4.1. Dataset
In this study, we used the following benchmark and freely available multitask datasets, whose main characteristics are summarized in
Table 3:
1. NUclear Receptor Activity dataset (NURA) is a recently published dataset [
19] containing information on nuclear receptor modulation by small molecules. Chemicals can bind to nuclear receptors by activating (agonist) or inhibiting (antagonist) the natural biological response. Hence, the NURA dataset contains annotations for binding, agonism, and antagonism activity for 15,206 molecules and 11 selected NRs. For each receptor and each activity type (agonism, binding, antagonism), a molecule can have one of the following annotations: (i) ‘active’; (ii) ‘weakly active’; (iii) ‘inactive’; (iv) ‘inconclusive’; and (v) ‘missing’. In this work, each bioactivity type for a given receptor was considered as a task. As in [
22], only active and inactive annotations were considered and tasks containing such annotations for less than 200 molecules were discarded. The considered dataset is therefore composed of a total of 14,963 chemicals annotated (as active or inactive) for at least one of the selected 30 tasks.
2. Tox21 dataset contains qualitative toxicity measurements on 12 biological targets (i.e., tasks) including nuclear receptors and stress response pathways. This dataset is curated by MoleculeNet as a benchmark for molecular machine learning [
20]. The 7831 compounds were pruned for disconnected structures obtaining a final dataset of 7586 molecules.
3. ClinTox dataset contains qualitative data of drugs approved by the FDA and those that have failed clinical trials for toxicity reasons (i.e., two tasks). Additionally, this dataset is curated by MoleculeNet [
20]. The 1478 compounds were pruned for disconnected structures obtaining a final dataset of 1472 molecules.
Molecules of each dataset were randomly split into the training (80%) and external test set (20%), trying to preserve the proportion between the classes (actives/inactives) for each task (stratified splitting). All analyses were performed in 3-fold cross-validation (i.e., for each architecture, three neural networks were trained on three different subsets of training data and performance was evaluated on the excluded data each time).
For each molecule, we computed extended connectivity fingerprints (ECFPs) [
23] as molecular descriptors and these were used as input variables for neural networks. ECFPs are binary vectors of predefined length that encode the presence/absence of atom-centered substructures through a hashing algorithm. In particular, we computed ECFPs with the following options: 1024 as the fingerprint length, two bits to encode each substructure, and a fragment radius comprised between 0 and two bonds.
4.2. Multitask Neural Network
The most common multitask networks in the literature are constituted by fully connected neural network layers trained on multiple tasks, where the output is shared among all learning tasks and then fed into individual classifiers. When some dependence relationships exist among the tasks, the model should learn a joint representation of these tasks and, thus benefit from an information boost [
2,
24]. Similar to single-task feedforward neural networks, the input vectors are mapped to the output vectors with repeated compositions of layers, which are constituted by neurons. When each neuron of a layer is connected to all the neurons of the following layer, the network is called dense or fully connected. The layers between input and output are called hidden layers. Each connection represents a weight, whereas each node represents a learning function
f that, in the feedforward phase, processes the information of the previous layer to be fed into the subsequent layer. In the backpropagation phase, each weight is adjusted according to the loss function and the optimization algorithm.
