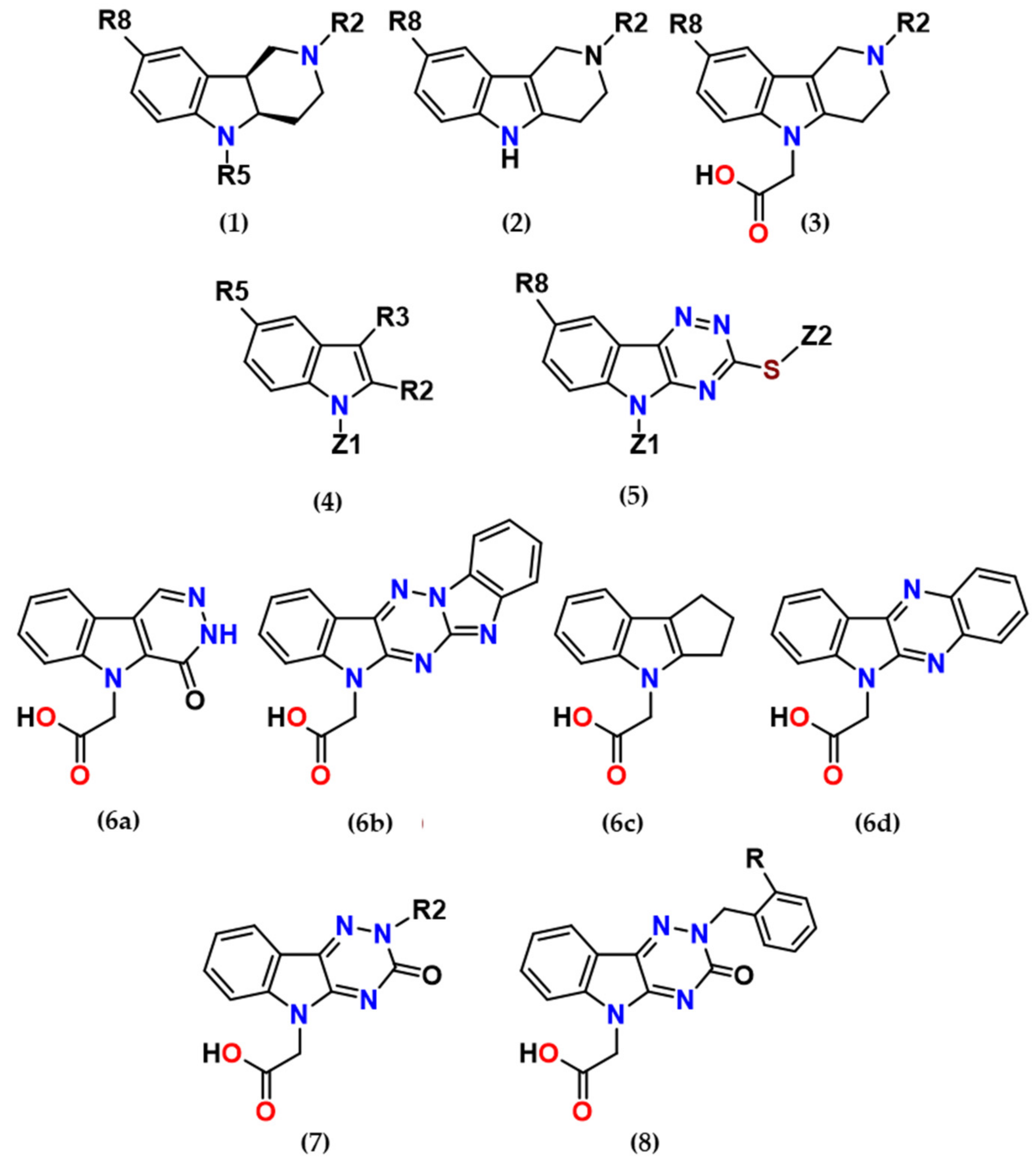
2.1. Inhibition of Aldo-Keto Reductases ALR2 and ALR1
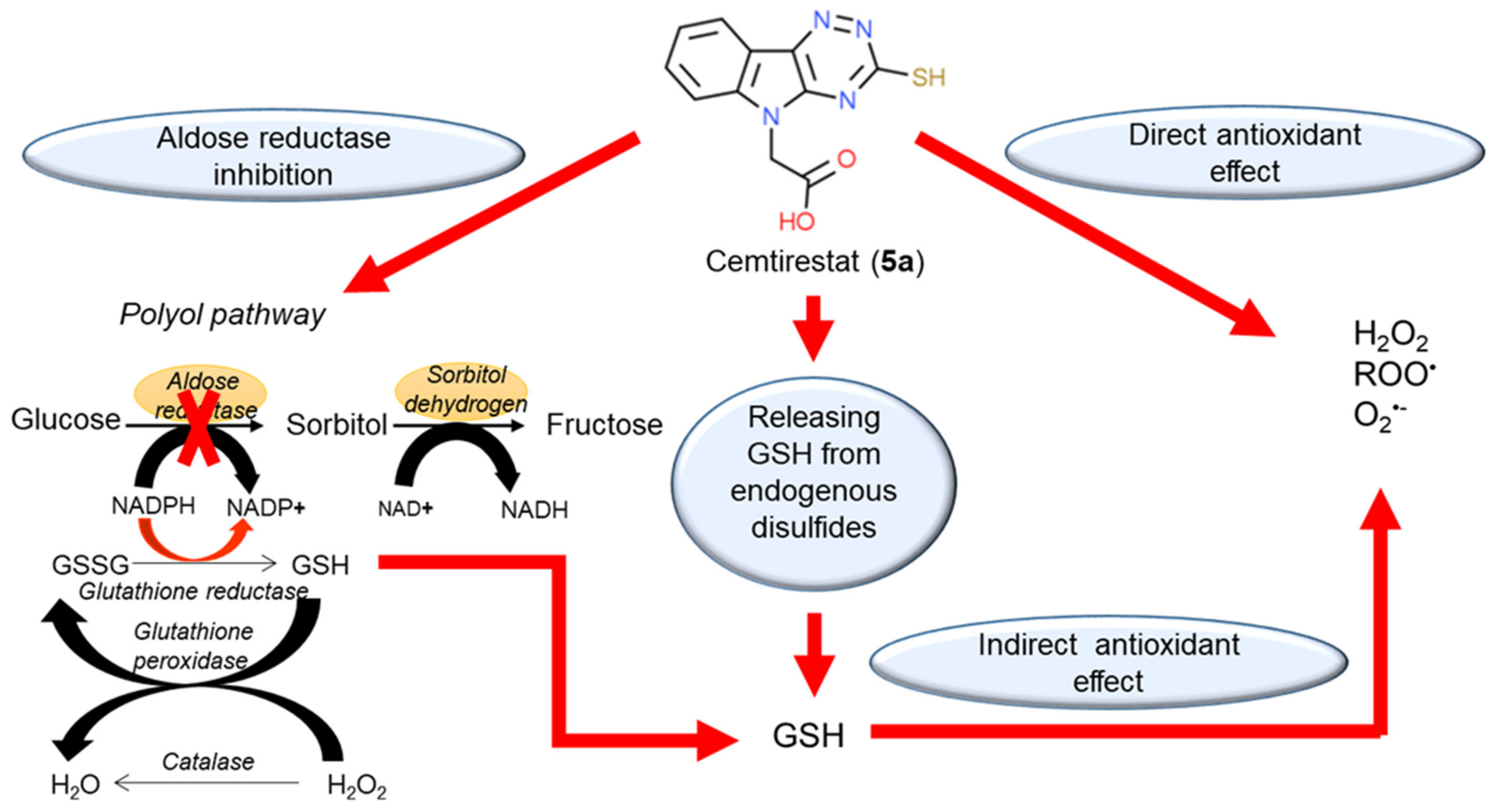
Aldose reductase (AR, ALR2, EC 1.1.1.21) is a member of the aldo-keto reductase (AKR) superfamily, which is composed of monomeric proteins with approximately 315–330 residues, and with a molecular weight of 36 kD. It is the first enzyme of the polyol pathway and converts glucose to sorbitol in the presence of NADPH. The pathway is terminated by the second enzyme, sorbitol dehydrogenase, which converts sorbitol to fructose with NAD
+ as a cofactor. In diabetic patients, the flux of glucose through the polyol pathway results in intracellular accumulation of sorbitol with consequent disruption of tissue osmotic homeostasis. The concurrent depletion of NADPH and the imbalance of the NADH/NAD
+ ratio eventually lead to oxidative stress. As such, these metabolic derangements have been found implicated in the pathophysiology of diabetic complications [
1,
2,
3,
4,
5,
6]. Another member of the AKR superfamily is aldehyde reductase (ALR1, EC 1.1.1.2). ALR1 is present in all tissues and is responsible for the reduction of toxic aldehydes. ALR1 and ALR2 share a high degree of amino acid sequence (~65%) and structural homology and their co-inhibition may result in undesired side effects [
12]. Thus, to test selectivity of novel ARIs, the closely related ALR1 is routinely employed.
Based on the premise that a bifunctional compound with joint antioxidant/aldose reductase inhibitory (AO/ARI) activities could be multifactorially beneficial, the first three series of novel derivatives (series 1 to 3), structurally based on the antioxidant drug stobadine [
1a, (4a
R,9b
S)-2,8-dimethyl-2,3,4,4a,5,9
b-hexahydro-1
H-pyrido[4,3-b]indole] [
31,
32,
33,
34,
35] were designed and synthesized.
