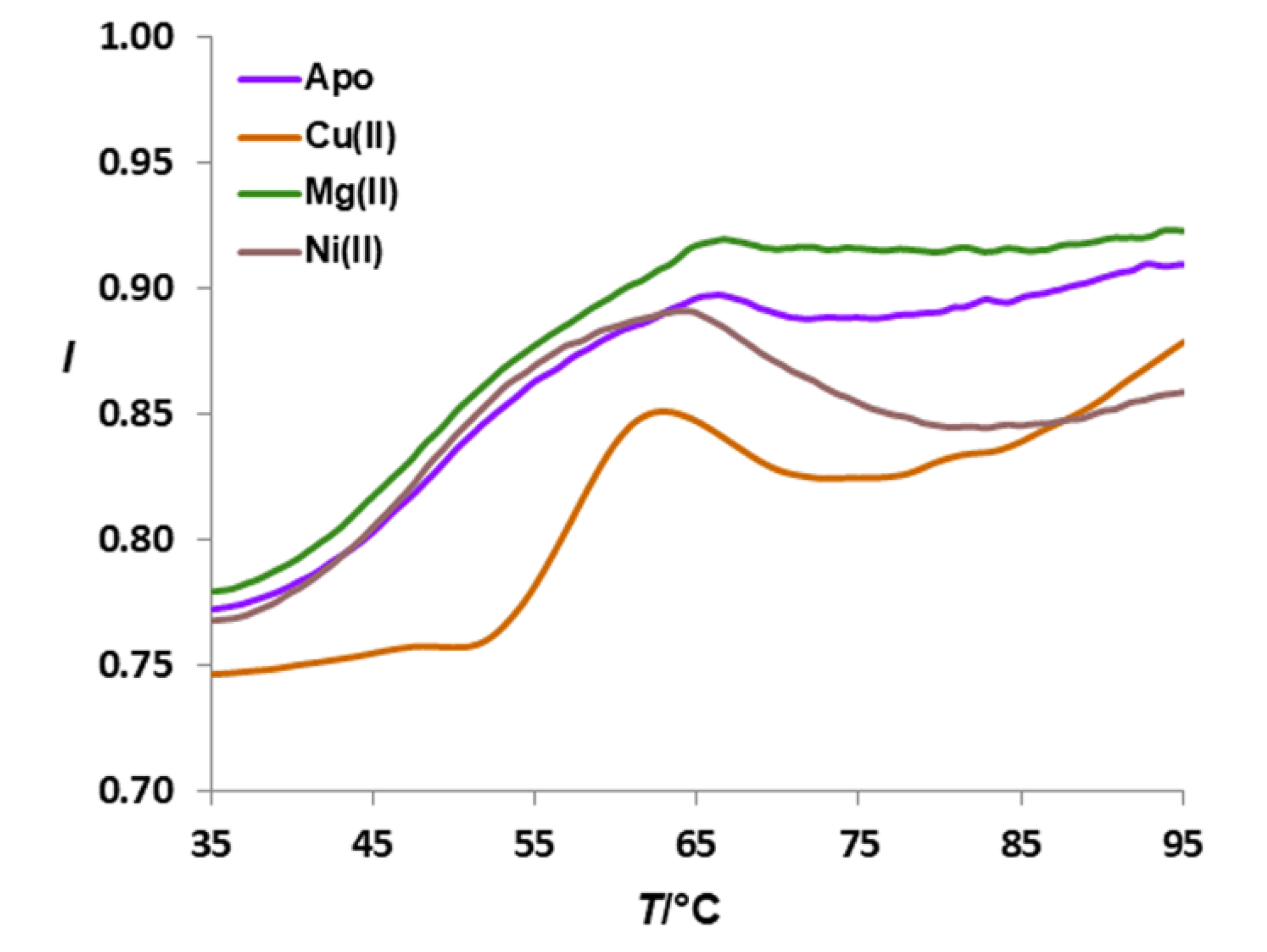
3.2. Size of HpCrdA in Solution
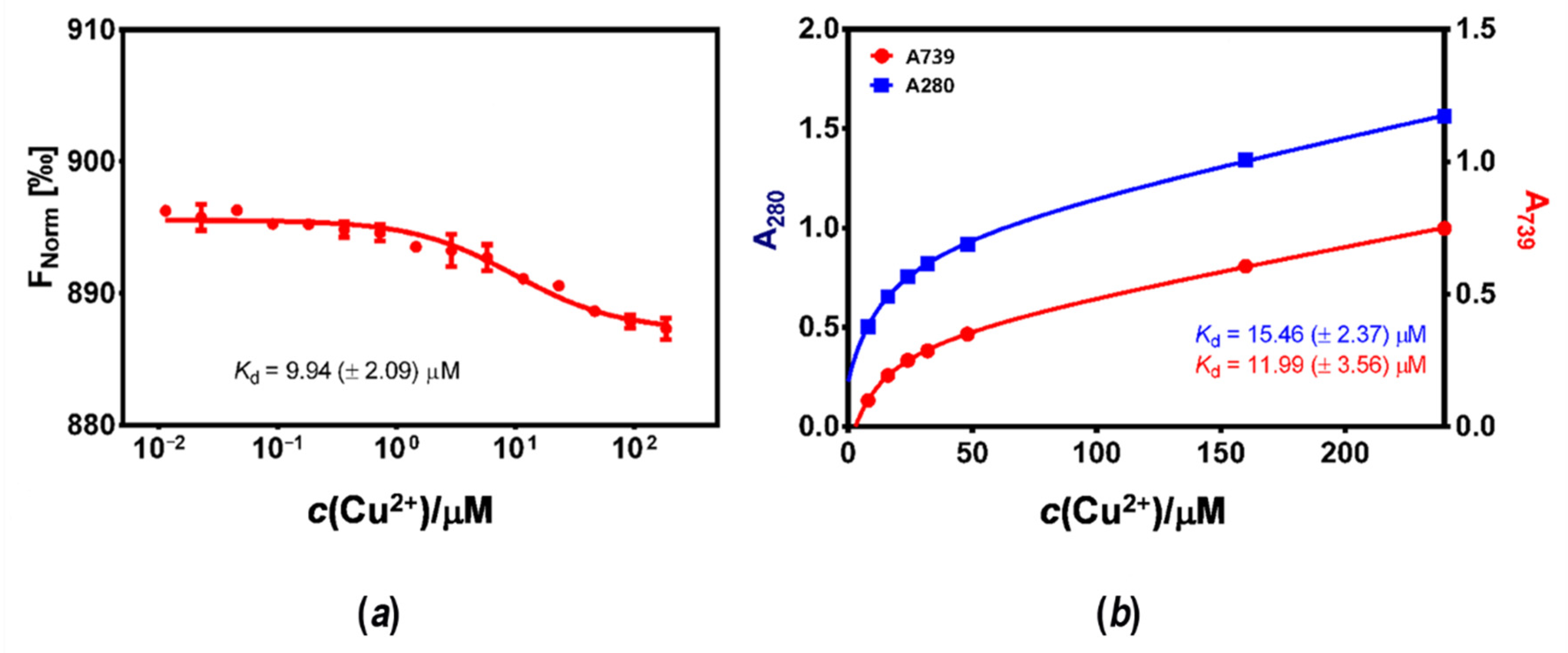
3.3. Affinity of HpCrdA towards Copper(II) Ions
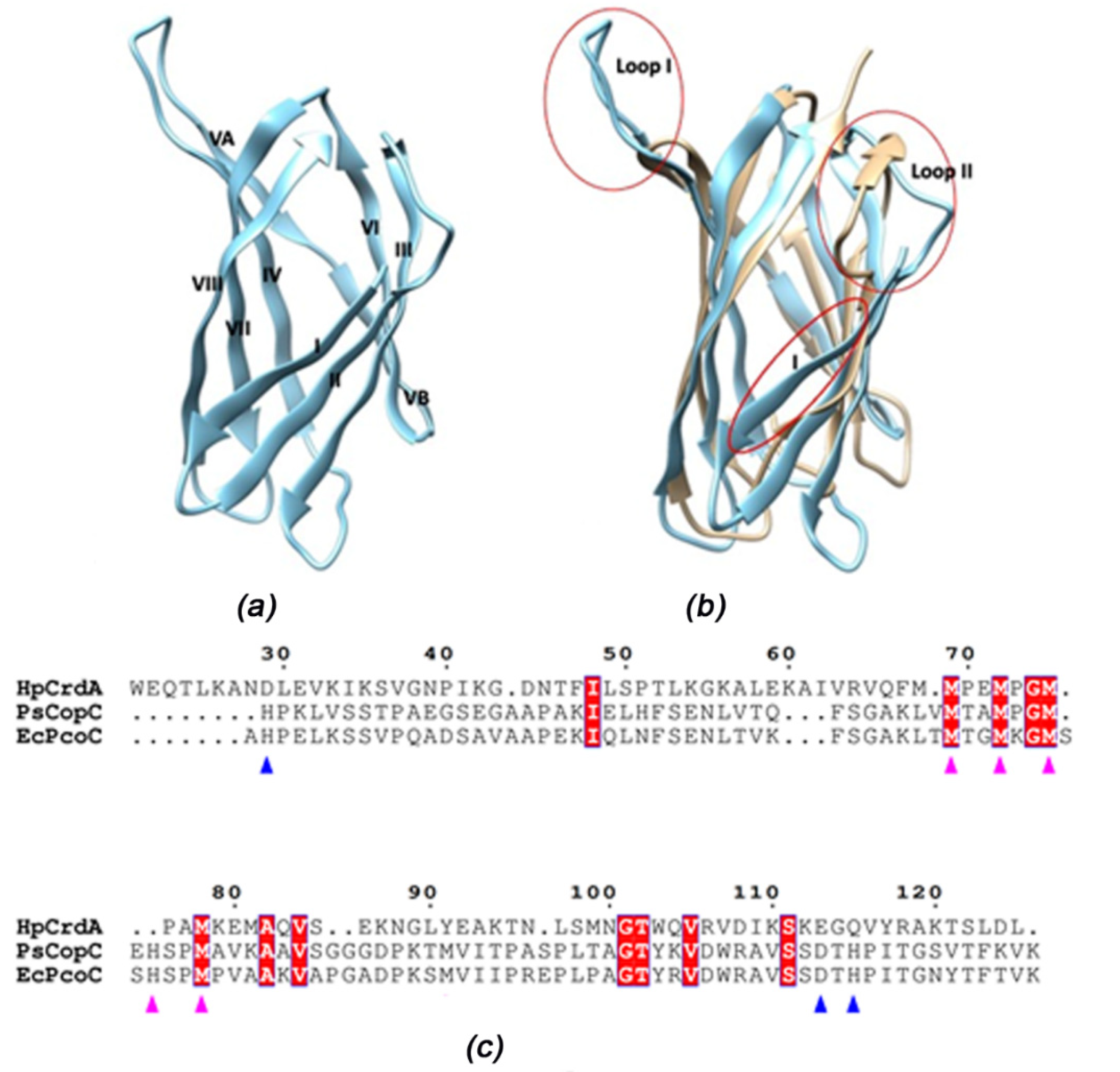
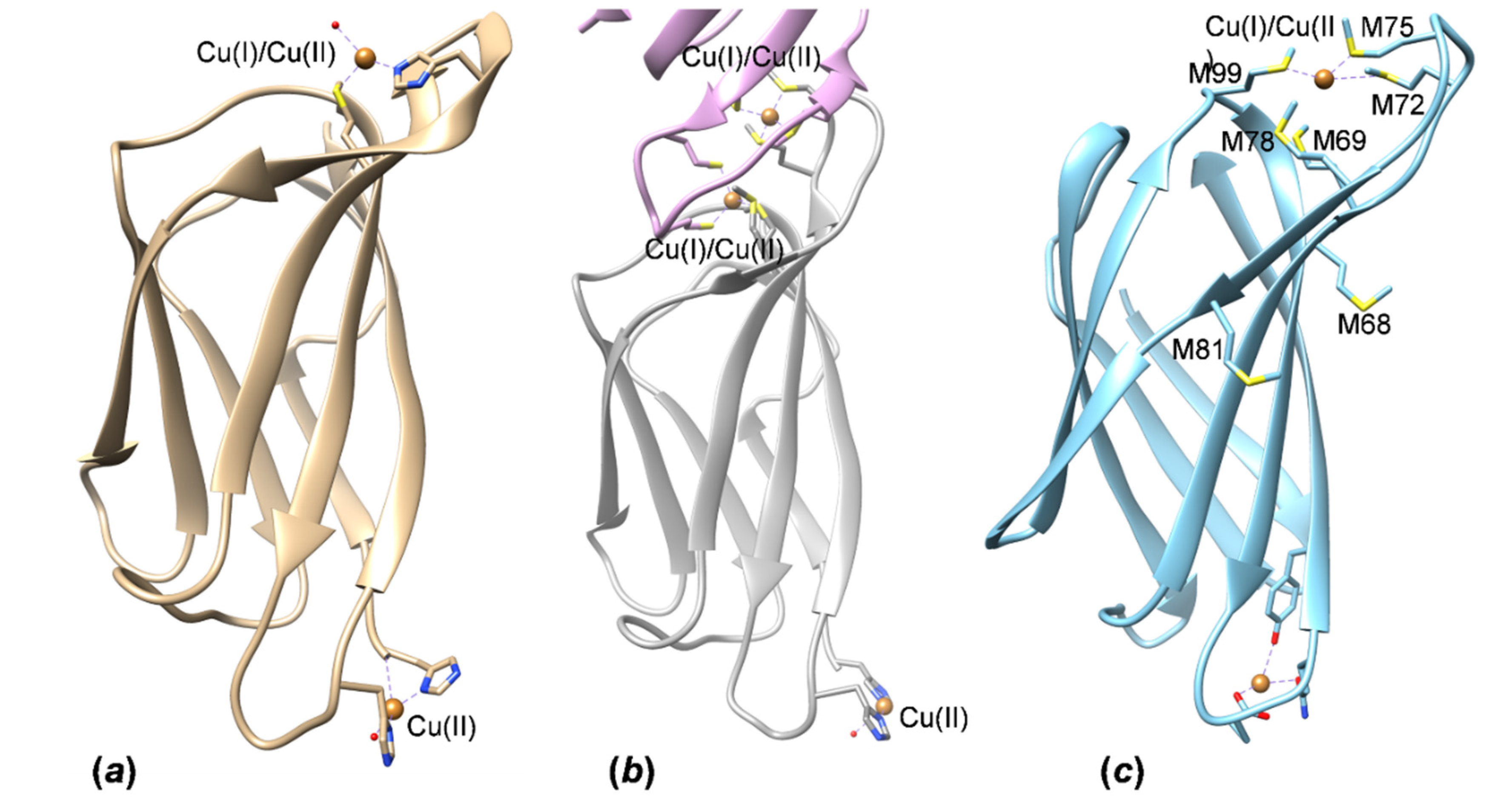
3.4. HpCrdA Modeling
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Bayle, D.; Wangler, S.; Weitzenegger, T.; Steinhilber, W.; Volz, J.; Przybylski, M.; Schafer, K.P.; Sachs, G.; Melchers, K. Properties of the P-type ATPases encoded by the copAP operons of Helicobacter pylori and Helicobacter felis. J. Bacteriol. 1998, 180, 317–329. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Waidner, B.; Melchers, K.; Ivanov, I.; Loferer, H.; Bensch, K.W.; Kist, M.; Bereswill, S. Identification by RNA profiling and mutational analysis of the novel copper resistance determinants CrdA (HP1326), CrdB (HP1327), and CzcB (HP1328) in Helicobacter pylori. J. Bacteriol. 2002, 184, 6700–6708. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gross, E.L. Plastocyanin: Structure and function. Photosynthesis research. Photosynth. Res. 1993, 37, 103–116. [Google Scholar] [CrossRef]