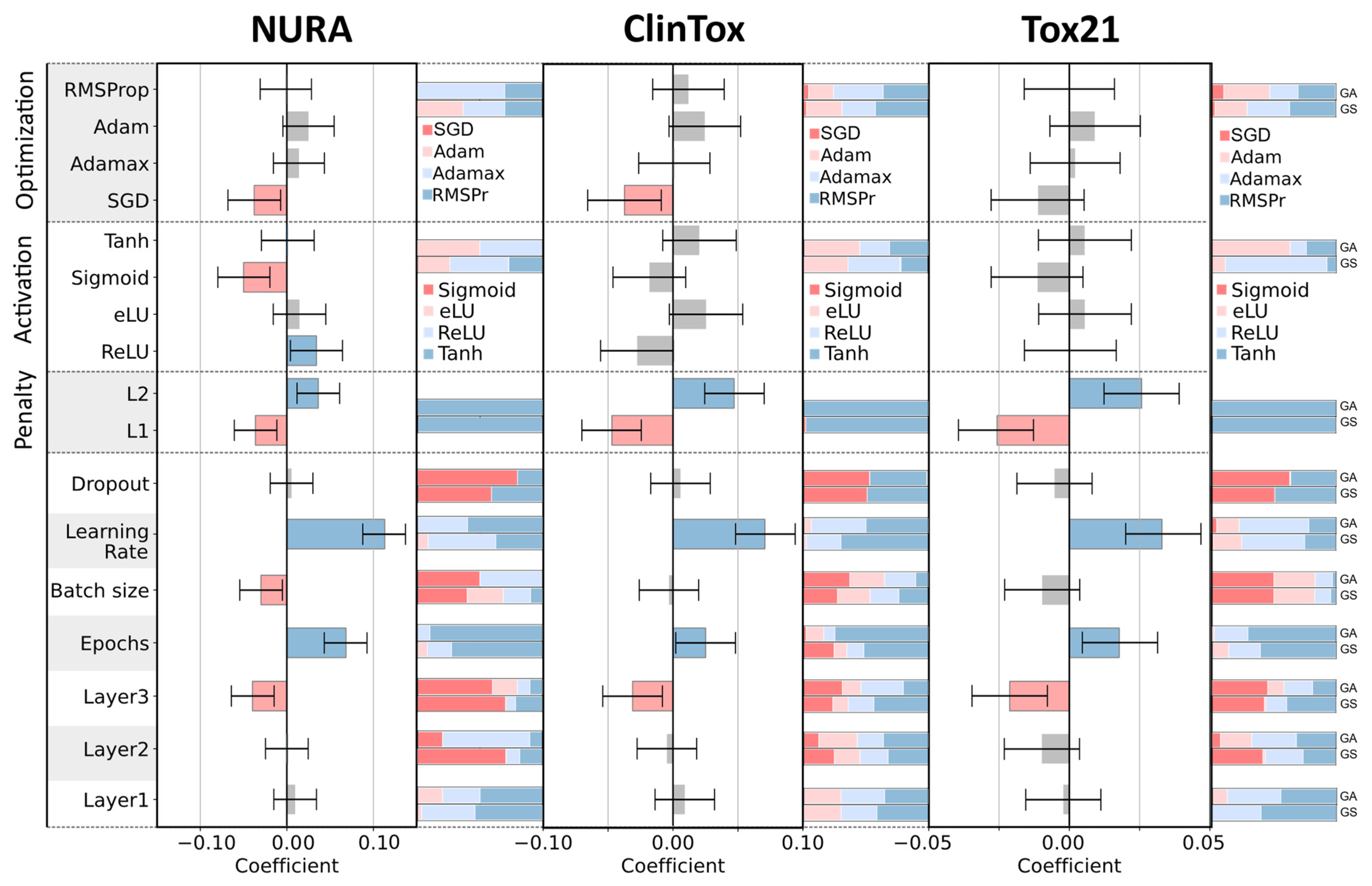
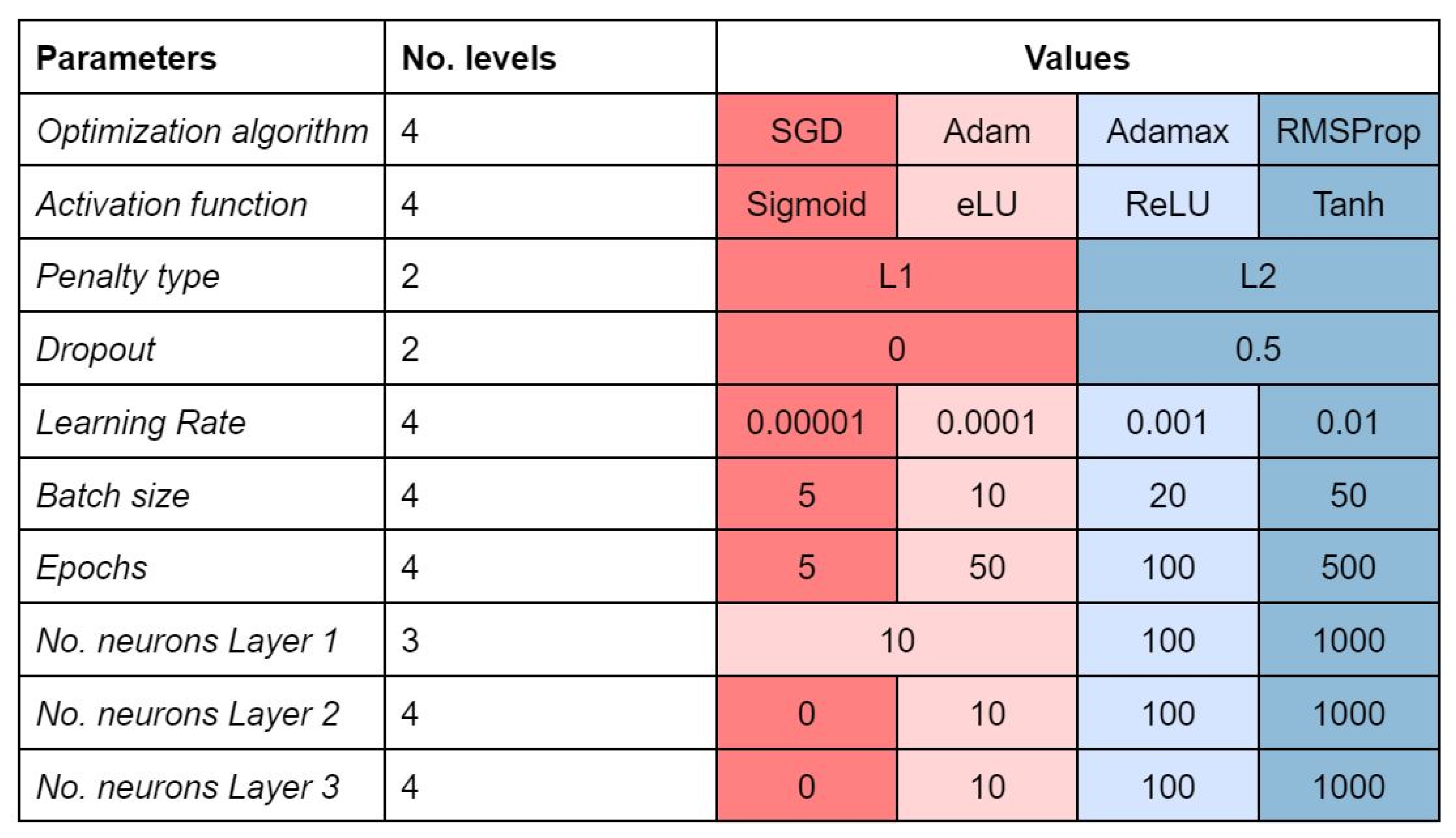
Figure 5 summarizes the hyperparameters we decided to tune.
Different types of activation functions exist in the literature; the most known are sigmoid, rectified linear unit (ReLU), hyperbolic tangent (Tanh), and exponential linear unit (eLU), which can be defined for an input
z as follows:
Neural network tuning implies setting a learning rate that determines the update of the weights in each iteration with respect to the gradient of the loss function. For computational and learning efficiency, in each training step (i.e., iteration), a subset of training examples called batch is used. When all the examples are seen by the model (i.e., after a number of iterations equal to the training set size divided by the batch size), a training epoch is completed.
Furthermore, several strategies called regularization techniques can improve the network’s generalizing ability and reduce overfitting. This is the case of dropout, where a percentage of randomly selected neurons are ignored during training, and the introduction of a penalty also called L1 or L2 regularization, on the basis of the chosen function.
In this work, we used the binary cross-entropy as loss function, which can handle multiple outputs and missing data. We considered both ‘shallow’ (i.e., only one hidden layer) and deep architectures up to three hidden layers, and neurons per layer varying between 0 and 1000. The output layer consists of as many nodes as tasks. The threshold of assignment for the output nodes was optimized on the basis of ROC curves for each task, that is, if the output of the neural network ensemble node is equal or lower than the threshold the compound is predicted inactive, otherwise active. We initialized the network weights randomly according to a truncated normal function with epochs varying from five to 500.
