Target Identification Using Homopharma and Network-Based Methods for Predicting Compounds Against Dengue Virus-Infected Cells
Abstract
1. Introduction
2. Results and Discussion
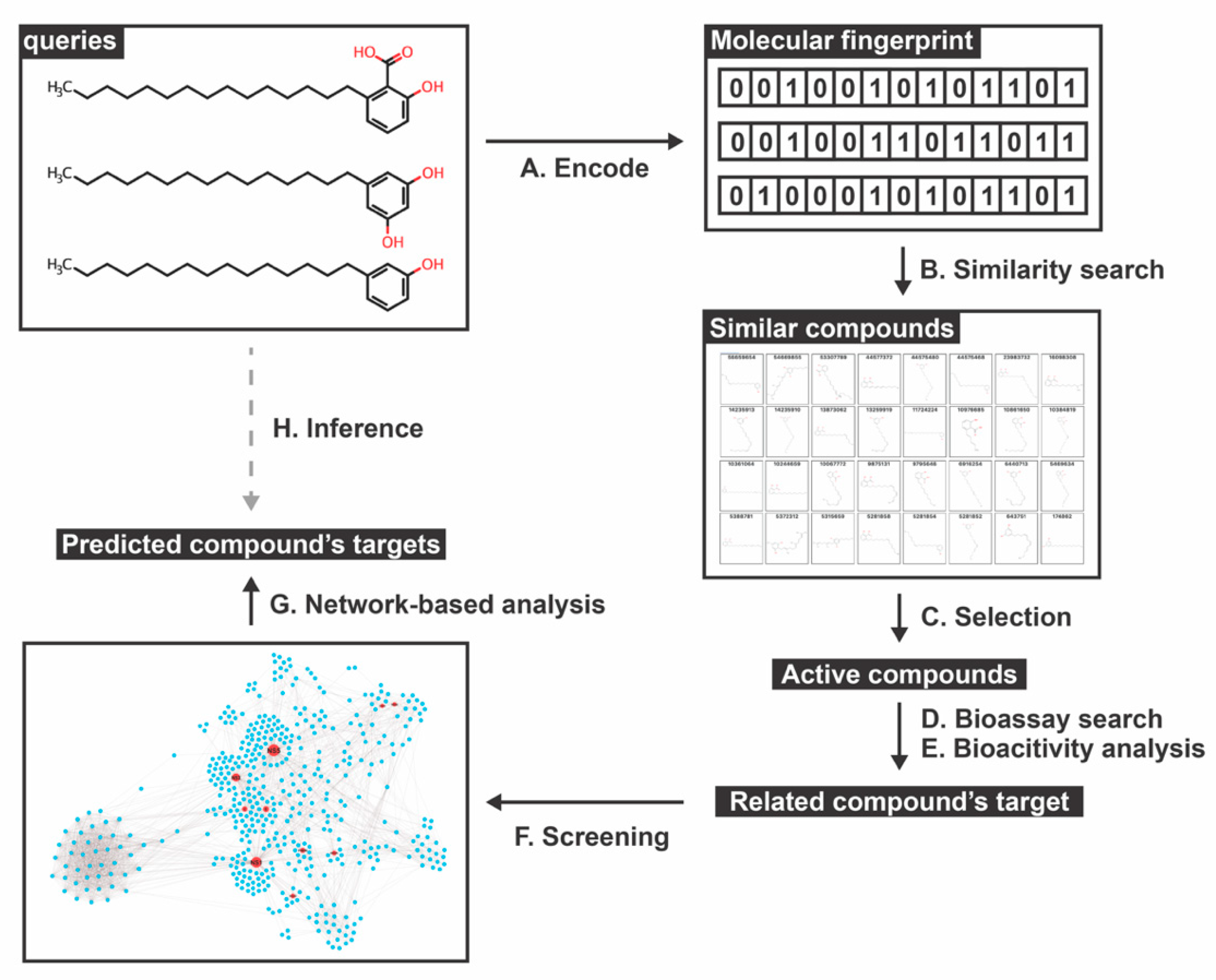
2.1. Similar Compounds of Phenolic Lipids for Predicting Target
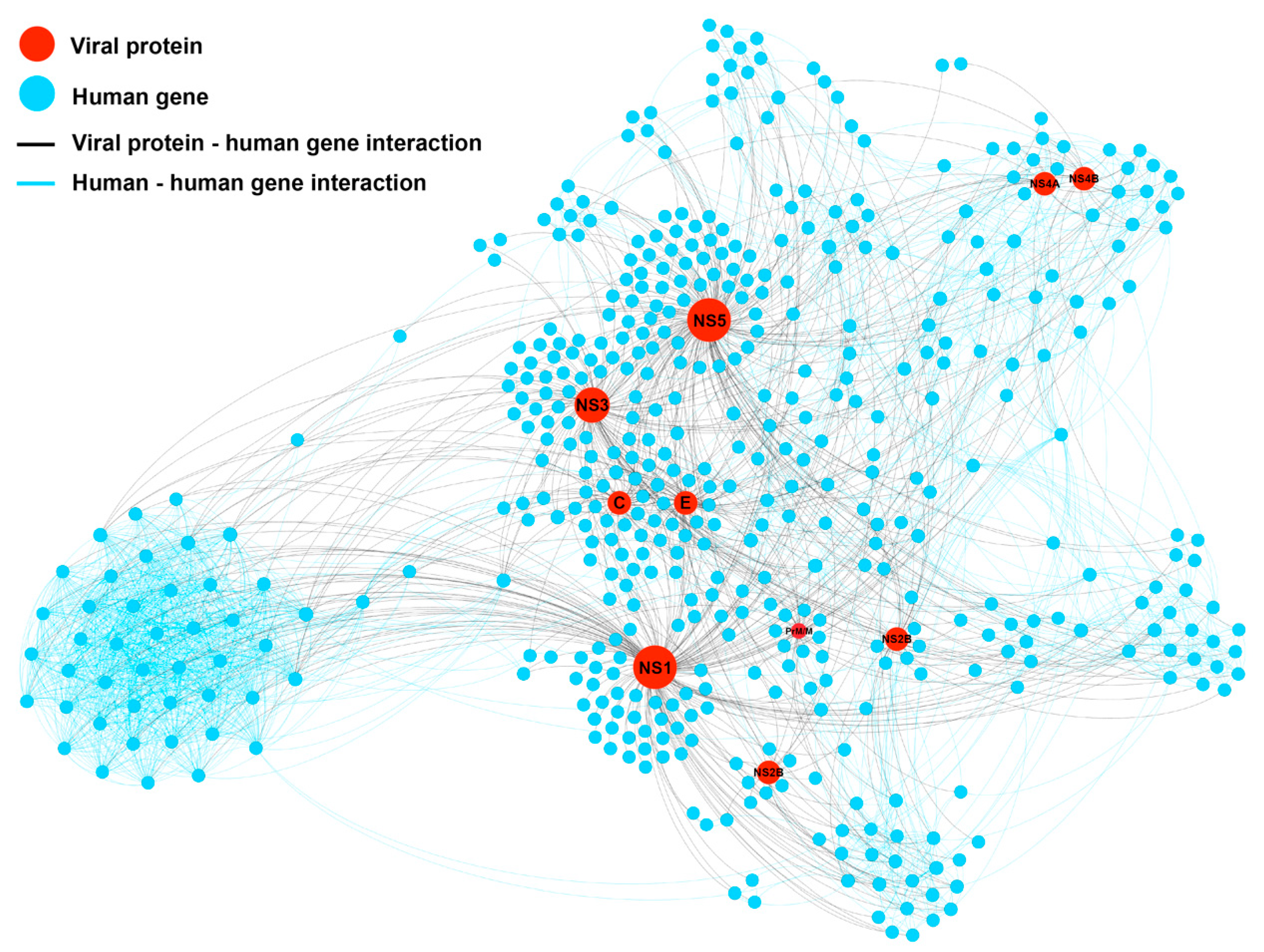
2.2. Network-Based Construction and Centrality Analysis
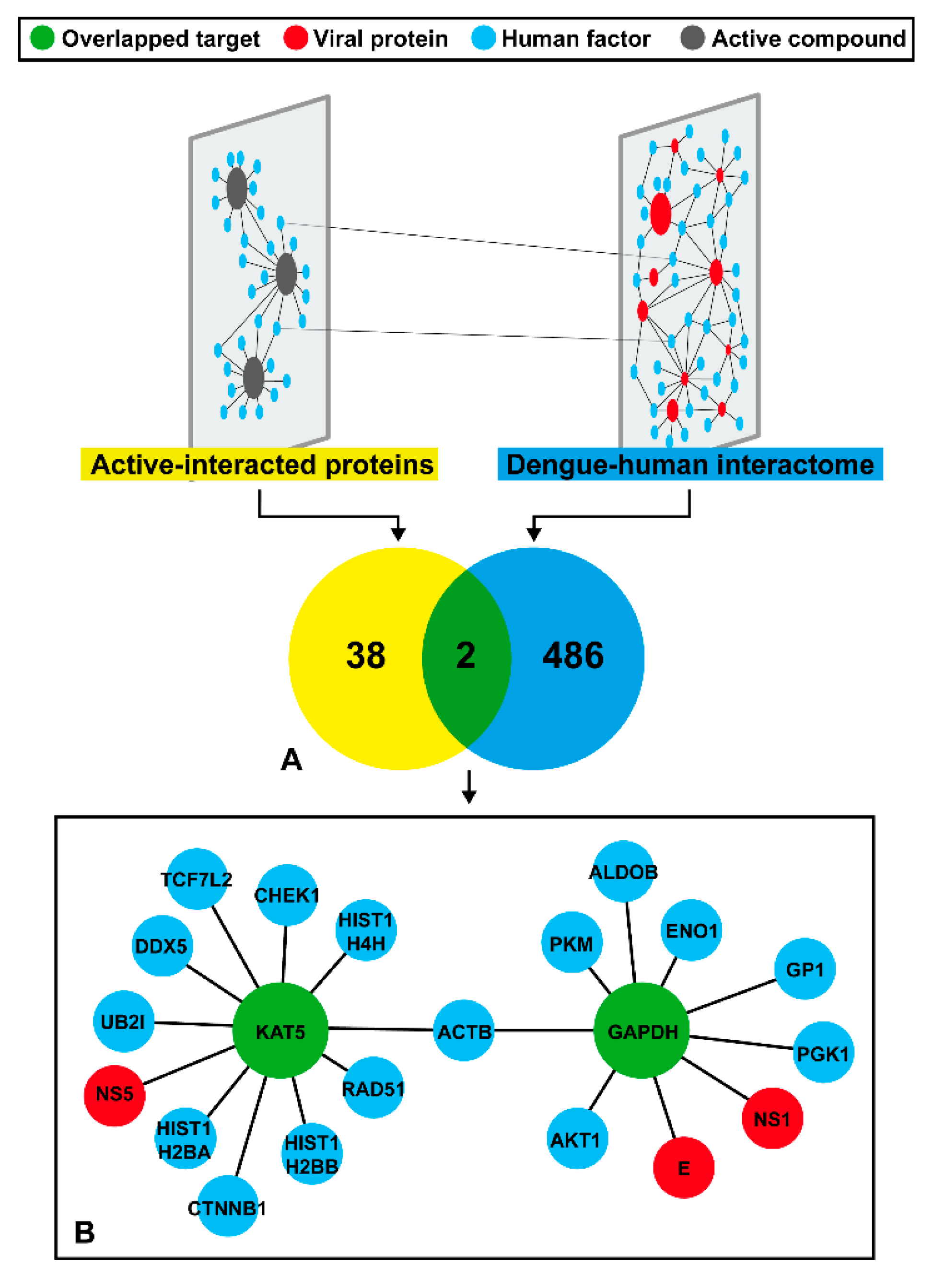
2.3. Prediction of DENV Related Target
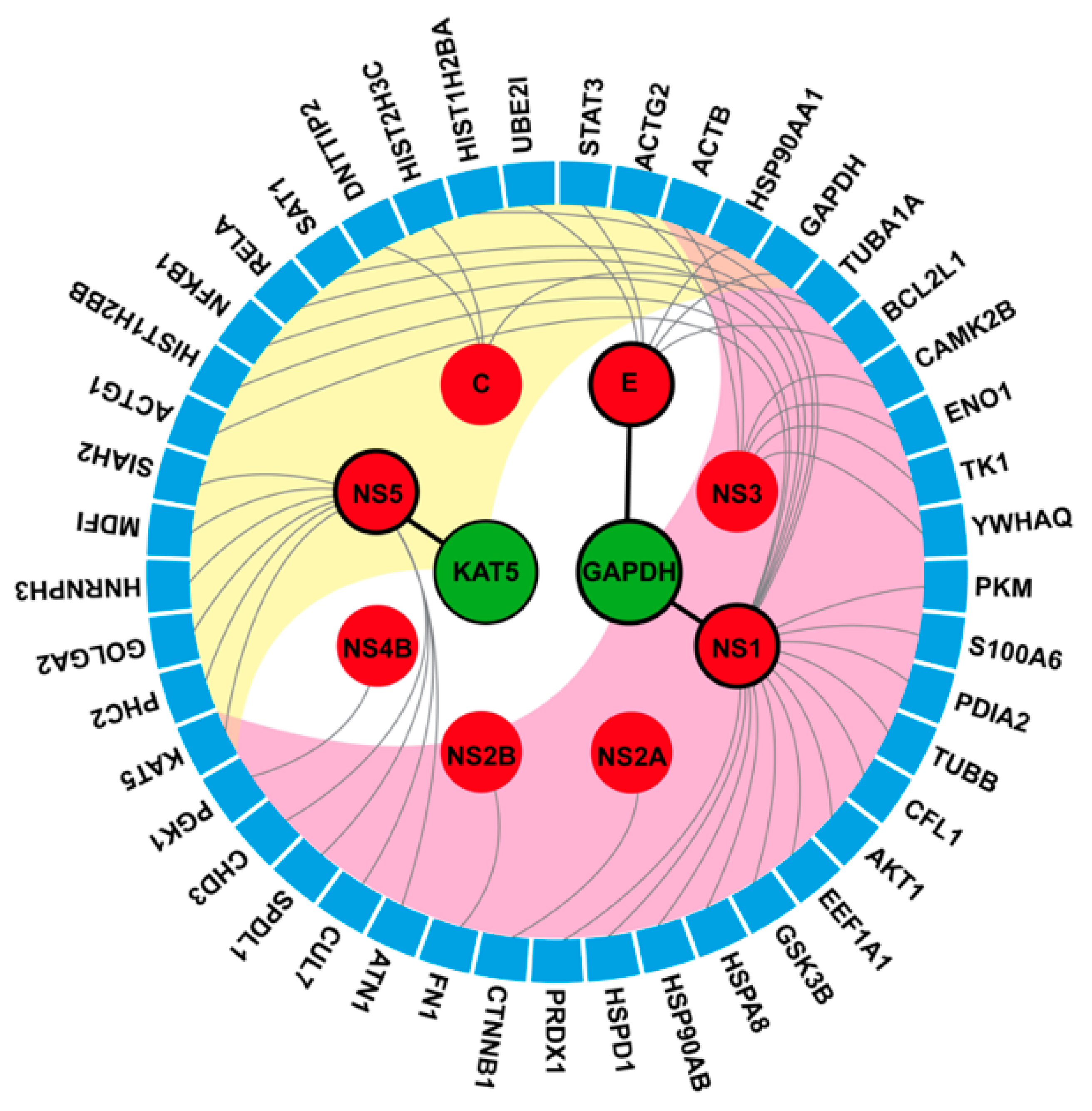
2.4. Viral Target for Phenolic Lipid Compounds
2.5. Human Protein Target Annotation
3. Materials and Methods
3.1. Data Collection
3.2. Related Target Evaluation
3.3. Network-Based Construction and Analysis
3.4. Identification of Targets Aassociated with DENV
3.5. Target Annotation
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wilder-Smith, A. Can dengue virus be sexually transmitted? J. Travel Med. 2019, 26, 1195–1982. [Google Scholar] [CrossRef] [PubMed]
- Gibbons, R.V.; Vaughn, D.W. Dengue: An escalating problem. BMJ 2002, 324, 1563–1566. [Google Scholar] [CrossRef]
- World Health Organisation. Dengue and Dengue Haemorrhagic Fever; World Health Organisation: Geneva, Switzerland, 2002. [Google Scholar]
- Guzman, M.G.; Gubler, D.J.; Izquierdo, A.; Martinez, E.; Halstead, S.B. Dengue infection. Nat. Rev. Dis. Primers 2016, 2, 16055. [Google Scholar] [CrossRef] [PubMed]
- Disease, T.V.-B. Dengue Situation in Thailand. 2017. Available online: http://www.thaivbd.org/n/dengues/view/587 (accessed on 5 March 2017).
