COVID-19: A Global Challenge with Old History, Epidemiology and Progress So Far
Abstract
1. Introduction
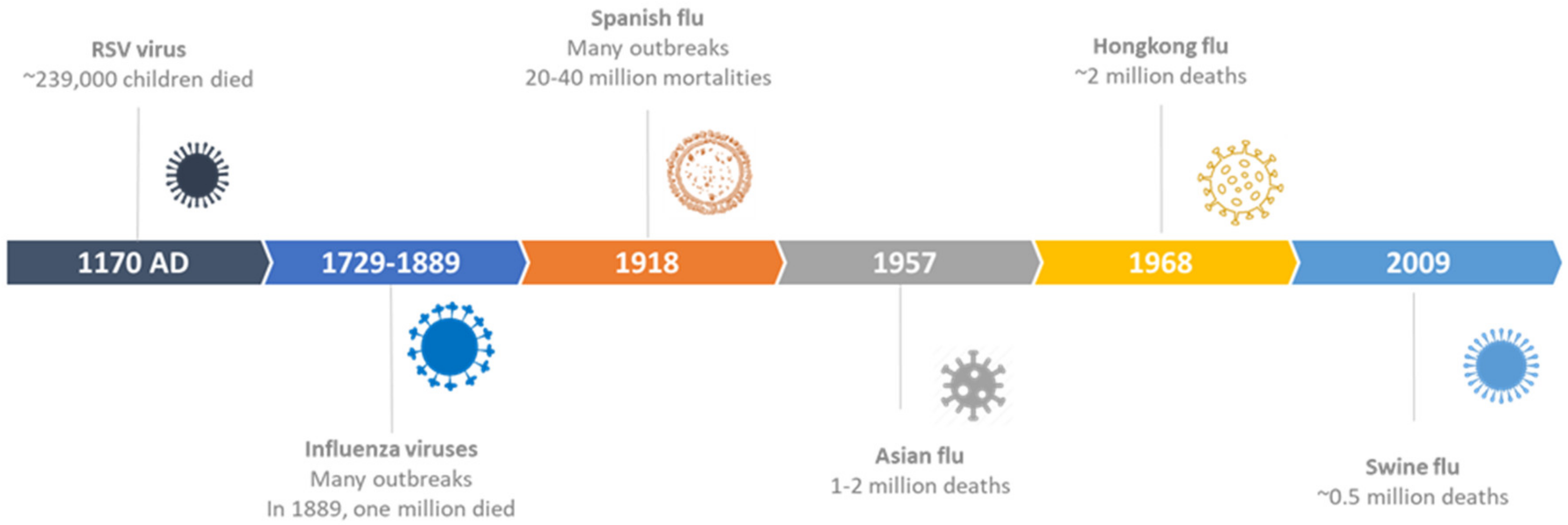
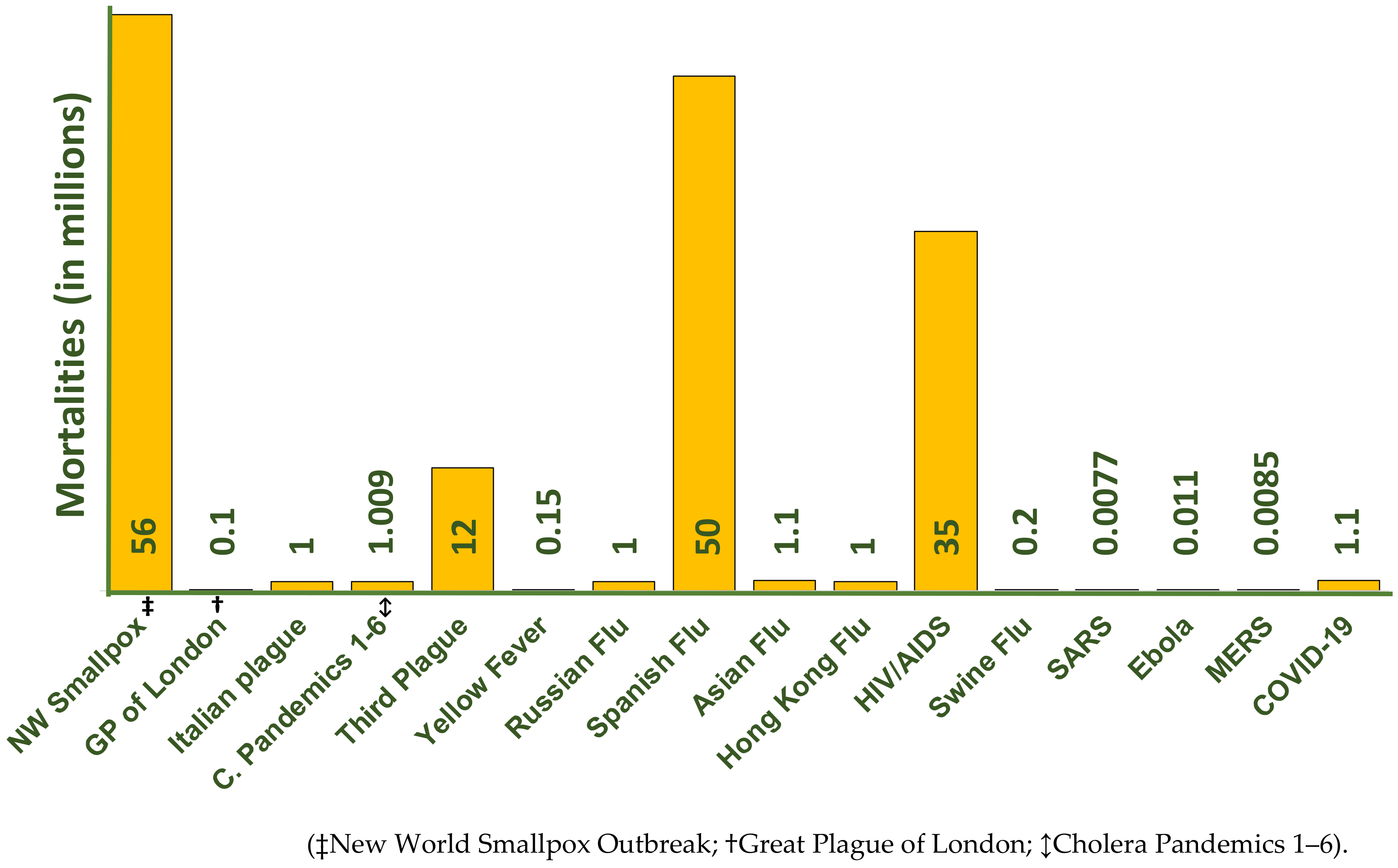
2. History of Respiratory Viruses Outbreaks
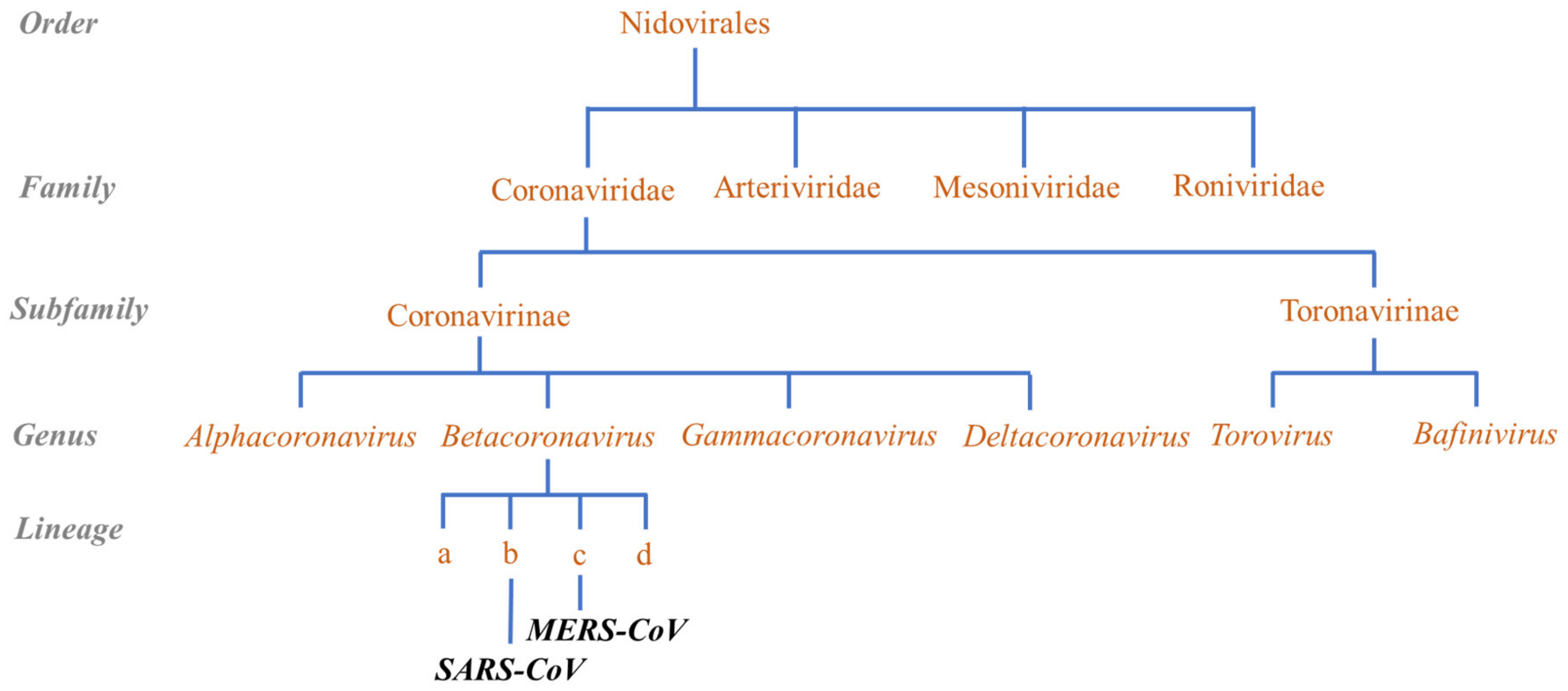
3. Coronaviruses and Earlier Outbreaks
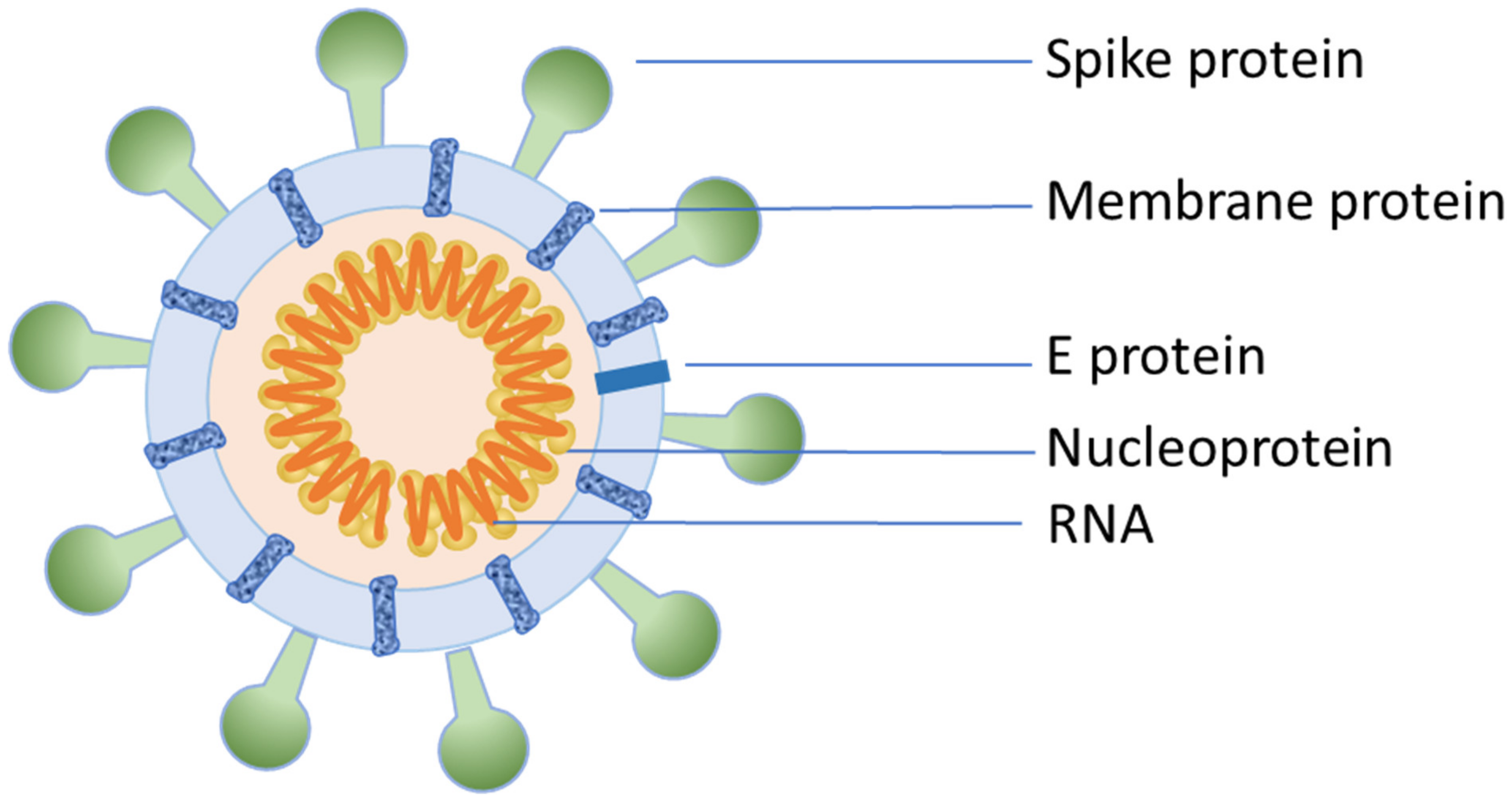
4. What Is New about COVID-19
5. Mode of Infection
6. Treatment of COVID-19 and Vaccine
7. Epidemiology of COVID-19
8. How COVID-19 Became a Pandemic
9. Some Statistics of COVID-19
10. Technical Problems in Containing the Virus
11. Were We Caught Unprepared for This Scale of Crisis?
12. Were Earlier Warnings Ignored?
13. Lessons for the Future: Proposed Strategies
Author Contributions
Funding
Conflicts of Interest
References
- Liao, Y.; Xu, B.; Wang, J.; Liu, X. A new method for assessing the risk of infectious disease outbreak. Sci. Rep. 2017, 7, 40084. [Google Scholar] [CrossRef]
- Grubaugh, N.D.; Ladner, J.T.; Lemey, P.; Pybus, O.G.; Rambaut, A.; Holmes, E.C.; Andersen, K.G. Tracking virus outbreaks in the twenty-first century. Nat. Microbiol. 2019, 4, 10–19. [Google Scholar] [CrossRef] [PubMed]
- Goenka, A.; Liu, L. Infectious diseases, human capital and economic growth. Econ. Theory 2019, 70, 1–47. [Google Scholar] [CrossRef]
- Hua, J.; Shaw, R. Corona virus (Covid-19) “infodemic” and emerging issues through a data lens: The case of china. Int. J. Environ. Res. Public Health 2020, 17, 2309. [Google Scholar] [CrossRef] [PubMed]
- McCloskey, B.; Heymann, D.L. SARS to novel coronavirus—Old lessons and new lessons. Epidemiol. Infect. 2020, 148, e22. [Google Scholar] [CrossRef]
- Nkengasong, J. China’s response to a novel coronavirus stands in stark contrast to the 2002 SARS outbreak response. Nat. Med. 2020, 26, 310–311. [Google Scholar] [CrossRef]
- Wang, C.; Horby, P.W.; Hayden, F.G.; Gao, G.F. A novel coronavirus outbreak of global health concern. Lancet 2020, 395, 470–473. [Google Scholar] [CrossRef]
- Zhou, M.; Zhang, X.; Qu, J. Coronavirus disease 2019 (COVID-19): A clinical update. Front. Med. 2020. [Google Scholar] [CrossRef]
- Jiang, S.; Xia, S.; Ying, T.; Lu, L. A novel coronavirus (2019-nCoV) causing pneumonia-associated respiratory syndrome. Cell. Mol. Immunol. 2020, 17, 554–554. [Google Scholar] [CrossRef]
- Yang, P.; Wang, X. COVID-19: A new challenge for human beings. Cell. Mol. Immunol. 2020, 17, 555–557. [Google Scholar] [CrossRef]
- Van Der Hoek, L.; Pyrc, K.; Jebbink, M.F.; Vermeulen-Oost, W.; Berkhout, R.J.; Wolthers, K.C.; Wertheim-van Dillen, P.M.; Kaandorp, J.; Spaargaren, J.; Berkhout, B. Identification of a new human coronavirus. Nat. Med. 2004, 10, 368–373. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Li, F.; Shi, Z.-L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 2019, 17, 181–192. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Novel Coronavirus ( 2019-nCoV): Situation Report, 3. 2020. Available online: https://apps.who.int/iris/bitstream/handle/10665/330762/nCoVsitrep23Jan2020-eng.pdf (accessed on 23 December 2020).
- Lupia, T.; Scabini, S.; Pinna, S.M.; Di Perri, G.; De Rosa, F.G.; Corcione, S. 2019-novel coronavirus outbreak: A new challenge. J. Glob. Antimicrob. Resist. 2020, 21, 22–27. [Google Scholar] [CrossRef] [PubMed]
- Rabi, F.A.; Al Zoubi, M.S.; Kasasbeh, G.A.; Salameh, D.M.; Al-Nasser, A.D. SARS-CoV-2 and coronavirus disease 2019: What we know so far. Pathogens 2020, 9, 231. [Google Scholar] [CrossRef] [PubMed]
- Walls, A.C.; Fiala, B.; Schäfer, A.; Wrenn, S.; Pham, M.N.; Murphy, M.; Tse, L.V.; Shehata, L.; O’Connor, M.A.; Chen, C.; et al. Elicitation of Potent Neutralizing Antibody Responses by Designed Protein Nanoparticle Vaccines for SARS-CoV-2. Cell 2020, 183, 1367–1382.e17. [Google Scholar] [CrossRef]
- Castagnoli, R.; Votto, M.; Licari, A.; Brambilla, I.; Bruno, R.; Perlini, S.; Rovida, F.; Baldanti, F.; Marseglia, G.L. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection in children and adolescents: A systematic review. JAMA Pediatrics 2020, 174, 882–889. [Google Scholar] [CrossRef] [PubMed]
- Gröndahl, B.; Puppe, W.; Hoppe, A.; Kühne, I.; Weigl, J.A.; Schmitt, H.-J. Rapid identification of nine microorganisms causing acute respiratory tract infections by single-tube multiplex reverse transcription-PCR: Feasibility study. J. Clin. Microbiol. 1999, 37, 1–7. [Google Scholar] [CrossRef]
