Fluorimetric Analysis of Five Amino Acids in Chocolate: Development and Validation
Abstract
:1. Introduction
2. Results
2.1. Optimization of the HPLC Conditions
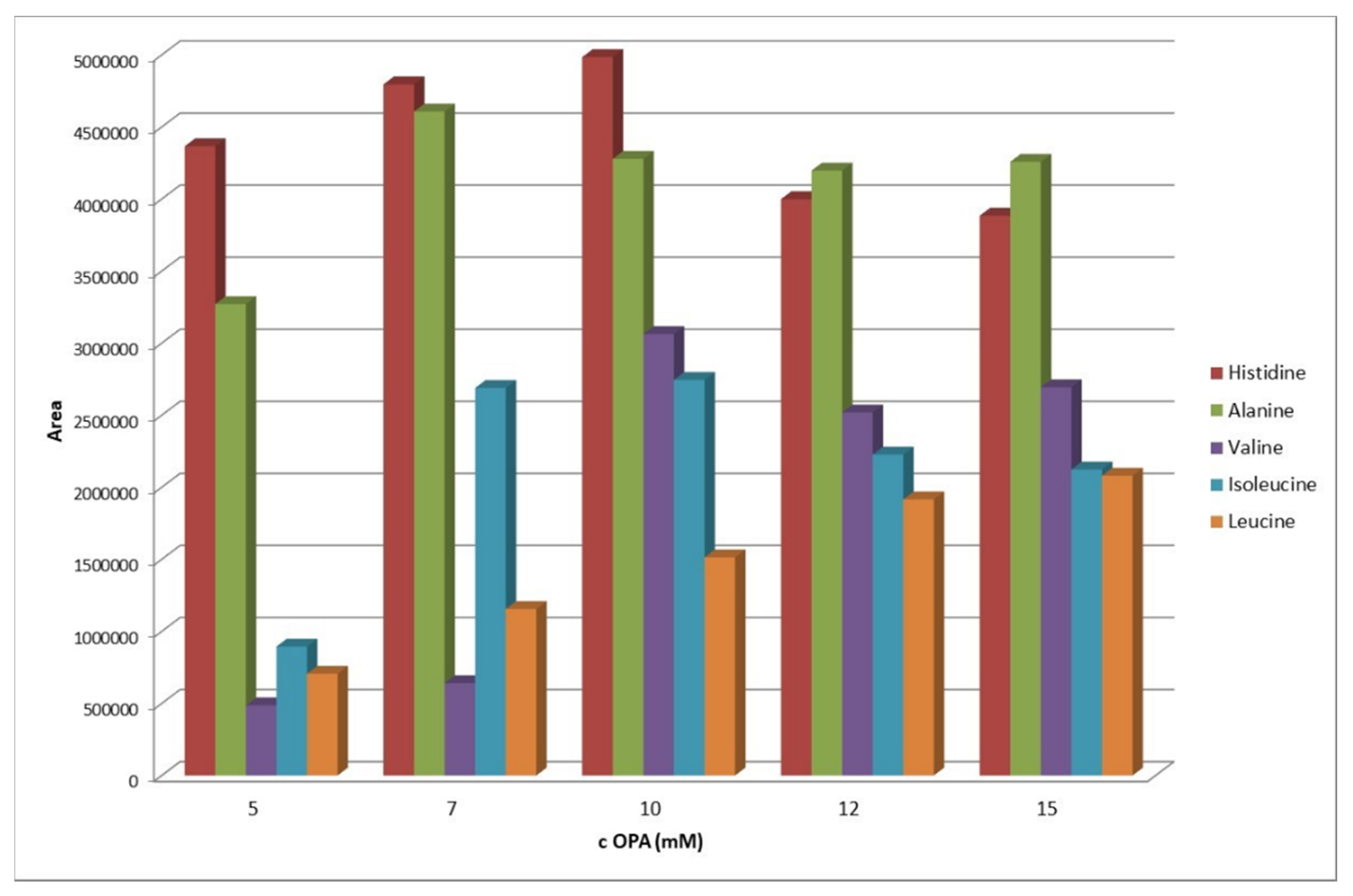
2.2. Optimization of the OPA-Amino Acids Derivatization Conditions
2.3. Stability of Amino Acids-OPA Derivatives
2.4. Extraction Recovery–Preliminary Experiments
2.5. Improvement of Trace Analysis
2.6. Method Validation
2.6.1. Selectivity of the Developed Chromatographic Method
2.6.2. Linearity, LODs, and LOQs
2.6.3. Accuracy and Precision
2.6.4. Robustness
3. Materials and Methods
3.1. Chemicals, Materials and Reagents
3.2. Instrumentation and Chromatographic Conditions
3.3. Derivatization Procedure
3.4. Preparation of the Chocolate Formulation
3.5. Sample Purification Procedures
3.5.1. Liquid–Solid Extraction (LSE)
3.5.2. Solid-Phase Extraction (SPE)
3.5.3. Trace Analysis of AAs in Chocolate
3.6. Validation of the Proposed Method
3.6.1. Specificity
3.6.2. Linearity
3.6.3. Limit of Detection (LOD) and Limit of Quantification (LOQ)
3.6.4. Precision and Accuracy
3.6.5. Robustness
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
Sample Availability
References
- The European Parliament and the Council of the European Union. Regulation (EU) No 609/2013 of the European Parliament and of the Council of 12 June 2013 on food intended for infants and young children, food for special medical purposes, and total diet replacement for weight control and repealing Council Directive 92/52/EEC, Commission Directives 96/8/EC, 1999/21/EC, 2006/125/EC and 2006/141/EC, Directive 2009/39/EC of the European Parliament and of the Council and Commission Regulations (EC) No 41/2009 and (EC) No 953/2009. Off. J. Eur. Union 2013, 181, 35–56. [Google Scholar]
- Oracz, J.; Nebesny, E.; Zyzelewicz, D.; Budryn, G.; Luzak, B. Bioavailability and metabolism of selected cocoa bioactive compounds: A comprehensive review. Crit. Rev. Food Sci. Nutr. 2020, 60, 1947–1985. [Google Scholar] [CrossRef]
- Oracz, J.; Nebesny, E.; Żyżelewicz, D. Identification and quantification of free and bound phenolic compounds contained in the high-molecular weight melanoidin fractions derived from two different types of cocoa beans by UHPLC-DAD-ESI-HR-MSn. Food Res. Int. 2019, 115, 135–149. [Google Scholar] [CrossRef] [PubMed]