- Rodriguez, R.C.; Carrasquilla, G.; Porras-Ramírez, A.; Galera-Gelvez, K.; Yescas, J.G.L.; Rueda-Gallardo, J.A. The Burden of Dengue and the Financial Cost to Colombia, 2010-2012. Am. J. Trop. Med. Hyg. 2016, 94, 1065–1072. [Google Scholar] [CrossRef] [PubMed]
- Yacoub, S.; Farrar, J. 15-Dengue, in Manson’s Tropical Infectious Diseases, 23th ed.; Farrar, J., Hotez, P.J., Junghanss, T., Kang, G., Lalloo, D., White, N.J., Eds.; W.B. Saunders: London, UK, 2014; pp. 162–170.e2. [Google Scholar]
- Halstead, S.; Wilder-Smith, A. Severe dengue in travellers: Pathogenesis, risk and clinical management. J. Travel Med. 2019, 26, 1195–1982. [Google Scholar] [CrossRef] [PubMed]
- Guzman, M.G.; Alvarez, M.; Halstead, S.B. Secondary infection as a risk factor for dengue hemorrhagic fever/dengue shock syndrome: An historical perspective and role of antibody-dependent enhancement of infection. Arch. Virol. 2013, 158, 1445–1459. [Google Scholar] [CrossRef]
- Rey, F.A.; Stiasny, K.; Vaney, M.; Dellarole, M.; Heinz, F.X. The bright and the dark side of human antibody responses to flaviviruses: Lessons for vaccine design. EMBO Rep. 2017, 19, 206–224. [Google Scholar] [CrossRef] [PubMed]
- Rothman, A.L. Immunology and Immunopathogenesis of Dengue Disease, in Advances in Virus Research; Academic Press: Cambridge, MA, USA, 2003; pp. 397–419. [Google Scholar]
- Normile, D. Dengue vaccine trial poses public health quandary. Sci. 2014, 345, 367–368. [Google Scholar] [CrossRef] [PubMed]
- The Lancet Infectious, D. The dengue vaccine dilemma. Lancet Infect. Dis. 2018, 18, 123. [Google Scholar] [CrossRef]
- Boonyasuppayakorn, S.; Reichert, E.D.; Manzano, M.; Nagarajan, K.; Padmanabhan, R. Amodiaquine, an antimalarial drug, inhibits dengue virus type 2 replication and infectivity. Antivir. Res. 2014, 106, 125–134. [Google Scholar] [CrossRef] [PubMed]
- Byrd, C.M.; Dai, N.; Grosenbach, U.W.; Berhanu, A.; Jones, K.F.; Cardwell, K.B.; Schneider, C.; Wineinger, K.A.; Page, J.M.; Harver, C.; et al. A Novel Inhibitor of Dengue Virus Replication That Targets the Capsid Protein. Antimicrob. Agents Chemother. 2012, 57, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Capeding, M.R.; Tran, N.H.; Hadinegoro, S.R.S.; Ismail, H.I.H.M.; Chotpitayasunondh, T.; Chua, M.N.; Luong, C.Q.; Rusmil, K.; Wirawan, D.N.; Nallusamy, R.; et al. Clinical efficacy and safety of a novel tetravalent dengue vaccine in healthy children in Asia: A phase 3, randomised, observer-masked, placebo-controlled trial. Lancet 2014, 384, 1358–1365. [Google Scholar] [CrossRef]
- Chew, M.-F.; Poh, K.-S.; Poh, C.-L. Peptides as Therapeutic Agents for Dengue Virus. Int. J. Med Sci. 2017, 14, 1342–1359. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.P. Dengue drug discovery: Progress, challenges and outlook. Antivir. Res. 2019, 163, 156–178. [Google Scholar] [CrossRef] [PubMed]
- Karlas, A.; Berre, S.; Couderc, T.; Varjak, M.; Braun, P.; Meyer, M.; Gangneux, N.; Karo-Astover, L.; Weege, F.; Raftery, M.; et al. A human genome-wide loss-of-function screen identifies effective chikungunya antiviral drugs. Nat. Commun. 2016, 7, 11320. [Google Scholar] [CrossRef] [PubMed]
- Wong, K.Z.; Chu, J.J.H. The Interplay of Viral and Host Factors in Chikungunya Virus Infection: Targets for Antiviral Strategies. Viruses 2018, 10, 294. [Google Scholar] [CrossRef]
- Kanyaboon, P.; Saelee, T.; Suroengrit, A.; Hengphasatporn, K.; Rungrotmongkol, T.; Chavasiri, W.; Boonyasuppayakorn, S. Cardol triene inhibits dengue infectivity by targeting kl loops and preventing envelope fusion. Sci. Rep. 2018, 8, 16643. [Google Scholar] [CrossRef]
- Teerasripreecha, D.; Phuwapraisirisan, P.; Puthong, S.; Kimura, K.; Okuyama, M.; Mori, H.; Kimura, A.; Chanchao, C. In vitro antiproliferative/cytotoxic activity on cancer cell lines of a cardanol and a cardol enriched from Thai Apis mellifera propolis. BMC Complement. Altern. Med. 2012, 12, 27. [Google Scholar] [CrossRef]
- Hundt, J.; Li, Z.; Liu, Q. The Inhibitory Effects of Anacardic Acid on Hepatitis C Virus Life Cycle. PLoS ONE 2015, 10, e0117514. [Google Scholar] [CrossRef]
- Chao, L.; Jang, J.; Johnson, A.; Nguyen, A.; Gray, N.S.; Yang, P.L.; Harrison, S.C. How small-molecule inhibitors of dengue-virus infection interfere with viral membrane fusion. eLife 2018, 7, e36461. [Google Scholar] [CrossRef]
- Coloma, J.; Harris, E. Broad and strong: The ultimate antibody to dengue virus. Nat. Immunol. 2015, 16, 135–137. [Google Scholar] [CrossRef] [PubMed]
