The Anticancer Peptide CIGB-552 Exerts Anti-Inflammatory and Anti-Angiogenic Effects through COMMD1
Abstract
1. Introduction
2. Results
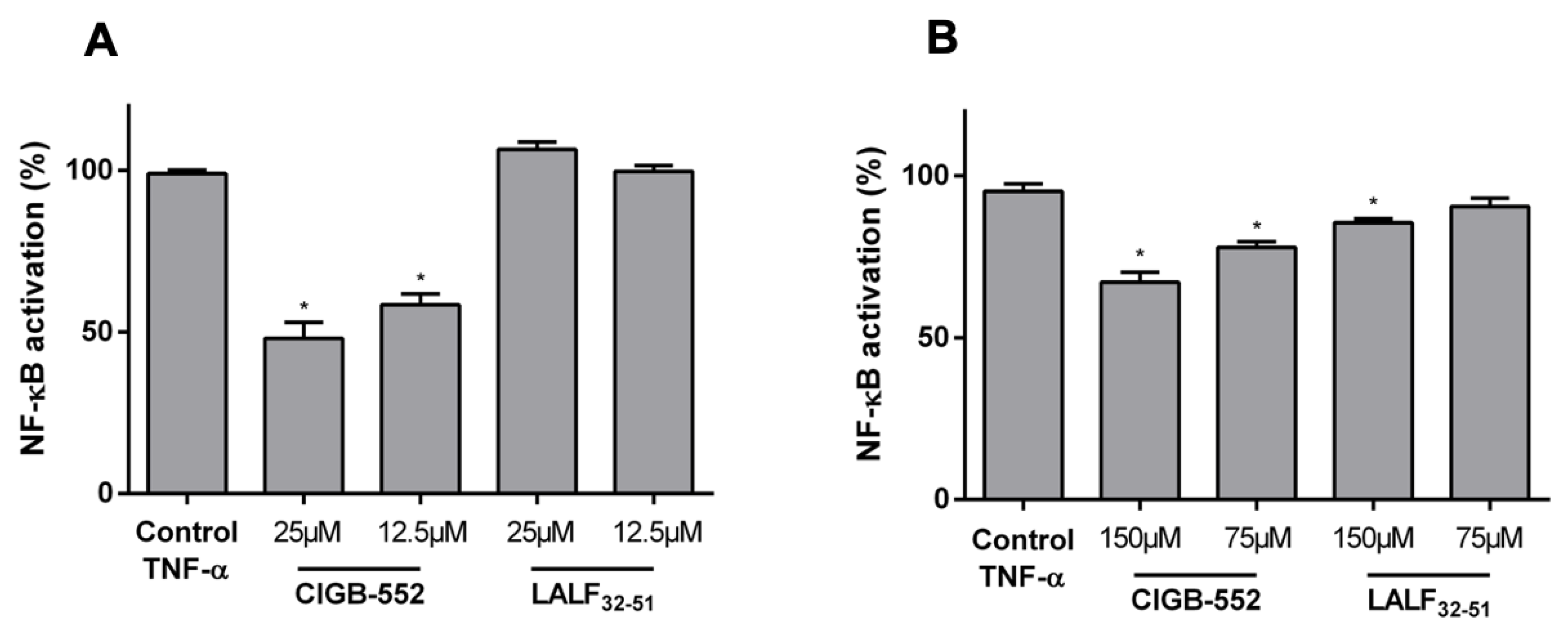
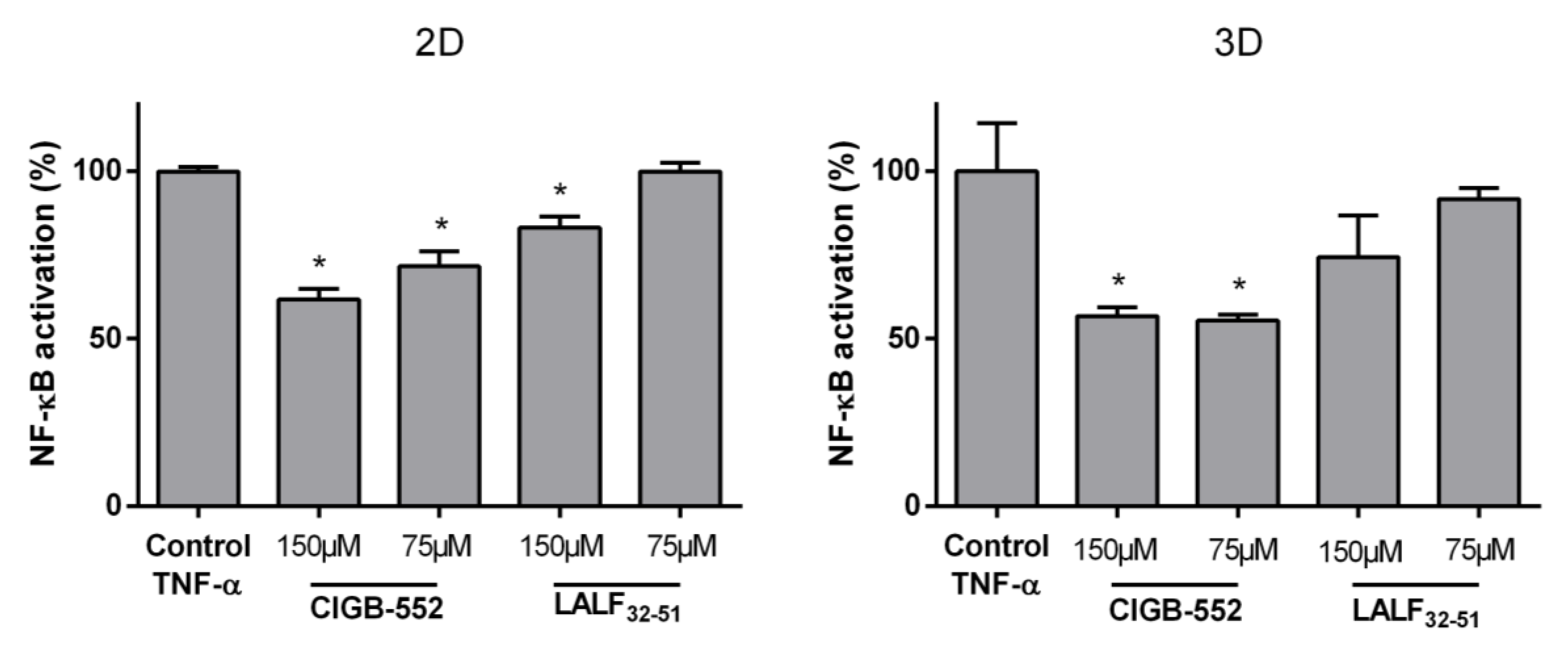
2.1. CIGB-552 but Not LALF32–51 Mediates NF-kB Activity in HT-29 and H460 Reporter Cell Lines
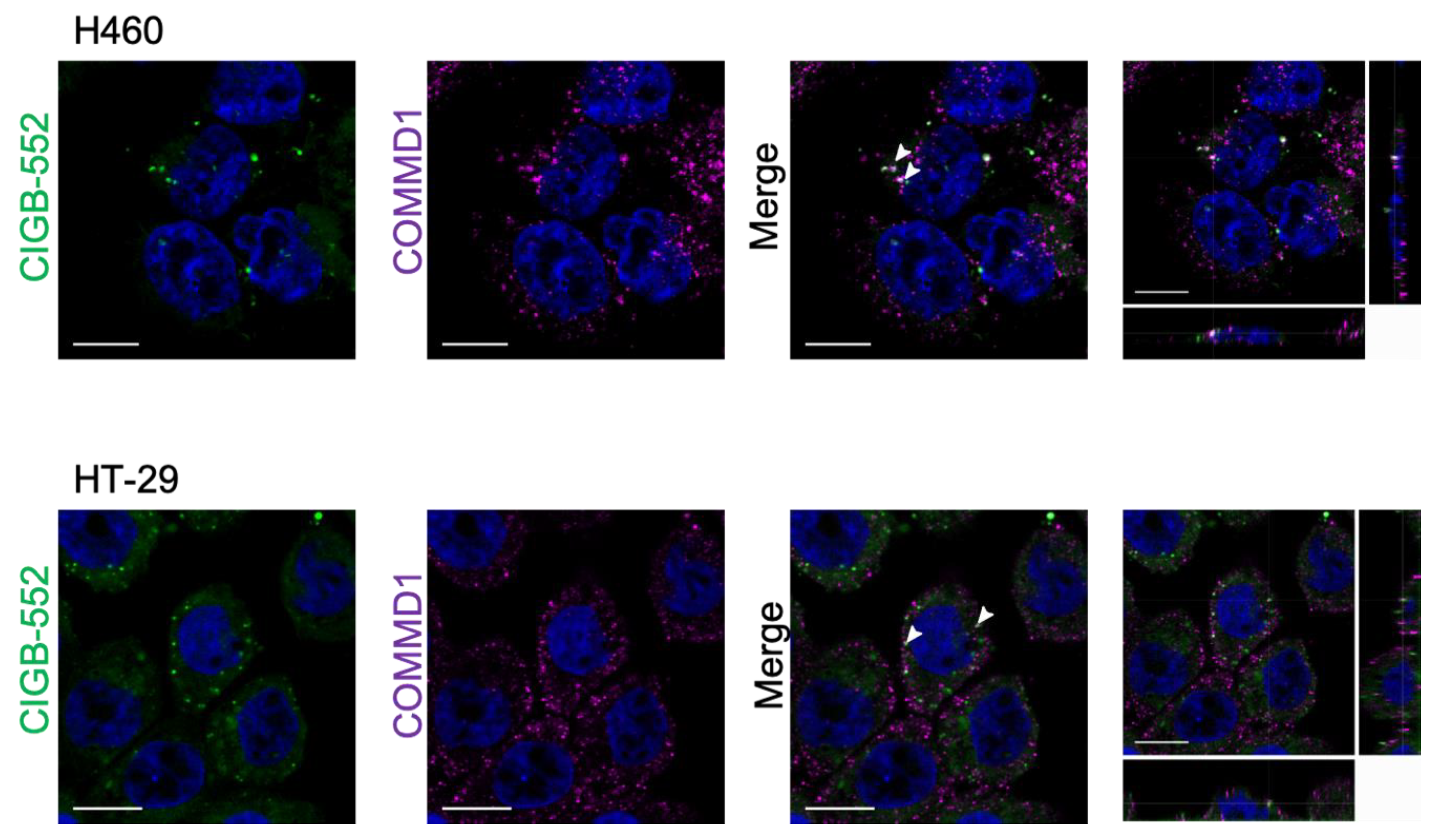
2.2. Interaction between CIGB-552 and COMMD1 by In Situ Immunodetection
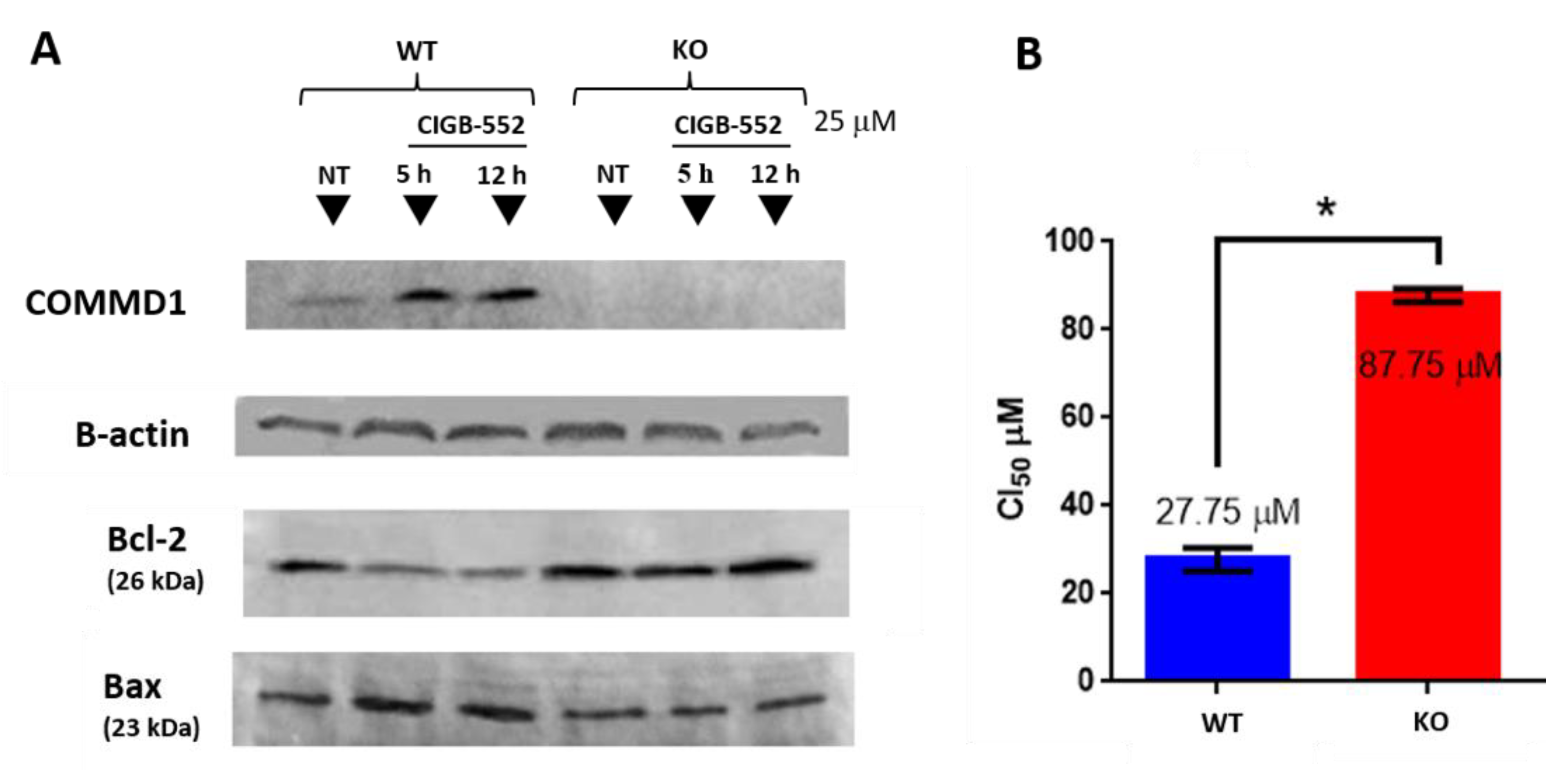
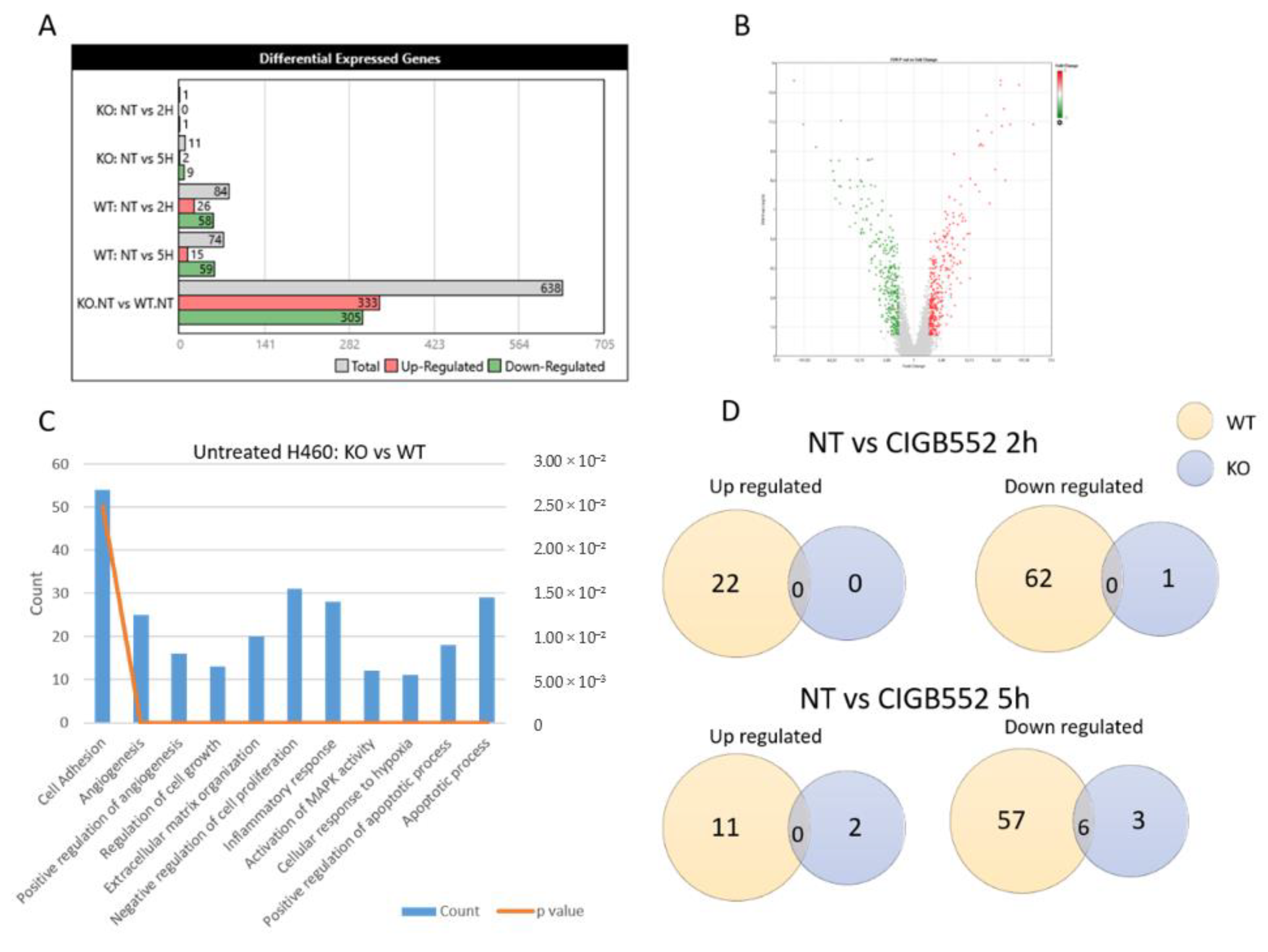
2.3. CIGB-552 Treatment in H460 WT and COMMD1 KO Cell Lines
2.4. CIGB-552 Requires COMMD1 to Exert Its Effect on Inflammation and Angiogenesis in the H460 Cell Line
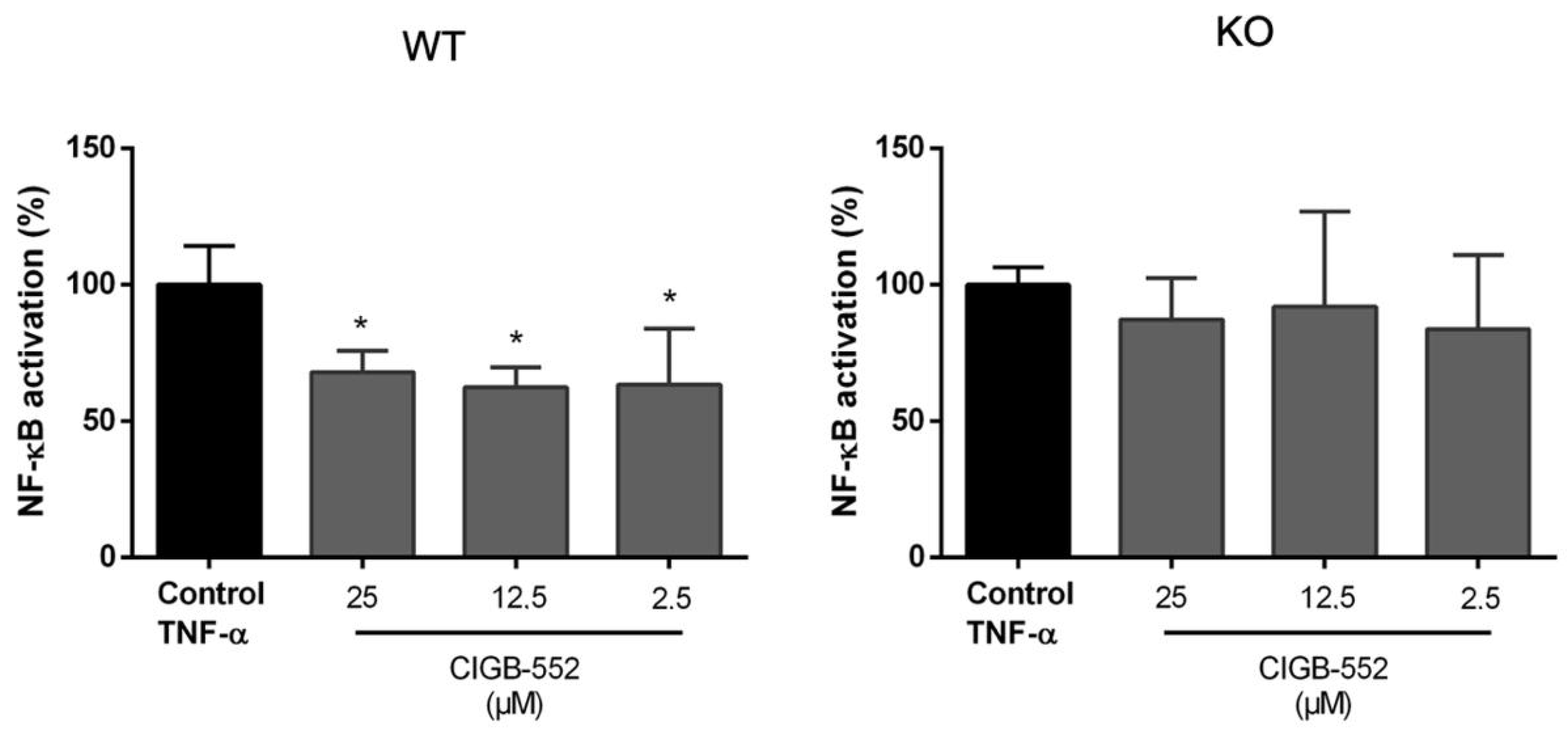
2.4.1. CIGB-552 Effects on NF-κB Transcriptional Activity Are Impaired in H460 COMMD1 KO Cells
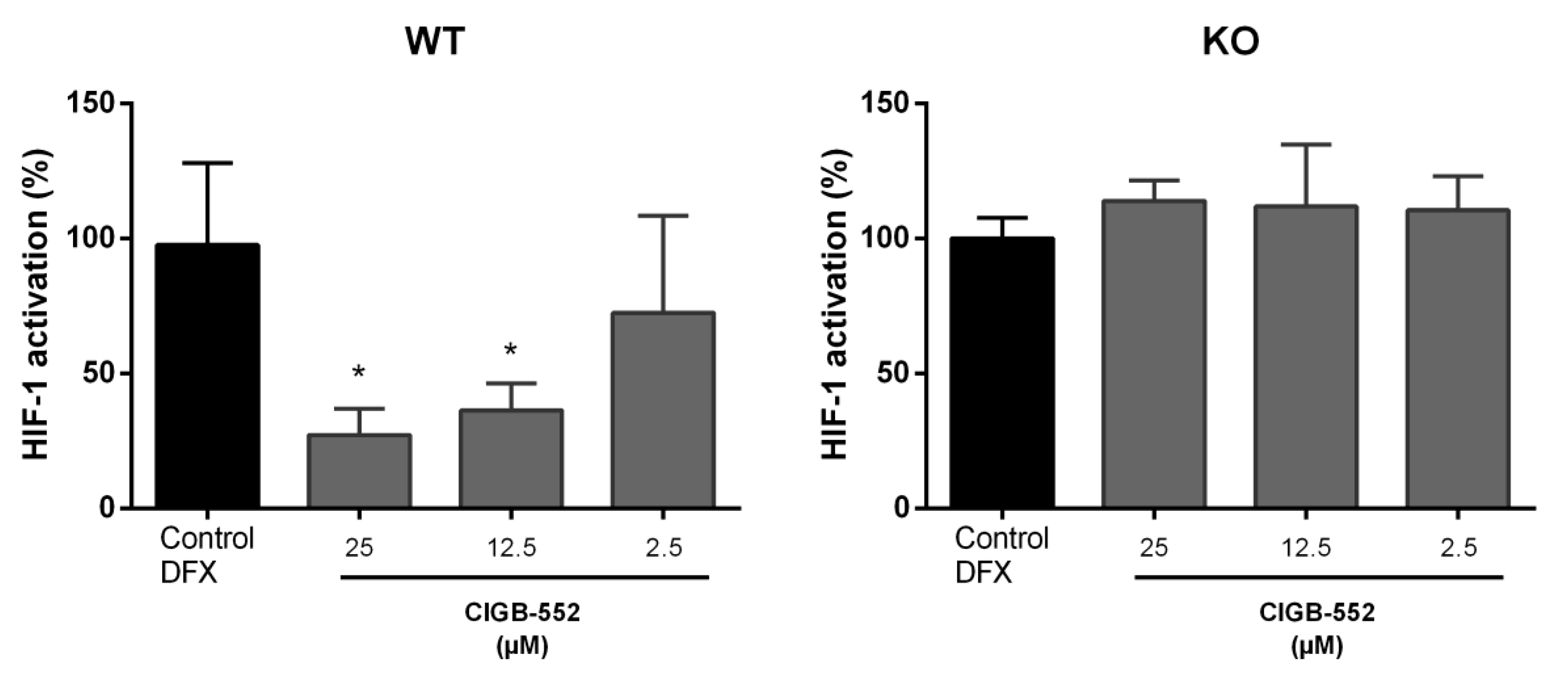
2.4.2. CIGB-552 Effects on HIF-1 Transcriptional Activity Are Impaired in H460 COMMD1 KO Cells
3. Discussion
4. Materials and Methods
4.1. Reagents and Chemicals
4.2. Peptide Synthesis
4.3. Cell Culture
4.4. NF-κB Activation Studies Using Stable Reporter Cell Lines
4.5. HT-29-NF-κB-hrGFP Spheroid Generation and Culture
4.6. Confocal Microscopy
4.7. COMMD1 Knock-out in H460 Cells by CRISPR-Cas9
4.8. HIF-1 and NF-kB Activation Studies in WT and COMMD1-KO H460 Cells
4.9. DNA Microarray
4.10. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Sample Availability
References
