Novel DNA Bis-Intercalator XR5944 as a Potent Anticancer Drug—Design and Mechanism of Action
Abstract
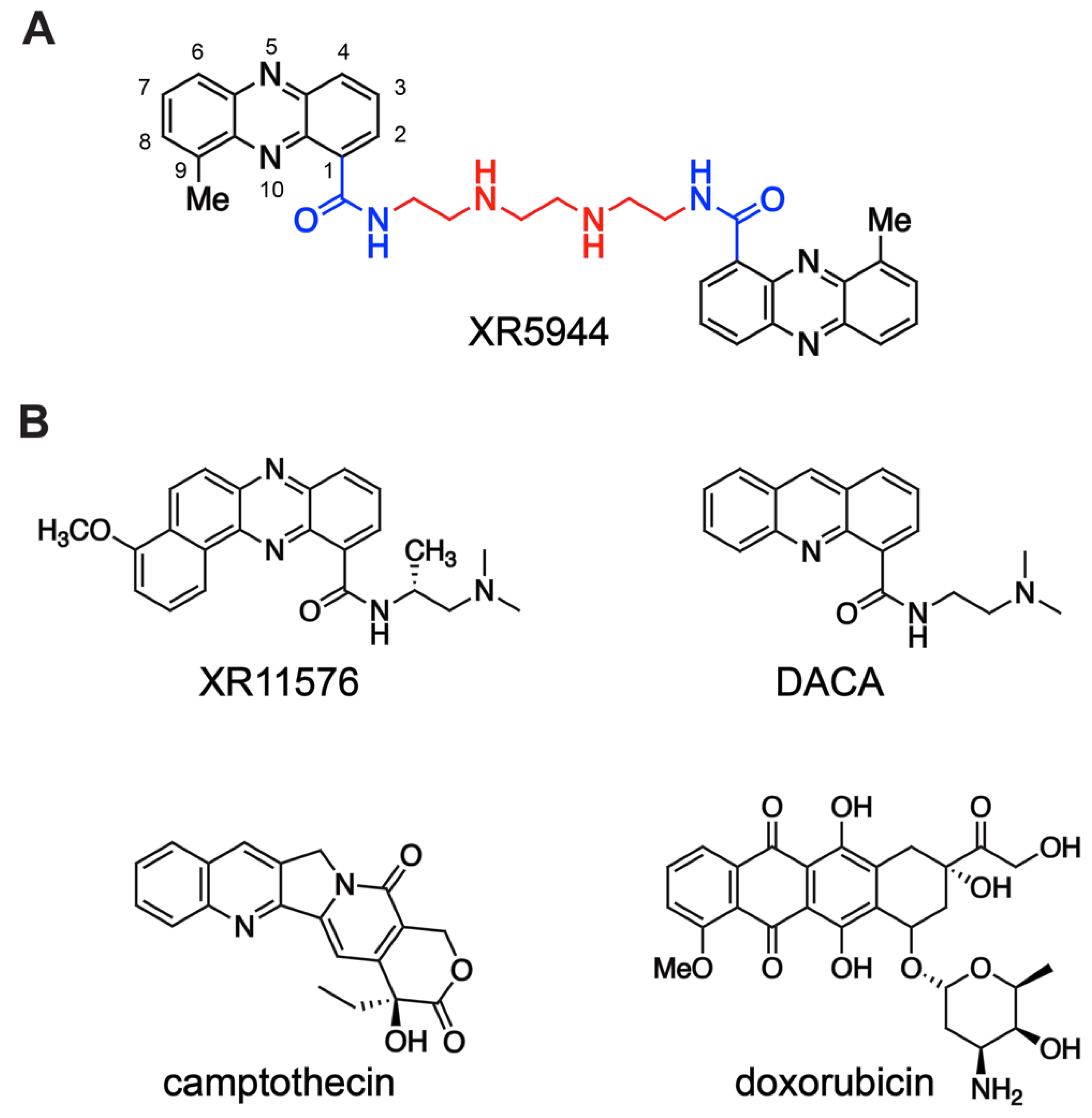
1. Introduction
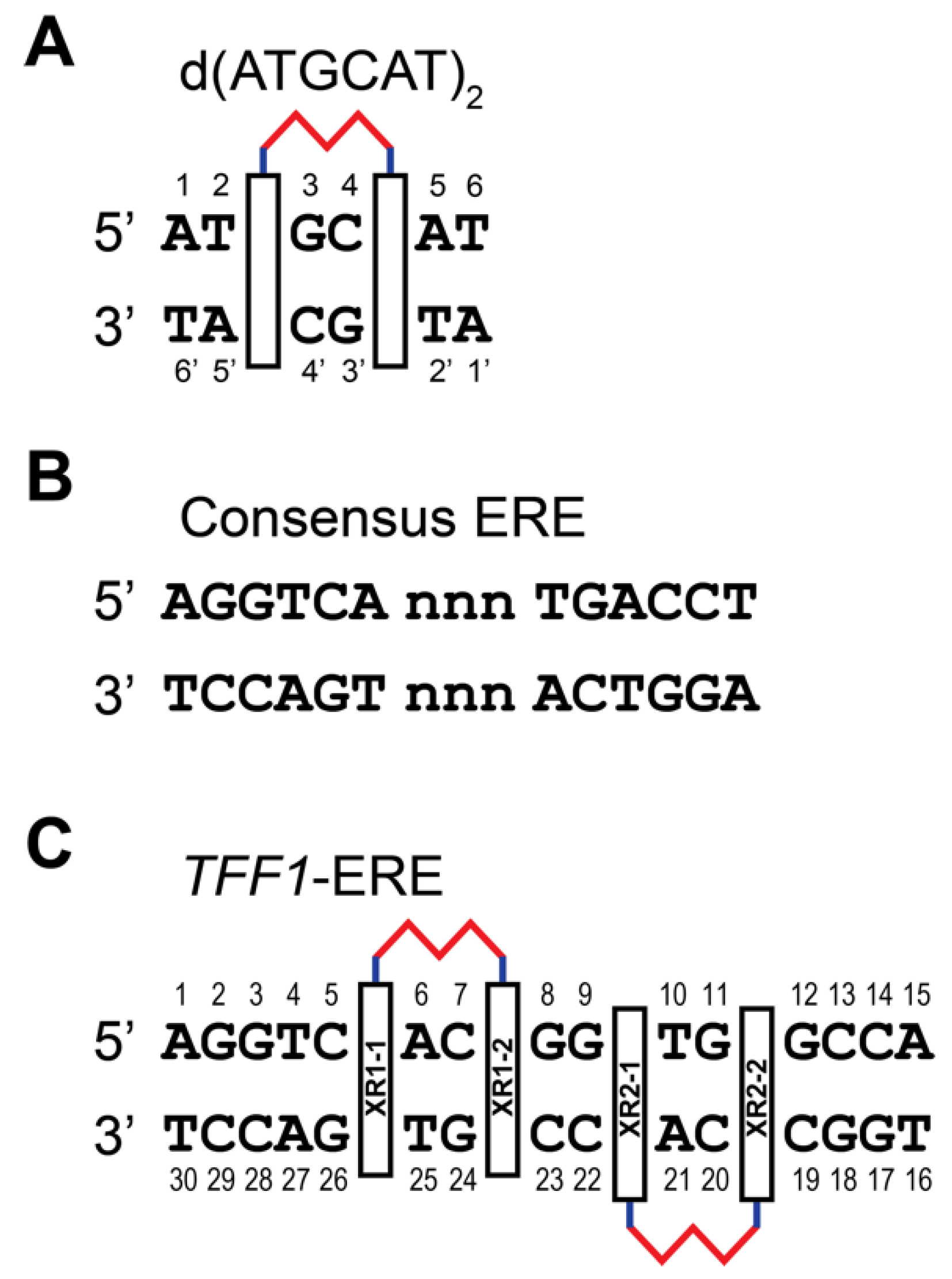
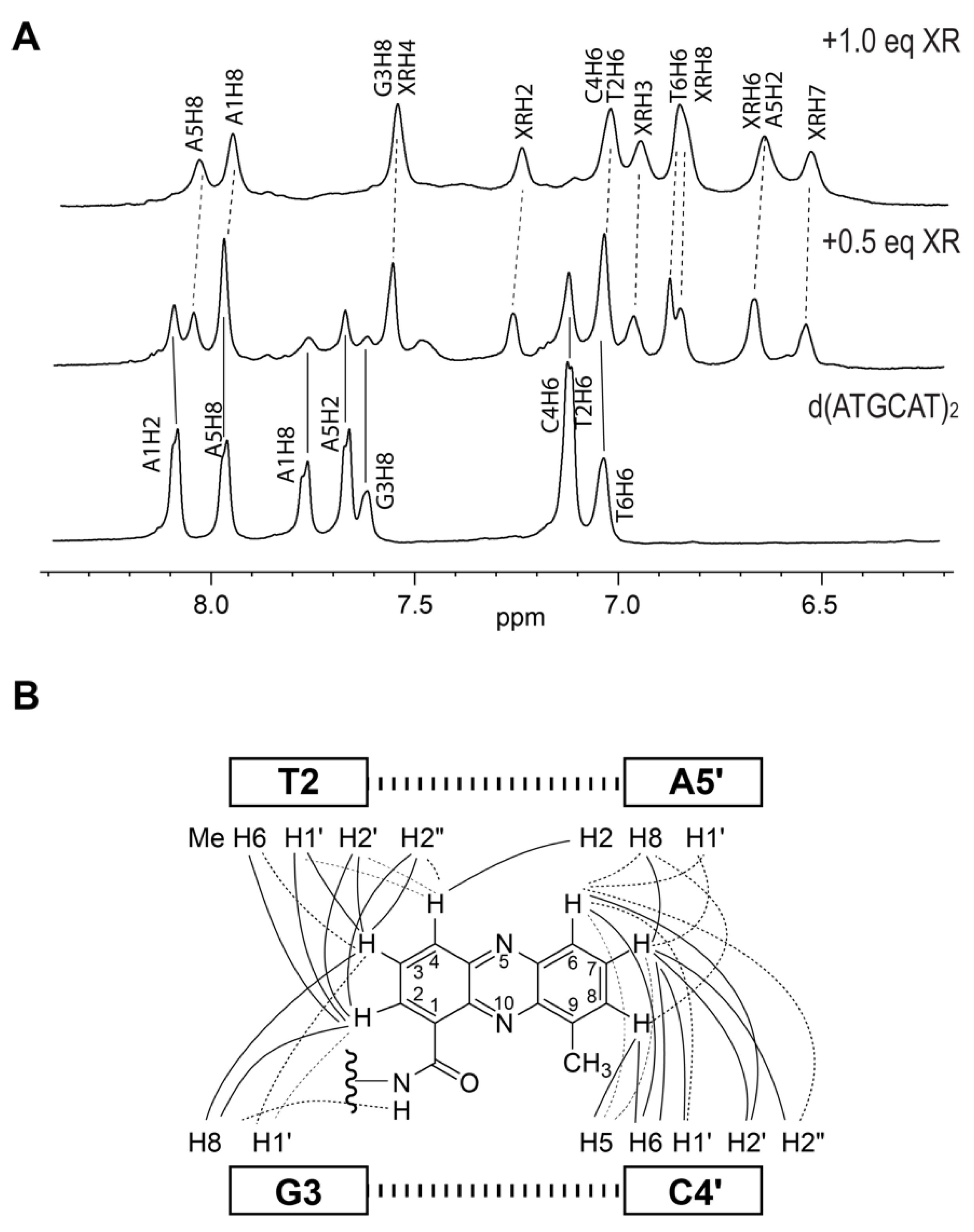
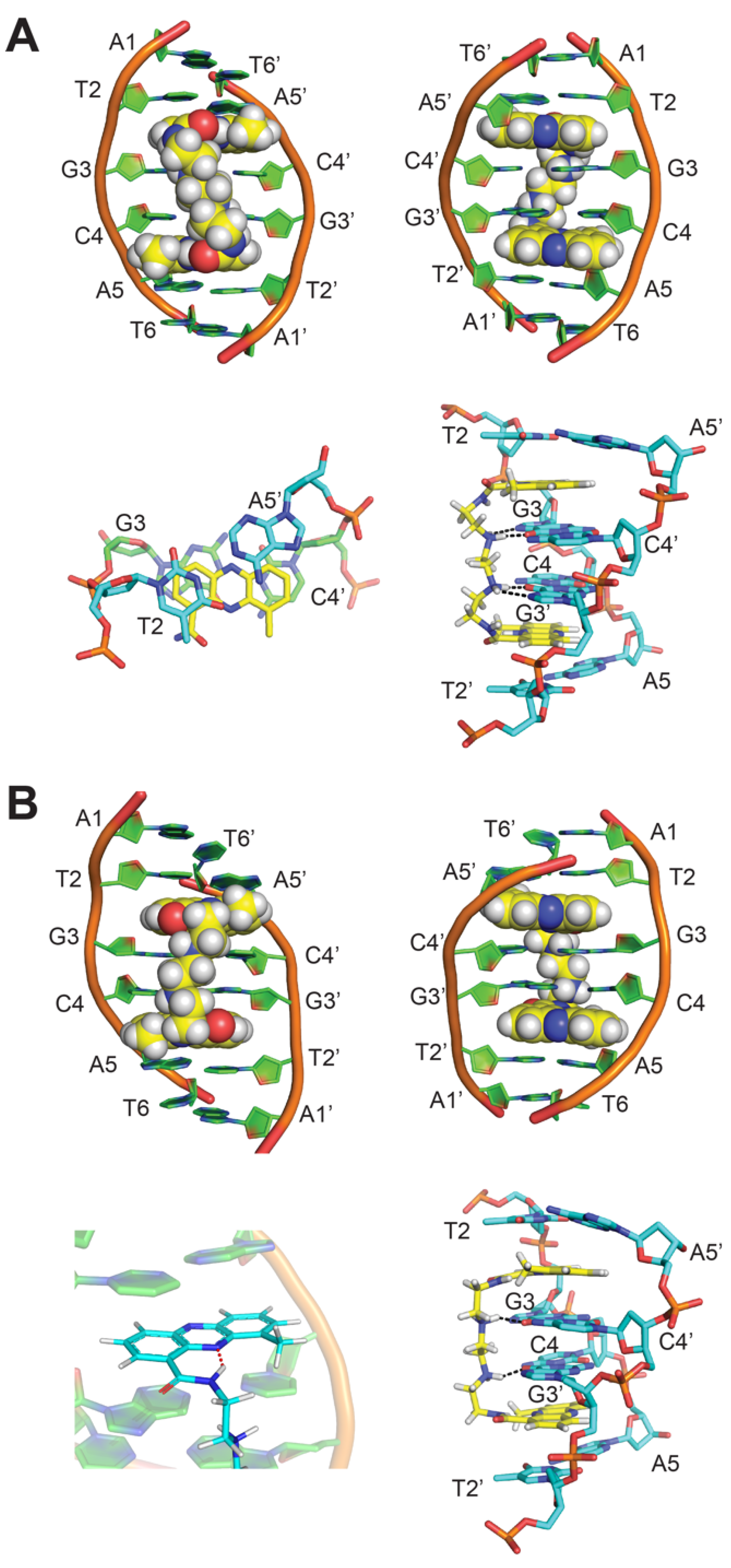
2. NMR Solution Structure of XR5944 in Complex with the d(ATGCAT)2 Duplex DNA
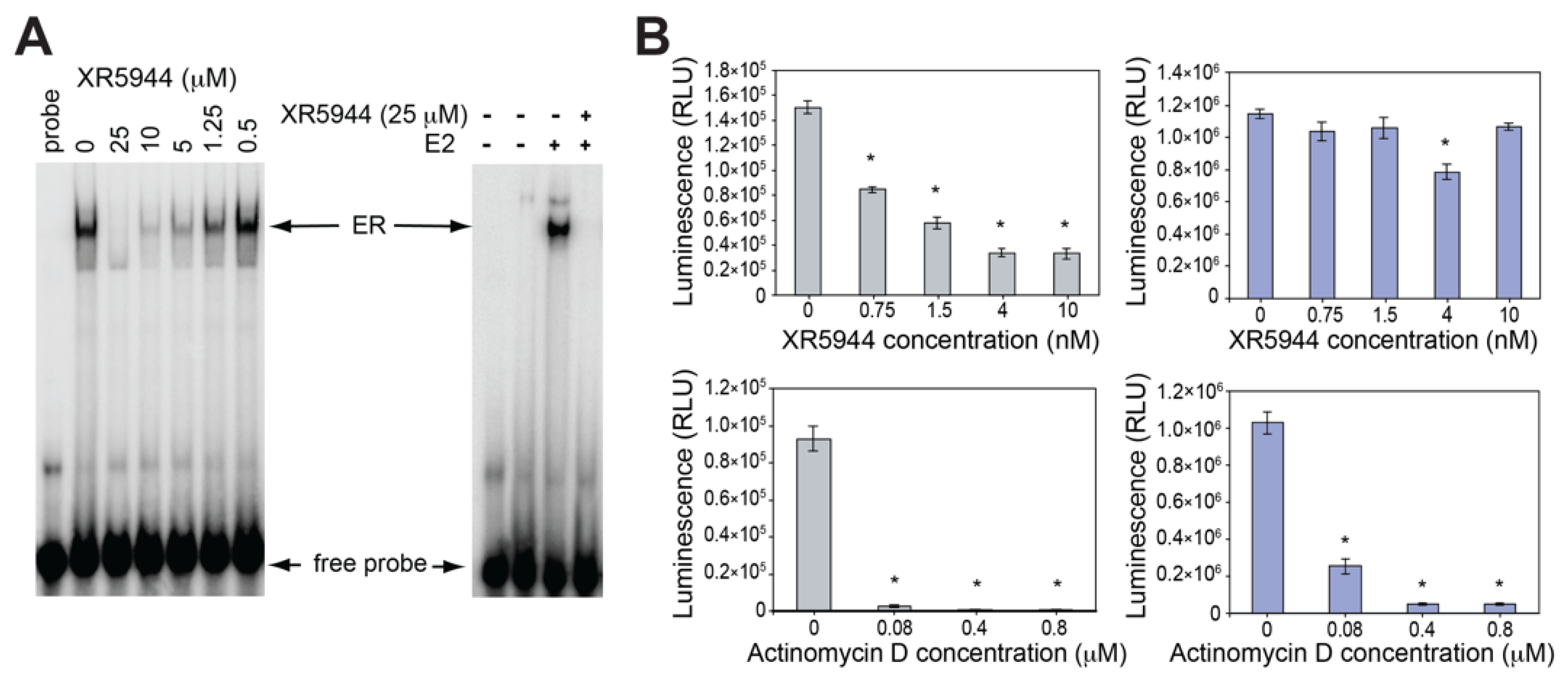
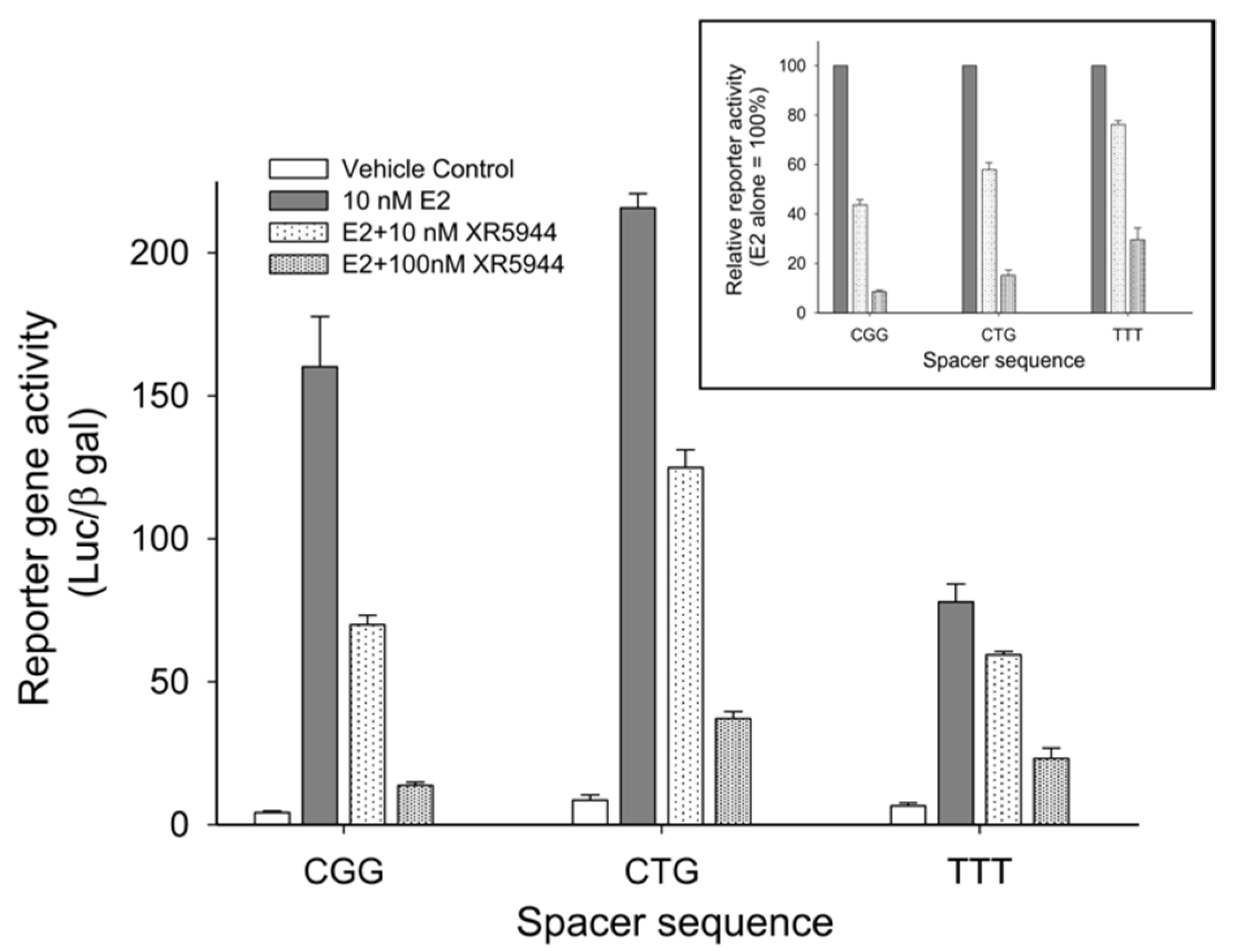
3. XR5944 Binds the ERE Sequence and Inhibits ERα-ERE Interaction
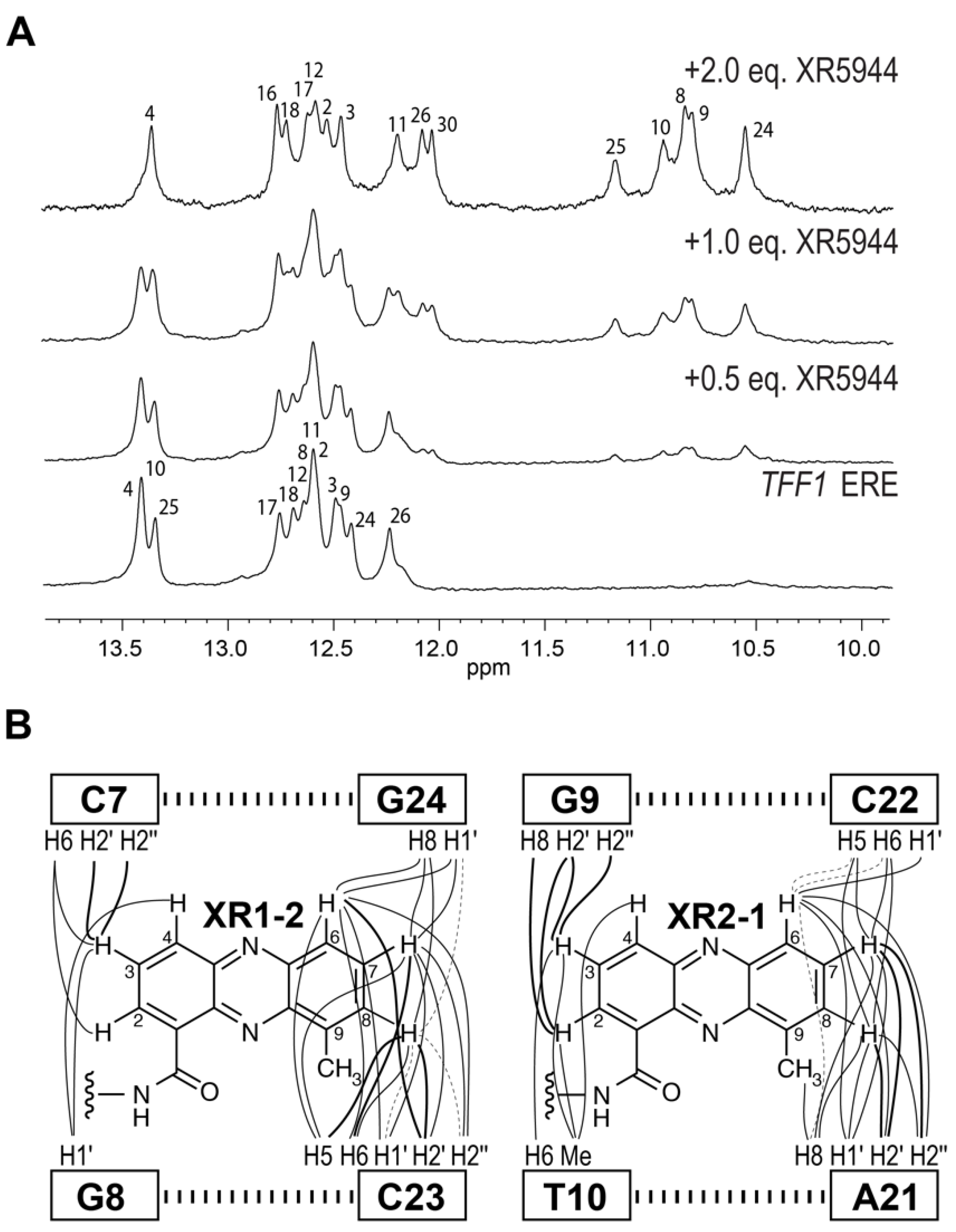
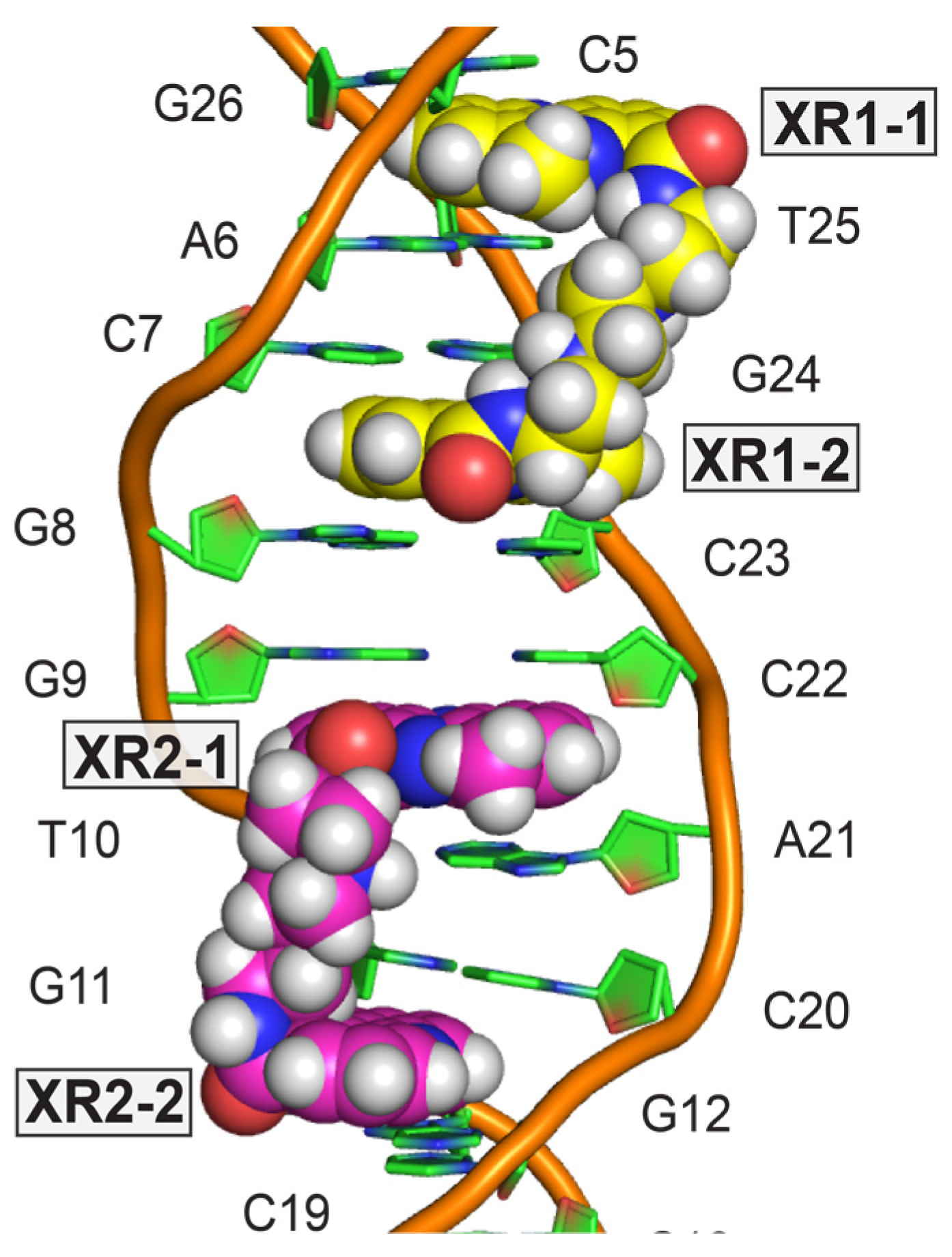
4. Solution Structure of 2:1 Complex of XR5944 with the Naturally Occurring ERE Sequence of TFF1 Gene
5. Disrupting ERE-ERα Complex Formation: A Potential Alternative to Antiestrogen Therapeutics
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Cuya, S.M.; Bjornsti, M.-A.; van Waardenburg, R.C.A.M. DNA Topoisomerase-Targeting Chemotherapeutics: What’s New? Cancer Chemother. Pharmacol. 2017, 80, 1–14. [Google Scholar] [CrossRef]
- Portugal, J. Challenging Transcription by DNA-Binding Antitumor Drugs. Biochem. Pharmacol. 2018, 155, 336–345. [Google Scholar] [CrossRef]
- Mokhtari, R.B.; Homayouni, T.S.; Baluch, N.; Morgatskaya, E.; Kumar, S.; Das, B.; Yeger, H. Combination Therapy in Combating Cancer. Oncotarget 2017, 8, 38022–38043. [Google Scholar] [CrossRef]
