A Review of Wine Authentication Using Spectroscopic Approaches in Combination with Chemometrics
Abstract
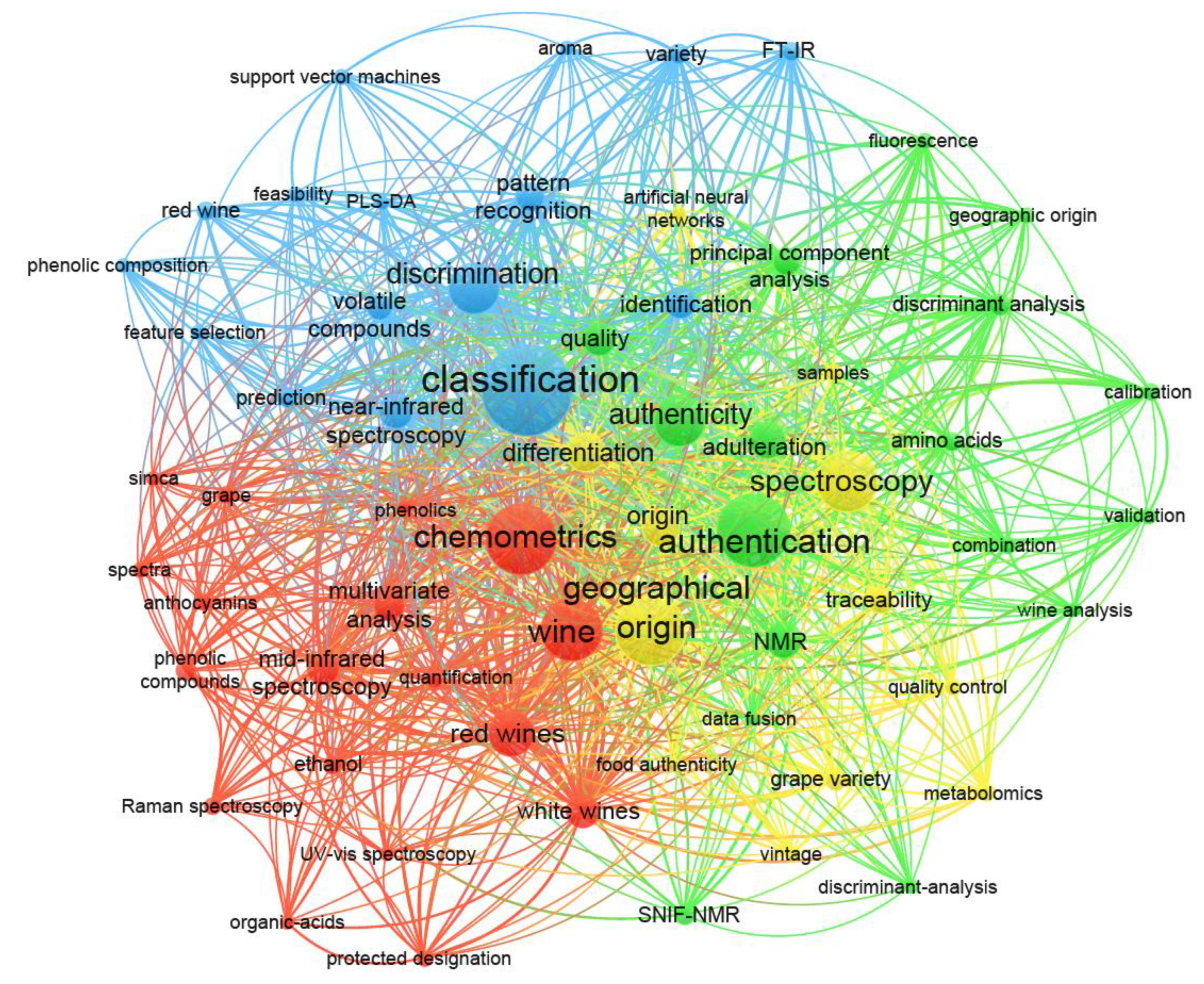
:1. Introduction
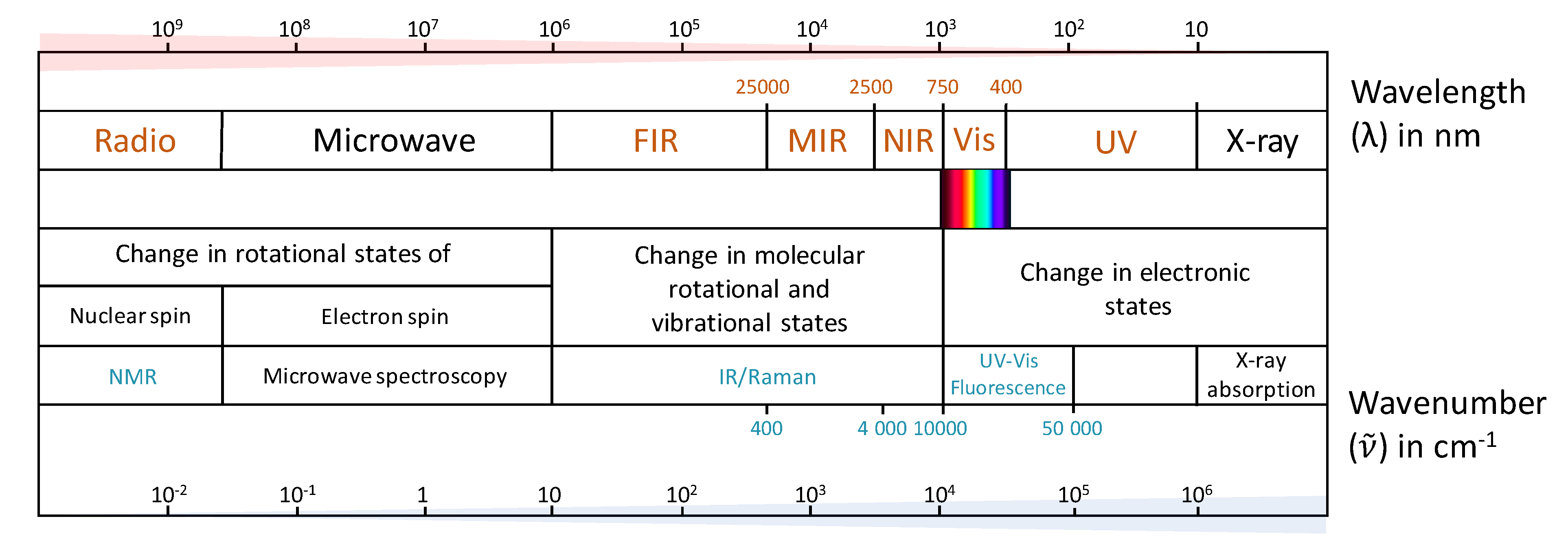
2. Spectroscopic Techniques Applied in Wine Authentication
2.1. UV–Vis Spectroscopy
2.2. IR Spectroscopy
2.3. Raman Spectroscopy
2.4. Fluorescence Spectroscopy
2.5. NMR Spectroscopy
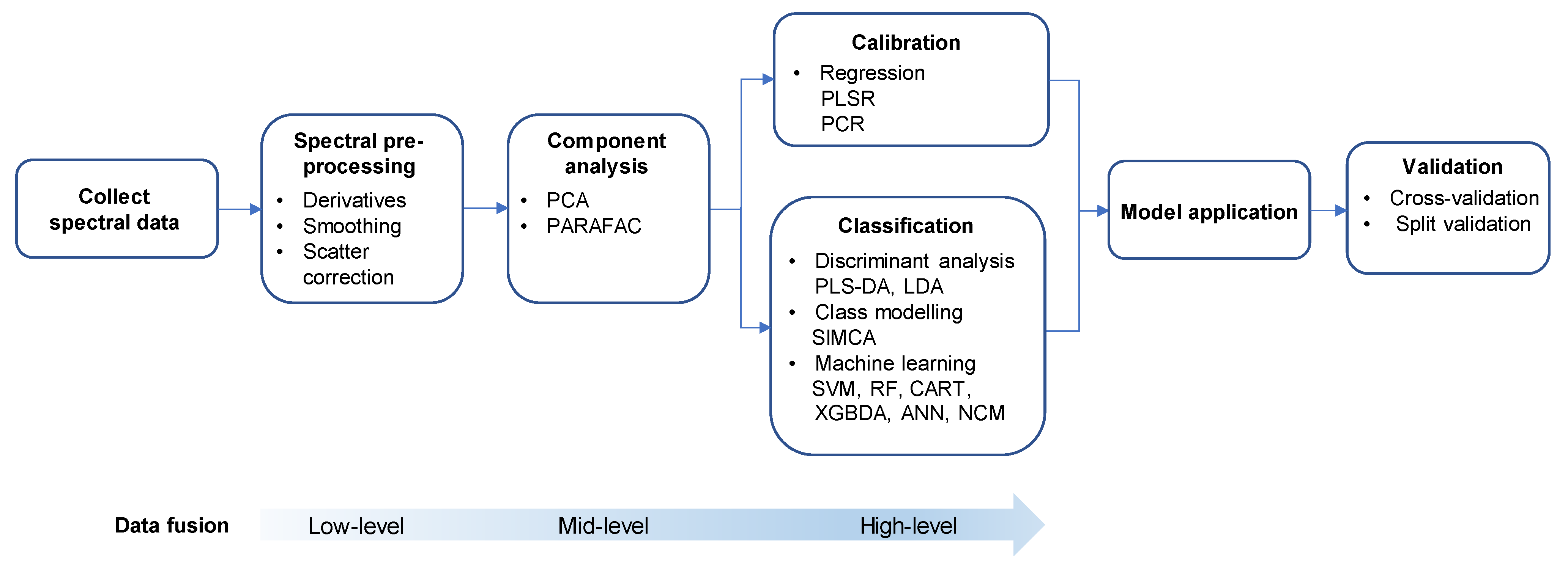
3. Application of Chemometrics for Modelling with Spectroscopic Data
Steps of Chemometric Analysis
4. Future Trends and Directions
Author Contributions
Funding
Conflicts of Interest
References
- Markoski, M.M.; Garavaglia, J.; Oliveira, A.; Olivaes, J.; Marcadenti, A. Molecular properties of red wine compounds and cardiometabolic benefits. Nutr. Metab. Insights 2016, 9, 51–57. [Google Scholar] [CrossRef]
- Waterhouse, A.L.; Sacks, G.L.; Jeffery, D.W. Introduction. In Understanding Wine Chemistry; Waterhouse, A.L., Sacks, G.L., Jeffery, D.W., Eds.; John Wiley & Sons, Ltd.: Chichester, UK, 2016. [Google Scholar]
- Médina, B.; Salagoïty, M.H.; Guyon, F.; Gaye, J.; Hubert, P.; Guillaume, F. Using new analytical approaches to verify the origin of wine. In New Analytical Approaches for Verifying the Origin of Food; Brereton, P., Ed.; Woodhead Publishing: Cambridge, UK, 2013; pp. 149–188. [Google Scholar]
- Waterhouse, A.L.; Sacks, G.L.; Jeffery, D.W. Authentication. In Understanding Wine Chemistry; Waterhouse, A.L., Sacks, G.L., Jeffery, D.W., Eds.; John Wiley & Sons, Ltd.: Chichester, UK, 2016; pp. 383–391. [Google Scholar]
- Popîrdă, A.; Luchian, C.E.; Cotea, V.V.; Colibaba, L.C.; Scutarașu, E.C.; Toader, A.M. A review of representative methods used in wine authentication. Agriculture 2021, 11, 225. [Google Scholar] [CrossRef]
- Arslan, M.; Tahir, H.E.; Zareef, M.; Shi, J.; Rakha, A.; Bilal, M.; Xiaowei, H.; Zhihua, L.; Xiaobo, Z. Recent trends in quality control, discrimination and authentication of alcoholic beverages using nondestructive instrumental techniques. Trends Food Sci. Technol. 2021, 107, 80–113. [Google Scholar] [CrossRef]
- Chapman, J.; Gangadoo, S.; Truong, V.K.; Cozzolino, D. Spectroscopic approaches for rapid beer and wine analysis. Curr. Opin. Food Sci. 2019, 28, 67–73. [Google Scholar] [CrossRef]
- Grijalva-Verdugo, C.; Hernández-Martínez, M.; Meza-Márquez, O.G.; Gallardo-Velázquez, T.; Osorio-Revilla, G. FT-MIR spectroscopy and multivariate analysis for determination of bioactive compounds and antioxidant capacity in Cabernet Sauvignon wines. CyTA-J. Food 2018, 16, 561–569. [Google Scholar] [CrossRef] [Green Version]
- Cayuela, J.A.; Puertas, B.; Cantos-Villar, E. Assessing wine sensory attributes using Vis/NIR. Eur. Food Res. Technol. 2017, 243, 941–953. [Google Scholar] [CrossRef]
- Teixeira dos Santos, C.A.; Pascoa, R.; Sarraguca, M.C.; Porto, P.; Cerdeira, A.L.; Gonzalez-Saiz, J.M.; Pizarro, C.; Lopes, J.A. Merging vibrational spectroscopic data for wine classification according to the geographic origin. Food Res. Int. 2017, 102, 504–510. [Google Scholar] [CrossRef]