- Bitko, V.; Musiyenko, A.; Shulyayeva, O.; Barik, S. Inhibition of respiratory viruses by nasally administered siRNA. Nat. Med. 2005, 11, 50–55. [Google Scholar] [CrossRef]
- Brundage, J.F. Interactions between influenza and bacterial respiratory pathogens: Implications for pandemic preparedness. Lancet Infect. Dis. 2006, 6, 303–312. [Google Scholar] [CrossRef]
- Bedford, J.; Enria, D.; Giesecke, J.; Heymann, D.L.; Ihekweazu, C.; Kobinger, G.; Lane, H.C.; Memish, Z.; Oh, M.-D.; Schuchat, A. COVID-19: Towards controlling of a pandemic. Lancet 2020, 395, 1015–1018. [Google Scholar] [CrossRef]
- Morens, D.M.; Folkers, G.K.; Fauci, A.S. What Is a Pandemic; The University of Chicago Press: Chicago, IL, USA, 2009. [Google Scholar]
- Blanken, M.O.; Rovers, M.M.; Molenaar, J.M.; Winkler-Seinstra, P.L.; Meijer, A.; Kimpen, J.L.; Bont, L. Respiratory syncytial virus and recurrent wheeze in healthy preterm infants. N. Engl. J. Med. 2013, 368, 1791–1799. [Google Scholar] [CrossRef] [PubMed]
- Acosta, P.L.; Caballero, M.T.; Polack, F.P. Brief history and characterization of enhanced respiratory syncytial virus disease. Clin. Vaccine Immunol. 2016, 23, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Rosner, F. Moses Maimonides’ treatise on asthma. Thorax 1981, 36, 245–251. [Google Scholar] [CrossRef] [PubMed]
- Zanone, S.M.; Krause, L.K.; Madhi, S.A.; Bassat, Q.; Jha, P.; Simões, E.A.; Wairagkar, N.; Polack, F.P. Challenges in estimating RSV-associated mortality rates. Lancet Respir. Med. 2016, 4, 345–347. [Google Scholar] [CrossRef]
- Lozano, R.; Naghavi, M.; Foreman, K.; Lim, S.; Shibuya, K.; Aboyans, V.; Abraham, J.; Adair, T.; Aggarwal, R.; Ahn, S.Y. Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: A systematic analysis for the Global Burden of Disease Study 2010. Lancet 2012, 380, 2095–2128. [Google Scholar] [CrossRef]
- Falsey, A.R.; Hennessey, P.A.; Formica, M.A.; Cox, C.; Walsh, E.E. Respiratory syncytial virus infection in elderly and high-risk adults. N. Engl. J. Med. 2005, 352, 1749–1759. [Google Scholar] [CrossRef] [PubMed]
- Agius, G.; Dindinaud, G.; Biggar, R.; Peyre, R.; Vaillant, V.; Ranger, S.; Poupet, J.; Cisse, M.; Castets, M. An epidemic of respiratory syncytial virus in elderly people: Clinical and serological findings. J. Med. Virol. 1990, 30, 117–127. [Google Scholar] [CrossRef]
- Falsey, A.R.; Treanor, J.J.; Betts, R.F.; Walsh, E.E. Viral respiratory infections in the institutionalized elderly: Clinical and epidemiologic findings. J. Am. Geriatr. Soc. 1992, 40, 115–119. [Google Scholar] [CrossRef]
- Zambon, M.C. The pathogenesis of influenza in humans. Rev. Med. Virol. 2001, 11, 227–241. [Google Scholar] [CrossRef]
- Zimmer, S.M.; Burke, D.S. Historical perspective—emergence of influenza A (H1N1) viruses. N. Engl. J. Med. 2009, 361, 279–285. [Google Scholar] [CrossRef]
- Webster, R. Influenza virus: Transmission between species and relevance to emergence of the next human pandemic. In Viral Zoonoses and Food of Animal Origin; Springer: Berlin/Heidelberg, Germany, 1997; pp. 105–113. [Google Scholar]
- Morens, D.M.; Taubenberger, J.K. Pandemic influenza: Certain uncertainties. Rev. Med. Virol. 2011, 21, 262–284. [Google Scholar] [CrossRef]
- Morens, D.M.; Taubenberger, J.K.; Fauci, A.S. The persistent legacy of the 1918 influenza virus. N. Engl. J. Med. 2009, 361, 225–229. [Google Scholar] [CrossRef]
- Kuszewski, K.; Brydak, L. The epidemiology and history of influenza. Biomed. Pharm. 2000, 54, 188–195. [Google Scholar] [CrossRef]
- Potter, C.W. Chronicle of influenza pandemics. Textb. Influenza 1998, 3–17. [Google Scholar]
- Taubenberger, J.; Morens, D. Pandemic influenza—Including a risk assessment of H5N1. Rev. Sci. Tech. 2009, 28, 187. [Google Scholar] [CrossRef] [PubMed]
- Pyle, G.F.; Patterson, K.D. Influenza diffusion in European history: Patterns and paradigms. Ecol. Dis. 1984, 2, 173–184. [Google Scholar]
- Patterson, K.D. Pandemic Influenza, 1700–1900: A Study in Historical Epidemiology; Rowman & Littlefield: Totowa, NJ, USA, 1986. [Google Scholar]
- Valleron, A.-J.; Cori, A.; Valtat, S.; Meurisse, S.; Carrat, F.; Boëlle, P.-Y. Transmissibility and geographic spread of the 1889 influenza pandemic. Proc. Natl. Acad. Sci. USA 2010, 107, 8778–8781. [Google Scholar] [CrossRef]