4.4. Optimization Strategies
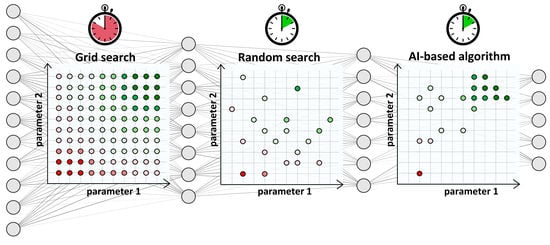
We compared three well-known strategies to tune the network hyperparameters: grid search, random search and genetic Algorithms [
8].
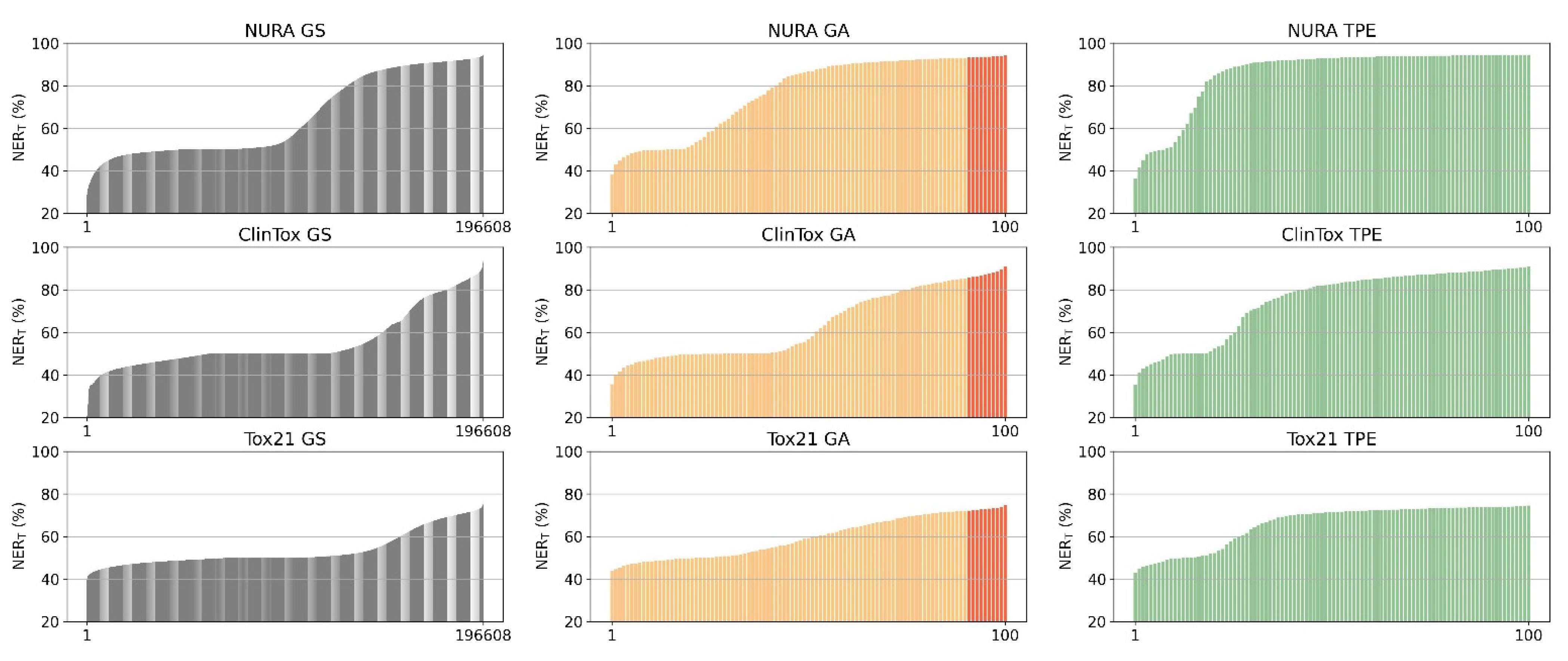
Grid search (GS) is the most computationally expensive and time-consuming approach since it explores all the possible combinations of hyperparameters at the selected levels in the search space. In our case study, 196,608 architectures were tested. GS suffers from high dimensional spaces, but can often be easily parallelized.