- Ferré, S.; González-Ruiz, V.; Guillarme, D.; Rudaz, S. Analytical strategies for the determination of amino acids: Past, present and future trends. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2019, 1132, 121819. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Xu, B.; Zheng, X.; Wan, X.; Zhang, X.; Wu, G.; Cong, Z. Developing an analytical method for free amino acids in atmospheric precipitation using gas chromatography coupled with mass spectrometry. Atmos. Res. 2021, 256, 105579. [Google Scholar] [CrossRef]
- Peace, R.W.; Gilani, G.S. Chromatographic determination of amino acids in foods. J. AOAC Int. 2005, 88, 877–887. [Google Scholar] [CrossRef] [Green Version]
- Dala-Paula, B.M.; Deus, V.L.; Tavano, O.L.; Gloria, M.B.A. In vitro bioaccessibility of amino acids and bioactive amines in 70% cocoa dark chocolate: What you eat and what you get. Food Chem. 2021, 343, 128397. [Google Scholar] [CrossRef]
- Dziagwa-Becker, M.M.; Marin Ramos, J.M.; Topolski, J.K.; Oleszek, W.A. Determination of free amino acids in plants by liquid chromatography coupled to tandem mass spectrometry (LC-MS/MS). Anal. Methods 2015, 7, 7574–7581. [Google Scholar] [CrossRef] [Green Version]
- Przybylska, A.; Gackowski, M.; Koba, M. Application of capillary electrophoresis to the analysis of bioactive compounds in herbal raw materials. Molecules 2021, 26, 2135. [Google Scholar] [CrossRef]
- Fekkes, D.; Voskuilen-Kooyman, A.; Jankie, R.; Huijmans, J. Precise analysis of primary amino acids in urine by an automated high-performance liquid chromatography method: Comparison with ion-exchange chromatography. J. Chromatogr. B Biomed. Sci. Appl. 2000, 744, 183–188. [Google Scholar] [CrossRef]
- Teerlink, T.; Van Leeuwen, P.A.M.; Houdijk, A. Plasma amino acids determined by liquid chromatography within 17 min. Clin. Chem. 1994, 40, 245–249. [Google Scholar] [CrossRef]
- Samardzic, K.; Rodgers, K.J. Cytotoxicity and mitochondrial dysfunction caused by the dietary supplement l-norvaline. Toxicol. Vitr. 2019, 56, 163–171. [Google Scholar] [CrossRef] [PubMed]
- Violi, J.P.; Bishop, D.P.; Padula, M.P.; Steele, J.R.; Rodgers, K.J. Considerations for amino acid analysis by liquid chromatography-tandem mass spectrometry: A tutorial review. TrAC-Trends Anal. Chem. 2020, 131, 116018. [Google Scholar] [CrossRef]