The parent drug stobadine (
1a,
Table 1) as an efficient reactive oxygen species (ROS) scavenger was extensively studied in multiple models of diabetic complications with the aim to attenuate the oxidative component of glucose toxicity. Indeed, under conditions of an experimental glycation model in vitro, compound
1a was found to protect bovine serum albumin against glyco-oxidative damage [
37,
38,
39]. Using a model of streptozotocin-diabetic rats in vivo, compound
1a was reported to attenuate pathological changes in diabetic cardiovascular system [
40,
41,
42,
43,
44], kidneys [
45,
46,
47,
48,
49,
50], eye lens [
51,
52], and retina [
53], vas deferens [
54], peripheral nerves [
55,
56], and brain [
57], to decrease matrix collagen cross-linking [
46,
58] and to reduce plasma cholesterol [
41] and triglyceride levels [
41,
42] in diabetic animals. Stobadine (
1a) treatment normalized calcium homeostasis in diabetic rat heart and liver [
42], produced a beneficial effect on leukocyte function [
59], inhibited doxorubicin-induced apoptosis [
60], and ameliorated alloxan toxicity in mice [
61,
62].
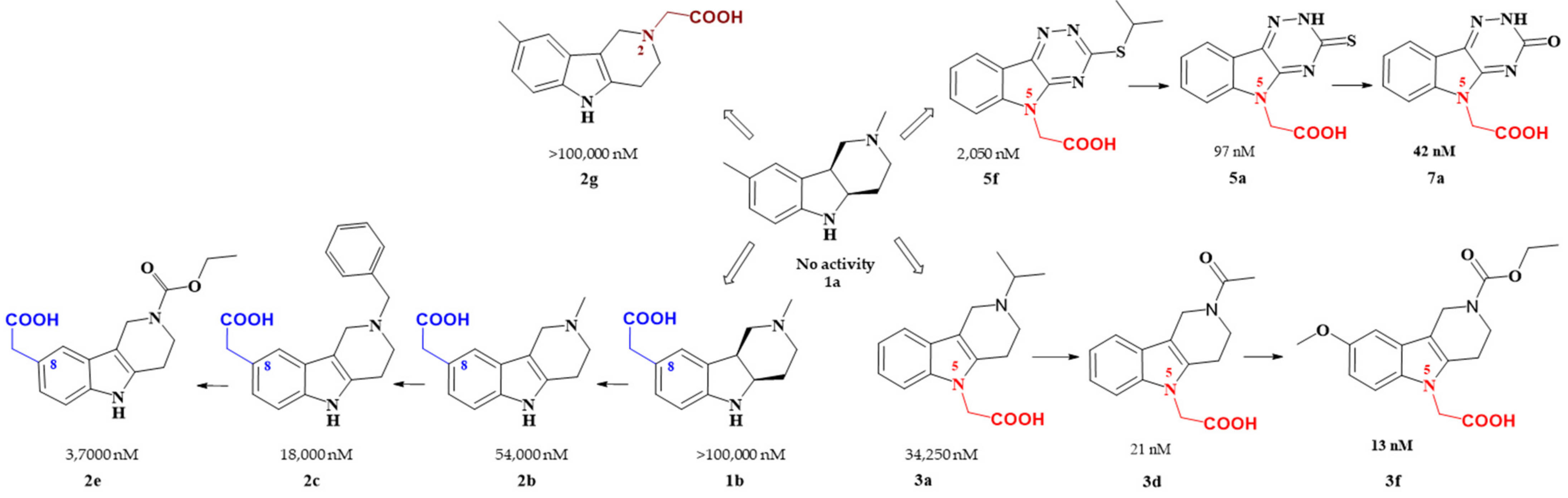
The rationale for designing of the novel multi-target directed/bifunctional drugs was based on an idea of endowing the stobadine scaffold bearing the antioxidant activity with a carboxymethyl functional group, a key pharmacophore of aldose reductase inhibitors. Hexahydropyridoindole stobadine (
1a) and its tetrahydro congener
2a, used as starting fragments of the drug design, were devoid of any ability to inhibit aldose reductase (
Table 1).
Introduction of the carboxymethyl group into the hexahydropyridoindole
1a at the position 8 resulted in compound
1b with a very mild inhibition of ALR2 [
I (%,100 µM) = 13%]. Linking the more lipophilic benzyl substituent into position 2 (compound
1c) did not affect the inhibition efficacy significantly. Yet inhibition activities in micromolar range were recorded for the unsaturated tetrahydropyridoindole congeners
2b–
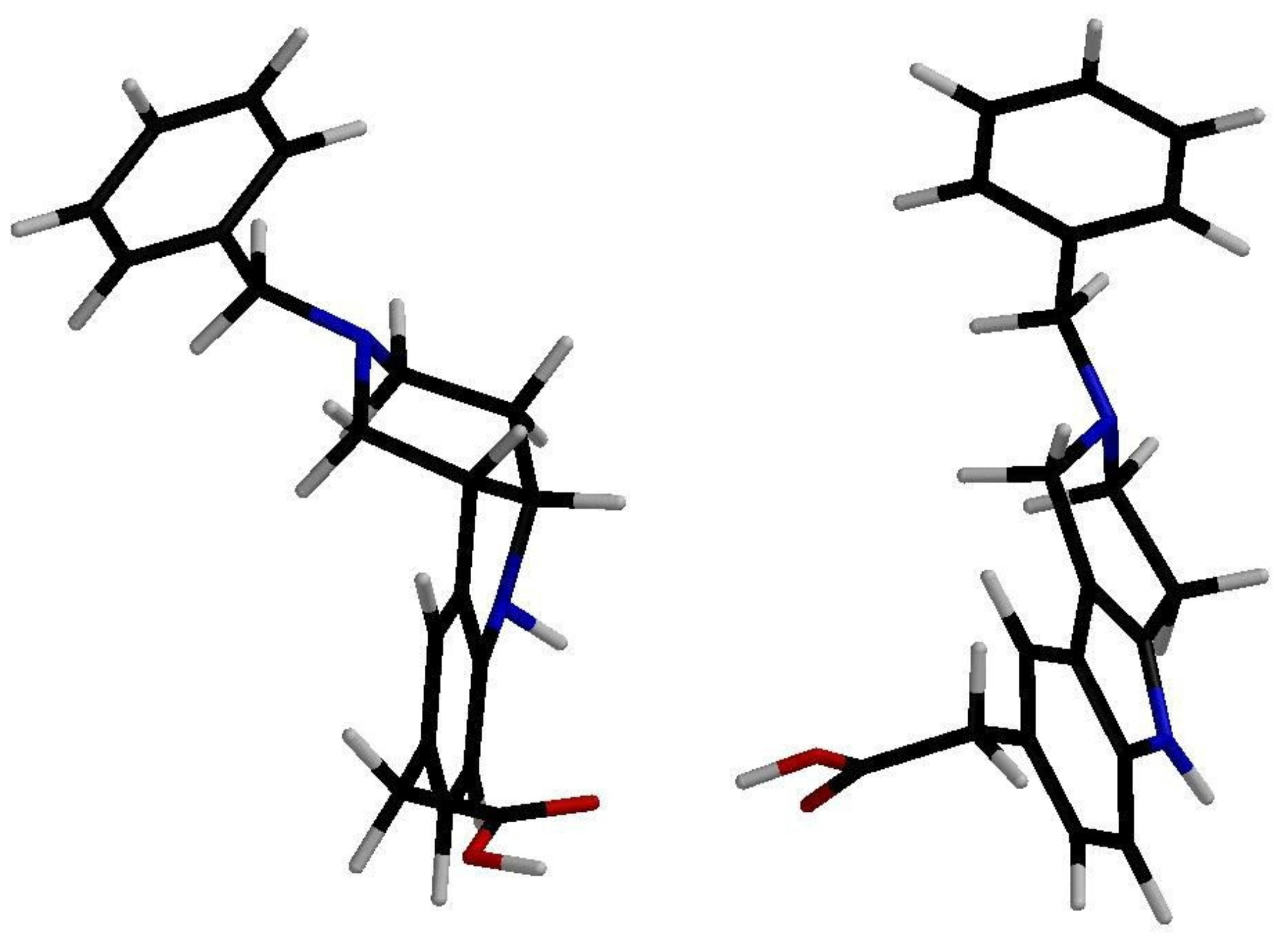
d. Visualization of low energy conformations of compounds
1c and
2c showed almost planar tricyclic moiety of the tetrahydropyridoindole
2c, contrasting with severe space distortion of the lipophilic heterocyclic backbone of the hexahydropyridoindole
1c (
Figure 2) [
63]. The presence of an extended aromatic planar region in the majority of potent ARIs is well documented as a crucial pharmacophoric element [
64,
65,
66].