Random search (RS) strategy avoids the complete selection of all combinations by a random selection of combinations and is more efficient than GS [
7]. We chose to randomly select a subset of GS combinations to limit the exploration to the levels of values reported in
Figure 5. The number of random combinations to test is user-defined, usually based on a trade-off between available computational time/power and satisfying performance. We chose to select a subset of one hundred combinations.
Genetic algorithms (GA) are heuristic stochastic evolutionary search algorithms based on sequential selections, combinations, and mutations simulating biological evolution. In other words, combinations of hyperparameters leading to a higher performance or fitness survive and have a higher chance to reproduce and generate children with a predefined mutation probability. Each generation consists of a population of chromosomes representing points in the search space. Each individual is represented as a binary vector. The considered algorithm consists of the following steps:
1. Randomly generate a population of 10 chromosomes;
2. Compute the fitness as overall NER (NERT) in 3-fold cross validation for each chromosome of the population;
3. Select two “parental” chromosomes on a roulette wheel basis (i.e., the probability to be selected is proportional to the fitness);
4. Generate two children with a recombination of the parental chromosomes and with a mutation probability for each bit equal to 10%;
5. Evaluate the fitness for the children and insert them into the population;
6. Exclude the two worst-performing chromosomes (i.e., those with the lowest NERT) of the population;
7. Repeat steps 3–7 for 42 times (or generations); and
8. At the 21st generation, replace the worst performing half of the population with randomly generated chromosomes (cataclysm or invasion simulation).
This process allows us to evaluate 100 combinations of hyperparameters.
Tree-structured Parzen estimator (TPE) is a sequential model-based optimization (SMBO) approach [
5]. SMBO methods sequentially construct models to approximate the performance of hyperparameters based on historical measurements, and then subsequently chooses new hyperparameters to test based on this model. The TPE approach models P(
x|
y) and P(
y) following the Bayesian rule, where
x represents hyperparameters and
y the associated quality score. P(
x|
y) is modeled by transforming the generative process of hyperparameters, replacing the distributions of the configuration prior with non-parametric densities. TPE consists of the following steps:
1. Define the domain of hyperparameter search space;
2. Create an objective function which takes in input the hyperparameters and outputs a score to be minimized or maximized;
3. Obtain a couple of observations (score) using randomly selected combinations of hyperparameters;
4. Sort the collected observations by score and divide them into two groups: group (x1) contains observations that gave the best scores and the second one (x2) all other observations;
5. Model two densities l(x1) and g(x2) using Parzen estimators (or kernel density estimators);
6. Raw sample hyperparameters from l(x1), evaluating them in terms of l(x1)/g(x2), and returning the set that yields the minimum value under l(x1)/g(x1) corresponding to the greatest expected improvement. These hyperparameters are then evaluated on the objective function;
7. Update the observation list from step 3; and
8. Repeat steps 4–7 with a fixed number of trials.
We set the total number of trials to 100 and the 3-fold cross-validation NERT as the score to maximize. We chose predefined values for each hyperparameters in order to constrain the possible combinations to the grid search space.