- Dai, J.; Punchihewa, C.; Mistry, P.; Ooi, A.T.; Yang, D. Novel DNA Bis-Intercalation by MLN944, a Potent Clinical Bisphenazine Anticancer Drug. J. Biol. Chem. 2004, 279, 46096–46103. [Google Scholar] [CrossRef]
- Byers, S.A.; Schafer, B.; Sappal, D.S.; Brown, J.; Price, D.H. The Antiproliferative Agent MLN944 Preferentially Inhibits Transcription. Mol. Cancer Ther. 2005, 4, 1260–1267. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Verborg, W.; Thomas, H.; Bissett, D.; Waterfall, J.; Steiner, J.; Cooper, M.; Rankin, E.M. First-into-Man Phase I and Pharmacokinetic Study of XR5944.14, a Novel Agent with a Unique Mechanism of Action. Br. J. Cancer 2007, 97, 844–850. [Google Scholar] [CrossRef][Green Version]
- Stewart, A.J.; Mistry, P.; Dangerfield, W.; Bootle, D.; Baker, M.; Kofler, B.; Okiji, S.; Baguley, B.C.; Denny, W.A.; Charlton, P.A. Antitumor Activity of XR5944, a Novel and Potent Topoisomerase Poison. Anti-Cancer Drugs 2001, 12, 359–367. [Google Scholar] [CrossRef] [PubMed]
- Gamage, S.A.; Spicer, J.A.; Finlay, G.J.; Stewart, A.J.; Charlton, P.; Baguley, B.C.; Denny, W.A. Dicationic Bis(9-Methylphenazine-1-Carboxamides): Relationships between Biological Activity and Linker Chain Structure for a Series of Potent Topoisomerase Targeted Anticancer Drugs. J. Med. Chem. 2001, 44, 1407–1415. [Google Scholar] [CrossRef] [PubMed]