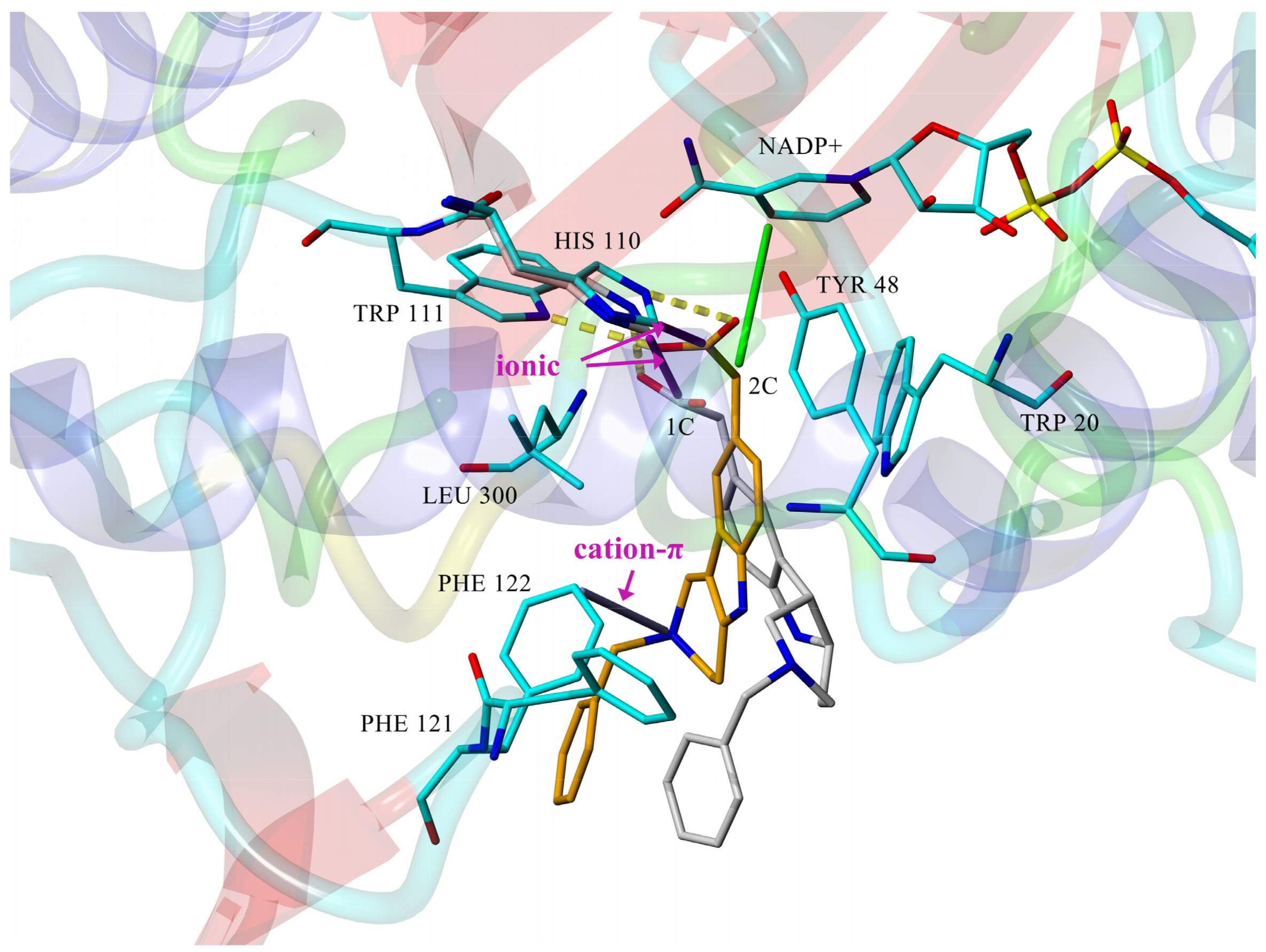
The computed stereoelectronic properties of compounds
1c and
2c along with a docking study into the ALR2-binding site (PDB: 1PWM) suggested an explanation for the higher inhibitory efficacies of the tetrahydropyridoindoles (
2) in comparison to the hexahydropyridoindoles (
1) shown in
Figure 3. Planar conformation of the tricyclic tetrahydropyridoindole moiety of
2c enabled closer contacts between the carboxymethyl pharmacophore and the key interaction partners His110 and NADP
+, in comparison with sterically distorted hexahydropyridoindole structure of compound
1c [
63]. Cation–π interaction between protonated
N2 nitrogen and benzene ring of Phe122 contributed to the stabilization of
2c-ALR2 complex, but also prevented
2c to adopt more advantageous position. Eliminating the basicity center at
N2 by introducing an acyl substituent in derivative
2e resulted in significant improvement of the inhibition efficacy [
68]. The selectivity of the tetrahydropyridoindoles
2b–
2d in relation to ALR1 was characterized by selectivity factors (SFs) in the range of 18 to 57. It is noteworthy that compound
2c retained ALR2 inhibitory activity even for the enzyme isolated from diabetic rats, with an IC
50 of 16.6 μM. The selectivity index of compound
2c even slightly increased from 18 to 21 in the enzyme preparations from diabetic rats [
69]. Compound
2c was reported [
70] to be selective in relation to the enzymes of the glycolytic pathway of glucose elimination.
Shifting the carboxymethyl pharmacophore from position 8 to position 2, yielded derivatives
2f–
h with markedly decreased inhibition efficacy (IC
50 >100 µM); therefore, this route of drug designing was not further followed [
67].
Transferring the carboxymethyl pharmacophore from position 8 to position 5, yielded derivatives (series
3) with markedly enhanced aldose reductase inhibition efficacy and selectivity (
Table 2) [
71]. Mild inhibition characterized by IC
50 in micromolar range was recorded for compound
3a with the isopropyl substituent in position 2. This alkylated tertiary nitrogen is characterized by a rather high basicity (pKa ~ 10, MarvinSketch Online 2016/ChemAxon), which ensures its complete protonation at physiological pH. The presence of a positive charge on the tertiary nitrogen, which predisposes these compounds to form double-charged zwitterionic species, has apparently a detrimental effect on AR inhibition efficacy. Similarly, only modest AR inhibition was recorded for structurally related zwitterionic 8-carboxymethylated pyridoindoles [
63,
72].
On the other hand, AR inhibition activity of the low basicity derivatives possessing an acyl or ethoxycarbonyl substituent on
N2 compounds
3b–
f is characterized with IC
50 values in low and medium nanomolar range (
Table 2). Based on SAR in this set of compounds, the flexible carbamate moiety of compounds
3e and
3f appears to fit better the enzyme-binding site contrary to alkyl– and aryl-acyl– substructures of compounds
3b–
d, respectively. In the latter series, the inhibition efficacy decreased with increasing bulkiness of the
N-substituent. The replacement of the methoxy group in position 8 of compound
3f by the more polar carboxylic group in compound
3e did not affect the resulting inhibition activity. Compounds
3b-
f revealed higher inhibition efficacy in comparison with the clinically used epalrestat. Compound
3b is presently undergoing clinical evaluation under the name of setipiprant for treatment of androgenic alopecia [
73]. On balance, these results establish the tetrahydropyridoindoles carboxymethylated at position 5 as a prospective scaffold for designing efficient AR inhibitors.
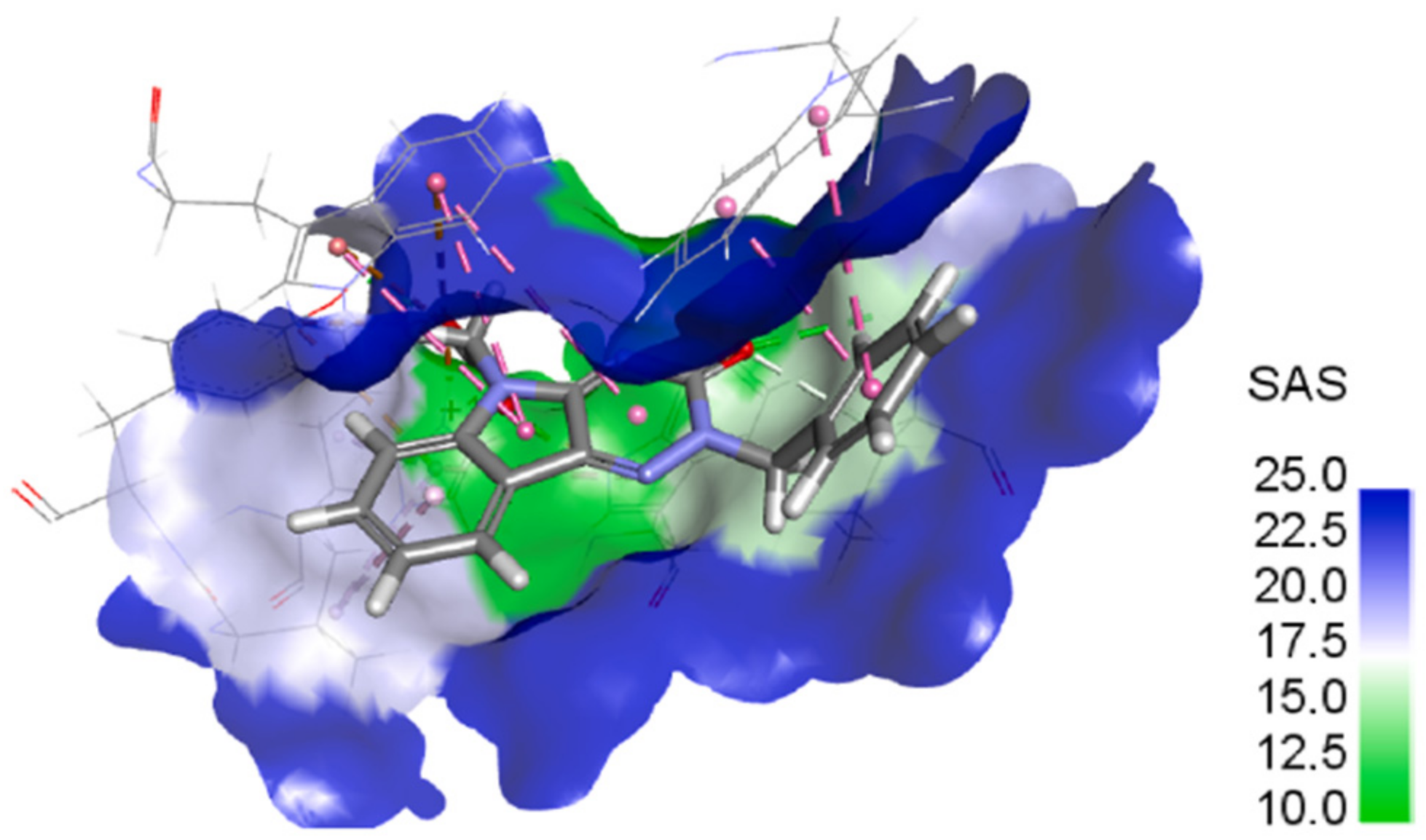
With the aim of analyzing possible interaction modes, human recombinant enzyme AKR1B1 in complex with structurally related lidorestat (PDB: 1z3n) was used for in silico docking followed by optimization of the resulting complexes in a water environment. The trial revealed several common features for the set of the most efficient inhibitors
3c–
f. The carboxylate group of these compounds and the one of lidorestat were found to align well. To avoid redundancy, details only for the derivative
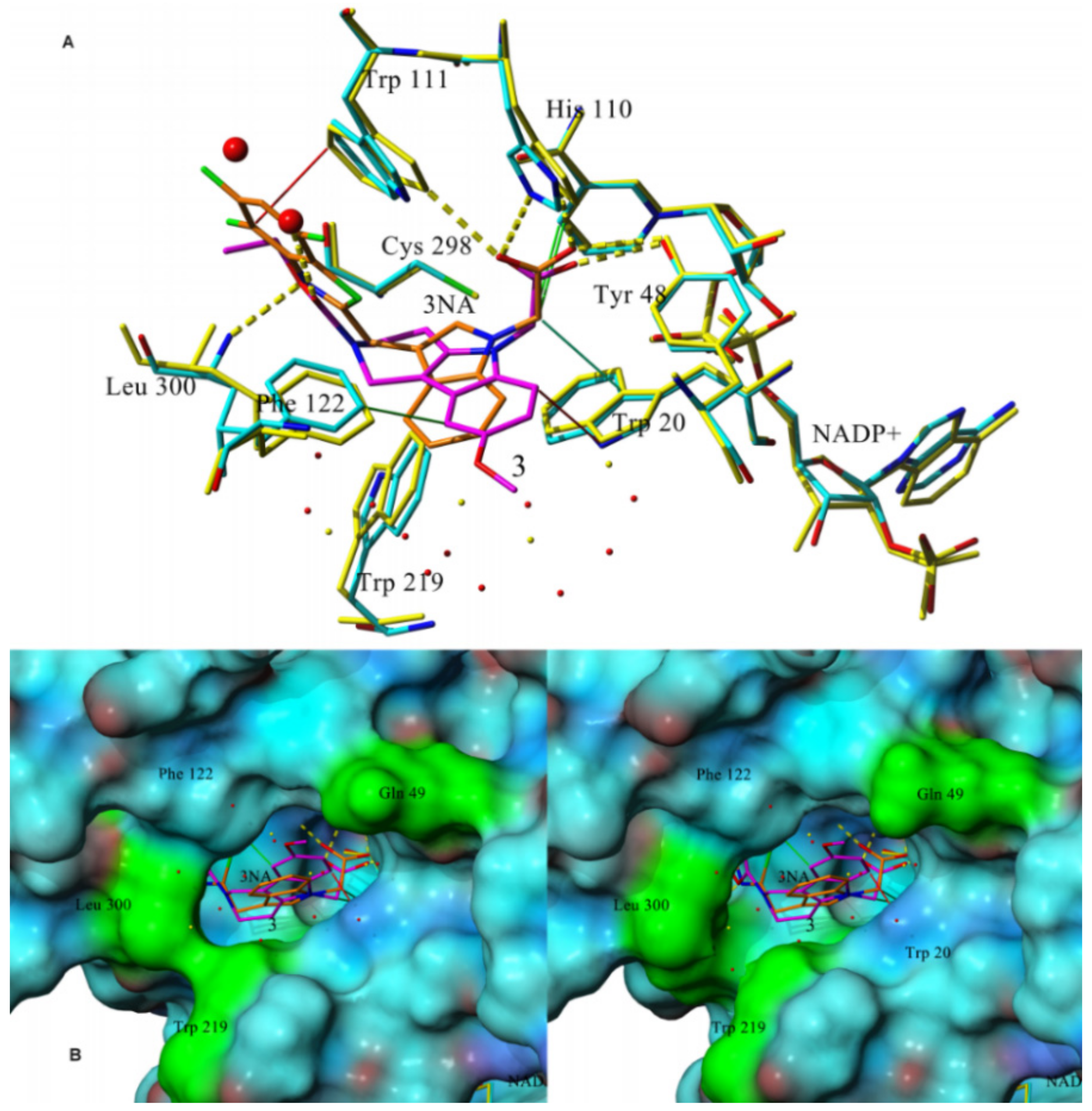
3f were reported. As shown in