- Crill, W.D.; Roehrig, J. Monoclonal Antibodies That Bind to Domain III of Dengue Virus E Glycoprotein Are the Most Efficient Blockers of Virus Adsorption to Vero Cells. J. Virol. 2001, 75, 7769–7773. [Google Scholar] [CrossRef] [PubMed]
- Cabarcas-Montalvo, M.; Maldonado-Rojas, W.; Montes-Grajales, D.; Bertel-Sevilla, A.; Wagner-Döbler, I.; Sztajer, H.; Reck, M.; Alarcón, M.C.F.; Ocazionez, R.; Olivero-Verbel, J. Discovery of antiviral molecules for dengue: In silico search and biological evaluation. Eur. J. Med. Chem. 2016, 110, 87–97. [Google Scholar] [CrossRef] [PubMed]
- Fernando, S.; Fernando, T.; Stefanik, M.; Eyer, L.; Růžek, D. An Approach for Zika Virus Inhibition Using Homology Structure of the Envelope Protein. Mol. Biotechnol. 2016, 58, 801–806. [Google Scholar] [CrossRef] [PubMed]
- Hughes, J.; Rees, S.; Kalindjian, S.; Philpott, K. Principles of early drug discovery. Br. J. Pharmacol. 2011, 162, 1239–1249. [Google Scholar] [CrossRef] [PubMed]
- Campillos, M.; Kuhn, M.; Gavin, A.-C.; Jensen, L.J.; Bork, P. Drug Target Identification Using Side-Effect Similarity. Sci. 2008, 321, 263–266. [Google Scholar] [CrossRef] [PubMed]
- Keiser, M.J.; Setola, V.; Irwin, J.J.; Laggner, C.; Abbas, A.I.; Hufeisen, S.J.; Jensen, N.H.; Kuijer, M.B.; Matos, R.C.; Tran, T.B.; et al. Predicting new molecular targets for known drugs. Nature 2009, 462, 175–181. [Google Scholar] [CrossRef]
- Leelananda, S.P.; Lindert, S. Computational methods in drug discovery. Beilstein J. Org. Chem. 2016, 12, 2694–2718. [Google Scholar] [CrossRef]
- Rognan, D. Docking Methods for Virtual Screening: Principles and Recent Advances, in Virtual Screening; Wiley-VCH Verlag GmbH & Co.: Hoboken, NJ, USA, 2011; pp. 153–176. [Google Scholar]
- Bredel, M.; Jacoby, E. Chemogenomics: An emerging strategy for rapid target and drug discovery. Nat. Rev. Genet. 2004, 5, 262–275. [Google Scholar] [CrossRef]
- Hengphasatporn, K.; Garon, A.; Wolschann, P.; Langer, T.; Shigeta, Y.; Huynh, T.; Chavasiri, W.; Saelee, T.; Boonyasuppayakorn, S.; Rungrotmongkol, T. Multiple Virtual Screening Strategies for the Discovery of Novel Compounds Active against Dengue Virus: A Hit Identification Study. Sci. Pharm. 2019, 88, 2. [Google Scholar] [CrossRef]
- Chiu, Y.-Y.; Tseng, J.-H.; Liu, K.-H.; Lin, C.-T.; Hsu, K.-C.; Yang, T.-P. Homopharma: A new concept for exploring the molecular binding mechanisms and drug repurposing. BMC Genom. 2014, 15, S8. [Google Scholar] [CrossRef] [PubMed]
- Lomenick, B.; Hao, R.; Jonai, N.; Chin, R.M.; Aghajan, M.; Warburton, S.; Wang, J.; Wu, R.P.; Gomez, F.; Loo, J.A.; et al. Target identification using drug affinity responsive target stability (DARTS). Proc. Natl. Acad. Sci. USA 2009, 106, 21984–21989. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Zhang, Q.; Chen, Z.; Xu, D.; Wang, Y. A network-based pathway-extending approach using DNA methylation and gene expression data to identify altered pathways. Sci. Rep. 2019, 9, 11853. [Google Scholar] [CrossRef] [PubMed]
- Yu, D.; Kim, M.S.; Xiao, G.; Hwang, T.H. Review of Biological Network Data and Its Applications. Genom. Informatics 2013, 11, 200–210. [Google Scholar] [CrossRef] [PubMed]
- Lo, Y.-C.; Senese, S.; Li, C.-M.; Hu, Q.; Huang, Y.; Damoiseaux, R.; Torres, J. Large-Scale Chemical Similarity Networks for Target Profiling of Compounds Identified in Cell-Based Chemical Screens. PLoS Comput. Boil. 2015, 11, e1004153. [Google Scholar] [CrossRef]
- Dey, L.; Mukhopadhyay, A. DenvInt: A database of protein–protein interactions between dengue virus and its hosts. PLOS Neglected Trop. Dis. 2017, 11, e0005879. [Google Scholar] [CrossRef] [PubMed]
- Plaimas, K.; Koenig, R. Identifying Antimalarial Drug Targets by Cellular Network Analysis. 2016. [Google Scholar]
- Kim, S. Getting the most out of PubChem for virtual screening. Expert Opin. Drug Discov. 2016, 11, 843–855. [Google Scholar] [CrossRef]
- Zhang, H.-Y. One-compound-multiple-targets strategy to combat Alzheimer’s disease. FEBS Lett. 2005, 579, 5260–5264. [Google Scholar] [CrossRef]
- Talevi, A. Multi-target pharmacology: Possibilities and limitations of the “skeleton key approach” from a medicinal chemist perspective. Front. Pharmacol. 2015, 6, 673. [Google Scholar] [CrossRef]
- Ramsay, R.R.; Nikolić, M.P.; Nikolic, K.M.; Uliassi, E.; Bolognesi, M.L. A perspective on multi-target drug discovery and design for complex diseases. Clin. Transl. Med. 2018, 7, 3. [Google Scholar] [CrossRef]