- Saunders-Hastings, P.R.; Krewski, D. Reviewing the history of pandemic influenza: Understanding patterns of emergence and transmission. Pathogens 2016, 5, 66. [Google Scholar] [CrossRef]
- Waring, J.I. A History of Medicine in South Carolina, 1670–1825; South Carolina Medical Association: Columbia, SC, USA, 1964; Volume 1. [Google Scholar]
- Crosby, A.W. America’s Forgotten Pandemic: The Influenza of 1918; Cambridge University Press: Cambridge, UK, 2003. [Google Scholar]
- Humphries, M.O. Paths of infection: The First World War and the origins of the 1918 influenza pandemic. War Hist. 2014, 21, 55–81. [Google Scholar] [CrossRef]
- Barry, J.M. The site of origin of the 1918 influenza pandemic and its public health implications. J. Transl. Med. 2004, 2, 1–4. [Google Scholar] [CrossRef]
- Morens, D.M.; Fauci, A.S. The 1918 influenza pandemic: Insights for the 21st century. J. Infect. Dis. 2007, 195, 1018–1028. [Google Scholar] [CrossRef] [PubMed]
- Shortridge, K.F. The 1918 ‘Spanish’flu: Pearls from swine? Nat. Med. 1999, 5, 384–385. [Google Scholar] [CrossRef] [PubMed]
- MacNeal, W.J. The influenza epidemic of 1918 in the American Expeditionary Forces in France and England. Arch. Intern. Med. 1919, 23, 657–688. [Google Scholar] [CrossRef]
- James, P. Chinese labour corps. World Chin. 2014, 4, 25–26. [Google Scholar]
- Van Epps, H.L. Influenza: Exposing the true killer. J. Exp. Med. 2006, 203, 803. [Google Scholar] [CrossRef]
- Markel, H.; Stern, A.M.; Navarro, J.A.; Michalsen, J.R.; Monto, A.S.; DiGiovanni, C., Jr. Nonpharmaceutical influenza mitigation strategies, US communities, 1918–1920 pandemic. Emerg. Infect. Dis. 2006, 12, 1961. [Google Scholar] [CrossRef]
- Fukumi, H. Summary report on the Asian influenza epidemic in Japan, 1957. Bull. WHO 1959, 20, 187. [Google Scholar]
- Dunn, F.L. Pandemic influenza in 1957: Review of international spread of new Asian strain. J. Am. Med. Assoc. 1958, 166, 1140–1148. [Google Scholar] [CrossRef]
- Henderson, D.A.; Courtney, B.; Inglesby, T.V.; Toner, E.; Nuzzo, J.B. Public health and medical responses to the 1957-58 influenza pandemic. Biosecur. Bioterrorism Biodef. Strategy Pract. Sci. 2009, 7, 265–273. [Google Scholar] [CrossRef]
- Jackson, C. History lessons: The Asian flu pandemic. Br. J. Gen. Pract. 2009, 59, 622–623. [Google Scholar] [CrossRef]
- Laver, W.; Air, G.; Dopheide, T.; Ward, C. Amino acid sequence changes in the haemagglutinin of A/Hong Kong (H3N2) influenza virus during the period 1968–1977. Nature 1980, 283, 454–457. [Google Scholar] [CrossRef] [PubMed]
- Cockburn, W.C.; Delon, P.; Ferreira, W. Origin and progress of the 1968–1969 Hong Kong influenza epidemic. Bull. WHO 1969, 41, 343. [Google Scholar]
- Norris, S. Pandemic Influenza; Library of Parliament/Bibliothèque du Parlement: Ottawa, ON, USA, 2015. [Google Scholar]
- Smith, G.J.; Vijaykrishna, D.; Bahl, J.; Lycett, S.J.; Worobey, M.; Pybus, O.G.; Ma, S.K.; Cheung, C.L.; Raghwani, J.; Bhatt, S. Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature 2009, 459, 1122–1125. [Google Scholar] [CrossRef] [PubMed]
- Gatherer, D. The 2009 H1N1 influenza outbreak in its historical context. J. Clin. Virol. 2009, 45, 174–178. [Google Scholar] [CrossRef]
- Viboud, C.; Miller, M.; Olson, D.R.; Osterholm, M.; Simonsen, L. Preliminary estimates of mortality and years of life lost associated with the 2009 A/H1N1 pandemic in the US and comparison with past influenza seasons. PLoS Curr. 2010, 2. [Google Scholar] [CrossRef]
- Dawood, F.S.; Iuliano, A.D.; Reed, C.; Meltzer, M.I.; Shay, D.K.; Cheng, P.-Y.; Bandaranayake, D.; Breiman, R.F.; Brooks, W.A.; Buchy, P. Estimated global mortality associated with the first 12 months of 2009 pandemic influenza A H1N1 virus circulation: A modelling study. Lancet Infect. Dis. 2012, 12, 687–695. [Google Scholar] [CrossRef]
- Chowell, G.; Echevarría-Zuno, S.; Viboud, C.; Simonsen, L.; Tamerius, J.; Miller, M.A.; Borja-Aburto, V.H. Characterizing the epidemiology of the 2009 influenza A/H1N1 pandemic in Mexico. PLoS Med. 2011, 8, e1000436. [Google Scholar] [CrossRef]
- Mishra, B. 2015 resurgence of influenza a (H1N1) 09: Smoldering pandemic in India? J. Glob. Infect. Dis. 2015, 7, 56. [Google Scholar] [CrossRef]
- Docea, A.O.; Tsatsakis, A.; Albulescu, D.; Cristea, O.; Zlatian, O.; Vinceti, M.; Moschos, S.A.; Tsoukalas, D.; Goumenou, M.; Drakoulis, N. A new threat from an old enemy: Re-emergence of coronavirus. Int. J. Mol. Med. 2020, 45, 1631–1643. [Google Scholar] [CrossRef]