- Di Nicolantonio, F.; Knight, L.A.; Whitehouse, P.A.; Mercer, S.J.; Sharma, S.; Charlton, P.A.; Norris, D.; Cree, I.A. The Ex Vivo Characterization of XR5944 (MLN944) against a Panel of Human Clinical Tumor Samples. Mol. Cancer Ther. 2004, 3, 1631–1637. [Google Scholar] [PubMed]
- Harris, S.M.; Mistry, P.; Freathy, C.; Brown, J.L.; Charlton, P.A. Antitumour Activity of XR5944 in Vitro and in Vivo in Combination with 5-Fluorouracil and Irinotecan in Colon Cancer Cell Lines. Br. J. Cancer 2005, 92, 722–728. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Harris, S.M.; Scott, J.A.; Brown, J.L.; Charlton, P.A.; Mistry, P. Preclinical Anti-Tumor Activity of XR5944 in Combination with Carboplatin or Doxorubicin in Non-Small-Cell Lung Carcinoma. Anti-Cancer Drugs 2005, 16, 945–951. [Google Scholar] [CrossRef]
- Millennium and Xenova Initiate Phase I Clinical Trial Of MLN944. Available online: https://www.takedaoncology.com/media/news-media/news-releases/Millennium-and-Xenova-Initiate-Phase-I-Clinical-Trial-Of-MLN-7341/Print (accessed on 17 June 2021).
- Atwell, G.J.; Cain, B.F.; Baguley, B.C.; Finlay, G.J.; Denny, W.A. Potential Antitumor Agents. Part 43. Synthesis and Biological Activity of Dibasic 9-Aminoacridine-4-Carboxamides, a New Class of Antitumor Agent. J. Med. Chem. 1984, 27, 1481–1485. [Google Scholar] [CrossRef]