- Rubino, J.T.; Franz, K.J. Coordination chemistry of copper proteins: How nature handles a toxic cargo for essential function. J. Inorg. Biochem. 2012, 107, 129–141. [Google Scholar] [CrossRef] [PubMed]
- Koch, M.; Velarde, M.; Harrison, M.D.; Echt, S.; Fischer, M.; Messerschmidt, A.; Dennison, C. Crystal structures of oxidized and reduced stellacyanin from horseradish roots. J. Am. Chem. Soc. 2005, 127, 158–166. [Google Scholar] [CrossRef]
- Tsukihara, T.; Aoyama, H.; Yamashita, E.; Tomizaki, T.; Yamaguchi, H.; Shinzawa-Itoh, K.; Nakashima, R.; Yaono, R.; Yoshikawa, S. Structures of metal sites of oxidized bovine heart cytochrome c oxidase at 2.8 Å. Science 1995, 269, 1069–1074. [Google Scholar] [CrossRef]
- Waidner, B.; Melchers, K.; Stähler, F.N.; Kist, M.; Bereswill, S. The Helicobacter pylori CrdRS two-component regulation system (HP1364/HP1365) is required for copper-mediated induction of the copper resistance determinant CrdA. J. Bacteriol. 2005, 187, 4683–4688. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.; Koay, M.; Maher, M.J.; Xiao, Z.; Wedd, A.G. Intermolecular transfer of copper ions from the CopC protein of Pseudomonas syringae. Crystal structures of fully loaded CuICuII forms. J. Am. Chem. Soc. 2006, 128, 5834–5850. [Google Scholar]