- Moore, S.; Stein, W.H. Photometric ninhydrin method for use in the chromatography of amino acids. J. Biol. Chem. 1948, 176, 367–388. [Google Scholar] [CrossRef]
- How, Z.T.; Busetti, F.; Linge, K.L.; Kristiana, I.; Joll, C.A.; Charrois, J.W.A. Analysis of free amino acids in natural waters by liquid chromatography-tandem mass spectrometry. J. Chromatogr. A 2014, 1370, 135–146. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Xu, R.; Hu, B.; Li, W.; Sun, Y.; Tu, Y.; Zeng, X. Analysis of free amino acids in Chinese teas and flower of tea plant by high performance liquid chromatography combined with solid-phase extraction. Food Chem. 2010, 123, 1259–1266. [Google Scholar] [CrossRef]
- Tang, X.; Gu, Y.; Nie, J.; Fan, S.; Wang, C. Quantification of amino acids in rat urine by solid-phase extraction and liquid chromatography/electrospray tandem mass spectrometry: Application to radiation injury rat model. J. Liq. Chromatogr. Relat. Technol. 2014, 37, 951–973. [Google Scholar] [CrossRef]
- Armstrong, M.; Jonscher, K.; Reisdorph, N.A. Analysis of 25 underivatized amino acids in human plasma using ion-pairing reversed-phase liquid chromatography/time-of-flight mass spectrometry. Rapid Commun. Mass Spectrom. 2007, 21, 2717–2726. [Google Scholar] [CrossRef]
- Bonvin, G.; Schappler, J.; Rudaz, S. Capillary electrophoresis-electrospray ionization-mass spectrometry interfaces: Fundamental concepts and technical developments. J. Chromatogr. A 2012, 1267, 17–31. [Google Scholar] [CrossRef]
- Bhawani, S.A.; Mohamad Ibrahim, M.N.; Sulaiman, O.; Hashim, R.; Mohammad, A.; Hena, S. Thin-layer chromatography of amino acids: A review. J. Liq. Chromatogr. Relat. Technol. 2012, 35, 1497. [Google Scholar] [CrossRef]
- D’Atri, V.; Fekete, S.; Clarke, A.; Veuthey, J.L.; Guillarme, D. Recent Advances in Chromatography for Pharmaceutical Analysis. Anal. Chem. 2019, 91, 210–239. [Google Scholar] [CrossRef] [Green Version]
- Corleto, K.A.; Singh, J.; Jayaprakasha, G.K.; Patil, B.S. A sensitive HPLC-FLD method combined with multivariate analysis for the determination of amino acids in L-citrulline rich vegetables. J. Food Drug Anal. 2019, 27, 717–728. [Google Scholar] [CrossRef] [Green Version]