- Cregan, S.; McDonagh, L.; Gao, Y.; Barr, M.P.; O’Byrne, K.J.; Finn, S.; Cuffe, S.; Gray, S.G. KAT5 (Tip60) is a potential therapeutic target in malignant pleural mesothelioma. Int. J. Oncol. 2016, 48, 1290–1296. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Zhang, M.-G.; Wang, X.-J.; Zhong, S.; Shao, Y.; Zhu, Y.; Shen, Z.-J. KAT5 and KAT6B are in positive regulation on cell proliferation of prostate cancer through PI3K-AKT signaling. Int. J. Clin. Exp. Pathol. 2013, 6, 2864–2871. [Google Scholar] [PubMed]
- Feng, F.-L.; Yu, Y.; Liu, C.; Zhang, B.-H.; Cheng, Q.-B.; Li, B.; Tan, W.-F.; Luo, X.-J.; Jiang, X.-Q. KAT5 silencing induces apoptosis of GBC-SD cells through p38MAPK-mediated upregulation of cleaved Casp9. Int. J. Clin. Exp. Pathol. 2013, 7, 80–91. [Google Scholar] [PubMed]
- Keiser, M.J.; Roth, B.L.; Armbruster, B.N.; Ernsberger, P.; Irwin, J.J.; Shoichet, B.K. Relating protein pharmacology by ligand chemistry. Nat. Biotechnol. 2007, 25, 197–206. [Google Scholar] [CrossRef] [PubMed]
- Lindenbach, B.D.; M, C. Rice Molecular Biology of Flaviviruses, in Advances in Virus Research; Academic Press: Cambridge, MA, USA, 2003; pp. 23–61. [Google Scholar]
- Mukhopadhyay, S.; Kuhn, R.J.; Rossmann, M.G. A structural perspective of the flavivirus life cycle. Nat. Rev. Genet. 2005, 3, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Garcia, L.; Padilla-S, L.; Castaño, J.C. Inhibitors compounds of the flavivirus replication process. Virol. J. 2017, 14, 95. [Google Scholar] [CrossRef] [PubMed]
- Mishra, B.; Raghuraman, R.; Agarwal, A.; Aduri, R. Finding small molecules with pan-serotype activity to target Dengue non-structural protein 1. VirusDisease 2019, 30, 477–489. [Google Scholar] [CrossRef]
- Qamar, M.; Mumtaz, A.; Naseem, R.; Ali, A.; Fatima, T.; Jabbar, T.; Ahmad, Z.; Ashfaq, U.A. Molecular Docking Based Screening of Plant Flavonoids as Dengue NS1 Inhibitors. Bioinformation 2014, 10, 460–465. [Google Scholar] [CrossRef]
- Zhang, C.; Feng, T.; Cheng, J.; Li, Y.; Yin, X.; Zeng, W.; Jin, X.; Li, Y.; Guo, F.; Jin, T. Structure of the NS5 methyltransferase from Zika virus and implications in inhibitor design. Biochem. Biophys. Res. Commun. 2017, 492, 624–630. [Google Scholar] [CrossRef]
- El Sahili, A.; Lescar, J. Dengue Virus Non-Structural Protein 5. Viruses 2017, 9, 91. [Google Scholar] [CrossRef]
- Srivarangkul, P.; Yuttithamnon, W.; Suroengrit, A.; Pankaew, S.; Hengphasatporn, K.; Rungrotmongkol, T.; Phuwapraisirisan, P.; Ruxrungtham, K.; Boonyasuppayakorn, S.; Phuwapriasirisan, P. A novel flavanone derivative inhibits dengue virus fusion and infectivity. Antivir. Res. 2018, 151, 27–38. [Google Scholar] [CrossRef] [PubMed]
- Tambunan, U.S.F.; Zahroh, H.; Parikesit, A.A.; Idrus, S.; Kerami, D. Screening Analogs of β-OG Pocket Binder as Fusion Inhibitor of Dengue Virus 2. Drug Target Insights 2015, 9, 33–49. [Google Scholar] [CrossRef] [PubMed]
- Yennamalli, R.M.; Subbarao, N.; Kampmann, T.; McGeary, R.; Young, P.; Kobe, B. Identification of novel target sites and an inhibitor of the dengue virus E protein. J. Comput. Mol. Des. 2009, 23, 333–341. [Google Scholar] [CrossRef] [PubMed]
- Jadav, S.S.; Kaptein, S.; Timiri, A.; De Burghgraeve, T.; Badavath, V.N.; Ganesan, R.; Sinha, B.N.; Neyts, J.; Leyssen, P.; Jayaprakash, V. Design, synthesis, optimization and antiviral activity of a class of hybrid dengue virus E protein inhibitors. Bioorganic Med. Chem. Lett. 2015, 25, 1747–1752. [Google Scholar] [CrossRef] [PubMed]
- Ismail, N.A.; Jusoh, S.A. Molecular Docking and Molecular Dynamics Simulation Studies to Predict Flavonoid Binding on the Surface of DENV2 E Protein. Interdiscip. Sci. Comput. Life Sci. 2016, 9, 499–511. [Google Scholar] [CrossRef]
- Sharma, N.; Murali, A.; Singh, S.K.; Giri, R. Epigallocatechin gallate, an active green tea compound inhibits the Zika virus entry into host cells via binding the envelope protein. Int. J. Boil. Macromol. 2017, 104, 1046–1054. [Google Scholar] [CrossRef]
- Hengphasatporn, K.; Kungwan, N.; Rungrotmongkol, T. Binding pattern and susceptibility of epigallocatechin gallate against envelope protein homodimer of Zika virus: A molecular dynamics study. J. Mol. Liq. 2019, 274, 140–147. [Google Scholar] [CrossRef]
- Wirawan, M.; Fibriansah, G.; Marzinek, J.K.; Lim, X.X.; Ng, T.-S.; Sim, A.Y.; Zhang, Q.; Kostyuchenko, V.; Shi, J.; Smith, S.A.; et al. Mechanism of Enhanced Immature Dengue Virus Attachment to Endosomal Membrane Induced by prM Antibody. Struct. 2019, 27, 253–267.e8. [Google Scholar] [CrossRef]
- Yasunaga, A.; Hanna, S.L.; Li, J.; Cho, H.; Rose, P.P.; Spiridigliozzi, A.; Gold, B.; Diamond, M.S.; Cherry, S. Genome-Wide RNAi Screen Identifies Broadly-Acting Host Factors That Inhibit Arbovirus Infection. PLOS Pathog. 2014, 10, 1003914. [Google Scholar] [CrossRef]