- Vallespi, M.G.; Fernandez, J.R.; Torrens, I.; Garcia, I.; Garay, H.; Mendoza, O.; Granadillo, M.; Falcon, V.; Acevedo, B.; Ubieta, R.; et al. Identification of a novel antitumor peptide based on the screening of an Ala-library derived from the LALF32–51 region. J. Pept. Sci. 2010, 16, 40–47. [Google Scholar] [CrossRef] [PubMed]
- Vallespi, M.G.; Glaria, L.A.; Reyes, O.; Garay, H.E.; Ferrero, J.; Araña, M.J. A Limulus antilipopolysaccharide factor-derived peptide exhibits a new immunological activity with potential applicability in infectious diseases. Clin. Diagn. Lab. Immunol. 2000, 7, 669–675. [Google Scholar] [CrossRef] [PubMed]
- Vallespi, M.G.; Alvarez-Obregón, J.C.; Rodriguez-Alonso, I.; Montero, T.; Garay, H.; Reyes, O.; Araña, M.J. A Limulus anti-LPS factor-derived peptide modulates cytokine gene expression and promotes resolution of bacterial acute infection in mice. Int. Immunopharmacol. 2003, 3, 247–256. [Google Scholar] [CrossRef]
- Fernández Massó, J.R.; Oliva Argüelles, B.; Tejeda, Y.; Astrada, S.; Garay, H.; Reyes, O.; Delgado-Roche, L.; Bollati-Fogolín, M.; Vallespí, M.G. The Antitumor Peptide CIGB-552 Increases COMMD1 and Inhibits Growth of Human Lung Cancer Cells. J. Amino Acids 2013, 2013, 251398. [Google Scholar] [CrossRef] [PubMed]
- Astrada, S.; Fernández Massó, J.; Vallespí, M.G.; Bollati-Fogolín, M.; Massó, J.R.F.; Vallespí, M.G.; Bollati-Fogolín, M. Cell penetrating capacity and internalization mechanisms used by the synthetic peptide CIGB-552 and its relationship with tumor cell line sensitivity. Molecules 2018, 23, 801. [Google Scholar] [CrossRef] [PubMed]
