In Silico and In Vivo Evaluation of SARS-CoV-2 Predicted Epitopes-Based Candidate Vaccine
Abstract
:1. Introduction
2. Results
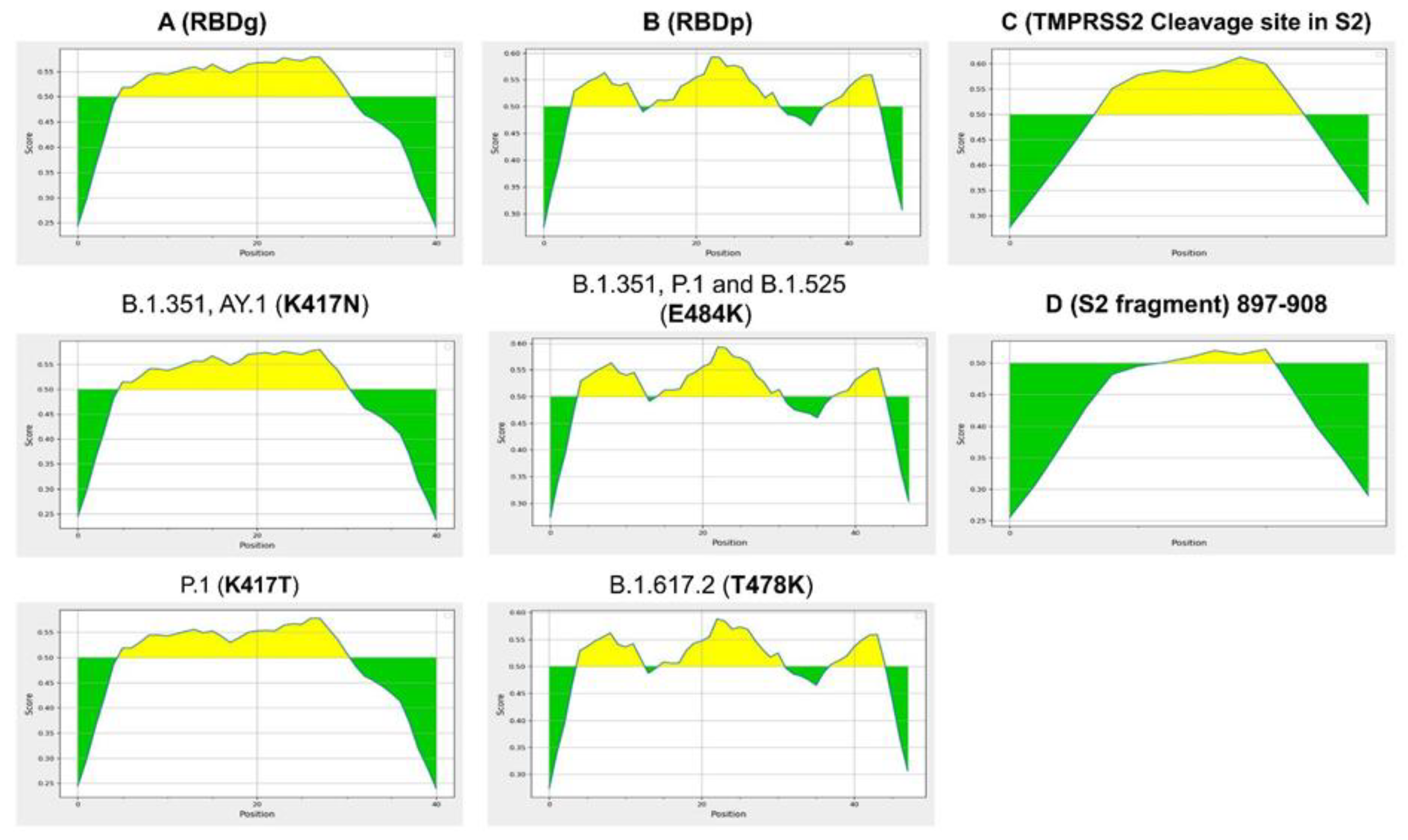
2.1. Computational Design of Neutralizing Epitopes
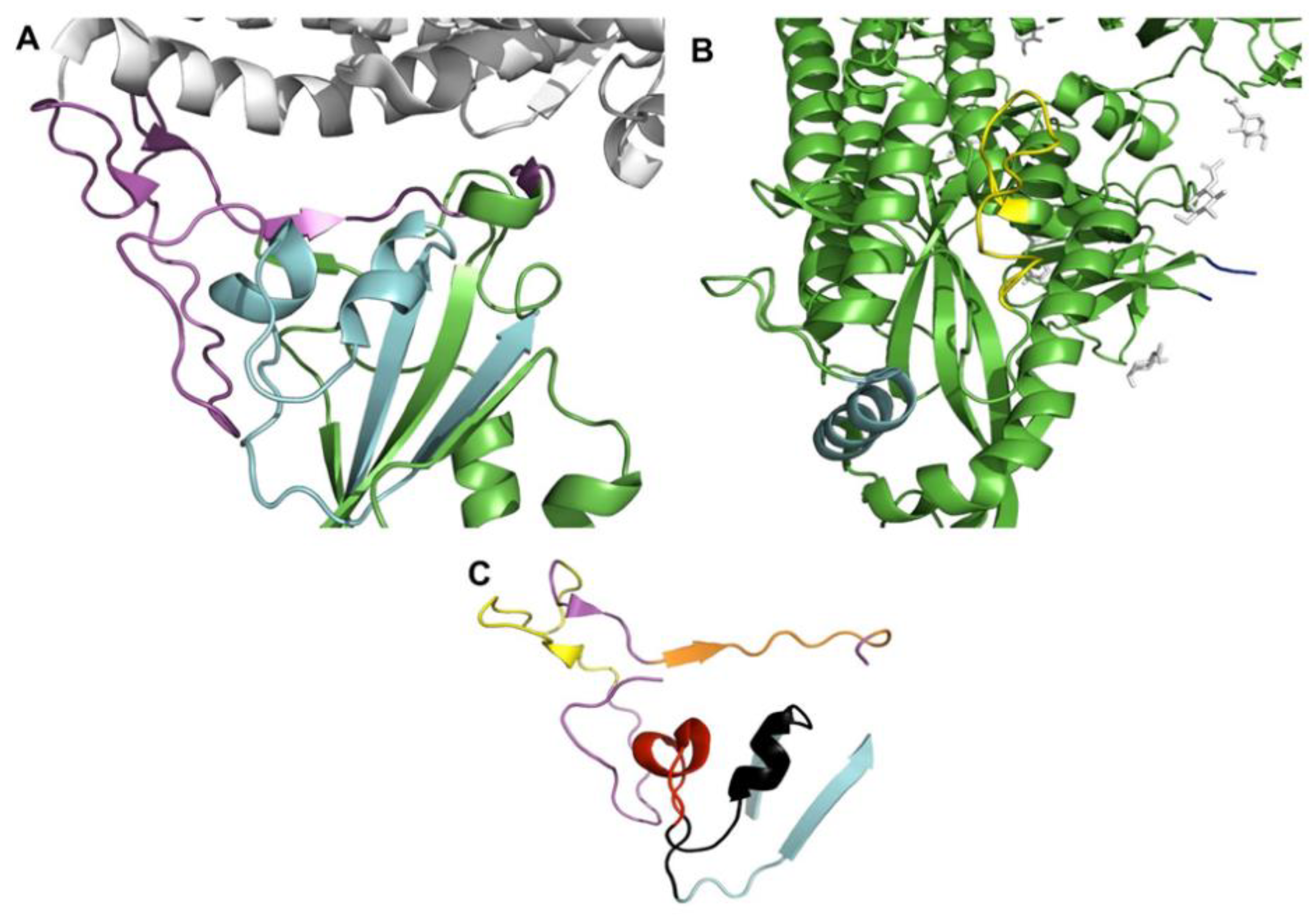
2.2. Structural Configuration of Spike ACE2 Interaction
2.3. Predicted Epitopes and VOCs
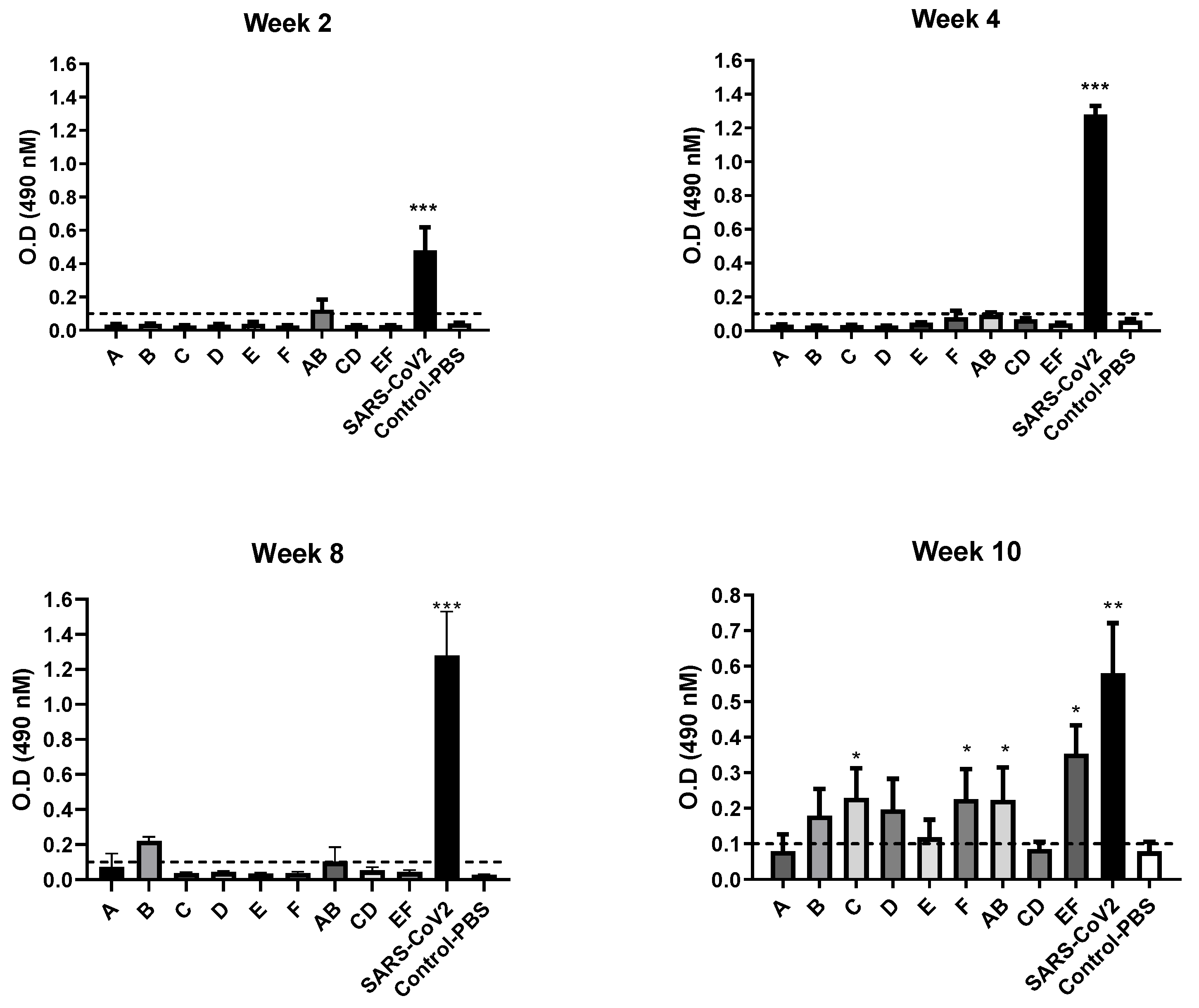
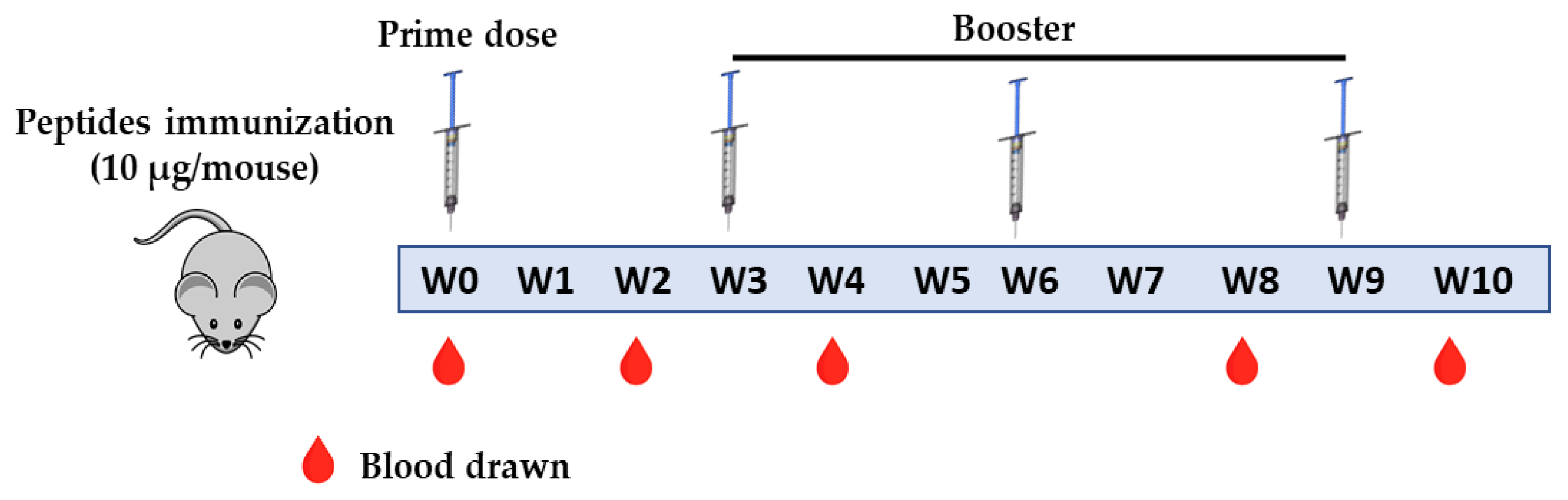
2.4. Mice Experiment
3. Discussion
4. Materials and Methods
4.1. Analysis of the Receptor Binding Domain for Epitope Prediction
4.2. Synthesis of the Peptides Panel
4.3. Preparation of Inactivated SARS-CoV-2 Antigen
4.4. Animal Experiment
4.5. ELISA Assay
4.6. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- WHO. Weekly Epidemiological Update on COVID-19—31 August 2021. Available online: https://www.who.int/publications/m/item/weekly-epidemiological-update-on-covid-19---31-august-2021 (accessed on 3 September 2021).
- Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species Severe acute respiratory syndrome-related coronavirus: Classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 2020, 5, 536–544. [Google Scholar] [CrossRef] [Green Version]
- Mostafa, A.; Kandeil, A.; Shehata, M.; El Shesheny, R.; Samy, A.M.; Kayali, G.; Ali, M.A. Middle East Respiratory Syndrome Coronavirus (MERS-CoV): State of the Science. Microorganisms 2020, 8, 991. [Google Scholar] [CrossRef]
- Dai, L.; Gao, G.F. Viral targets for vaccines against COVID-19. Nat. Rev. Immunol. 2021, 21, 73–82. [Google Scholar] [CrossRef]