- Yang, S.H.; Liu, M.L.; Tien, C.F.; Chou, S.J.; Chang, R.Y. Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) interaction with 3′ ends of Japanese encephalitis virus RNA and colocalization with the viral NS5 protein. J. Biomed. Sci. 2009, 16, 40. [Google Scholar] [CrossRef]
- Silva, E.M.; Conde, J.N.; Allonso, D.; Ventura, G.T.; Coelho, D.R.; Carneiro, P.H.; Silva, M.L.; Paes, M.V.; Rabelo, K.; Weissmuller, G.; et al. Dengue virus nonstructural 3 protein interacts directly with human glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and reduces its glycolytic activity. Sci. Rep. 2019, 9, 2651. [Google Scholar] [CrossRef] [PubMed]
- Raj, M.; Langley, M.; McArthur, S.J.; Jean, F. Moonlighting glycolytic enzyme glyceraldehyde-3-phosphate dehydrogenase (GAPDH) is required for efficient hepatitis C virus and dengue virus infections in human Huh-7.5.1 cells. J. Gen. Virol. 2017, 98, 977–991. [Google Scholar] [CrossRef] [PubMed]
- Yi, M.; Schultz, D.E.; Lemon, S.M. Functional Significance of the Interaction of Hepatitis A Virus RNA with Glyceraldehyde 3-Phosphate Dehydrogenase (GAPDH): Opposing Effects of GAPDH and Polypyrimidine Tract Binding Protein on Internal Ribosome Entry Site Function. J. Virol. 2000, 74, 6459–6468. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Xie, X.; Xia, H.; Zou, J.; Huang, L.; Popov, V.L.; Chen, X.; Shi, P.-Y. Zika Virus NS2A-Mediated Virion Assembly. mBio 2019, 10, e02375-19. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Gao, W.; Li, J.; Wu, W.; Jiu, Y. The Role of Host Cytoskeleton in Flavivirus Infection. Virol. Sin. 2019, 34, 30–41. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.E.; Cherupanakkal, C.; Catherine, M.; Kadhiravan, T.; Parameswaran, N.; Rajendiran, S.; Pillai, A.B. Endogenous gene selection for relative quantification PCR and IL6 transcript levels in the PBMC’s of severe and non-severe dengue cases. BMC Res. Notes 2018, 11, 550. [Google Scholar] [CrossRef]
- Srisutthisamphan, K.; Jirakanwisal, K.; Ramphan, S.; Tongluan, N.; Kuadkitkan, A.; Smith, D.R. Hsp90 interacts with multiple dengue virus 2 proteins. Sci. Rep. 2018, 8, 4308. [Google Scholar] [CrossRef]
- Kim, S.; Thiessen, P.A.; Bolton, E.; Chen, J.; Fu, G.; Gindulyte, A.; Han, L.; He, J.; He, S.; Shoemaker, B.A.; et al. PubChem Substance and Compound databases. Nucleic Acids Res. 2015, 44, D1202–D1213. [Google Scholar] [CrossRef]
- Bajusz, D.; Rácz, A.; Héberger, K. Why is Tanimoto index an appropriate choice for fingerprint-based similarity calculations? J. Chemin. 2015, 7, 20. [Google Scholar] [CrossRef]
- Cereto-Massagué, A.; Ojeda, M.J.; Valls, C.; Mulero, M.; Garcia-Vallvé, S.; Pujadas, G. Molecular fingerprint similarity search in virtual screening. Methods 2015, 71, 58–63. [Google Scholar] [CrossRef]
- Anastasiu, D.; Karypis, G. Efficient identification of Tanimoto nearest neighbors. Int. J. Data Sci. Anal. 2017, 4, 153–172. [Google Scholar] [CrossRef]
- Backman, T.; Cao, Y.; Girke, T. ChemMine tools: An online service for analyzing and clustering small molecules. Nucleic Acids Res. 2011, 39, W486–W491. [Google Scholar] [CrossRef] [PubMed]
- Liang, L.; Ma, C.; Du, T.; Zhao, Y.; Zhao, X.; Liu, M.; Wang, Z.; Lin, J. Bioactivity-explorer: A web application for interactive visualization and exploration of bioactivity data. J. Cheminform. 2019, 11, 47. [Google Scholar] [CrossRef] [PubMed]
- Bento, A.P.S.F.F.; Gaulton, A.; Hersey, A.; Bellis, L.; Chambers, J.; Davies, M.; A Kruger, F.; Light, Y.; Mak, L.; McGlinchey, S.; et al. The ChEMBL bioactivity database: An update. Nucleic Acids Res. 2013, 42, D1083–D1090. [Google Scholar] [CrossRef]
- Huang, R.; Xia, M. Editorial: Tox21 Challenge to Build Predictive Models of Nuclear Receptor and Stress Response Pathways As Mediated by Exposure to Environmental Toxicants and Drugs. Front. Environ. Sci. 2017, 5, 906. [Google Scholar] [CrossRef]
- Gilson, M.K.; Liu, T.; Baitaluk, M.; Nicola, G.; Hwang, L.; Chong, J. BindingDB in 2015: A public database for medicinal chemistry, computational chemistry and systems pharmacology. Nucleic Acids Res. 2015, 44, D1045–D1053. [Google Scholar] [CrossRef] [PubMed]
- Wadman, M. National prescription for drug development. Nat. Biotechnol. 2012, 30, 309–312. [Google Scholar] [CrossRef]
- Wang, Y.; Xiao, J.; Suzek, T.; Zhang, J.; Wang, J.; Zhou, Z.; Han, L.; Karapetyan, K.; Dracheva, S.; Shoemaker, B.A.; et al. PubChem’s BioAssay Database. Nucleic Acids Res. 2011, 40, D400–D412. [Google Scholar] [CrossRef]