- Vallespi, M.G.; Colas, M.; Garay, H.; Reyes, O.; Araña, M.J. Differential regulation of Th1/Th2 in relevant tissues for sepsis pathogenesis with a Limulus anti-LPS factor-derived peptide increases survival in Gram-positive sepsis. Int. Immunopharmacol. 2004, 4, 1343–1351. [Google Scholar] [CrossRef]
- Vallespí, M.G.; Pimentel, G.; Cabrales-Rico, A.; Garza, J.; Oliva, B.; Mendoza, O.; Gomez, Y.; Basaco, T.; Sánchez, I.; Calderón, C.; et al. Antitumor efficacy, pharmacokinetic and biodistribution studies of the anticancer peptide CIGB-552 in mouse models. J. Pept. Sci. 2014, 20, 850–859. [Google Scholar] [CrossRef]
- Van De Sluis, B.; Mao, X.; Zhai, Y.; Groot, A.J.; Vermeulen, J.F.; Van Der Wall, E.; Van Diest, P.J.; Hofker, M.H.; Wijmenga, C.; Klomp, L.W.; et al. COMMD1 disrupts HIF-1α/β dimerization and inhibits human tumor cell invasion. J. Clin. Investig. 2010, 120, 2119–2130. [Google Scholar] [CrossRef]
- Bartuzi, P.; Hofker, M.H.; van de Sluis, B. Tuning NF-κB activity: A touch of COMMD proteins. Biochim. Biophys. Acta-Mol. Basis Dis. 2013, 1832, 2315–2321. [Google Scholar] [CrossRef]
- Riera-Romo, M. COMMD1: A Multifunctional Regulatory Protein. J. Cell. Biochem. 2018, 119, 34–51. [Google Scholar] [CrossRef]