- Ranaweera, R.K.R.; Gilmore, A.M.; Capone, D.L.; Bastian, S.E.P.; Jeffery, D.W. Authentication of the geographical origin of Australian Cabernet Sauvignon wines using spectrofluorometric and multi-element analyses with multivariate statistical modelling. Food Chem. 2021, 335, 127592. [Google Scholar] [CrossRef] [PubMed]
- van Eck, N.J.; Waltman, L. Software survey: VOSviewer, a computer program for bibliometric mapping. Scientometrics 2010, 84, 523–538. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Christoph, N.; Hermann, A.; Wachter, H. 25 years authentication of wine with stable isotope analysis in the European Union—Review and outlook. In 38th World Congress of Vine and Wine; Jean-Marie, A., Ed.; BIO Web of Conferences; EDP Sciences: Yulis, France, 2015; Volume 5. [Google Scholar]
- Ranaweera, R.K.R.; Souza Gonzaga, L.; Capone, D.L.; Bastian, S.E.P.; Jeffery, D.W. 3.33—Authenticity and traceability in the wine industry: From analytical chemistry to consumer perceptions. Compr. Foodomics 2021, 3, 452–480. [Google Scholar] [CrossRef]
- Hofmann, A. Spectroscopic techniques. In Principles and Techniques of Biochemistry and Molecular Biology, 7th ed.; Walker, J., Wilson, K., Eds.; Cambridge University Press: Cambridge, UK, 2010; pp. 477–521. [Google Scholar]
- Aleixandre-Tudo, J.L.; Buica, A.; Nieuwoudt, H.; Aleixandre, J.L.; du Toit, W. Spectrophotometric analysis of phenolic compounds in grapes and wines. J. Agric. Food Chem. 2017, 65, 4009–4026. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Wang, H.; Zhan, J.; Huang, W. Review of recent UV–Vis and infrared spectroscopy researches on wine detection and discrimination. Appl. Spectrosc. Rev. 2018, 53, 65–86. [Google Scholar] [CrossRef]
- Acevedo, F.J.; Jiménez, J.; Maldonado, S.; Domínguez, E.; Narváez, A. Classification of wines produced in specific regions by UV−visible spectroscopy combined with support vector machines. J. Agric. Food Chem. 2007, 55, 6842–6849. [Google Scholar] [CrossRef]
- Azcarate, S.M.; Cantarelli, M.Á.; Pellerano, R.G.; Marchevsky, E.J.; Camiña, J.M. Classification of Argentinean Sauvignon blanc wines by UV spectroscopy and chemometric methods. J. Food Sci. 2013, 78, C432–C436. [Google Scholar] [CrossRef]
- Urbano, M.; Luque de Castro, M.D.; Pérez, P.M.; García-Olmo, J.; Gómez-Nieto, M.A. Ultraviolet–visible spectroscopy and pattern recognition methods for differentiation and classification of wines. Food Chem. 2006, 97, 166–175. [Google Scholar] [CrossRef]
- Philippidis, A.; Poulakis, E.; Kontzedaki, R.; Orfanakis, E.; Symianaki, A.; Zoumi, A.; Velegrakis, M. Application of ultraviolet-visible absorption spectroscopy with machine learning techniques for the classification of Cretan wines. Foods 2021, 10, 9. [Google Scholar] [CrossRef]
- Tan, J.; Li, R.; Jiang, Z.-T.; Zhang, Y.; Hou, Y.-M.; Wang, Y.-R.; Wu, X.; Gong, L. Geographical classification of Chinese Cabernet Sauvignon wines by data fusion of ultraviolet–visible and synchronous fluorescence spectroscopies: The combined use of multiple wavelength differences. Aust. J. Grape Wine Res. 2016, 22, 358–365. [Google Scholar] [CrossRef]
- Cozzolino, D.; Cynkar, W.; Shah, N.; Smith, P.A. Can spectroscopy geographically classify Sauvignon Blanc wines from Australia and New Zealand? Food Chem. 2011, 126, 673–678. [Google Scholar] [CrossRef]