- Danielsen, M.; Nebel, C.; Dalsgaard, T.K. Simultaneous determination of L- And D-amino acids in proteins: A sensitive method using hydrolysis in deuterated acid and liquid chromatography–tandem mass spectrometry analysis. Foods 2020, 9, 309. [Google Scholar] [CrossRef] [Green Version]
- Zhou, W.; Wang, Y.; Yang, F.; Dong, Q.; Wang, H.; Hu, N. Rapid determination of amino acids of nitraria tangutorum bobr. From the qinghai-tibet plateau using HPLC-FLD-MS/MS and a highly selective and sensitive pre-column derivatization method. Molecules 2019, 24, 1665. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rutherfurd, S.M.; Gilani, G.S. Amino Acid Analysis. Curr. Protoc. Protein Sci. 2009. [Google Scholar] [CrossRef] [PubMed]
- Dong, M.W. Modern HPLC for Practicing Scientists; John Wiley & Sons, Inc: Hoboken, NJ, USA, 2006; pp. 1–286. [Google Scholar]
- Christofi, M.; Markopoulou, C.K.; Tzanavaras, P.D.; Zacharis, C.K. UHPLC-fluorescence method for the determination of trace levels of hydrazine in allopurinol and its formulations: Validation using total-error concept. J. Pharm. Biomed. Anal. 2020, 187, 113354. [Google Scholar] [CrossRef] [PubMed]
- AlRabiah, H.; Attia, S.M.; Al-Shakliah, N.S.; Mostafa, G.A.E. Development and validation of an HPLC-UV detection Assay for the determination of clonidine in mouse plasma and its application to a pharmacokinetic study. Molecules 2020, 25, 4109. [Google Scholar] [CrossRef]
- ICH. Validation of Analytical Procedures: Text and Methodology (Q2 R1); European Medicines Agency: Amsterdam, The Netherlands, 1995; CPMP/ICH/381/95.
- National Research Council. Amino Acids. In Recommended Dietary Allowances, 10th ed.; The National Academies Press: Washington, DC, USA, 1989; Chapter 6. [Google Scholar] [CrossRef]
- Reeds, P.J.; Garlick, P.J. Protein and amino acid requirements and the composition of complementary foods. J. Nutr. 2003, 133, 2953–2961. [Google Scholar] [CrossRef]
- U.S. Department of Health and Human Services Food and Drug Administration. Analytical Procedures and Methods Validation for Drugs and Biologics Guidance for Industry Analytical Procedures and Methods Validation for Drugs and Biologics Guidance for Industry; FDA: Silver Spring, MD, USA, 2015.