- Muller, P.A.J.; Van de Sluis, B.; Groot, A.J.; Verbeek, D.; Vonk, W.I.M.; Maine, G.N.; Burstein, E.; Wijmenga, C.; Vooijs, M.; Reits, E.; et al. Nuclear-Cytosolic Transport of COMMD1 Regulates NF-κB and HIF-1 Activity. Traffic 2009, 10, 514–527. [Google Scholar] [CrossRef] [PubMed]
- Semenza, G.L. Targeting HIF-1 for cancer therapy. Nat. Rev. Cancer 2003, 3, 721–732. [Google Scholar] [CrossRef] [PubMed]
- Perkins, N.D. The diverse and complex roles of NF-κB subunits in cancer. Nat. Rev. Cancer 2012, 12, 121–132. [Google Scholar] [CrossRef] [PubMed]
- Núñez de Villavicencio-Díaz, T.; Ramos Gómez, Y.; Oliva Argüelles, B.; Fernández Masso, J.R.; Rodríguez-Ulloa, A.; Cruz García, Y.; Guirola-Cruz, O.; Perez-Riverol, Y.; Javier González, L.; Tiscornia, I.; et al. Comparative proteomics analysis of the antitumor effect of CIGB-552 peptide in HT-29 colon adenocarcinoma cells. J. Proteomics 2015, 126, 163–171. [Google Scholar] [CrossRef]
- Daghero, H.; Pagotto, R.; Vallespí, M.G.; Bollati-Fogolín, M. Generation of stable reporter breast and lung cancer cell lines for NF-κB activation studies. J. Biotechnol. 2019, 301, 79–87. [Google Scholar] [CrossRef]
- Mastropietro, G.; Tiscornia, I.; Perelmuter, K.; Astrada, S.; Bollati-fogolín, M. HT-29 and Caco-2 reporter cell lines for functional studies of nuclear factor kappa B activation. Mediators Inflamm. 2015, 2015, 13. [Google Scholar] [CrossRef]