- Cozzolino, D.; Cynkar, W.; Shah, N.; Smith, P. Technical solutions for analysis of grape juice, must, and wine: The role of infrared spectroscopy and chemometrics. Anal. Bioanal. Chem. 2011, 401, 1475–1484. [Google Scholar] [CrossRef]
- Aleixandre-Tudó, J.L.; Castelló-Cogollos, L.; Aleixandre, J.L.; Aleixandre-Benavent, R. Bibliometric insights into the spectroscopy research field: A food science and technology case study. Appl. Spectrosc. Rev. 2020, 55, 873–906. [Google Scholar] [CrossRef]
- Bevin, C.J.; Dambergs, R.G.; Fergusson, A.J.; Cozzolino, D. Varietal discrimination of Australian wines by means of mid-infrared spectroscopy and multivariate analysis. Anal. Chim. Acta 2008, 621, 19–23. [Google Scholar] [CrossRef]
- Geană, E.-I.; Ciucure, C.T.; Apetrei, C.; Artem, V. Application of spectroscopic UV–Vis and FT-IR screening techniques coupled with multivariate statistical analysis for red wine authentication: Varietal and vintage year discrimination. Molecules 2019, 24, 4166. [Google Scholar] [CrossRef] [Green Version]
- Rodriguez-Saona, L.E.; Allendorf, M.E. Use of FTIR for rapid authentication and detection of adulteration of food. Annu. Rev. Food Sci. Technol. 2011, 2, 467–483. [Google Scholar] [CrossRef]
- Cozzolino, D. The role of visible and infrared spectroscopy combined with chemometrics to measure phenolic compounds in grape and wine samples. Molecules 2015, 20, 726–737. [Google Scholar] [CrossRef] [Green Version]
- Martelo-Vidal, M.J.; Domínguez-Agis, F.; Vázquez, M. Ultraviolet/visible/near-infrared spectral analysis and chemometric tools for the discrimination of wines between subzones inside a controlled designation of origin: A case study of Rías Baixas. Aust. J. Grape Wine Res. 2013, 19, 62–67. [Google Scholar] [CrossRef]
- Hu, X.-Z.; Liu, S.-Q.; Li, X.-H.; Wang, C.-X.; Ni, X.-L.; Liu, X.; Wang, Y.; Liu, Y.; Xu, C.-H. Geographical origin traceability of Cabernet Sauvignon wines based on Infrared fingerprint technology combined with chemometrics. Sci. Rep. 2019, 9, 8256. [Google Scholar] [CrossRef]
- Patz, C.D.; Blieke, A.; Ristow, R.; Dietrich, H. Application of FT-MIR spectrometry in wine analysis. Anal. Chim. Acta 2004, 513, 81–89. [Google Scholar] [CrossRef]
- Ríos-Reina, R.; Camiña, J.M.; Callejón, R.M.; Azcarate, S.M. Spectralprint techniques for wine and vinegar characterization, authentication and quality control: Advances and projections. TrAC Trends Anal. Chem. 2021, 134, 116121. [Google Scholar] [CrossRef]
- Magdas, D.A.; Guyon, F.; Feher, I.; Pinzaru, S.C. Wine discrimination based on chemometric analysis of untargeted markers using FT-Raman spectroscopy. Food Control. 2018, 85, 385–391. [Google Scholar] [CrossRef]
- Magdas, D.A.; Cinta Pinzaru, S.; Guyon, F.; Feher, I.; Cozar, B.I. Application of SERS technique in white wines discrimination. Food Control. 2018, 92, 30–36. [Google Scholar] [CrossRef]
- Deneva, V.; Bakardzhiyski, I.; Bambalov, K.; Antonova, D.; Tsobanova, D.; Bambalov, V.; Cozzolino, D.; Antonov, L. Using Raman spectroscopy as a fast tool to classify and analyze Bulgarian wines—A feasibility study. Molecules 2020, 25, 170. [Google Scholar] [CrossRef] [Green Version]