- Lawton, T.J.; Kenney, G.E.; Hurley, J.D.; Rosenzweig, A.C. The CopC family: Structural and bioinformatic insights into a diverse group of periplasmic copper binding proteins. Biochemistry 2016, 55, 2278–2290. [Google Scholar] [CrossRef] [Green Version]
- Udagedara, S.R.; Wijekoon, C.J.K.; Xiao, Z.; Wedd, A.G.; Maher, M.J. The crystal structure of the CopC protein from Pseudomonas fluorescens reveals amended classifications for the CopC protein family. J. Inorg. Biochem. 2019, 195, 194–200. [Google Scholar] [CrossRef]
- Osipov, E.M.; Lilina, A.V.; Tsallagov, S.I.; Safonova, T.N.; Sorokin, D.Y.; Tikhonova, T.V.; Popov, V.O. Structure of the flavocytochrome c sulfide dehydrogenase associated with the copper-binding protein CopC from the haloalkaliphilic sulfur-oxidizing bacterium Thioalkalivibrio paradoxus ARh 1. Acta Crystallogr. D 2018, 74, 632–642. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brander, S.; Horvath, I.; Ipsen, J.Ø.; Peciulyte, A.; Olsson, L.; Hernández-Rollán, C.; Nørholm, M.H.H.; Mossin, S.; Leggio, L.L.; Probst, C.; et al. Biochemical evidence of both copper chelation and oxygenase activity at the histidine brace. Sci. Rep. 2020, 10, 16369. [Google Scholar] [CrossRef] [PubMed]