- Riedl, A.; Schlederer, M.; Pudelko, K.; Stadler, M.; Walter, S.; Unterleuthner, D.; Unger, C.; Kramer, N.; Hengstschläger, M.; Kenner, L.; et al. Comparison of cancer cells in 2D vs 3D culture reveals differences in AKT-mTOR-S6K signaling and drug responses. J. Cell Sci. 2017, 130, 203–218. [Google Scholar] [CrossRef]
- Astrada, S.; Gomez, Y.; Barrera, E.; Obal, G.; Pritsch, O.; Pantano, S.; Vallesp�, M.G.; Bollati-Fogol�n, M. Comparative analysis reveals amino acids critical for anticancer activity of peptide CIGB-552. J. Pept. Sci. 2016, 22, 711–722. [Google Scholar] [CrossRef]
- Tong, X.; Xie, D.; O’Kelly, J.; Miller, C.W.; Muller-Tidow, C.; Koeffler, H.P. Cyr61, a Member of CCN Family, Is a Tumor Suppressor in Non-Small Cell Lung Cancer. J. Biol. Chem. 2001, 276, 47709–47714. [Google Scholar] [CrossRef]
- Chien, W.; Yin, D.; Gui, D.; Mori, A.; Frank, J.M.; Said, J.; Kusuanco, D.; Marchevsky, A.; McKenna, R.; Koeffler, H.P. Suppression of cell proliferation and signaling transduction by connective tissue growth factor in non-small cell lung cancer cells. Mol. Cancer Res. 2006, 4, 591–598. [Google Scholar] [CrossRef]
- Liu, C.; Rangnekar, V.M.; Adamson, E.; Mercola, D. Suppression of growth and transformation and induction of apoptosis by EGR-1. Cancer Gene Ther. 1998, 5, 3–28. [Google Scholar] [PubMed]
- Sakaue, M.; Adachi, H.; Dawson, M.; Jetten, A.M. Induction of Egr-1 expression by the retinoid AHPN in human lung carcinoma cells is dependent on activated ERK1/2. Cell Death Differ. 2001, 8, 411–424. [Google Scholar] [CrossRef] [PubMed]