- Harrison, A.G.; Lin, T.; Wang, P. Mechanisms of SARS-CoV-2 Transmission and Pathogenesis. Trends Immunol. 2020, 41, 1100–1115. [Google Scholar] [CrossRef]
- Tai, W.; He, L.; Zhang, X.; Pu, J.; Voronin, D.; Jiang, S.; Zhou, Y.; Du, L. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: Implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol. Immunol. 2020, 17, 613–620. [Google Scholar] [CrossRef] [Green Version]
- WHO. COVID-19 Vaccine Tracker and Landscape. Available online: https://www.who.int/publications/m/item/draft-landscape-of-covid-19-candidate-vaccines (accessed on 6 October 2021).
- Abduljaleel, Z.; Al-Allaf, F.A.; Aziz, S.A. Peptides-based vaccine against SARS-nCoV-2 antigenic fragmented synthetic epitopes recognized by T cell and beta-cell initiation of specific antibodies to fight the infection. Biodes. Manuf. 2021, 4, 490–505. [Google Scholar] [CrossRef]
- Saha, R.; Ghosh, P.; Burra, V. Designing a next generation multi-epitope based peptide vaccine candidate against SARS-CoV-2 using computational approaches. 3 Biotech 2021, 11, 47. [Google Scholar] [CrossRef]
- Singh, A.; Thakur, M.; Sharma, L.K.; Chandra, K. Designing a multi-epitope peptide based vaccine against SARS-CoV-2. Sci. Rep. 2020, 10, 16219. [Google Scholar] [CrossRef] [PubMed]
- Grifoni, A.; Sidney, J.; Zhang, Y.; Scheuermann, R.H.; Peters, B.; Sette, A. A Sequence Homology and Bioinformatic Approach Can Predict Candidate Targets for Immune Responses to SARS-CoV-2. Cell Host Microbe 2020, 27, 671–680.e2. [Google Scholar] [CrossRef] [PubMed]