- Shabir, G.A. Validation of high-performance liquid chromatography methods for pharmaceutical analysis. J. Chromatogr. A 2003, 987, 57–66. [Google Scholar] [CrossRef]
- Miller, J.N.; Miller, J.C. Statistics and Chemometrics for Analytical Chemistry, 6th ed.; Pearson Education: London, UK, 2005. [Google Scholar]

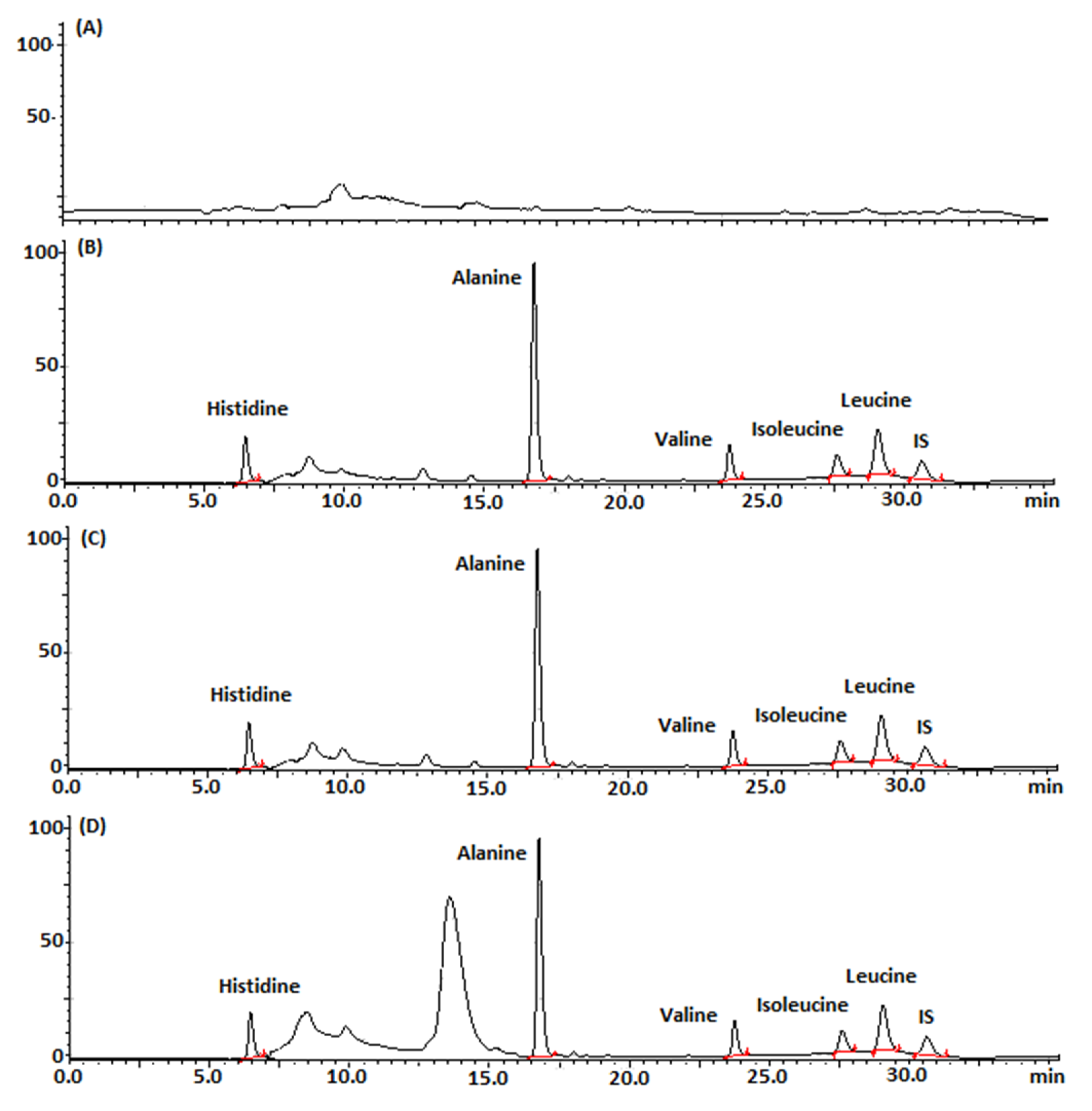
| Parameters | Retention Time (min) | k’ | Resolution | Tailing Factor |
|---|---|---|---|---|
| Histidine | 6.45 | - | - | 1.217 |
| Alanine | 16.78 | 1.599 | 31.518 | 1.209 |
| Valine | 23.75 | 2.683 | 20.922 | 1.203 |
| Isoleucine | 27.60 | 3.280 | 9.486 | 1.193 |
| Leucine | 29.07 | 3.505 | 2.831 | 1.207 |
| IS | 30.65 | 3.749 | 2.695 | 1.198 |
| Standard | C (Added) (μg mL−1) | Observed (ng mL−1) | Recovery (%) |
|---|---|---|---|
| Mean ± SD | Mean ± SD | ||
| Histidine | 0.09 | 0.09 ± 0.00 | 97.8 ± 0.2 |
| 3.00 | 2.97 ± 0.01 | 99.0 ± 0.4 | |
| 5.00 | 4.92 ± 0.06 | 98.3 ± 1.1 | |
| Average (n = 9) | 98.4 ± 0.9 | ||
| Confidence interval (95%) | 97.5–99.3 | ||
| Alanine | 0.02 | 0.02 ± 0.00 | 98.5 ± 0.5 |
| 1.00 | 1.01 ± 0.01 | 100.6 ± 0.7 | |
| 2.00 | 2.00 ± 0.04 | 99.9 ± 1.2 | |
| Average (n = 9) | 99.6 ± 1.7 | ||
| Confidence interval (95%) | 97.9–101.4 | ||
| Valine | 0.03 | 0.03 ± 0.00 | 99.1 ± 0.5 |
| 5.00 | 4.92 ± 0.02 | 98.4 ± 0.4 | |
| 8.00 | 7.86 ± 0.01 | 98.2 ± 0.1 | |
| Average (n = 9) | 98.6 ± 0.6 | ||
| Confidence interval (95%) | 98.0–99.2 | ||
| Isoleucine | 0.03 | 0.03 ± 0.00 | 98.3 ± 0.4 |
| 5.00 | 4.94 ± 0.01 | 98.8 ± 0.1 | |
| 8.00 | 7.90 ± 0.04 | 98.8 ± 0.5 | |
| Average (n = 9) | 98.6 ± 0.5 | ||
| Confidence interval (95%) | 98.2–99.1 | ||
| Leucine | 0.06 | 0.06 ± 0.00 | 99.1 ± 0.2 |
| 10.00 | 10.05 ± 0.09 | 100.5 ± 0.9 | |
| 15.00 | 15.17 ± 0.11 | 101.1 ± 0.7 | |
| Average (n = 9) | 100.2 ± 1.1 | ||
| Confidence interval (95%) | 99.1–101.4 | ||
| Standard | C (Added) (μg mL−1) | SPE/Diol | SPE/NH2 | ||