- Chapman, N.R.; Perkins, N.D. Inhibition of the RelA(p65) NF-kappaB subunit by Egr-1. J. Biol. Chem. 2000, 275, 4719–4725. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.J.; Tsai, J.W.; Hsieh, K.C.; Yang, Y.C.; Chen, Y.J.; Huang, M.S.; Yuan, S.S. Id1 promotes lung cancer cell proliferation and tumor growth through Akt-related pathway. Cancer Lett. 2011, 307, 191–199. [Google Scholar] [CrossRef] [PubMed]
- Gautschi, O.; Tepper, C.G.; Purnell, P.R.; Izumiya, Y.; Evans, C.P.; Green, T.P.; Desprez, P.Y.; Lara, P.N.; Gandara, D.R.; Mack, P.C.; et al. Regulation of Id1 expression by Src: Implications for targeting of the bone morphogenetic protein pathway in cancer. Cancer Res. 2008, 68, 2250–2258. [Google Scholar] [CrossRef]
- Maine, G.N.; Mao, X.; Komarck, C.M.; Burstein, E. COMMD1 promotes the ubiquitination of NF-κB subunits through a cullin-containing ubiquitin ligase. EMBO J. 2007, 26, 436–447. [Google Scholar] [CrossRef]
- Van De Sluis, B.; Muller, P.; Duran, K.; Chen, A.; Groot, A.J.; Klomp, L.W.; Liu, P.P.; Wijmenga, C. Increased Activity of Hypoxia-Inducible Factor 1 Is Associated with Early Embryonic Lethality in Commd1 Null Mice. Mol. Cell. Biol. 2007, 27, 4142–4156. [Google Scholar] [CrossRef]