- Tarek, M.; Elhefnawi, M.; Maricato, J.T.; Diaz, R.S.; Shytaj, I.L.; Savarino, A. Custommune: A web tool to design personalized and population-targeted vaccine epitopes. J. medRxiv 2020. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Kruger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef] [PubMed]
- Wu, K.; Werner, A.P.; Moliva, J.I.; Koch, M.; Choi, A.; Stewart-Jones, G.B.E.; Bennett, H.; Boyoglu-Barnum, S.; Shi, W.; Graham, B.S.; et al. mRNA-1273 vaccine induces neutralizing antibodies against spike mutants from global SARS-CoV-2 variants. bioRxiv 2021. [Google Scholar] [CrossRef]
- Corey, L.; Mascola, J.R.; Fauci, A.S.; Collins, F.S. A strategic approach to COVID-19 vaccine R&D. Science 2020, 368, 948–950. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Shi, W.; Joyce, M.G.; Modjarrad, K.; Zhang, Y.; Leung, K.; Lees, C.R.; Zhou, T.; Yassine, H.M.; Kanekiyo, M.; et al. Evaluation of candidate vaccine approaches for MERS-CoV. Nat. Commun. 2015, 6, 7712. [Google Scholar] [CrossRef]
- Yang, H.; Kim, D.S. Peptide Immunotherapy in Vaccine Development: From Epitope to Adjuvant. Adv. Protein Chem. Struct. Biol. 2015, 99, 1–14. [Google Scholar] [CrossRef]
- Guo, J.P.; Petric, M.; Campbell, W.; McGeer, P.L. SARS corona virus peptides recognized by antibodies in the sera of convalescent cases. Virology 2004, 324, 251–256. [Google Scholar] [CrossRef] [Green Version]
- Requena, D.; Medico, A.; Chacon, R.D.; Ramirez, M.; Marin-Sanchez, O. Identification of Novel Candidate Epitopes on SARS-CoV-2 Proteins for South America: A Review of HLA Frequencies by Country. Front. Immunol. 2020, 11, 2008. [Google Scholar] [CrossRef]