- Atwell, G.J.; Rewcastle, G.W.; Baguley, B.C.; Denny, W.A. Potential Antitumor Agents. 50. In Vivo Solid-Tumor Activity of Derivatives of N-[2-(Dimethylamino)Ethyl]Acridine-4-Carboxamide. J. Med. Chem. 1987, 30, 664–669. [Google Scholar] [CrossRef] [PubMed]
- Rewcastle, G.W.; Denny, W.A.; Baguley, B.C. Potential Antitumor Agents. 51. Synthesis and Antitumor Activity of Substituted Phenazine-1-Carboxamides. J. Med. Chem. 1987, 30, 843–851. [Google Scholar] [CrossRef]
- Palmer, B.D.; Rewcastle, G.W.; Atwell, G.J.; Baguley, B.C.; Denny, W.A. Potential Antitumor Agents. 54. Chromophore Requirements for in Vivo Antitumor Activity among the General Class of Linear Tricyclic Carboxamides. J. Med. Chem. 1988, 31, 707–712. [Google Scholar] [CrossRef] [PubMed]
- Spicer, J.A.; Gamage, S.A.; Atwell, G.J.; Finlay, G.J.; Baguley, B.C.; Denny, W.A. Dimeric Analogues of Non-Cationic Tricyclic Aromatic Carboxamides Are a New Class of Cytotoxic Agents. Anticancer Drug Des. 1999, 14, 281–289. [Google Scholar]
- Spicer, J.A.; Gamage, S.A.; Rewcastle, G.W.; Finlay, G.J.; Bridewell, D.J.A.; Baguley, B.C.; Denny, W.A. Bis(Phenazine-1-Carboxamides): Structure−Activity Relationships for a New Class of Dual Topoisomerase I/II-Directed Anticancer Drugs. J. Med. Chem. 2000, 43, 1350–1358. [Google Scholar] [CrossRef]