- Wijekoon, C.J.K.; Young, T.R.; Wedd, A.G.; Xiao, Z. CopC protein from Pseudomonas fluorescens SBW25 features a conserved novel high-affinity Cu (II) binding site. Inorg. Chem. 2015, 54, 2950–2959. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, M.; Udagedara, S.R.; Sze, C.M.; Ryan, T.M.; Howlett, G.J.; Xiao, Z.; Wedd, A.G. PcoE—A metal sponge expressed to the periplasm of copper resistance Escherichia coli. Implication of its function role in copper resistance. J. Inorg. Biochem. 2015, 115, 186–197. [Google Scholar]
- Bagai, I.; Rensing, C.; Blackburn, N.J.; McEvoy, M.M. Direct metal transfer between periplasmic proteins identifies a bacterial copper chaperone. Biochemistry 2008, 47, 11408–11414. [Google Scholar] [CrossRef] [Green Version]
- Kittleson, J.T.; Loftin, I.R.; Hausrath, A.C.; Engelhardt, K.P.; Rensing, C.; McEvoy, M.M. Periplasmic metal-resistance protein CusF exhibits high affinity and specificity for both CuI and AgI. Biochemistry 2006, 45, 11096–11102. [Google Scholar] [CrossRef]
- Lee, L.J.; Barret, J.A.; Poole, R.K. Genome-wide transcriptional response of chemostat-cultured Escherichia coli to zinc. J. Bacteriol. 2005, 187, 1124–1134. [Google Scholar] [CrossRef] [Green Version]
- Loftin, I.R.; Franke, S.; Blackburn, N.J.; McEvoy, M.M. Unusual Cu(I)/Ag(I) coordination of Escherichia coli CusF as revealed by atomic resolution crystallography and X-ray absorption spectroscopy. Prot. Sci. 2007, 16, 2287–2293. [Google Scholar] [CrossRef] [Green Version]
- Loftin, I.R.; Blackburn, N.J. Tryptophan Cu (I)–π interaction fine-tunes the metal binding properties of the bacterial metallochaperone CusF. J. Biol. Inorg. Chem. 2009, 14, 905–912. [Google Scholar] [CrossRef] [Green Version]
- Pulić, I.; Cendron, L.; Salamina, M.; de Laureto, P.P.; Matković-Čalogović, D.; Zanotti, G. Crystal structure of truncated FlgD from the human pathogen Helicobacter pylori. J. Struct. Biol. 2016, 194, 147–155. [Google Scholar] [CrossRef]
- Loconte, V.; Kekez, I.; Matković-Čalogović, D.; Zanotti, G. Structural characterization of FlgE2 protein from Helicobacter pylori hook. FEBS J. 2017, 284, 4328–4342. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- LipoP Bioinformatic Tool. Available online: http://www.cbs.dtu.dk/services/LipoP/ (accessed on 4 September 2016).