- Jensen, L.J.; Kuhn, M.; Stark, M.; Chaffron, S.; Creevey, C.J.; Muller, J.; Doerks, T.; Julien, P.; Roth, A.; Simonovic, M.; et al. STRING 8–a global view on proteins and their functional interactions in 630 organisms. Nucleic Acids Res. 2008, 37, D412–D416. [Google Scholar] [CrossRef]
- Bastian, M.; Heymann, S.; Jacomy, M. Gephi: An Open Source Software for Exploring and Manipulating Networks. In Proceedings of the Third International AAAI Conference on Weblogs and Social Media, San Jose, CA, USA, 17–20 May 2009. [Google Scholar]
- Rodrigues, F.A. Network Centrality: An. Introduction, in A Mathematical Modeling Approach from Nonlinear Dynamics to Complex Systems; Macau, E.E.N., Ed.; Springer: Berlin/Heidelberg, Germany, 2019; pp. 177–196. [Google Scholar]
- Plaimas, K.; Eils, R.; König, R. Identifying essential genes in bacterial metabolic networks with machine learning methods. BMC Syst. Boil. 2010, 4, 56. [Google Scholar] [CrossRef]
- Yu, H.; Kim, P.M.; Sprecher, E.; Trifonov, V.; Gerstein, M. The Importance of Bottlenecks in Protein Networks: Correlation with Gene Essentiality and Expression Dynamics. PLoS Comput. Boil. 2007, 3, e59. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Wang, Q.; Wang, T. Drug Target Protein-Protein Interaction Networks: A Systematic Perspective. BioMed Res. Int. 2017, 2017, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Barabasi, A.-L.; Oltvai, Z.N. Network biology: Understanding the cell’s functional organization. Nat. Rev. Genet. 2004, 5, 101–113. [Google Scholar] [CrossRef] [PubMed]
- Kolountzakis, M.N.; Miller, G.; Peng, R.; Tsourakakis, C.E. Efficient Triangle Counting in Large Graphs via Degree-Based Vertex Partitioning. In Algorithms and Models for the Web-Graph; Springer: Berlin/Heidelberg, Germany, 2010. [Google Scholar]
- UniProt Consortium; The UniProt Consortium UniProt: A hub for protein information. Nucleic Acids Res. 2014, 43, D204–D212.
- Rappaport, N.; Nativ, N.; Stelzer, G.; Twik, M.; Guan-Golan, Y.; Stein, T.I.; Bahir, I.; Belinky, F.; Morrey, C.P.; Safran, M.; et al. MalaCards: An integrated compendium for diseases and their annotation. Database 2013, 2013, bat018. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples are not available from the authors. |


| Phenolic Lipid Compounds | Similar Compounds |
|---|---|
| Anacardic acid | 223 |
| Cardol | 311 |
| Cardanol | 447 |
| Property | DenvIntS Network (Whole Network) |
|---|---|
| Nodes | 488 |
| Edges | 2523 |
| Average degree | 10.340 |
| Nodes per edge | 0.193 |
| Diameter | 5 |
| Average clustering coefficient | 0.45 |
| Average path length | 2.842 |
| Graph density | 0.021 |
| Node Property (In Average) | All Nodes | Human Protein Nodes | DENV Protein Nodes |
|---|---|---|---|
| Degree | 10.340 | 9.142 | 67.600 |
| Eigencentrality | 0.116 | 0.113 | 0.238 |
| CC | 0.357 | 0.356 | 0.410 |
| BC | 0.004 | 0.002 | 0.108 |
| Clustering coefficient | 0.315 | 0.320 | 0.054 |
| Number of triangles | 83.195 | 82.797 | 102.200 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hengphasatporn, K.; Plaimas, K.; Suratanee, A.; Wongsriphisant, P.; Yang, J.-M.; Shigeta, Y.; Chavasiri, W.; Boonyasuppayakorn, S.; Rungrotmongkol, T. Target Identification Using Homopharma and Network-Based Methods for Predicting Compounds Against Dengue Virus-Infected Cells. Molecules 2020, 25, 1883. https://doi.org/10.3390/molecules25081883
Hengphasatporn K, Plaimas K, Suratanee A, Wongsriphisant P, Yang J-M, Shigeta Y, Chavasiri W, Boonyasuppayakorn S, Rungrotmongkol T. Target Identification Using Homopharma and Network-Based Methods for Predicting Compounds Against Dengue Virus-Infected Cells. Molecules. 2020; 25(8):1883. https://doi.org/10.3390/molecules25081883
Chicago/Turabian StyleHengphasatporn, Kowit, Kitiporn Plaimas, Apichat Suratanee, Peemapat Wongsriphisant, Jinn-Moon Yang, Yasuteru Shigeta, Warinthorn Chavasiri, Siwaporn Boonyasuppayakorn, and Thanyada Rungrotmongkol. 2020. "Target Identification Using Homopharma and Network-Based Methods for Predicting Compounds Against Dengue Virus-Infected Cells" Molecules 25, no. 8: 1883. https://doi.org/10.3390/molecules25081883
APA StyleHengphasatporn, K., Plaimas, K., Suratanee, A., Wongsriphisant, P., Yang, J.-M., Shigeta, Y., Chavasiri, W., Boonyasuppayakorn, S., & Rungrotmongkol, T. (2020). Target Identification Using Homopharma and Network-Based Methods for Predicting Compounds Against Dengue Virus-Infected Cells. Molecules, 25(8), 1883. https://doi.org/10.3390/molecules25081883