- Wang, S.; Miller, W.; Milton, J.; Vicker, N.; Stewart, A.; Charlton, P.; Mistry, P.; Hardick, D.; Denny, W.A. Structure–Activity Relationships for Analogues of the Phenazine-Based Dual Topoisomerase I/II Inhibitor XR11576. Bioorganic Med. Chem. Lett. 2002, 12, 415–418. [Google Scholar] [CrossRef]
- Finlay, G.J.; Riou, J.-F.; Baguley, B.C. From Amsacrine to DACA (N-[2-(Dimethylamino)Ethyl]Acridine-4-Carboxamide): Selectivity for Topoisomerases I and II among Acridine Derivatives. Eur. J. Cancer 1996, 32, 708–714. [Google Scholar] [CrossRef]
- Sappal, D.S.; McClendon, A.K.; Fleming, J.A.; Thoroddsen, V.; Connolly, K.; Reimer, C.; Blackman, R.K.; Bulawa, C.E.; Osheroff, N.; Charlton, P.; et al. Biological Characterization of MLN944: A Potent DNA Binding Agent. Mol. Cancer Ther. 2004, 3, 47–58. [Google Scholar] [PubMed]
- Serobian, A.; Thomas, D.S.; Ball, G.E.; Denny, W.A.; Wakelin, L.P.G. The Solution Structure of Bis(Phenazine-1-Carboxamide)-DNA Complexes: MLN 944 Binding Corrected and Extended: MLN 944 Binding Corrected and Extended. Biopolymers 2014, 101, 1099–1113. [Google Scholar] [CrossRef] [PubMed]