- King, A.M.; Lefkowitz, E.; Adams, M.J.; Carstens, E.B. Virus Taxonomy: Ninth Report of the International Committee on Taxonomy of Viruses; Elsevier: Amsterdam, The Netherlands, 2011; Volume 9. [Google Scholar]
- Fehr, A.R.; Perlman, S. Coronaviruses: An overview of their replication and pathogenesis. In Coronaviruses; Springer: Berlin/Heidelberg, Germany, 2015; pp. 1–23. [Google Scholar]
- Van Doremalen, N.; Bushmaker, T.; Morris, D.H.; Holbrook, M.G.; Gamble, A.; Williamson, B.N.; Tamin, A.; Harcourt, J.L.; Thornburg, N.J.; Gerber, S.I. Aerosol and surface stability of SARS-CoV-2 as compared with SARS-CoV-1. N. Engl. J. Med. 2020, 382, 1564–1567. [Google Scholar] [CrossRef]
- Geller, C.; Varbanov, M.; Duval, R.E. Human coronaviruses: Insights into environmental resistance and its influence on the development of new antiseptic strategies. Viruses 2012, 4, 3044–3068. [Google Scholar] [CrossRef] [PubMed]
- Anthony, S.J.; Johnson, C.K.; Greig, D.J.; Kramer, S.; Che, X.; Wells, H.; Hicks, A.L.; Joly, D.O.; Wolfe, N.D.; Daszak, P. Global patterns in coronavirus diversity. Virus Evol. 2017, 3, 1–15. [Google Scholar] [CrossRef] [PubMed]
- McIntosh, K.; Becker, W.B.; Chanock, R.M. Growth in suckling-mouse brain of “IBV-like” viruses from patients with upper respiratory tract disease. Proc. Natl. Acad. Sci. USA 1967, 58, 2268. [Google Scholar] [CrossRef] [PubMed]
- Kaye, H.; Marsh, H.; Dowdle, W. Seroepidemiologic survey of coronavirus (strain OC 43) related infections in a children’s population. Am. J. Epidemiol. 1971, 94, 43–49. [Google Scholar] [CrossRef] [PubMed]
- De Wit, E.; Van Doremalen, N.; Falzarano, D.; Munster, V.J. SARS and MERS: Recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 2016, 14, 523. [Google Scholar] [CrossRef]
- Drexler, J.F.; Corman, V.M.; Drosten, C. Ecology, evolution and classification of bat coronaviruses in the aftermath of SARS. Antivir. Res. 2014, 101, 45–56. [Google Scholar] [CrossRef]
- Petrosillo, N.; Viceconte, G.; Ergonul, O.; Ippolito, G.; Petersen, E. COVID-19, SARS and MERS: Are they closely related? Clin. Microbiol. Infect. 2020, 26, 729–734. [Google Scholar] [CrossRef]
- Zhong, N.; Zheng, B.; Li, Y.; Poon, L.; Xie, Z.; Chan, K.; Li, P.; Tan, S.; Chang, Q.; Xie, J. Epidemiology and cause of severe acute respiratory syndrome (SARS) in Guangdong, People’s Republic of China, in February, 2003. Lancet 2003, 362, 1353–1358. [Google Scholar] [CrossRef]
- Guan, Y.; Peiris, J.; Zheng, B.; Poon, L.; Chan, K.; Zeng, F.; Chan, C.; Chan, M.; Chen, J.; Chow, K. Molecular epidemiology of the novel coronavirus that causes severe acute respiratory syndrome. Lancet 2004, 363, 99–104. [Google Scholar] [CrossRef]
- Drosten, C.; Günther, S.; Preiser, W.; Van Der Werf, S.; Brodt, H.-R.; Becker, S.; Rabenau, H.; Panning, M.; Kolesnikova, L.; Fouchier, R.A. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 2003, 348, 1967–1976. [Google Scholar] [CrossRef]
- Huang, Y. The SARS Epidemic and Its Aftermath in China: A Political Perspective; Learning from SARS: Preparing for the Next Disease Outbreak: Workshop Summary; National Academies Press: Washington, DC, USA, 2004; pp. 116–136. [Google Scholar]
- Anderson, R.M.; Fraser, C.; Ghani, A.C.; Donnelly, C.A.; Riley, S.; Ferguson, N.M.; Leung, G.M.; Lam, T.H.; Hedley, A.J. Epidemiology, transmission dynamics and control of SARS: The 2002–2003 epidemic. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2004, 359, 1091–1105. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Epidemic and Pandemic Alert Response: Summary of probably SARS cases with onset of illness from 1 November 2002 to 31 July 2003. Based Data 2003, 31. [Google Scholar]
- Wang, M.; Yan, M.; Xu, H.; Liang, W.; Kan, B.; Zheng, B.; Chen, H.; Zheng, H.; Xu, Y.; Zhang, E. SARS-CoV infection in a restaurant from palm civet. Emerg. Infect. Dis. 2005, 11, 1860. [Google Scholar] [CrossRef] [PubMed]
- Chang, H.-J.; Huang, N.; Lee, C.-H.; Hsu, Y.-J.; Hsieh, C.-J.; Chou, Y.-J. The impact of the SARS epidemic on the utilization of medical services: SARS and the fear of SARS. Am. J. Public Health 2004, 94, 562–564. [Google Scholar] [CrossRef]
- Zaki, A.M.; Van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.; Fouchier, R.A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef]