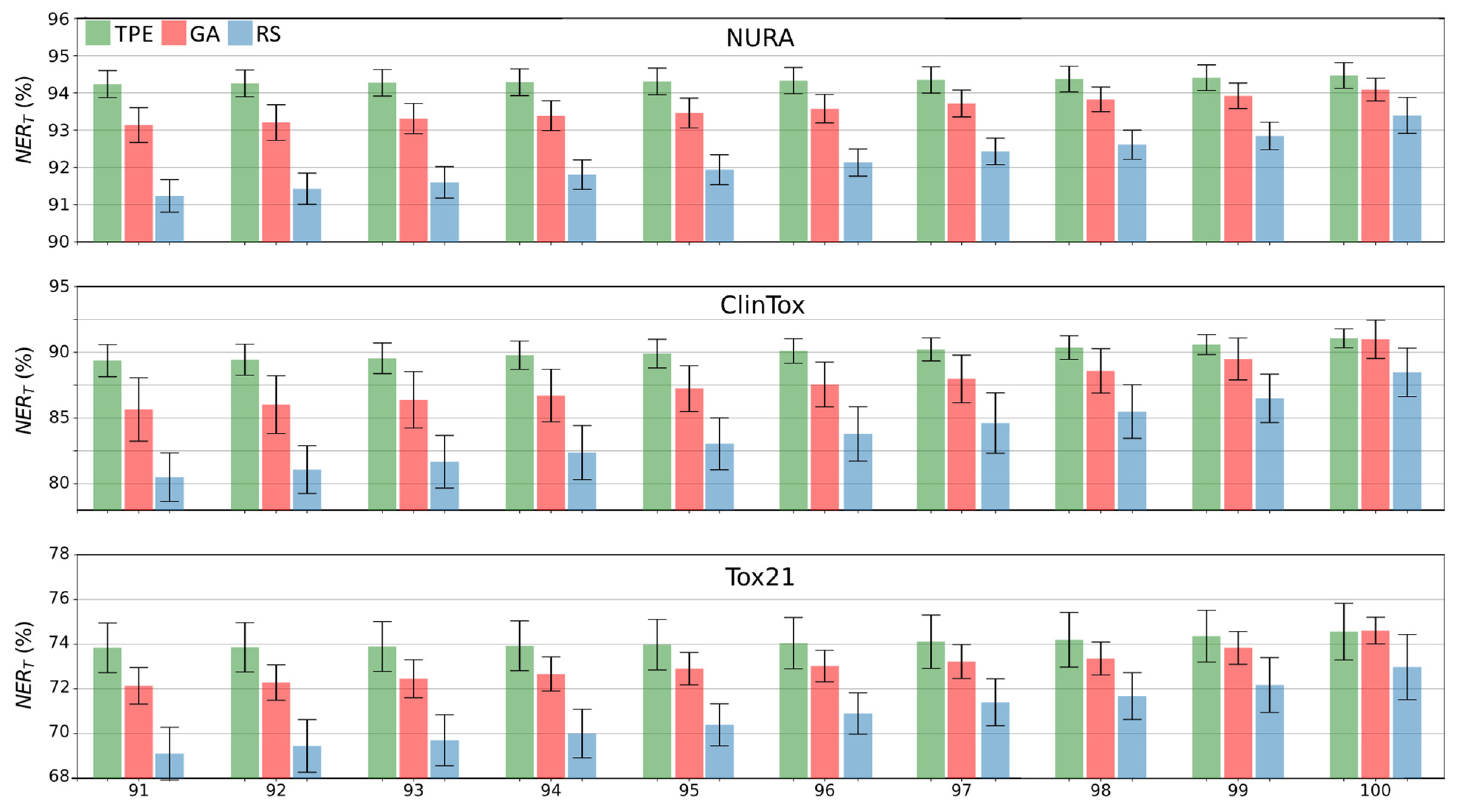
RS, GA, and TPE strategies were independently repeated ten times to guarantee robustness of the results and to measure their uncertainty.
Experimental design (or design of experiments, DoE) aims at maximizing the information on an investigated system or process and at the same time minimizing the experimental effort; this is achieved by a rational experimental plan to obtain the best knowledge of the system [
21]. In this framework, DoE was used to obtain quantitative relationships between architectures and the performance of multitask networks, thus, to understand how the choice of hyperparameter values can directly influence the modeling.
The D-optimal design is a particular kind of experimental design [
26], which is commonly used when (i) the shape of the analyzed domain is not regular; (ii) there are qualitative factors together with quantitative ones; and (iii) the number of experimental runs has to be reduced. This was the case with the application of DoE on the hyperparameters in this case study, where both qualitative and quantitative factors were present and, being the number of considered hyperparameters large, we had to reduce the computational effort in terms of neural network architectures tested.