- Mitra, D.; Pandey, J.; Jain, A.; Swaroop, S. In silico design of multi-epitope-based peptide vaccine against SARS-CoV-2 using its spike protein. J. Biomol. Struct. Dyn. 2021, 1–14. [Google Scholar] [CrossRef]
- Alam, A.; Khan, A.; Imam, N.; Siddiqui, M.F.; Waseem, M.; Malik, M.Z.; Ishrat, R. Design of an epitope-based peptide vaccine against the SARS-CoV-2: A vaccine-informatics approach. Brief. Bioinform. 2021, 22, 1309–1323. [Google Scholar] [CrossRef]
- Rahman, M.S.; Hoque, M.N.; Islam, M.R.; Akter, S.; Rubayet Ul Alam, A.S.M.; Siddique, M.A.; Saha, O.; Rahaman, M.M.; Sultana, M.; Crandall, K.A.; et al. Epitope-based chimeric peptide vaccine design against S, M and E proteins of SARS-CoV-2, the etiologic agent of COVID-19 pandemic: An in silico approach. PeerJ 2020, 8, e9572. [Google Scholar] [CrossRef]
- Xu, C.; Wang, Y.; Liu, C.; Zhang, C.; Han, W.; Hong, X.; Wang, Y.; Hong, Q.; Wang, S.; Zhao, Q.; et al. Conformational dynamics of SARS-CoV-2 trimeric spike glycoprotein in complex with receptor ACE2 revealed by cryo-EM. Sci. Adv. 2021, 7. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [Green Version]
- Jespersen, M.C.; Peters, B.; Nielsen, M.; Marcatili, P. BepiPred-2.0: Improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Res. 2017, 45, W24–W29. [Google Scholar] [CrossRef] [Green Version]
- Dhanda, S.K.; Vir, P.; Raghava, G.P. Designing of interferon-gamma inducing MHC class-II binders. Biol. Direct. 2013, 8, 30. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, A.C.J.; Powell, J.E.; Tregear, G.W.; Niall, H.D.; Stevens, V.C. A Method for Preparing Beta-Hcg Cooh Peptide-Carrier Conjugates of Predictable Composition. Mol. Immunol. 1980, 17, 749–756. [Google Scholar] [CrossRef]