- Hijawi, B.; Abdallat, M.; Sayaydeh, A.; Alqasrawi, S.; Haddadin, A.; Jaarour, N.; El Sheikh, S.; Alsanouri, T. Novel coronavirus infections in Jordan, April 2012: Epidemiological findings from a retrospective investigation. EMHJ-East. Mediterr. Health J. 2013, 19, S12–S18. [Google Scholar] [CrossRef]
- Ki, M. 2015 MERS outbreak in Korea: Hospital-to-hospital transmission. Epidemiol. Health 2015, 37, e2015033. [Google Scholar] [CrossRef]
- Whitelaw, S.; Mamas, M.; Topol, E.; Van Spall, H. Applications of digital technology in COVID-19 pandemic planning and response. Lancet Digit. Health 2020, 2, e435–e440. [Google Scholar] [CrossRef]
- Reeves, T.; Samy, A.M.; Peterson, A.T. MERS-CoV geography and ecology in the Middle East: Analyses of reported camel exposures and a preliminary risk map. BMC Res. Notes 2015, 8, 801. [Google Scholar] [CrossRef]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- Yao, T.T.; Qian, J.D.; Zhu, W.Y.; Wang, Y.; Wang, G.Q. A systematic review of lopinavir therapy for SARS coronavirus and MERS coronavirus—A possible reference for coronavirus disease-19 treatment option. J. Med. Virol. 2020, 92, 556–563. [Google Scholar] [CrossRef] [PubMed]
- Yao, H.; Song, Y.; Chen, Y.; Wu, N.; Xu, J.; Sun, C.; Zhang, J.; Weng, T.; Zhang, Z.; Wu, Z.; et al. Molecular Architecture of the SARS-CoV-2 Virus. Cell 2020, 183, 730–738.e13. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-T.; Landeras-Bueno, S.; Hsieh, L.-E.; Terada, Y.; Kim, K.; Ley, K.; Shresta, S.; Saphire, E.O.; Regla-Nava, J.A. Spiking Pandemic Potential: Structural and Immunological Aspects of SARS-CoV-2. Trends Microbiol. 2020, 28, 605–618. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef]
- Wang, H.; Yang, P.; Liu, K.; Guo, F.; Zhang, Y.; Zhang, G.; Jiang, C. SARS coronavirus entry into host cells through a novel clathrin-and caveolae-independent endocytic pathway. Cell Res. 2008, 18, 290–301. [Google Scholar] [CrossRef] [PubMed]
- Fani, M.; Teimoori, A.; Ghafari, S. Comparison of the COVID-2019 (SARS-CoV-2) pathogenesis with SARS-CoV and MERS-CoV infections. Future Virol. 2020. [Google Scholar] [CrossRef]
- Wu, Y.; Wang, F.; Shen, C.; Peng, W.; Li, D.; Zhao, C.; Li, Z.; Li, S.; Bi, Y.; Yang, Y.; et al. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science 2020, 368, 1274–1278. [Google Scholar] [CrossRef]
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef]
- Xu, Z.; Shi, L.; Wang, Y.; Zhang, J.; Huang, L.; Zhang, C.; Liu, S.; Zhao, P.; Liu, H.; Zhu, L. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 2020, 8, 420–422. [Google Scholar] [CrossRef]
- Weiss, S.R.; Navas-Martin, S. Coronavirus pathogenesis and the emerging pathogen severe acute respiratory syndrome coronavirus. Microbiol. Mol. Biol. Rev. 2005, 69, 635–664. [Google Scholar] [CrossRef]
- Mahmoud, I.S.; Jarrar, Y.B.; Alshaer, W.; Ismail, S. SARS-CoV-2 entry in host cells-multiple targets for treatment and prevention. Biochimie 2020, 175, 93–98. [Google Scholar] [CrossRef]
- Seah, I.; Agrawal, R. Can the coronavirus disease 2019 (COVID-19) affect the eyes? A review of coronaviruses and ocular implications in humans and animals. Ocul. Immunol. Inflamm. 2020, 28, 391–395. [Google Scholar] [CrossRef] [PubMed]
- Paules, C.I.; Marston, H.D.; Fauci, A.S. Coronavirus infections—More than just the common cold. JAMA 2020, 323, 707–708. [Google Scholar] [CrossRef] [PubMed]
- Shaffer, L. 15 drugs being tested to treat COVID-19 and how they would work. Nat. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; De Clercq, E. Therapeutic Options for the 2019 Novel Coronavirus (2019-nCoV); Nature Publishing Group: Berlin, Germany, 2020. [Google Scholar]
- Sheahan, T.P.; Sims, A.C.; Graham, R.L.; Menachery, V.D.; Gralinski, L.E.; Case, J.B.; Leist, S.R.; Pyrc, K.; Feng, J.Y.; Trantcheva, I. Broad-spectrum antiviral GS-5734 inhibits both epidemic and zoonotic coronaviruses. Sci. Transl. Med. 2017, 9, 396. [Google Scholar] [CrossRef]
- Agostini, M.L.; Andres, E.L.; Sims, A.C.; Graham, R.L.; Sheahan, T.P.; Lu, X.; Smith, E.C.; Case, J.B.; Feng, J.Y.; Jordan, R. Coronavirus susceptibility to the antiviral remdesivir (GS-5734) is mediated by the viral polymerase and the proofreading exoribonuclease. MBio 2018, 9, e00221-18. [Google Scholar] [CrossRef] [PubMed]