- Hu, Y.; Liu, J.; Huang, H. Recent agents targeting HIF-1α for cancer therapy. J. Cell. Biochem. 2013, 114, 498–509. [Google Scholar] [CrossRef]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Jauset, T.; Beaulieu, M.E. Bioactive cell penetrating peptides and proteins in cancer: A bright future ahead. Curr. Opin. Pharmacol. 2019, 47, 133–140. [Google Scholar] [CrossRef]
- Tafani, M.; Pucci, B.; Russo, A.; Schito, L.; Pellegrini, L.; Perrone, G.A.; Villanova, L.; Salvatori, L.; Ravenna, L.; Petrangeli, E.; et al. Modulators of HIF1α and NFkB in cancer treatment: Is it a rational approach for controlling malignant progression? Front. Pharmacol. 2013, 4, 13. [Google Scholar] [CrossRef] [PubMed]
- Pampaloni, F.; Reynaud, E.G.; Stelzer, E.H.K. The third dimension bridges the gap between cell culture and live tissue. Nat. Rev. Mol. Cell Biol. 2007, 8, 839–845. [Google Scholar] [CrossRef] [PubMed]
- Hirschhaeuser, F.; Menne, H.; Dittfeld, C.; West, J.; Mueller-Klieser, W.; Kunz-Schughart, L.A. Multicellular tumor spheroids: An underestimated tool is catching up again. J. Biotechnol. 2010, 148, 3–15. [Google Scholar] [CrossRef] [PubMed]
- Bhakat, K.K.; Mantha, A.K.; Mitra, S. Transcriptional regulatory functions of mammalian AP-endonuclease (APE1/Ref-1), an essential multifunctional protein. Antioxid. Redox Signal. 2009, 11, 621–637. [Google Scholar] [CrossRef] [PubMed]
- Ryan, J.A. Cell Cloning by Serial Dilution on 96 Well Plates Protocol. Available online: https://www.corning.com/catalog/cls/documents/protocols/Single_cell_cloning_protocol.pdf (accessed on 23 November 2020).