- Bohm, G.; Muhr, R.; Jaenicke, R. Quantitative analysis of protein far UV circular dichroism spectra by neural networks. Protein Eng. 1992, 5, 191–195. [Google Scholar] [CrossRef] [PubMed]
- GraphPad Prism Version 6.01 for Windows; GraphPad Software: San Diego, CA, USA, 2012; Available online: www.graphpad.com (accessed on 18 February 2018).
- Kelly, S.M.; Price, N.C. The application of circular dichroism to studies of protein folding and unfolding. Biochim. Biophys. Acta. 1997, 1338, 161–185. [Google Scholar] [CrossRef]
- Wernimont, A.K.; Huffman, D.L.; Finney, L.A.; Demeler, B.; O’Halloran, T.V.; Rosenzweig, A.C. Crystal structure and dimerization equilibria of PcoC, a methionine-rich copper resistance protein from Escherichia coli. J. Biol. Inorg. Chem. 2003, 8, 185–194. [Google Scholar] [CrossRef]
- Jumper, J. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Varadi, M. AlphaFold Protein Structure Database: Massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucl. Acids Res. 2022, 50, D439–D444. [Google Scholar] [CrossRef]
- Arnesano, F.; Banci, L.; Bertini, I.; Mangani, S.; Thompsett, A.R. A redox switch in CopC: An intriguing copper trafficking protein that binds copper (I) and copper (II) at different sites. Proc. Natl. Acad. Sci. USA 2003, 100, 3814–3819. [Google Scholar] [CrossRef] [Green Version]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [Green Version]
- Robert, X.; Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucl. Acids Res. 2014, 42, W320–W324. [Google Scholar] [CrossRef] [Green Version]
- Harding, M.M. The architecture of metal coordination groups in proteins. Acta Crystallogr. D 2004, 60, 849–859. [Google Scholar] [CrossRef] [Green Version]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kekez, I.; Faletar, M.; Kekez, M.; Cendron, L.; Wright, M.; Zanotti, G.; Matković-Čalogović, D. Copper Binding and Oligomerization Studies of the Metal Resistance Determinant CrdA from Helicobacter pylori. Molecules 2022, 27, 3387. https://doi.org/10.3390/molecules27113387
Kekez I, Faletar M, Kekez M, Cendron L, Wright M, Zanotti G, Matković-Čalogović D. Copper Binding and Oligomerization Studies of the Metal Resistance Determinant CrdA from Helicobacter pylori. Molecules. 2022; 27(11):3387. https://doi.org/10.3390/molecules27113387
Chicago/Turabian StyleKekez, Ivana, Mihovil Faletar, Mario Kekez, Laura Cendron, Maya Wright, Giuseppe Zanotti, and Dubravka Matković-Čalogović. 2022. "Copper Binding and Oligomerization Studies of the Metal Resistance Determinant CrdA from Helicobacter pylori" Molecules 27, no. 11: 3387. https://doi.org/10.3390/molecules27113387
APA StyleKekez, I., Faletar, M., Kekez, M., Cendron, L., Wright, M., Zanotti, G., & Matković-Čalogović, D. (2022). Copper Binding and Oligomerization Studies of the Metal Resistance Determinant CrdA from Helicobacter pylori. Molecules, 27(11), 3387. https://doi.org/10.3390/molecules27113387