- Magdas, D.A.; Cozar, B.I.; Feher, I.; Guyon, F.; Dehelean, A.; Cinta Pinzaru, S. Testing the limits of FT-Raman spectroscopy for wine authentication: Cultivar, geographical origin, vintage and terroir effect influence. Sci. Rep. 2019, 9, 19954. [Google Scholar] [CrossRef]
- Zanuttin, F.; Gurian, E.; Ignat, I.; Fornasaro, S.; Calabretti, A.; Bigot, G.; Bonifacio, A. Characterization of white wines from north-eastern Italy with surface-enhanced Raman spectroscopy. Talanta 2019, 203, 99–105. [Google Scholar] [CrossRef]
- de Almeida, M.P.; Leopold, N.; Franco, R.; Pereira, E. Expedite SERS fingerprinting of Portuguese white wines using plasmonic silver nanostars. Front. Chem. 2019, 7. [Google Scholar] [CrossRef]
- Xu, Y.; Zhong, P.; Jiang, A.; Shen, X.; Li, X.; Xu, Z.; Shen, Y.; Sun, Y.; Lei, H. Raman spectroscopy coupled with chemometrics for food authentication: A. review. TrAC Trends Anal. Chem. 2020, 131, 116017. [Google Scholar] [CrossRef]
- Sádecká, J.; Tothova, J. Fluorescence spectroscopy and chemometrics in the food classification: A review. Czech. J. Food Sci. 2007, 25, 159–174. [Google Scholar] [CrossRef] [Green Version]
- Airado-Rodríguez, D.; Durán-Merás, I.; Galeano-Díaz, T.; Wold, J.P. Front-face fluorescence spectroscopy: A new tool for control in the wine industry. J. Food Compos. Anal. 2011, 24, 257–264. [Google Scholar] [CrossRef]
- Airado-Rodríguez, D.; Galeano-Díaz, T.; Durán-Merás, I.; Wold, J.P. Usefulness of fluorescence excitation−emission matrices in combination with PARAFAC, as fingerprints of red wines. J. Agric. Food Chem. 2009, 57, 1711–1720. [Google Scholar] [CrossRef]
- Coelho, C.; Aron, A.; Roullier-Gall, C.; Gonsior, M.; Schmitt-Kopplin, P.; Gougeon, R.D. Fluorescence fingerprinting of bottled white wines can reveal memories related to sulfur dioxide treatments of the must. Anal. Chem. 2015, 87, 8132–8137. [Google Scholar] [CrossRef]
- Dankowska, A. Advances in fluorescence emission spectroscopy for food authenticity testing. In Advances in Food Authenticity Testing; Woodhead Publishing: Cambridge, UK, 2016; pp. 117–145. [Google Scholar]
- Sádecká, J.; Jakubíková, M. Varietal classification of white wines by fluorescence spectroscopy. J. Food Sci. Technol. 2020, 57, 2545–2553. [Google Scholar] [CrossRef] [PubMed]
- Suciu, R.-C.; Zarbo, L.; Guyon, F.; Magdas, D.A. Application of fluorescence spectroscopy using classical right angle technique in white wines classification. Sci. Rep. 2019, 9, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Ranaweera, R.K.R.; Gilmore, A.M.; Capone, D.L.; Bastian, S.E.P.; Jeffery, D.W. Spectrofluorometric analysis combined with machine learning for geographical and varietal authentication, and prediction of phenolic compound concentrations in red wine. Food Chem. 2021, 130149. [Google Scholar] [CrossRef] [PubMed]
- Martin, G.J.; Martin, M.L.; Mabon, F.; Michon, M.J. A new method for the identification of the origin of ethanols in grain and fruit spirits: High-field quantitative deuterium nuclear magnetic resonance at the natural abundance level. J. Agric. Food Chem. 1983, 31, 311–315. [Google Scholar] [CrossRef]