Figure 4A, the carboxylate group is directed to the main residual trio of the “anion binding pocket“, Tyr48, His110, and Trp111.
The methylene group of the carboxymethyl substituent creates hydrophobic interactions with NADP
+ with the exception of the least efficient inhibitor
3a, the structure of which was, similarly as for
2c, captured in cation–π interaction with Phe122 keeping it too far from NADP
+. By their overall positions, compound
3f and lidorestat take up approximately the same space in the binding site with several specific differences. The trifluorobenzothiazole part of lidorestat creates strong π–π interactions with Trp111 and Phe122, leaving no space available for water molecules in the binding pocket. However, compound
3f binds firmly to Leu300, leaving enough space in the cavity for two water molecules and creating H-bond with one of them. Water environment plays an important role in thermodynamic balance of ligand-enzyme interactions [
74]. No relevant interactions were observable for the 8-methoxy moiety, which is in agreement with the experimental findings showing no difference between the inhibition efficacies of compounds
3e and
3f. Compound
3f and lidorestat induced different water arrangement near the cavity access. As shown in the fitted molecular surface (
Figure 4B), lidorestat induced a narrower access to the cavity, keeping amino acids Trp219 and Leu300 closer in comparison with the complex of
3f. The amino group of Gln49 (colored in green in
Figure 4B) is perpendicular to the surface of the protein in the complex with lidorestat (
Figure 4B left), while for
3f this group is parallel to the surface (
Figure 4B right) [
71].
To test selectivity, we used the comparison to the closely related aldehyde reductase (ALR1). All compounds
3a–
f were found to be less active inhibitors of ALR1 compared to ALR2 (
Table 2). The corresponding selectivity factors calculated for the most efficient compounds
3e and
3f were found to be 381 and 792, respectively. Methoxy substituent in position 8 of compound
3f thus ensured higher selectivity than the corresponding carboxyl group in compound
3e. This effect may be obviously assigned to the higher affinity to ALR1 of compound
3e bearing the carboxyl group, since the alterations of the 8-metoxy vs. 8-carboxyl substituents did not affect ALR2 inhibition. In silico study revealed a strong interaction of compound
3f with Phe122, Trp219, Leu300, and Tyr309 from the “specificity pocket”, set of residues in ALR2 which are not conserved in ALR1 [
71]. The highest selectivity was recorded for compound
3d with the corresponding selectivity factor exceeding 1000.
One of the most efficient inhibitors, compound
3f, was tested for inhibition of human recombinant AKR1B1 and AKR1B10, yielding IC
50 values 84 nM and 9434 nM, respectively, pointing to high selectivity relative to AKR1B10. Zopolrestat used as reference gave for AKR1B1 the IC
50 value of 25 nM, while for AKR1B10 it was inactive. According to the docking study, selectivity of compound
3f towards AKR1B10 is related to the fact that in Cys298 is replaced by hydroxylated cysteine Cso299, which creates H-bond with the acetate group of compound
3f and keeps it too far from NADP
+ to form the hydrophobic interaction with nicotinamide ring [
71].
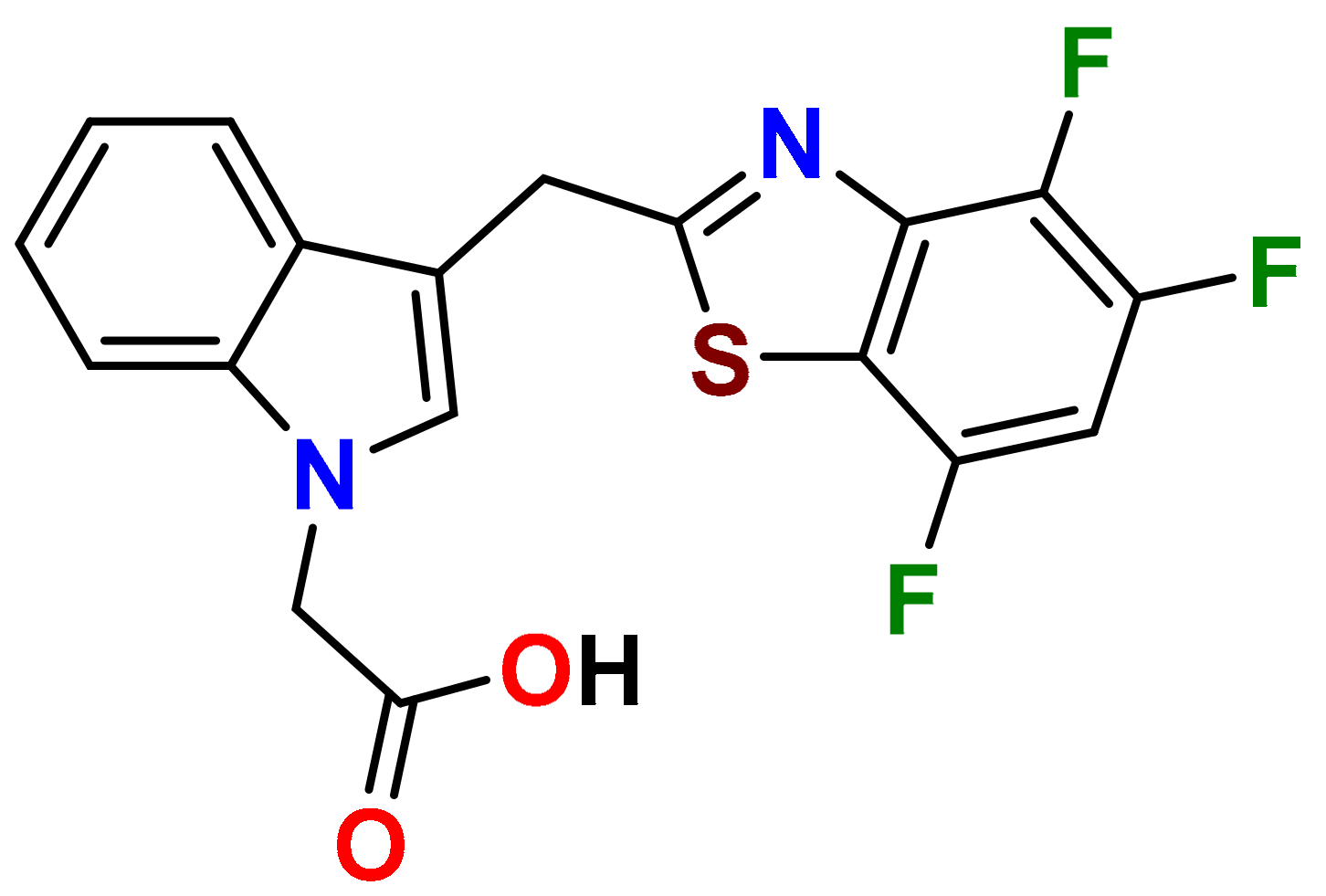
In our persistent search for highly efficient and selective ARIs, we were in the next stage of drug design inspired by lidorestat (
Figure 5), one of the most efficient inhibitors of aldose reductase with reported IC
50 value in low nanomolar region [
36]. Chemically, lidorestat is a derivative of indol-1-yl acetic acid. Despite the failure of lidorestat in clinical trials and based on encouraging highly efficient derivatives
3c–
f, we still had believed that indol-1-yl acetic acid moiety was a promising starting fragment for drug design. Indeed, in our preliminary study [
75], indol-1-yl acetic acid
4a was identified as an ARI with a mild efficacy. We decided to employ this compound as a starting structural moiety (hit) in a fragment based drug design. By applying this approach to the virtual screening of ChemSpider database, three series (
4 to
6) of indol-1-yl acetic acid derivatives, as summarized in
Table 3,
Table 4 and
Table 5, were identified as promising ARIs and subjected to experimental SAR study for AR inhibition [
76].