- Snijders, A.; Benaissatrouw, B.J.; Oosterlaken, T.A.M.; Puijk, W.C.; Posthumus, W.P.A.; Meloen, R.H.; Boere, W.A.M.; Oosting, J.D.; Kraaijeveld, C.A.; Snippe, H. Identification of Linear Epitopes on Semliki Forest Virus E2 Membrane-Protein and Their Effectiveness as a Synthetic Peptide Vaccine. J. Gen. Virol. 1991, 72, 557–565. [Google Scholar] [CrossRef]
- Nagata, T.; Toyota, T.; Ishigaki, H.; Ichihashi, T.; Kajino, K.; Kashima, Y.; Itoh, Y.; Mori, M.; Oda, H.; Yamamura, H.; et al. Peptides coupled to the surface of a kind of liposome protect infection of influenza viruses. Vaccine 2007, 25, 4914–4921. [Google Scholar] [CrossRef] [PubMed]
- Kandeil, A.; Mostafa, A.; Hegazy, R.R.; El-Shesheny, R.; El Taweel, A.; Gomaa, M.R.; Shehata, M.; Elbaset, M.A.; Kayed, A.E.; Mahmoud, S.H.; et al. Immunogenicity and Safety of an Inactivated SARS-CoV-2 Vaccine: Preclinical Studies. Vaccines 2021, 9, 214. [Google Scholar] [CrossRef]
- Classen, D.C.; Morningstar, J.M.; Shanley, J.D. Detection of antibody to murine cytomegalovirus by enzyme-linked immunosorbent and indirect immunofluorescence assays. J. Clin. Microbiol. 1987, 25, 600–604. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Short Peptide Names | Neutralizing Epitope Sequence | Location | SARS-CoV-2 Variants | IFNepitope Score |
|---|---|---|---|---|
| A | QTGKIADYNYK | RBDg * | Wuhan_Reference B.1.1.7, A.23.1 B.1.525, B.1.617.2 | 0.23869643 |
| B | TEIYQAGSTPCNGVEG | RBDp * | Wuhan_Reference B.1.1.7, A.23.1 | 0.256374 |
| C | LQSYGFQPT | RBDp * | Wuhan_Reference P.1, B.1.351, B.1.1.7 A.23.1, B.1.525, B.1.617.2 | 0.25635466 |