- Solovyev, P.A.; Fauhl-Hassek, C.; Riedl, J.; Esslinger, S.; Bontempo, L.; Camin, F. NMR spectroscopy in wine authentication: An official control perspective. Compr. Rev. Food Sci. Food Saf. 2021, 20, 2040–2062. [Google Scholar] [CrossRef] [PubMed]
- Viskić, M.; Bandić, L.M.; Korenika, A.-M.J.; Jeromel, A. NMR in the service of wine differentiation. Foods 2021, 10, 120. [Google Scholar] [CrossRef]
- Godelmann, R.; Fang, F.; Humpfer, E.; Schütz, B.; Bansbach, M.; Schäfer, H.; Spraul, M. Targeted and nontargeted wine analysis by 1H NMR spectroscopy combined with multivariate statistical analysis. Differentiation of important parameters: Grape variety, geographical origin, year of vintage. J. Agric. Food Chem. 2013, 61, 5610–5619. [Google Scholar] [CrossRef]
- Fan, S.; Zhong, Q.; Fauhl-Hassek, C.; Pfister, M.K.H.; Horn, B.; Huang, Z.B. Classification of Chinese wine varieties using 1H NMR spectroscopy combined with multivariate statistical analysis. Food Control 2018, 88, 113–122. [Google Scholar] [CrossRef]
- Mascellani, A.; Hoca, G.; Babisz, M.; Krska, P.; Kloucek, P.; Havlik, J. 1H NMR chemometric models for classification of Czech wine type and variety. Food Chem. 2021, 339, 127852. [Google Scholar] [CrossRef] [PubMed]
- Gougeon, L.; Da Costa, G.; Le Mao, I.; Ma, W.; Teissedre, P.-L.; Guyon, F.; Richard, T. Wine analysis and authenticity using 1H-NMR metabolomics data: Application to Chinese wines. Food Anal. Methods 2018, 11, 3425–3434. [Google Scholar] [CrossRef]
- Crook, A.A.; Zamora-Olivares, D.; Bhinderwala, F.; Woods, J.; Winkler, M.; Rivera, S.; Shannon, C.E.; Wagner, H.R.; Zhuang, D.L.; Lynch, J.E.; et al. Combination of two analytical techniques improves wine classification by vineyard, region, and vintage. Food Chem. 2021, 354, 129531. [Google Scholar] [CrossRef]
- Sáenz Navajas, M.-P.; Jeffery, D.W. Perspectives on wines of provenance: Sensory typicality, quality, and authenticity. ACS Food Sci. Technol. 2021, 1, 986–992. [Google Scholar] [CrossRef]
- Rodionova, O.; Kucheryavskiy, S.; Pomerantsev, A. Efficient tools for principal component analysis of complex data— A tutorial. Chemom. Intell. Lab. Syst. 2021, 213, 104304. [Google Scholar] [CrossRef]
- Cozzolino, D.; Power, A.; Chapman, J. Interpreting and reporting principal component analysis in food science analysis and beyond. Food Anal. Methods 2019, 12, 2469–2473. [Google Scholar] [CrossRef]
- Brito, A.L.B.; Brito, L.R.; Honorato, F.A.; Pontes, M.J.C.; Pontes, L.F.B.L. Classification of cereal bars using near infrared spectroscopy and linear discriminant analysis. Food Res. Int. 2013, 51, 924–928. [Google Scholar] [CrossRef] [Green Version]
- Rodionova, O.Y.; Titova, A.V.; Pomerantsev, A.L. Discriminant analysis is an inappropriate method of authentication. TrAC Trends Anal. Chem. 2016, 78, 17–22. [Google Scholar] [CrossRef]
- Ríos-Reina, R.; Elcoroaristizabal, S.; Ocaña-González, J.A.; García-González, D.L.; Amigo, J.M.; Callejón, R.M. Characterization and authentication of Spanish PDO wine vinegars using multidimensional fluorescence and chemometrics. Food Chem. 2017, 230, 108–116. [Google Scholar] [CrossRef] [Green Version]
- Xu, Y.; Zomer, S.; Brereton, R.G. Support vector machines: A recent method for classification in chemometrics. Crit. Rev. Anal. Chem. 2006, 36, 177–188. [Google Scholar] [CrossRef]