For comparison, two indol-1-yl propionic acid congeners (compounds 4b and 4i) were included in the experimental sample set. In addition, two structural variants lacking the acetic moiety on indolyl skeleton were incorporated (compounds 5b and 5c).
As shown in
Table 3, unsubstituted indol-1-yl acetic acid (
4a) inhibited ALR2 in low micromolar range. Insertion of a substituent at position 3 in compounds
4e–
j, resulted in reduced inhibitory power towards ALR2. Introduction of additional methyl substituent in position 2, did not affect the inhibition significantly, as shown in the case of compound
4f [
76].
On the other hand, combination of –CHO and –COCH
3 substituents in position 3 with methyl in position 2 for compounds
4k and
4l, respectively, resulted in marked increase in inhibition efficacy as documented by substantial decrease of IC
50 values (23- to 35-fold, respectively) in comparison with unsubstituted indol-1-yl acetic acid (
4a) [
71]. In compound
4m, the presence of benzyloxy group in position 5 resulted in about 10-fold improvement of inhibition when compared with the unsubstituted indol-1-yl acetic acid (
4a), based on the experimental IC
50 values [
77].
Profoundly increased inhibition was recorded for the thioxo triazine derivatives
5a and
5g, with IC
50 values in submicromolar range (
Table 4). Values of IC
50 in low micromolar range were recorded for the remaining thioxotriazine derivatives
5d–
f [76]. The strongest inhibition of ALR2 was found for
5a (cemtirestat, CMTI) with IC
50 = 97 nM. The structure–activity relationship in the series of
5 and
6 points to the necessity of a concurrent presence of both, the carboxymethyl group in position 5 and the terminal sulfur to achieve best affinity. Shifting of the carboxymethyl group from indole nitrogen to sulfur in compound
5c or its absence in compound
5b resulted in a loss of activity.
High resolution X-ray crystallography of the human recombinant AKR1B1 enzyme complexed with cemtirestat (
5a, PDB: 4QX4) demonstrated a unique pattern of cemtirestat binding, with the specificity pocket closed, contrary to the interaction of the structurally related lidorestat [
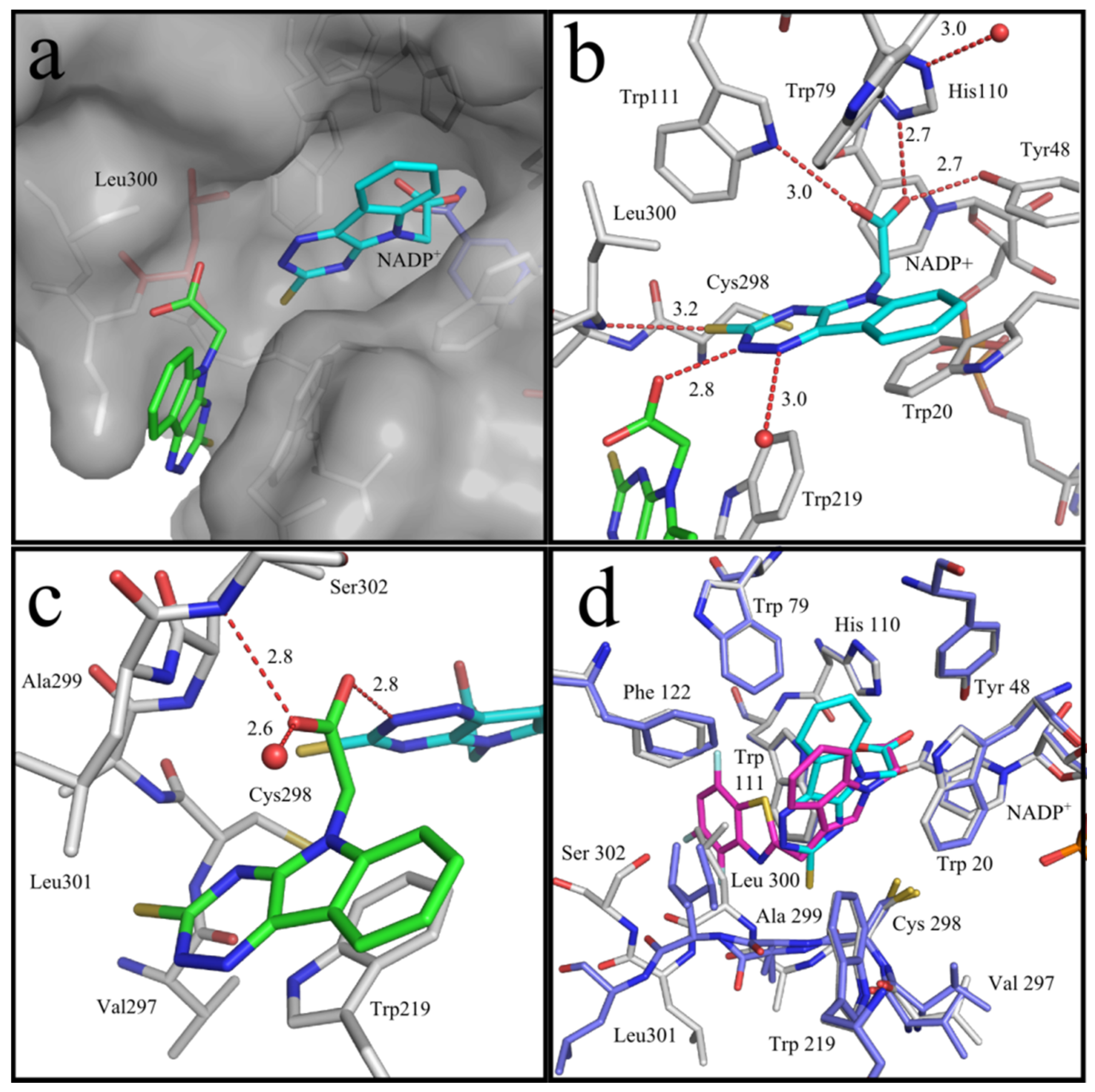
76]. As shown in greater detail in
Figure 6a, two molecules of the inhibitor
5a were visible in the determined difference electron density, one in the entrance of the binding pocket and the second one as expected inside the binding pocket. The oxygen atom of the carboxylate group of the first inhibitor molecule
5a and
N2 of the second one form an H-bond (2.8 Å,
Figure 6b). The proton is most likely contributed by the nitrogen of the triazine ring of the second molecule of compound
5a, as we assume that the carboxylate was deprotonated under the applied conditions. The other carboxylate oxygen atom of the first molecule of compound
5a is in H-bond distance to the backbone nitrogen of Ser302 (2.8 Å). Furthermore, the aromatic core skeleton makes a face-to-face π-stacking to Trp219 (3.4 Å,
Figure 6c).
Figure 6d depicts the superposition of compound
5a with lidorestat in the binding pocket of AKR1B1. In contrast to compound
5a, lidorestat opens the specificity pocket. Molecular dynamics simulations corroborated [
78] the above results of the X-ray crystallographic assay.
The other polycyclic derivatives
6a–
d revealed moderate inhibition of ALR2 characterized by IC
50 values in low micromolar range (
Table 5).
All tested compounds of the series
4 to
6 were found to be less active on ALR1 compared to ALR2. It should be noted that the most selective compound
5a, with a selectivity factor above 400, is also the most potent ARI in the series
4 to
6 (
Table 3,
Table 4 and
Table 5).
Considering excellent “lead-likeness” of cemtirestat (
5a) [
76], we proceeded with optimization of its thioxotriazinoindole scaffold by replacing sulfur with oxygen, with the aim to improve the inhibitory efficacy and selectivity. Based on preliminary molecular modeling and docking calculations, a series of 2-(3-oxo-2
H-[
1,
2,
4]triazino[5,6-
b]indol-5(3
H)-yl)acetic acid derivatives
7a–
d was proposed, synthesized and their AR inhibitory efficacy and selectivity determined [
79].
The preliminary molecular modeling and docking study on protein conformer from PDB: 4QX4 pointed to preferable fitting of the O-derivative into the inhibitor-binding site of AR. Indeed, the O-variant of cemtirestat (5a), oxotriazinoindole, compound 7a (OTI), demonstrated significantly increased inhibition efficacy characterized by IC50 values 2-3 times lower compared to compound 5a, depending on the solvent used. SAR evaluation in the series of novel OTI derivatives revealed that the presence of simple substituents at N2 position of compound 7a decreased their ALR2 inhibition efficacy. The most remarkable decrease was recorded for compound 7c with the lipophilic benzyl substituent.