- Omrani, A.S.; Saad, M.M.; Baig, K.; Bahloul, A.; Abdul-Matin, M.; Alaidaroos, A.Y.; Almakhlafi, G.A.; Albarrak, M.M.; Memish, Z.A.; Albarrak, A.M. Ribavirin and interferon alfa-2a for severe Middle East respiratory syndrome coronavirus infection: A retrospective cohort study. Lancet Infect. Dis. 2014, 14, 1090–1095. [Google Scholar] [CrossRef]
- Khalili, J.S.; Zhu, H.; Mak, N.S.A.; Yan, Y.; Zhu, Y. Novel coronavirus treatment with ribavirin: Groundwork for an evaluation concerning COVID-19. J. Med. Virol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Biot, C.; Daher, W.; Chavain, N.; Fandeur, T.; Khalife, J.; Dive, D.; De Clercq, E. Design and synthesis of hydroxyferroquine derivatives with antimalarial and antiviral activities. J. Med. Chem. 2006, 49, 2845–2849. [Google Scholar] [CrossRef]
- Vincent, M.J.; Bergeron, E.; Benjannet, S.; Erickson, B.R.; Rollin, P.E.; Ksiazek, T.G.; Seidah, N.G.; Nichol, S.T. Chloroquine is a potent inhibitor of SARS coronavirus infection and spread. Virol. J. 2005, 2, 1–10. [Google Scholar] [CrossRef]
- Savarino, A.; Boelaert, J.R.; Cassone, A.; Majori, G.; Cauda, R. Effects of chloroquine on viral infections: An old drug against today’s diseases. Lancet Infect. Dis. 2003, 3, 722–727. [Google Scholar] [CrossRef]
- Geleris, J.; Sun, Y.; Platt, J.; Zucker, J.; Baldwin, M.; Hripcsak, G.; Labella, A.; Manson, D.K.; Kubin, C.; Barr, R.G. Observational study of hydroxychloroquine in hospitalized patients with Covid-19. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Boulware, D.R.; Pullen, M.F.; Bangdiwala, A.S.; Pastick, K.A.; Lofgren, S.M.; Okafor, E.C.; Skipper, C.P.; Nascene, A.A.; Nicol, M.R.; Abassi, M. A randomized trial of hydroxychloroquine as postexposure prophylaxis for Covid-19. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Zha, L.; Li, S.; Pan, L.; Tefsen, B.; Li, Y.; French, N.; Chen, L.; Yang, G.; Villanueva, E.V. Corticosteroid treatment of patients with coronavirus disease 2019 (COVID-19). Med. J. Aust. 2020, 212, 416–420. [Google Scholar] [CrossRef] [PubMed]
- Viner, R.M.; Russell, S.J.; Croker, H.; Packer, J.; Ward, J.; Stansfield, C.; Mytton, O.; Bonell, C.; Booy, R. School closure and management practices during coronavirus outbreaks including COVID-19: A rapid systematic review. Lancet Child Adolesc. Health 2020, 4, 397–404. [Google Scholar] [CrossRef]
- Cohen, J. The Race Is on for Antibodies That Stop the New Coronavirus; American Association for the Advancement of Science: Washington, DC, USA, 2020. [Google Scholar]
- Pang, J.; Wang, M.X.; Ang, I.Y.H.; Tan, S.H.X.; Lewis, R.F.; Chen, J.I.-P.; Gutierrez, R.A.; Gwee, S.X.W.; Chua, P.E.Y.; Yang, Q. Potential rapid diagnostics, vaccine and therapeutics for 2019 novel coronavirus (2019-nCoV): A systematic review. J. Clin. Med. 2020, 9, 623. [Google Scholar] [CrossRef]
- Lurie, N.; Saville, M.; Hatchett, R.; Halton, J. Developing Covid-19 vaccines at pandemic speed. N. Engl. J. Med. 2020, 382, 1969–1973. [Google Scholar] [CrossRef]
- Su, S.; Wong, G.; Shi, W.; Liu, J.; Lai, A.C.; Zhou, J.; Liu, W.; Bi, Y.; Gao, G.F. Epidemiology, genetic recombination, and pathogenesis of coronaviruses. Trends Microbiol. 2016, 24, 490–502. [Google Scholar] [CrossRef]
- Xu, J.; Zhao, S.; Teng, T.; Abdalla, A.E.; Zhu, W.; Xie, L.; Wang, Y.; Guo, X. Systematic comparison of two animal-to-human transmitted human coronaviruses: SARS-CoV-2 and SARS-CoV. Viruses 2020, 12, 244. [Google Scholar] [CrossRef]
- Guan, W.-J.; Ni, Z.-Y.; Hu, Y.; Liang, W.-H.; Ou, C.-Q.; He, J.-X.; Liu, L.; Shan, H.; Lei, C.-L.; Hui, D.S. Clinical characteristics of coronavirus disease 2019 in China. N. Engl. J. Med. 2020, 382, 1708–1720. [Google Scholar] [CrossRef]
- Shereen, M.A.; Khan, S.; Kazmi, A.; Bashir, N.; Siddique, R. COVID-19 infection: Origin, transmission, and characteristics of human coronaviruses. J. Adv. Res. 2020, 24, 91–98. [Google Scholar] [CrossRef]
- Chan, J.F.-W.; Yuan, S.; Kok, K.-H.; To, K.K.-W.; Chu, H.; Yang, J.; Xing, F.; Liu, J.; Yip, C.C.-Y.; Poon, R.W.-S. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: A study of a family cluster. Lancet 2020, 395, 514–523. [Google Scholar] [CrossRef]
- Liu, J.; Liao, X.; Qian, S.; Yuan, J.; Wang, F.; Liu, Y.; Wang, Z.; Wang, F.-S.; Liu, L.; Zhang, Z. Community Transmission of Severe Acute Respiratory Syndrome Coronavirus 2, Shenzhen, China. 2020. Available online: https://stacks.cdc.gov/view/cdc/88896/cdc_88896_DS1.pdf (accessed on 23 December 2020).