|---|---|---|---|---|---|
| Observed (ng mL−1) | Recovery (%) | Observed (ng mL−1) | Recovery (%) | ||
| Mean ± SD | Mean ± SD | Mean ± SD | Mean ± SD | ||
| Histidine | 0.09 | 0.09 ± 0.00 | 98.5 ± 0.7 | 0.09 ± 0.00 | 96.1 ± 0.2 |
| 3.00 | 2.93 ± 0.02 | 97.7 ± 0.5 | 2.92 ± 0.01 | 97.2 ± 0.2 | |
| 5.00 | 4.87 ± 0.01 | 97.4 ± 0.2 | 4.88 ± 0.01 | 97.5 ± 0.2 | |
| Average (n = 9) | 97.8 ± 0.7 | Average (n = 9) | 96.9 ± 0.7 | ||
| Confidence interval (95%) | 97.1–98.6 | Confidence interval (95%) | 96.2–97.6 | ||
| Alanine | 0.02 | 0.02 ± 0.00 | 98.9 ± 0.1 | 0.02 ± 0.00 | 96.7 ± 0.4 |
| 1.00 | 1.01 ± 0.01 | 100.9 ± 0.8 | 1.01 ± 0.01 | 101.3 ± 1.3 | |
| 2.00 | 2.03 ± 0.00 | 101.7 ± 0.2 | 2.06 ± 0.03 | 102.9 ± 1.3 | |
| Average (n = 9) | 100.5 ± 1.3 | Average (n = 9) | 100.3 ± 3.0 | ||
| Confidence interval (95%) | 99.2–101.8 | Confidence interval (95%) | 97.3–103.3 | ||
| Valine | 0.03 | 0.03 ± 0.00 | 99.1 ± 0.3 | 0.03 ± 0.00 | 97.1 ± 0.2 |
| 5.00 | 4.96 ± 0.03 | 99.2 ± 0.5 | 4.95 ± 0.03 | 99.1 ± 0.5 | |
| 8.00 | 7.99 ± 0.02 | 99.9 ± 0.2 | 8.03 ± 0.08 | 100.4 ± 0.9 | |
| Average (n = 9) | 99.4 ± 0.5 | Average (n = 9) | 98.9 ± 1.6 | ||
| Confidence interval (95%) | 98.9–100.0 | Confidence interval (95%) | 97.3–100.4 | ||
| Isoleucine | 0.03 | 0.03 ± 0.00 | 98.0 ± 0.1 | 0.03 ± 0.00 | 97.0 ± 0.6 |
| 5.00 | 4.90 ± 0.02 | 98.0 ± 0.5 | 4.85 ± 0.02 | 96.9 ± 0.3 | |
| 8.00 | 7.83 ± 0.05 | 97.9 ± 0.6 | 7.73 ± 0.02 | 96.7 ± 0.2 | |
| Average (n = 9) | 98.0 ± 0.5 | Average (n = 9) | 96.9 ± 0.5 | ||
| Confidence interval (95%) | 97.5–98.4 | Confidence interval (95%) | 96.4–97.3 | ||
| Leucine | 0.06 | 0.06 ± 0.00 | 100.4 ± 1.3 | 0.06 ± 0.00 | 96.5 ± 0.4 |
| 10.00 | 10.07 ± 0.08 | 100.7 ± 0.8 | 9.82 ± 0.04 | 98.2 ± 0.4 | |
| 15.00 | 15.10 ± 0.23 | 100.6 ± 1.5 | 14.85 ± 0.06 | 99.0 ± 0.4 | |
| Average (n = 9) | 100.6 ± 1.3 | Average (n = 9) | 97.9 ± 1.2 | ||