- Jiménez-Carvelo, A.M.; González-Casado, A.; Bagur-González, M.G.; Cuadros-Rodríguez, L. Alternative data mining/machine learning methods for the analytical evaluation of food quality and authenticity—A review. Food Res. Int. 2019, 122, 25–39. [Google Scholar] [CrossRef]
- Borràs, E.; Ferré, J.; Boqué, R.; Mestres, M.; Aceña, L.; Busto, O. Data fusion methodologies for food and beverage authentication and quality assessment—A review. Anal. Chim. Acta 2015, 891, 1–14. [Google Scholar] [CrossRef]
- Murru, C.; Chimeno-Trinchet, C.; Díaz-García, M.E.; Badía-Laíño, R.; Fernández-González, A. Artificial Neural Network and Attenuated Total Reflectance-Fourier Transform Infrared Spectroscopy to identify the chemical variables related to ripeness and variety classification of grapes for protected. Designation of origin wine production. Comput. Electron. Agric. 2019, 164, 104922. [Google Scholar] [CrossRef]
- Cosme, F.; Milheiro, J.; Pires, J.; Guerra-Gomes, F.I.; Filipe-Ribeiro, L.; Nunes, F.M. Authentication of Douro DO monovarietal red wines based on anthocyanin profile: Comparison of partial least squares—Discriminant analysis, decision trees and artificial neural networks. Food Control. 2021, 125, 107979. [Google Scholar] [CrossRef]
- Kruzlicova, D.; Mocak, J.; Balla, B.; Petka, J.; Farkova, M.; Havel, J. Classification of Slovak white wines using artificial neural networks and discriminant techniques. Food Chem. 2009, 112, 1046–1052. [Google Scholar] [CrossRef]
- Engel, J.; Gerretzen, J.; Szymańska, E.; Jansen, J.J.; Downey, G.; Blanchet, L.; Buydens, L.M.C. Breaking with trends in pre-processing? TrAC Trends Anal. Chem. 2013, 50, 96–106. [Google Scholar] [CrossRef]
- Callao, M.P.; Ruisanchez, I. An overview of multivariate qualitative methods for food fraud detection. Food Control. 2018, 86, 283–293. [Google Scholar] [CrossRef]
- Moghaddam, D.D.; Rahmati, O.; Panahi, M.; Tiefenbacher, J.; Darabi, H.; Haghizadeh, A.; Haghighi, A.T.; Nalivan, O.A.; Tien Bui, D. The effect of sample size on different machine learning models for groundwater potential mapping in mountain bedrock aquifers. CATENA 2020, 187, 104421. [Google Scholar] [CrossRef]
- Xu, Y.; Goodacre, R. On splitting training and validation set: A comparative study of cross-validation, bootstrap and systematic sampling for estimating the generalization performance of supervised learning. J. Anal. Test. 2018, 2, 249–262. [Google Scholar] [CrossRef] [Green Version]
- Augustine, M.P.; Harley, S.J.; Lim, V.; Stucky, P. An Authentication Device for Full Intact Wine Bottles. U.S. Patent 9,488,599, 8 November 2016. [Google Scholar]
| Spectroscopic Technique | Spectral Range | Parameters for Authentication | Classification Method 1 | Remark | Reference |
|---|---|---|---|---|---|
| UV–Vis | 200–800 nm | Geographical origin (Spanish denomination of origin) | SVM | Correct classification rates above 96% | [18] |
| UV–Vis | 200–500 nm | Geographical origin of Argentinian regions | PCA, LDA, PLS-DA | Correct classification with LDA and PLS-DA methods of 100% | [19] |