- World Health Organization. Infection Prevention and Control of Epidemic-and Pandemic-Prone Acute Respiratory Infections in Health Care; World Health Organization: Geneva, Switzerland, 2014. [Google Scholar]
- World Health Organization. Modes of Transmission of Virus Causing COVID-19: Implications for IPC Precaution Recommendations: Scientific Brief, 27 March 2020; World Health Organization: Geneva, Switzerland, 2020. [Google Scholar]
- Peng, Y.; Zhou, Y.H. Is novel coronavirus disease (COVID-19) transmitted through conjunctiva? J. Med. Virol. 2020, 92, 1408–1409. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.W.X.; Tan, Y.K.; Chia, P.Y.; Lee, T.H.; Ng, O.T.; Wong, M.S.Y.; Marimuthu, K. Air, surface environmental, and personal protective equipment contamination by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) from a symptomatic patient. JAMA 2020, 323, 1610–1612. [Google Scholar] [CrossRef] [PubMed]
- Zheng, J. SARS-CoV-2: An emerging coronavirus that causes a global threat. Int. J. Biol. Sci. 2020, 16, 1678. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.T.; Leung, K.; Bushman, M.; Kishore, N.; Niehus, R.; de Salazar, P.M.; Cowling, B.J.; Lipsitch, M.; Leung, G.M. Estimating clinical severity of COVID-19 from the transmission dynamics in Wuhan, China. Nat. Med. 2020, 26, 506–510. [Google Scholar] [CrossRef] [PubMed]
- Whitworth, J. COVID-19: A fast evolving pandemic. Trans. R. Soc. Trop. Med. Hyg. 2020, 114, 241. [Google Scholar] [CrossRef]
- Worldometer, C.C. Worldometer. 2020. Available online: https://www.worldometers.info/ (accessed on 23 December 2020).
- Roser, M.; Ritchie, H.; Ortiz-Ospina, E.; Hasell, J. Coronavirus Disease (COVID-19)–Statistics and Research. Our World Data. 2020. Available online: https://www.sipotra.it/wp-content/uploads/2020/03/Coronavirus-Disease-COVID-19-%E2%80%93-Statistics-and-Research.pdf (accessed on 23 December 2020).
- Munster, V.J.; Koopmans, M.; van Doremalen, N.; van Riel, D.; de Wit, E. A novel coronavirus emerging in China—key questions for impact assessment. N. Engl. J. Med. 2020, 382, 692–694. [Google Scholar] [CrossRef]
- Chang, D.; Lin, M.; Wei, L.; Xie, L.; Zhu, G.; Cruz, C.S.D.; Sharma, L. Epidemiologic and clinical characteristics of novel coronavirus infections involving 13 patients outside Wuhan, China. JAMA 2020, 323, 1092–1093. [Google Scholar] [CrossRef]
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus—Infected pneumonia in Wuhan, China. JAMA 2020, 323, 1061–1069. [Google Scholar] [CrossRef]
- Zhao, S.; Lin, Q.; Ran, J.; Musa, S.S.; Yang, G.; Wang, W.; Lou, Y.; Gao, D.; Yang, L.; He, D. Preliminary estimation of the basic reproduction number of novel coronavirus (2019-nCoV) in China, from 2019 to 2020: A data-driven analysis in the early phase of the outbreak. Int. J. Infect. Dis. 2020, 92, 214–217. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Gayle, A.A.; Wilder-Smith, A.; Rocklöv, J. The reproductive number of COVID-19 is higher compared to SARS coronavirus. J. Travel Med. 2020, 27, taaa021. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.T.; Leung, K.; Leung, G.M. Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: A modelling study. Lancet 2020, 395, 689–697. [Google Scholar] [CrossRef]
- Imai, N.; Cori, A.; Dorigatti, I.; Baguelin, M.; Donnelly, C.A.; Riley, S.; Ferguson, N.M. Report 3: Transmissibility of 2019-nCoV. 2020. Available online: https://fpmag.net/wp-content/uploads/2020/01/Imperial-2019-nCoV-transmissibility.pdf (accessed on 23 December 2020).
- Backer, J.A.; Klinkenberg, D.; Wallinga, J. Incubation period of 2019 novel coronavirus (2019-nCoV) infections among travellers from Wuhan, China, 20–28 January 2020. Eurosurveillance 2020, 25, 2000062. [Google Scholar] [CrossRef] [PubMed]
- Lai, C.-C.; Shih, T.-P.; Ko, W.-C.; Tang, H.-J.; Hsueh, P.-R. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and corona virus disease-2019 (COVID-19): The epidemic and the challenges. Int. J. Antimicrob. Agents 2020, 2020, 105924. [Google Scholar] [CrossRef] [PubMed]
- Desai, A.N.; Aronoff, D.M. Masks and coronavirus disease 2019 (covid-19). JAMA 2020. [Google Scholar] [CrossRef] [PubMed]
- Bai, Y.; Yao, L.; Wei, T.; Tian, F.; Jin, D.-Y.; Chen, L.; Wang, M. Presumed asymptomatic carrier transmission of COVID-19. JAMA 2020, 323, 1406–1407. [Google Scholar] [CrossRef]
- Feng, S.; Shen, C.; Xia, N.; Song, W.; Fan, M.; Cowling, B.J. Rational use of face masks in the COVID-19 pandemic. Lancet Respir. Med. 2020, 8, 434–436. [Google Scholar] [CrossRef]
- World Health Organization. Advice on the Use of Masks in the Context of COVID-19: Interim Guidance, 5 June 2020; World Health Organization: Geneva, Switzerland, 2020. [Google Scholar]
- Lyu, W.; Wehby, G.L. Community Use of Face Masks And COVID-19: Evidence From A Natural Experiment Of State Mandates In The US. Health Aff. 2020, 39, 1419–1425. [Google Scholar] [CrossRef]
- Greenhalgh, T.; Schmid, M.B.; Czypionka, T.; Bassler, D.; Gruer, L. Face masks for the public during the covid-19 crisis. BMJ 2020, 369, m1435. [Google Scholar] [CrossRef]
- Federal Ministry of Health Group. Daily Updates on the Coronavirus: Is Wearing a Surgical Mask, as Protection Against Acute Respiratory Infections, Useful for Members of the General Public; Federal Ministry of Health Group: Bonn, Germany, 2020.