| Confidence interval (95%) | 99.2–101.9 | Confidence interval (95%) | 96.7–99.1 | ||
| Parameter | Optimal Value | Selected Value |
|---|---|---|
| Chocolate weight (mg) | 848.2 | 850.0 |
| Centrifugation time (min) | 18.3 | 20.0 |
| Freezing time (min) | 43.3 | 45.0 |
| Ultrasonic time (min) | 20.8 | 20.0 |
| Initial volume of diluent (mL) | 6.5 | 5.0 |
| pH (initial) diluent | 5.0 | 5.0 |
| Volume of sample on SPE (mL) | 2.2 | 2.0 |
| Elution volume on SPE (μL) | 300.0 | 300.0 |
| Standard | C (Added) (ng mL−1) | Observed (ng mL−1) | Recovery (%) |
|---|---|---|---|
| Mean ± SD | Mean ± SD | ||
| Histidine | 20.0 | 18.7 ± 0.1 | 93.7 ± 0.6 |
| 100.0 | 93.3 ± 0.4 | 93.3 ± 0.4 | |
| 500.0 | 460.2 ± 2.5 | 92.0 ± 0.5 | |
| Average (n = 9) | 93.0 ± 0.9 | ||
| Confidence interval (95%) | 92.1–93.9 | ||
| Alanine | 20.0 | 18.2 ± 0.1 | 91.2 ± 0.4 |
| 100.0 | 90.3 ± 0.6 | 90.3 ± 0.6 | |
| 500.0 | 446.2 ± 0.9 | 89.2 ± 0.2 | |
| Average (n = 9) | 90.2 ± 1.0 | ||
| Confidence interval (95%) | 89.3–91.2 | ||
| Valine | 30.0 | 27.2 ± 0.3 | 90.8 ± 0.8 |
| 200.0 | 179.9 ± 1.2 | 90.0 ± 0.6 | |
| 1000.0 | 885.0 ± 7.5 | 88.5 ± 0.7 | |
| Average (n = 9) | 89.7 ± 1.3 | ||
| Confidence interval (95%) | 88.5–91.0 | ||
| Isoleucine | 100.0 | 95.1 ± 0.3 | 95.1 ± 0.3 |
| 200.0 | 187.5 ± 0.4 | 93.7 ± 0.2 | |
| 1000.0 | 938.3 ± 0.9 | 93.8 ± 0.1 | |
| Average (n = 9) | 94.2 ± 0.7 | ||
| Confidence interval (95%) | 93.5–94.9 | ||
| Leucine | 50.0 | 48.0 ± 0.1 | 96.0 ± 0.2 |
| 200.0 | 191.5 ± 1.4 | 95.7 ± 0.7 | |
| 1000.0 | 949.7 ± 1.7 | 95.0 ± 0.2 | |
| Average (n = 9) | 95.6 ± 0.7 | ||
| Confidence interval (95%) | 94.9–96.2 | ||
| %Recovery | |||||
|---|---|---|---|---|---|
| Histidine | Alanine | Valine | Isoleucine | Leucine | |
| Predicted Value–LSE | 90.6 | 89.5 | 86.4 | 87.0 | 89.4 |
| Predicted Value–SPE | 87.1 | 84.0 | 84.8 | 81.2 | 86.6 |
| Predicted value (average) | 88.9 | 86.8 | 85.6 | 84.1 | 88.0 |
| Experimental value (average) | 93.0 | 90.2 | 89.7 | 94.2 | 95.6 |