- Adams, A.; Guss, J.M.; Collyer, C.A.; Denny, W.A.; Wakelin, L.P. A Novel Form of Intercalation Involving Four DNA Duplexes in an Acridine-4-Carboxamide Complex of d(CGTACG)(2). Nucleic Acids Res. 2000, 28, 4244–4253. [Google Scholar] [CrossRef][Green Version]
- Todd, A.K.; Adams, A.; Thorpe, J.H.; Denny, W.A.; Wakelin, L.P.G.; Cardin, C.J. Major Groove Binding and ‘DNA-Induced’ Fit in the Intercalation of a Derivative of the Mixed Topoisomerase I/II Poison N -(2-(Dimethylamino)Ethyl)Acridine-4- Carboxamide (DACA) into DNA: X-Ray Structure Complexed to d(CG(5-BrU)ACG) 2 at 1.3-Å Resolution. J. Med. Chem. 1999, 42, 536–540. [Google Scholar] [CrossRef]
- Bhaduri, S.; Ranjan, N.; Arya, D.P. An Overview of Recent Advances in Duplex DNA Recognition by Small Molecules. Beilstein J. Org. Chem. 2018, 14, 1051–1086. [Google Scholar] [CrossRef]
- Rohs, R.; Jin, X.; West, S.M.; Joshi, R.; Honig, B.; Mann, R.S. Origins of Specificity in Protein-DNA Recognition. Annu. Rev. Biochem. 2010, 79, 233–269. [Google Scholar] [CrossRef]
- MacGregor, J.I.; Jordan, V.C. Basic Guide to the Mechanisms of Antiestrogen Action. Pharmacol. Rev. 1998, 50, 151–196. [Google Scholar]
- Björnström, L.; Sjöberg, M. Mechanisms of Estrogen Receptor Signaling: Convergence of Genomic and Nongenomic Actions on Target Genes. J. Mol. Endocrinol. 2005, 19, 833–842. [Google Scholar] [CrossRef]