| Peptide | Sequence | Properties | H460 1 (μM) | HT-29 1 (μM) | Ref. |
|---|---|---|---|---|---|
| LALF32–51 | HYRIKPTFRRLKWKYKGKFW | AMP/IMP/CPP | 460 ± 25 | 549 ± 25 | [3,6] |
| L-2 | HARIKPTFRRLKWKYKGKFW | ACP/CCP | 57 ± 6 2 | ND 3 | [1] |
| CIGB-552 | Ac-HARIKpTFRRlKWKYKGKFW | ACP/CCP | 23 ± 8 2 | 166 ± 66 2 | [4,5] |
| Gene Symbol | Description | Fold Change |
|---|---|---|
| Angiogenesis | ||
| VEGFC | vascular endothelial growth factor C | −3.67 |
| CTGF | connective tissue growth factor | −2.98 |
| GATA6 | GATA binding protein 6 | 4.64 |
| STC1 | stanniocalcin 1 | 2.72 |
| PTGS2 | prostaglandin-endoperoxide synthase 2 (cyclooxygenase 2) | 9.22 |
| ANPEP | alanyl aminopeptidase, membrane (ANP, CD13) | 2.90 |
| C5AR1 | complement component 5a receptor 1 | 2.57 |
| Inflammation | ||
| IL1A | interleukin 1 alpha | 2.33 |
| IL1B | interleukin 1 beta | 6.1 |
| CXCL1 | chemokine (C-X-C motif) ligand 1 | 2.30 |
| CXCL5 | chemokine (C-X-C motif) ligand 5 | 2.12 |
| IL6R | interleukin 6 receptor | 3.62 |
| NFKBIA | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 2.53 |
| NFKBID | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, delta | 3.42 |
| Gene Symbol | Description | Fold Change |
|---|---|---|
| COMMD1 KO | ||
| EGR1 | early growth response 1 | −90.15 |
| ID1 | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 2.87 |
| CYR61 | cysteine-rich, angiogenic inducer, 61 | −2.32 |
| CTGF | connective tissue growth factor | −2.22 |
| WT | ||
| EGR1 | early growth response 1 | −704.56 |
| ID1 | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 2.97 |
| MAP3K14 | mitogen-activated protein kinase kinase kinase 14 | 3.01 |
| ZFP36 | ZFP36 Ring Finger Protein (Tristetraprolin) | −6.33 |
| NFKBIA | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | −2.90 |
| BCL2L11 | BCL2-like 11 (apoptosis facilitator) | −2.14 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Daghero, H.; Fernández Massó, J.R.; Astrada, S.; Guerra Vallespí, M.; Bollati-Fogolín, M. The Anticancer Peptide CIGB-552 Exerts Anti-Inflammatory and Anti-Angiogenic Effects through COMMD1. Molecules 2021, 26, 152. https://doi.org/10.3390/molecules26010152
Daghero H, Fernández Massó JR, Astrada S, Guerra Vallespí M, Bollati-Fogolín M. The Anticancer Peptide CIGB-552 Exerts Anti-Inflammatory and Anti-Angiogenic Effects through COMMD1. Molecules. 2021; 26(1):152. https://doi.org/10.3390/molecules26010152
Chicago/Turabian StyleDaghero, Hellen, Julio Raúl Fernández Massó, Soledad Astrada, Maribel Guerra Vallespí, and Mariela Bollati-Fogolín. 2021. "The Anticancer Peptide CIGB-552 Exerts Anti-Inflammatory and Anti-Angiogenic Effects through COMMD1" Molecules 26, no. 1: 152. https://doi.org/10.3390/molecules26010152
APA StyleDaghero, H., Fernández Massó, J. R., Astrada, S., Guerra Vallespí, M., & Bollati-Fogolín, M. (2021). The Anticancer Peptide CIGB-552 Exerts Anti-Inflammatory and Anti-Angiogenic Effects through COMMD1. Molecules, 26(1), 152. https://doi.org/10.3390/molecules26010152