| %Prediction Error | 4.4 | 3.8 | 4.6 | 10.7 | 7.9 |
| Standard | Calibration Range (μg mL−1) | Regression Equation | r2 | LOD (μg mL−1) | LOQ (μg mL−1) |
|---|---|---|---|---|---|
| Histidine | 0.02–5.0 | 27.9x − 0.23 | 0.9992 | 0.01 | 0.02 |
| Alanine | 0.01–2.0 | 95.96x − 1.89 | 0.9991 | 0.002 | 0.01 |
| Valine | 0.01–8.0 | 18.67x − 0.06 | 0.9996 | 0.002 | 0.01 |
| Isoleucine | 0.01–8.0 | 18.96x − 0.13 | 0.9997 | 0.004 | 0.01 |
| Leucine | 0.03–15.0 | 10.15x − 0.04 | 0.9991 | 0.01 | 0.03 |
| Amino Acids | Recommended Dosage (mg)/Day | Quantity (mg)/ Formulation |
|---|---|---|
| Leucine | 1700 | 540 |
| Isoleucine | 850 | 270 |
| Valine | 850 | 270 |
| Histidine | 520 | 175 |
| Alanine | 750 | 250 |
| Milk chocolate (excipient.) | - | 4.6 g |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Synaridou, M.S.; Tsamis, V.; Sidiropoulou, G.; Zacharis, C.K.; Panderi, I.; Markopoulou, C.K. Fluorimetric Analysis of Five Amino Acids in Chocolate: Development and Validation. Molecules 2021, 26, 4325. https://doi.org/10.3390/molecules26144325
Synaridou MS, Tsamis V, Sidiropoulou G, Zacharis CK, Panderi I, Markopoulou CK. Fluorimetric Analysis of Five Amino Acids in Chocolate: Development and Validation. Molecules. 2021; 26(14):4325. https://doi.org/10.3390/molecules26144325
Chicago/Turabian StyleSynaridou, Maria S., Vasilis Tsamis, Georgia Sidiropoulou, Constantinos K. Zacharis, Irene Panderi, and Catherine K. Markopoulou. 2021. "Fluorimetric Analysis of Five Amino Acids in Chocolate: Development and Validation" Molecules 26, no. 14: 4325. https://doi.org/10.3390/molecules26144325
APA StyleSynaridou, M. S., Tsamis, V., Sidiropoulou, G., Zacharis, C. K., Panderi, I., & Markopoulou, C. K. (2021). Fluorimetric Analysis of Five Amino Acids in Chocolate: Development and Validation. Molecules, 26(14), 4325. https://doi.org/10.3390/molecules26144325