- Punchihewa, C.; De Alba, A.; Sidell, N.; Yang, D. XR5944: A Potent Inhibitor of Estrogen Receptors. Mol. Cancer. Ther. 2007, 6, 213–219. [Google Scholar] [CrossRef]
- Sidell, N.; Mathad, R.I.; Shu, F.; Zhang, Z.; Kallen, C.B.; Yang, D. Intercalation of XR5944 with the Estrogen Response Element Is Modulated by the Tri-Nucleotide Spacer Sequence between Half-Sites. J. Steroid Biochem. Mol. Biol. 2011, 124, 121–127. [Google Scholar] [CrossRef] [PubMed]
- Shu, F.; Sidell, N.; Yang, D.; Kallen, C.B. The Tri-Nucleotide Spacer Sequence between Estrogen Response Element Half-Sites Is Conserved and Modulates ERα-Mediated Transcriptional Responses. J. Steroid Biochem. Mol. Biol. 2010, 120, 172–179. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lin, C.; Mathad, R.I.; Zhang, Z.; Sidell, N.; Yang, D. Solution Structure of a 2:1 Complex of Anticancer Drug XR5944 with TFF1 Estrogen Response Element: Insights into DNA Recognition by a Bis-Intercalator. Nucleic Acids Res. 2014, 42, 6012–6024. [Google Scholar] [CrossRef] [PubMed]
- Clark, G.M. Prognostic and Predictive Factors for Breast Cancer. Breast Cancer 1995, 2, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Hua, H.; Zhang, H.; Kong, Q.; Jiang, Y. Mechanisms for Estrogen Receptor Expression in Human Cancer. Exp. Hematol. Oncol. 2018, 7, 24. [Google Scholar] [CrossRef] [PubMed]