All compounds evaluated were less efficient inhibitors of ALR1 compared to ALR2. With exception of compound
7b, the ALR1 IC
50 values of the OTI derivatives (
7a,
7c, and
7d) were found to be over 100 µM. For these derivatives, the percentage inhibition I(%) at 100 µM concentration of the inhibitor was determined. Estimates of the particular selectivity factors calculated for the most efficient inhibitors
7a and
7d were found to be >2381 and >1177, respectively. It is striking to observe the enormous increase of the selectivity factor in the couple cemtirestat (
5a) vs. its O isostere
7a (from 422 to >2381). Molecular docking into the binding site of ALR1 offered a feasible explanation: cation–π interaction of protonated Arg312 in ALR1 with the aromatic ring of O-derivative
7a supported by two H-bonds was found to keep this molecule out of reach of NADP
+ cofactor in contrast to more tightly attached cemtirestat
5a (Figure S52 in [
79]). In addition, a hydrophobic interaction between the methylene residue of an acetate group of compound
7a and
5a, respectively, and the nicotinamide ring of NADP
+ should also be taken into consideration [
74]. Based on the IC
50 values shown in
Table 6, compound
7a inhibits ALR2 more efficiently (IC
50 = 42 nM) than the reference inhibitor epalrestat (IC
50 = 227 nM,
Table 2).
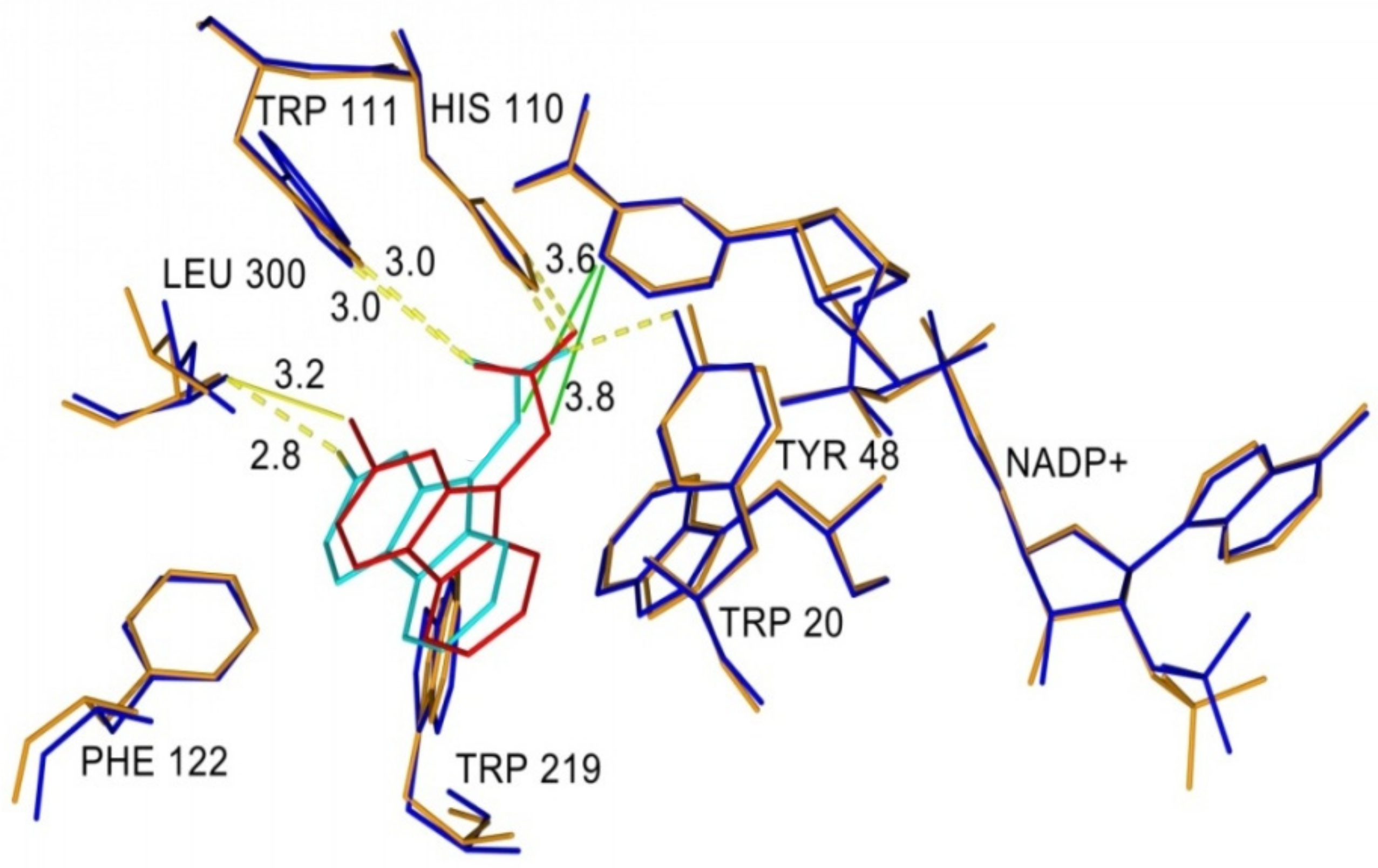
Figure 7 shows superposition of cemtirestat (
5a) and its O-analogue
7a in the inhibitor-binding site of AKR1B1. Polar carboxymethyl groups of both compounds were found to align well. Slight distortion of the fused planar aromatic system was apparently caused by higher rotational flexibility of the carboxymethyl moiety of compound
7a (in cyan) owing to less bulky oxygen compared to the sulfur of the original thioxotriazinoindole
5a (in red, van der Waals radius of O is 1.4 Å, while for S it is 1.9 Å). As a result, compound 7a was able to create stronger H-bond with Leu300 (contact distance 2.8 Å vs. 3.2 Å) and gave more favorable hydrophobic interaction (2.0 kJ/mol for compound
7a vs. 0.6 kJ/mol for cemtirestat
5a) with NADP+ (contact distance 3.6 Å vs. 3.8 Å). Compound
7a interacts with the residues of the specificity pocket Phe122 and Leu300 while letting the specificity pocket closed, which is in line with the previously published crystal structure of AKR1B1 complexed with cemtirestat (
5a, PDB: 4QX4). These in silico findings provided an explanation of better fitting of the O-derivative
7a into the binding site of AR when compared with its parent molecule of cemtirestat (
5a).
In the next stage, we focused on the development of novel oxotriazinoindole inhibitors of aldose reductase designed to fit an unoccupied ALR2 pocket shown in PDB: 4QX4 surrounded by the amino acid residues Trp219, Ala299, and Leu301. In order to utilize this pocket for additional ligand–enzyme interactions, novel
N-benzyl(oxotriazinoindole) derivatives
8a–
d have been designed and developed (
Table 7) [
80].
Molecules
8a–
d were tested for their ability to inhibit the reduction of
D,L-glyceraldehyde using ALR2 isolated from the rat eye lenses. Unsubstituted benzyl analogue
7c was used as a reference inhibitor. To assess selectivity, we used a structurally related detoxification enzyme (an antitarget), aldehyde reductase (ALR1), isolated from the rat kidneys. We found out that all substituted
N-(benzyl) derivatives
8a–
d exhibited from 2- to 6-fold better inhibitory efficacy than unsubstituted analogue
7c. In addition, they also revealed low inhibition of ALR1, which resulted in good (compounds
8b–
d) and excellent (compound
8a) enzyme selectivity (
Table 7).
The results confirmed the proposed additional interactions of substituted
N-(benzyl) derivatives
8a–
d within the interactive pocket of ALR2. The best ALR2 inhibition (IC
50 = 76 nM) and selectivity relative to ALR1 (SF = 1316) was obtained for compound
8a containing a
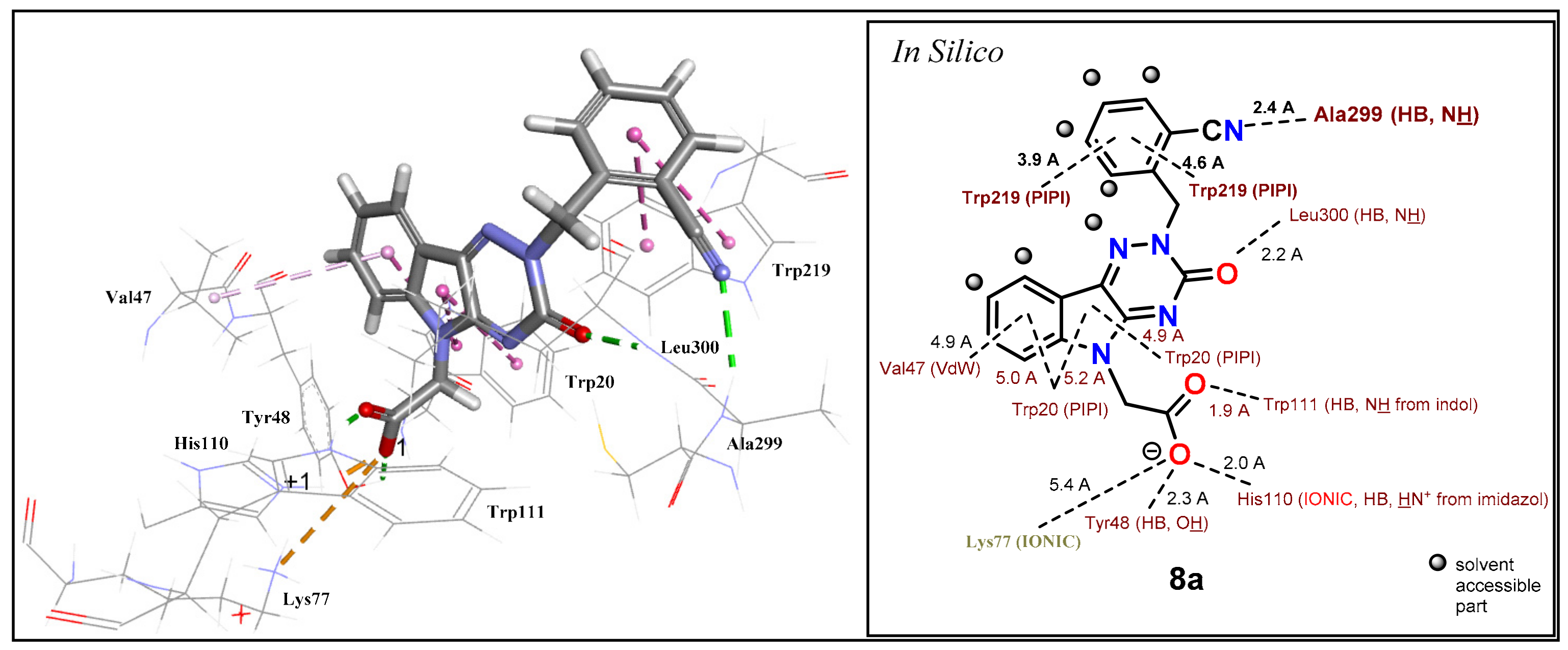
N-(2-cyanobenzyl) group. Predicted binding position of compound
8a in an active site of ALR2 shows an H-bond of a cyano group with Ala299 from backbone (2.4 Å). Moreover, the aromatic ring of a benzyl moiety forms two π–π interactions with Trp219 (3.9 and 4.6 Å) (
Figure 8).