| D | IRGDEVRQIAPGQTGKIADYNYKLPD | RBDg * | Wuhan_Reference B.1.1.7, A.23.1, B.1.525, B.1.617.2 | 0.22687041 |
| E | FSQILPDPSKPSKRS | TMPRSS-2 Cleavage site in S2 * | Wuhan_Reference P.1, B.1.351, B.1.1.7 A.23.1, B.1.525, B.1.617.2 | 3.6814886 |
| F | PFAMQMAYRFNG | S2 fragment | Wuhan_Reference P.1, B.1.351, B.1.1.7 A.23.1, B.1.525, B.1.617.2 | 3.5004815 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shehata, M.M.; Mahmoud, S.H.; Tarek, M.; Al-Karmalawy, A.A.; Mahmoud, A.; Mostafa, A.; M. Elhefnawi, M.; Ali, M.A. In Silico and In Vivo Evaluation of SARS-CoV-2 Predicted Epitopes-Based Candidate Vaccine. Molecules 2021, 26, 6182. https://doi.org/10.3390/molecules26206182
Shehata MM, Mahmoud SH, Tarek M, Al-Karmalawy AA, Mahmoud A, Mostafa A, M. Elhefnawi M, Ali MA. In Silico and In Vivo Evaluation of SARS-CoV-2 Predicted Epitopes-Based Candidate Vaccine. Molecules. 2021; 26(20):6182. https://doi.org/10.3390/molecules26206182
Chicago/Turabian StyleShehata, Mahmoud M., Sara H. Mahmoud, Mohammad Tarek, Ahmed A. Al-Karmalawy, Amal Mahmoud, Ahmed Mostafa, Mahmoud M. Elhefnawi, and Mohamed A. Ali. 2021. "In Silico and In Vivo Evaluation of SARS-CoV-2 Predicted Epitopes-Based Candidate Vaccine" Molecules 26, no. 20: 6182. https://doi.org/10.3390/molecules26206182
APA StyleShehata, M. M., Mahmoud, S. H., Tarek, M., Al-Karmalawy, A. A., Mahmoud, A., Mostafa, A., M. Elhefnawi, M., & Ali, M. A. (2021). In Silico and In Vivo Evaluation of SARS-CoV-2 Predicted Epitopes-Based Candidate Vaccine. Molecules, 26(20), 6182. https://doi.org/10.3390/molecules26206182