- Brufsky, A.M.; Dickler, M.N. Estrogen Receptor-Positive Breast Cancer: Exploiting Signaling Pathways Implicated in Endocrine Resistance. The Oncol. 2018, 23, 528–539. [Google Scholar] [CrossRef] [PubMed]
- Bouker, K.B.; Wang, Y.; Xuan, J.; Clarke, R. Antiestrogen Resistance and the Application of Systems Biology. Drug Discov. Today Dis. Mech. 2012, 9, e11–e17. [Google Scholar] [CrossRef][Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Buric, A.J.; Dickerhoff, J.; Yang, D. Novel DNA Bis-Intercalator XR5944 as a Potent Anticancer Drug—Design and Mechanism of Action. Molecules 2021, 26, 4132. https://doi.org/10.3390/molecules26144132
Buric AJ, Dickerhoff J, Yang D. Novel DNA Bis-Intercalator XR5944 as a Potent Anticancer Drug—Design and Mechanism of Action. Molecules. 2021; 26(14):4132. https://doi.org/10.3390/molecules26144132
Chicago/Turabian StyleBuric, Adam J., Jonathan Dickerhoff, and Danzhou Yang. 2021. "Novel DNA Bis-Intercalator XR5944 as a Potent Anticancer Drug—Design and Mechanism of Action" Molecules 26, no. 14: 4132. https://doi.org/10.3390/molecules26144132
APA StyleBuric, A. J., Dickerhoff, J., & Yang, D. (2021). Novel DNA Bis-Intercalator XR5944 as a Potent Anticancer Drug—Design and Mechanism of Action. Molecules, 26(14), 4132. https://doi.org/10.3390/molecules26144132