In spite of the predicted similar positions and interactions of compounds
8d and
8a, the derivative
8d exhibited about 3-times lower inhibitory activity characterized with IC
50 = 244 nM. The additional reason for the lower activity of compound
8d could be the desolvation and the rotation penalty caused by a –CH
2OH group. Desolvation penalty effect is a subject of recent publication [
81].
Derivatives
8b,c (–CONH
2, –COOH, respectively) revealed 2- to 3-fold lower inhibitory activity in comparison to compound
8a (–CN). Besides desolvation penalty, the lower activity of a carbamoyl derivative
8b could be also caused by conformational penalty of –CONH
2 group. The symmetry of the delocalized carboxylate group in
8c (–COO
-) exhibited better inhibition activity than –CONH
2 in compound
8b and –CH
2OH in compound
8d. In addition, the outer part of the studied binding pocket and the benzyl group are well water accessible (
Figure 9) and, as a consequence, the benzyl group could provide two conformers.
The first conformer allows the formation of the predicted H-bond with an orientation of a polar group inside the pocket and the second conformer prefers an orientation towards the solvent. Consequently, very polar groups would not be oriented inside the pocket, where the predicted H-bonds could be formed, but owing to solvation, they would remain oriented towards a water environment out of the pocket. Therefore, significant H-bond with Ala299 or Leu301 would not be formed as predicted and thus the least solvated derivative 8a (–CN) revealed the highest inhibitory activity.
Table 8 summarizes the human recombinant enzyme AKR1B1 inhibition data of the most promising ARIs representing the four leading scaffolds. The inhibition efficacies and selectivities of AKR1B1 are compared with those of rat lens ALR2.
2.2. Antioxidant Activity
As mentioned above, the hexahydropyridoindole scaffold of stobadine (
1a) and its tetrahydro-congener
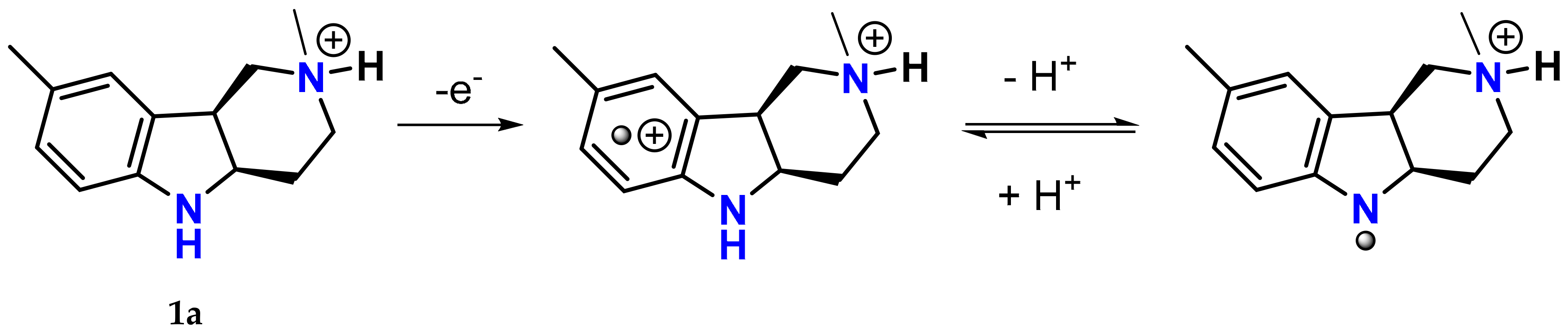
2a had been used as core structures in designing bifunctional agents combining antioxidant activity with aldose reductase inhibitory action. The antioxidant and free radical scavenging ability of stobadine stems from ability of the indole to form a resonance-stabilized nitrogen-centered radical, which is formed after one-electron removal followed by deprotonation (
Figure 10) [
82]. It has been well documented that structural alterations in the close proximity of the indole nitrogen, affecting its hydrogen donating ability, are crucial for the free radical scavenging efficiency [
83,
84].
Aromatization of the hexahydropyridoindole skeleton of stobadine (
1a) into its tetrahydro-congener
2a significantly lowered the antioxidant activity, while acetylation of the indole nitrogen in compound
1e completely abolished the ability to scavenge free radicals [
83]. The results are fully in agreement with the notion that the antioxidant activity of stobadine and the related pyridoindoles is mediated via the indole nitrogen center [
82,
84,
85].
As shown in
Table 9, in agreement with our previously published results [
83,
84], stobadine (
1a) and its carboxymethylated hexahydropyridoindole derivatives
1b and
1c rapidly reacted with DPPH, their antiradical activities being from 4 to 8 times higher than those of their tetrahydropyridoindole analogues
2a–
d in which nitrogen lone pair is involved in the indole aromatic system. In the series of hexahydropyridoindoles
1a–
c, the presence of the carboxymethyl group in position 8, decreased the DPPH scavenging ability markedly. On the other hand, analogous carboxymethyl substitution in the tetrahydro- congeners
2a–
d did not affect the antioxidant activity significantly. Notably, substitution of the methyl group in position 8 by the more electron-donating methoxy substituent yielding compound
1d, with augmented electron density on the aromatic ring, markedly increased the free radical scavenging activity compared to the parent stobadine (
1a). The initial rate of DPPH decolorization by compound
1d was comparable with that of equimolar trolox. The presence of the carbamate moiety in position 2 decreased the basicity of the nitrogen, without affecting the intrinsic chemical reactivity with free radicals [
86]. However, it had profound consequences for the bioavailability of the drug as discussed below (
Section 2.3). Acetylation of the indole nitrogen (position 5) of the parent stobadine (
1a) yielded a completely non-active derivative
1e (
Table 9). This finding is in the line with the above-mentioned mechanism of free radical scavenging action of the hexahyropyridoindoles according to which the presence of a free hydrogen on the indole nitrogen (position 5) is a prerequisite for the antiradical activity.
Introducing the carboxymethyl pharmacophore into indole nitrogen (position N5), yielded derivatives
3a–
f with appreciable aldose reductase inhibition efficacy as shown in
Table 2, yet with abolished antioxidant activity as reported in [
67,
71].
Similarly, in the series
4, indol-1-yl acetic acid (
4a) was devoid of any antioxidant activity as shown in
Table 10. On the other hand, in the case of the indol-3-yl acetic acid derivatives with unsubstituted nitrogen
4c,
4d, the antiradical activity higher than that of the structurally related reference antioxidant melatonin was recorded. The electron donor substituent -OCH
3 in the more efficient derivative
4d facilitates the delocalization of the unpaired electron of the intermediate indolyl radical originating after free radical encounter [
75].
In the series
5, the antiradical activity of the most efficient AR inhibitor cemtirestat (
5a) markedly exceeded that of equimolar melatonin (
Table 10). On placing an isopropyl group on the S atom resulting in compound
5f, thus preventing thione–thiol tautomerism and deprotonation of the –SH group, the antiradical activity of compound
5f was completely lost [
87].
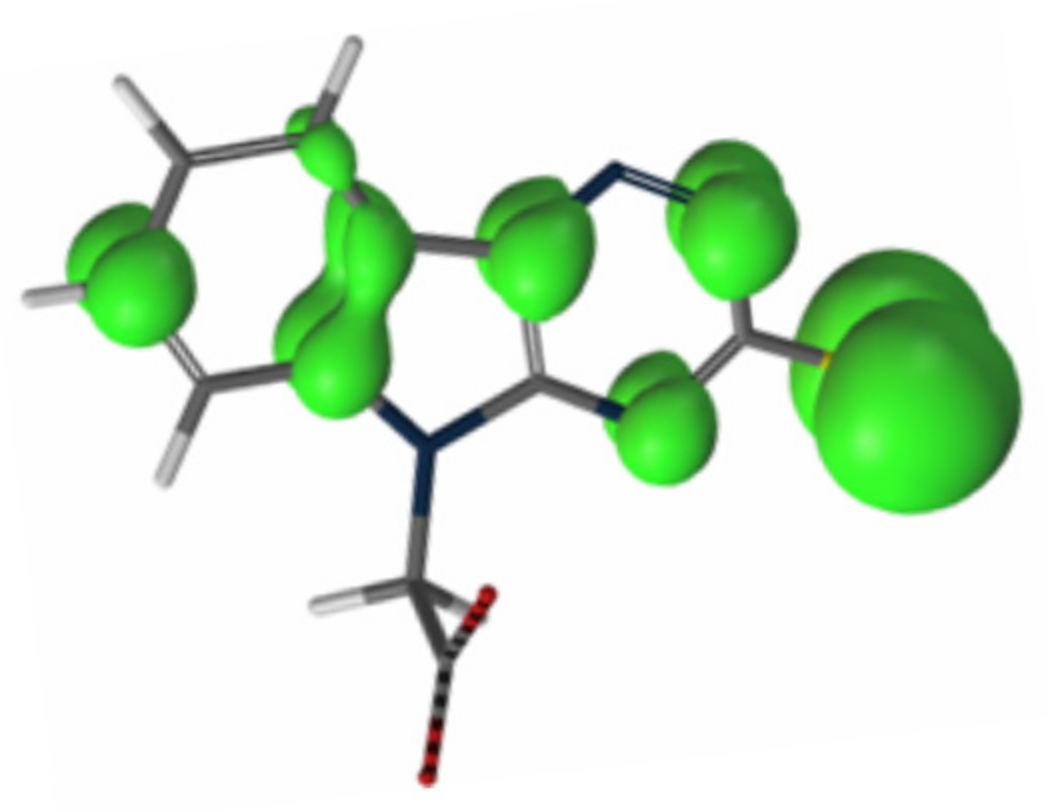
Taking into account the above findings and spin density calculations, an S-centered charge localization in one electron-oxidized radical of cemtirestat (
5a) was suggested as shown in
Figure 11 [
88]. The total stoichiometry of DPPH scavenging by cemtirestat (
5a) was determined. Under the experimental conditions used, one molecule of compound
5a was found to quench about 1.5 DPPH radicals [
87].