- Tindale, L.; Coombe, M.; Stockdale, J.E.; Garlock, E.; Lau, W.Y.V.; Saraswat, M.; Lee, Y.-H.B.; Zhang, L.; Chen, D.; Wallinga, J. Transmission interval estimates suggest pre-symptomatic spread of COVID-19. MedRxiv 2020. [Google Scholar] [CrossRef]
- Wang, Y.; Tian, H.; Zhang, L.; Zhang, M.; Guo, D.; Wu, W.; Zhang, X.; Kan, G.L.; Jia, L.; Huo, D. Reduction of secondary transmission of SARS-CoV-2 in households by face mask use, disinfection and social distancing: A cohort study in Beijing, China. BMJ Glob. Health 2020, 5, e002794. [Google Scholar] [CrossRef] [PubMed]
- Geldsetzer, P. Use of rapid online surveys to assess People’s perceptions during infectious disease outbreaks: A cross-sectional survey on COVID-19. J. Med. Internet Res. 2020, 22, e18790. [Google Scholar] [CrossRef] [PubMed]
- Qian, X.; Ren, R.; Wang, Y.; Guo, Y.; Fang, J.; Wu, Z.-D.; Liu, P.-L.; Han, T.-R. Fighting against the common enemy of COVID-19: A practice of building a community with a shared future for mankind. Infect. Dis. Poverty 2020, 9, 1–6. [Google Scholar] [CrossRef]
- World Health Organization. Social Stigma Associated with COVID-19. A Guide to Preventing and Addressing Social Stigma; World Health Organization: Geneva, Switzerland, 2020. [Google Scholar]
- Person, B.; Sy, F.; Holton, K.; Govert, B.; Liang, A. Fear and stigma: The epidemic within the SARS outbreak. Emerg. Infect. Dis. 2004, 10, 358. [Google Scholar] [CrossRef]
- Bruns, D.P.; Kraguljac, N.V.; Bruns, T.R. COVID-19: Facts, cultural considerations, and risk of stigmatization. J. Transcult. Nurs. 2020, 31, 326. [Google Scholar] [CrossRef]
- McCauley, M.; Minsky, S.; Viswanath, K. The H1N1 pandemic: Media frames, stigmatization and coping. BMC Public Health 2013, 13, 1116. [Google Scholar] [CrossRef]
- Shultz, J.M.; Cooper, J.L.; Baingana, F.; Oquendo, M.A.; Espinel, Z.; Althouse, B.M.; Marcelin, L.H.; Towers, S.; Espinola, M.; McCoy, C.B. The role of fear-related behaviors in the 2013–2016 West Africa Ebola virus disease outbreak. Curr. Psychiatry Rep. 2016, 18, 104. [Google Scholar] [CrossRef]
- Brooks, S.K.; Webster, R.K.; Smith, L.E.; Woodland, L.; Wessely, S.; Greenberg, N.; Rubin, G.J. The psychological impact of quarantine and how to reduce it: Rapid review of the evidence. Lancet 2020, 395, 912–920. [Google Scholar] [CrossRef]
- Belshe, R.B. The origins of pandemic influenza—Lessons from the 1918 virus. N. Engl. J. Med. 2005, 353, 2209–2211. [Google Scholar] [CrossRef]
- Brownstein, J.S.; Freifeld, C.C.; Madoff, L.C. Digital disease detection—Harnessing the Web for public health surveillance. N. Engl. J. Med. 2009, 360, 2153. [Google Scholar] [CrossRef] [PubMed]
- Jiang, F.; Jiang, Y.; Zhi, H.; Dong, Y.; Li, H.; Ma, S.; Wang, Y.; Dong, Q.; Shen, H.; Wang, Y. Artificial intelligence in healthcare: Past, present and future. Stroke Vasc. Neurol. 2017, 2, 230–243. [Google Scholar] [CrossRef]
- Snacken, R.; Kendal, A.P.; Haaheim, L.R.; Wood, J.M. The next influenza pandemic: Lessons from Hong Kong, 1997. Emerg. Infect. Dis. 1999, 5, 195. [Google Scholar] [CrossRef]
- Fineberg, H.V. Pandemic preparedness and response—Lessons from the H1N1 influenza of 2009. N. Engl. J. Med. 2014, 370, 1335–1342. [Google Scholar] [CrossRef] [PubMed]
- Poland, G.A. Another coronavirus, another epidemic, another warning. Vaccine 2020, 38, 5. [Google Scholar] [CrossRef] [PubMed]
- Perico, L.; Benigni, A.; Casiraghi, F.; Ng, L.F.P.; Renia, L.; Remuzzi, G. Immunity, endothelial injury and complement-induced coagulopathy in COVID-19. Nat. Rev. Nephrol. 2021, 17, 46–64. [Google Scholar] [CrossRef] [PubMed]

| S.No. | Country | Total Cases | Total Deaths | Total Recovered | Active Cases * | Serious Critical * |
|---|---|---|---|---|---|---|
| 1 | USA | 9,802,374 | 239,842 | 6,293,132 | 3,269,400 | 18,045 |
| 2 | India | 8,364,086 | 124,354 | 7,711,809 | 527,923 | 8944 |
| 3 | Brazil | 5,590,941 | 161,170 | 5,064,344 | 365,427 | 8318 |
| 4 | Russia | 1,712,858 | 29,509 | 1,279,169 | 404,180 | 2300 |
| 5 | France | 1,543,321 | 38,674 | 122,662 | 1,381,985 | 4089 |
| 6 | Spain | 1,356,798 | 38,118 | N/A | N/A | 2786 |
| 7 | Argentina | 1,205,928 | 32,520 | 1,017,647 | 155,761 | 4816 |
| 8 | Colombia | 1,108,084 | 32,013 | 1,002,202 | 73,869 | 2376 |
| 9 | UK | 1,099,059 | 47,742 | N/A | N/A | 1142 |
| 10 | Mexico | 943,630 | 93,228 | 697,402 | 153,000 | 2838 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Khan, M.; Adil, S.F.; Alkhathlan, H.Z.; Tahir, M.N.; Saif, S.; Khan, M.; Khan, S.T. COVID-19: A Global Challenge with Old History, Epidemiology and Progress So Far. Molecules 2021, 26, 39. https://doi.org/10.3390/molecules26010039
Khan M, Adil SF, Alkhathlan HZ, Tahir MN, Saif S, Khan M, Khan ST. COVID-19: A Global Challenge with Old History, Epidemiology and Progress So Far. Molecules. 2021; 26(1):39. https://doi.org/10.3390/molecules26010039
Chicago/Turabian StyleKhan, Mujeeb, Syed F. Adil, Hamad Z. Alkhathlan, Muhammad N. Tahir, Sadia Saif, Merajuddin Khan, and Shams T. Khan. 2021. "COVID-19: A Global Challenge with Old History, Epidemiology and Progress So Far" Molecules 26, no. 1: 39. https://doi.org/10.3390/molecules26010039
APA StyleKhan, M., Adil, S. F., Alkhathlan, H. Z., Tahir, M. N., Saif, S., Khan, M., & Khan, S. T. (2021). COVID-19: A Global Challenge with Old History, Epidemiology and Progress So Far. Molecules, 26(1), 39. https://doi.org/10.3390/molecules26010039