| UV–Vis | 300–800 nm | Discrimination by origin, grape variety and ageing process | PCA, SIMCA | Correct classification of 90% for geographical origin, and 75% for variety and ageing process | [20] |
| UV–Vis | 240–700 nm | Discrimination according to grape variety, ageing process and barrel/container type | OPLS-DA | Correct classification of 97% for variety, 73% for ageing process and 100% for container type | [21] |
| Spectroscopic Technique | Spectral Range | Parameters for Authentication | Classification Method 1 | Remark | Reference |
|---|---|---|---|---|---|
| MIR | 5012–926 cm−1 | Discrimination of red and white varieties from Australian regions | PCA, LDA | Correct classification of red varieties, 96% and white varieties, 94% | [26] |
| UV–Vis, NIR and MIR | 400–2500 nm (UV–Vis and NIR) and 4000–400 cm−1 (MIR) | Geographical origin of Sauvignon blanc wines from Australia and New Zealand | PCA, SIMCA, PLS-DA | Correct classification using PLS-DA with: UV–Vis, 67%; NIR, 76%; MIR, 90%; and combined IR spectra, 93% | [23] |
| UV–Vis/NIR | 190–2500 nm | Discrimination of white wines (Albariño cultivar) from Rías Baixas subzones in Spain | PCA, LDA, SIMCA, SVM | Correct classification using: LDA, 86%; SIMCA, 56%; and SVM, 84% | [30] |
| NIR and MIR | 1750–1000 cm−1 and 4555–4353 cm−1 | Geographical origin of Cabernet Sauvignon wines from Australia, Chile, and China | PCA, SIMCA, DA | Correct classification using: SIMCA, 97%, 97%, and 92% for Australian, Chilean, and Chinese wines; and DA, 86%, 85%, and 77%, respectively. | [31] |
| Spectroscopic Technique | Spectral Region | Parameters for Authentication | Classification Method 1 | Remark | Reference |
|---|---|---|---|---|---|
| FT-Raman | 1700–0 cm−1 (Stokes), −1000–0 cm−1 (anti-Stokes) (laser emitting at 1064 nm) | Discrimination of wines geographically, varietally, and by vintage | LDA | Correct classification of: variety, 84%; geographical origin, 100%; and vintage, 95% | [37] |
| SERS | 3350–200 cm−1 (laser emitting at 532 nm) | Discrimination of wines geographically (Romanian and French and different Romanian regions), varietally, and by vintage | LDA | Correct classification of: variety, 90%; geographical origin, 83% among Romanian wines and 100% between countries; and vintage, 90% | [35] |
| SERS | 1600–450 cm−1 (laser emitting at 785 nm) | Discrimination of wines according to variety and producer | PCA, SIMCA | Correct classification of: variety, 87%; and producer, 93% | [38] |
| Spectroscopic Technique | Spectral Region | Parameters for Authentication | Classification Method 1 | Remark | Reference |
| Synchronous fluorescence spectroscopy | λex = 250–350 nm and λem = 250–500 nm | Discrimination of white wines according to variety in Tokaj (Slovakia) | PCA, LDA | Correct classification of variety, 100% | [46] |
| Total fluorescence spectroscopy | EEM λex =240–800 nm and λem 242–824 nm | Discrimination of Cabernet Sauvignon wines from Australia and Bordeaux, France | SVMDA XGBDA | Correct classification of geographical origin using: XGBDA, 100%; and SVMDA, 85% | [11] |
| Total fluorescence spectroscopy | EEM λex = 250–500 nm and λem 275–600 nm | Discrimination of white wine from Romania and France for geographical origin and variety | PARAFAC, SIMCA | Correct classification of: variety, 97%; and geographical origin, 98% | [47] |