As shown in
Table 10, the oxygen isosteric congeners
7a and
7d of cemtirestat (
5a) affected the kinetics of DPPH decolorization only marginally, pointing to their decreased free radical scavenging ability compared to that of sulfur- containing cemtirestat (
5a). Yet the free radical scavenging efficacy of compounds
7a and
7d still exceeded that of the equimolar melatonin used as a standard.
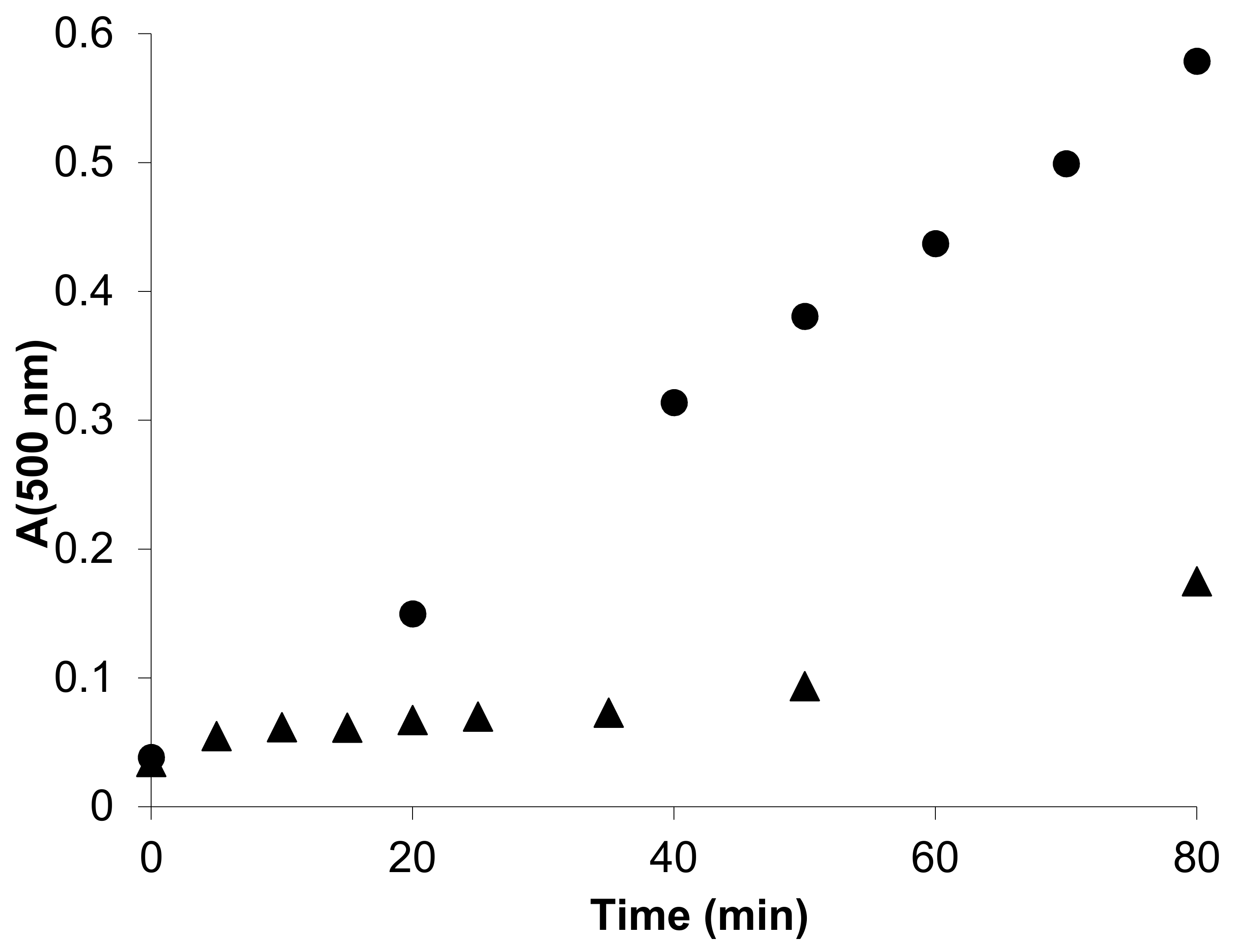
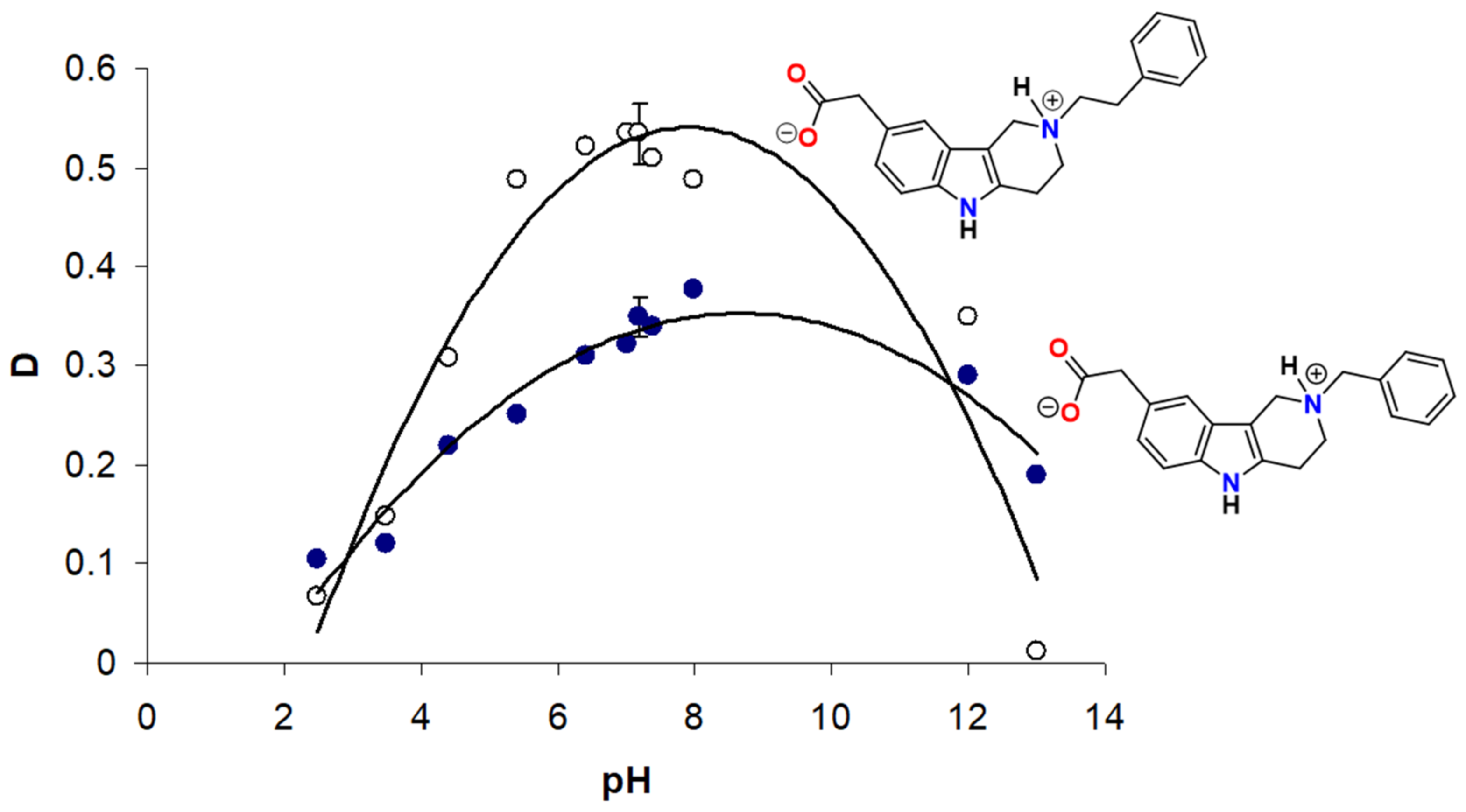
Unilamellar 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) liposomes were used as model membranes to study an overall antioxidant activity. Peroxidation of liposomes was induced by the water-soluble radical generator 2,2′-azobis(2-methylpropionamidine) dihydrochloride (AAPH) which simulates an attack by free radicals from the aqueous region. The carboxymethylated tetrahydropyridoindole
2c effectively suppressed oxidation and gave a distinct lag phase in the lipid peroxide accumulation curve (
Figure 12). As shown in
Table 11, the overall antioxidant efficiency, based on IC
50 values, of the carboxymethylated congener
2c, is significantly lower in comparison with the parent compound
2a in spite of the fact that the intrinsic antiradical activity determined in the DPPH test is similar for both compounds (
Table 9). The lipophilicity drop caused by the presence of the polar carboxymethyl group in
2c is obviously responsible for the marked decrease in the overall antioxidant efficacy in liposomes. A similar situation was recorded for the hexahydropyridoindole congeners
1a vs.
1c.
In the homogeneous model system of DPPH in ethanol, the antioxidant activity of a compound is determined by its intrinsic chemical reactivity with the radicals. In membranes, however, the apparent reactivity may be different since it is affected by the distribution ratio of the antioxidant between water and lipid compartments [
63].