| Total fluorescence spectroscopy | EEM λex =240–700 nm and λem 242–824 nm | Discrimination of red wine varieties from different Australian regions for variety and geographical origin | XGBDA | Correct classification of: variety, 100%; and geographical origin, 99.7% | [48] |
| Spectroscopic Technique | Spectral Range | Parameters for Authentication | Classification Method 1 | Remark | Reference |
|---|---|---|---|---|---|
| 1H NMR | 0.5–9.5 ppm | Discrimination of wines geographically (German wine regions), varietally, and by vintage | PCA, LDA, NCM | Correct classification of: variety, 95%; geographical origin, 89%; and vintage, 96–97% | [52] |
| 1H NMR | 0.8–9.7 ppm | Varietal differentiation of red and white wines produced in different regions in China | PCA, LDA | Correct classification of: red wines, 83%; and white wines, 94% | [53] |
| 1H NMR | 0.0–10.0 ppm | Varietal differentiation of red and white wines produced in Czech Republic | PCA, RF | Correct classification of: most varieties, ~70%; and type of wine, 92% | [54] |
| Technique | Chemical Marker | Advantages | Disadvantages |
|---|---|---|---|
| UV–Vis | Hydroxybenzoic acids, hydroxycinnamic acids, flavan-3-ols, flavonols, and anthocyanin glucosides | Simple analysis, low cost, small volume | Difficulty in identifying specific analytes |
| IR | Organic acids, alcohols, reducing sugars, and polyphenols | Rapid, simple, qualitative and quantitative analysis | Sensitive to pH and temperature, high interference of water (NIR) |
| Raman | Organic acids, alcohols, sugars, phenolics | Rapid, small volume, low impact of water | Weak signals, extensive pre-processing requirements |
| Fluorescence | Phenolics, pigments, vitamins, amino acids | Rapid, sensitive and selective, qualitative and quantitative analysis | Extensive pre-processing requirements, quenching effect |
| NMR | Phenolics, alcohols, organic acids, amino acids, sugars | Rapid, selective, repeatable and reproducible | Costly equipment, experienced analyst required |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ranaweera, R.K.R.; Capone, D.L.; Bastian, S.E.P.; Cozzolino, D.; Jeffery, D.W. A Review of Wine Authentication Using Spectroscopic Approaches in Combination with Chemometrics. Molecules 2021, 26, 4334. https://doi.org/10.3390/molecules26144334
Ranaweera RKR, Capone DL, Bastian SEP, Cozzolino D, Jeffery DW. A Review of Wine Authentication Using Spectroscopic Approaches in Combination with Chemometrics. Molecules. 2021; 26(14):4334. https://doi.org/10.3390/molecules26144334
Chicago/Turabian StyleRanaweera, Ranaweera K. R., Dimitra L. Capone, Susan E. P. Bastian, Daniel Cozzolino, and David W. Jeffery. 2021. "A Review of Wine Authentication Using Spectroscopic Approaches in Combination with Chemometrics" Molecules 26, no. 14: 4334. https://doi.org/10.3390/molecules26144334
APA StyleRanaweera, R. K. R., Capone, D. L., Bastian, S. E. P., Cozzolino, D., & Jeffery, D. W. (2021). A Review of Wine Authentication Using Spectroscopic Approaches in Combination with Chemometrics. Molecules, 26(14), 4334. https://doi.org/10.3390/molecules26144334







