Quinolinecarboxamides Inhibit the Replication of the Bovine Viral Diarrhea Virus by Targeting a Hot Spot for the Inhibition of Pestivirus Replication in the RNA-Dependent RNA Polymerase
Abstract
1. Introduction
2. Materials and Methods
2.1. Compounds
2.2. Cells and Viruses
2.3. Antiviral Assay
2.4. Cytotoxicity Assay
2.5. 3D QSAR Model
2.6. Isolation of TO502-2403 res /CSFCII res TO505-6180 res /CSFCIres BVDV
2.7. RNA Isolation
2.8. RT-qPCR
2.9. Sequencing
2.10. RNA-Dependent RNA Polymerase Assay
2.11. Replication Complex Assay
2.12. Molecular Modeling
3. Results
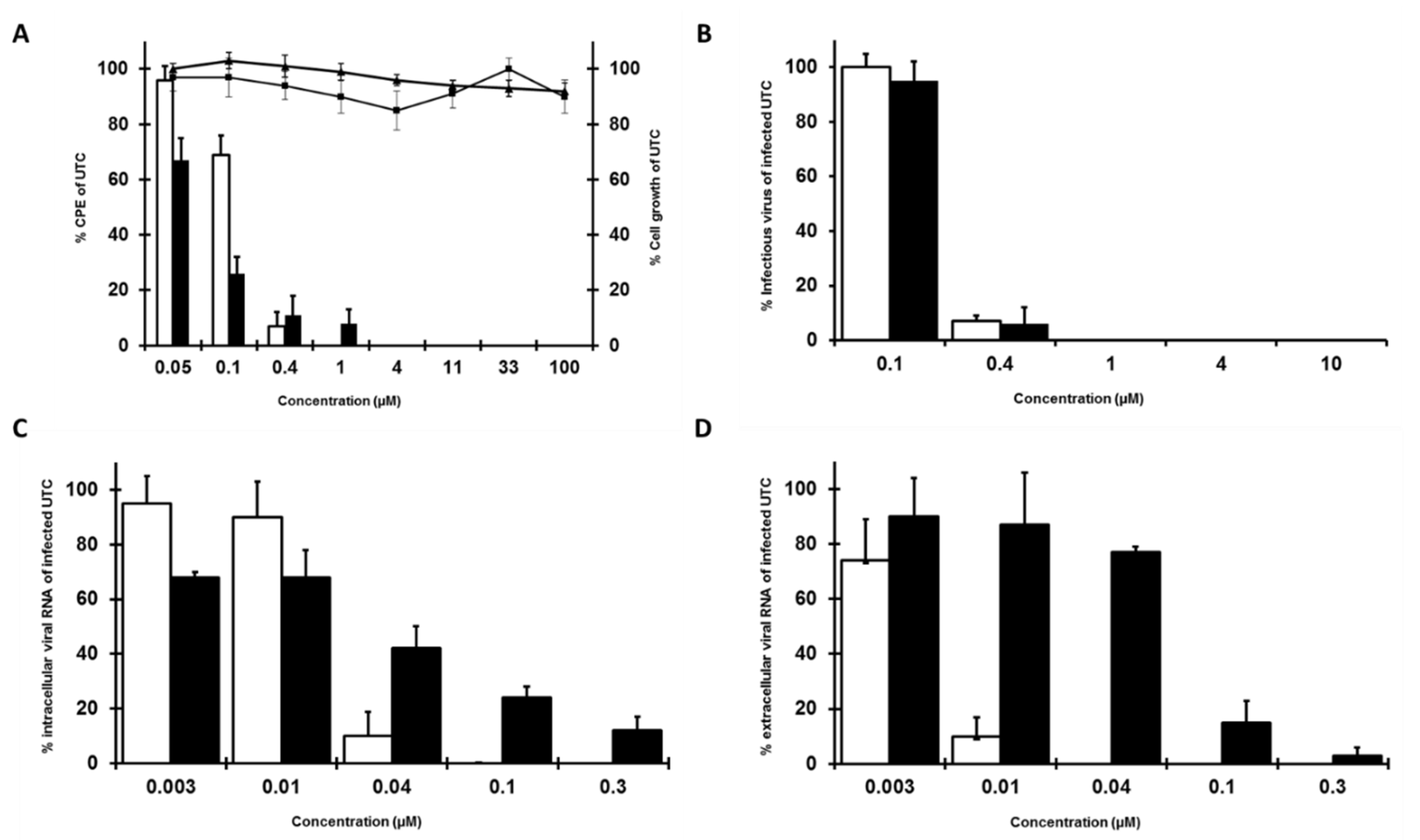
3.1. In Vitro Antiviral Activity of Quinolinecarboxamides
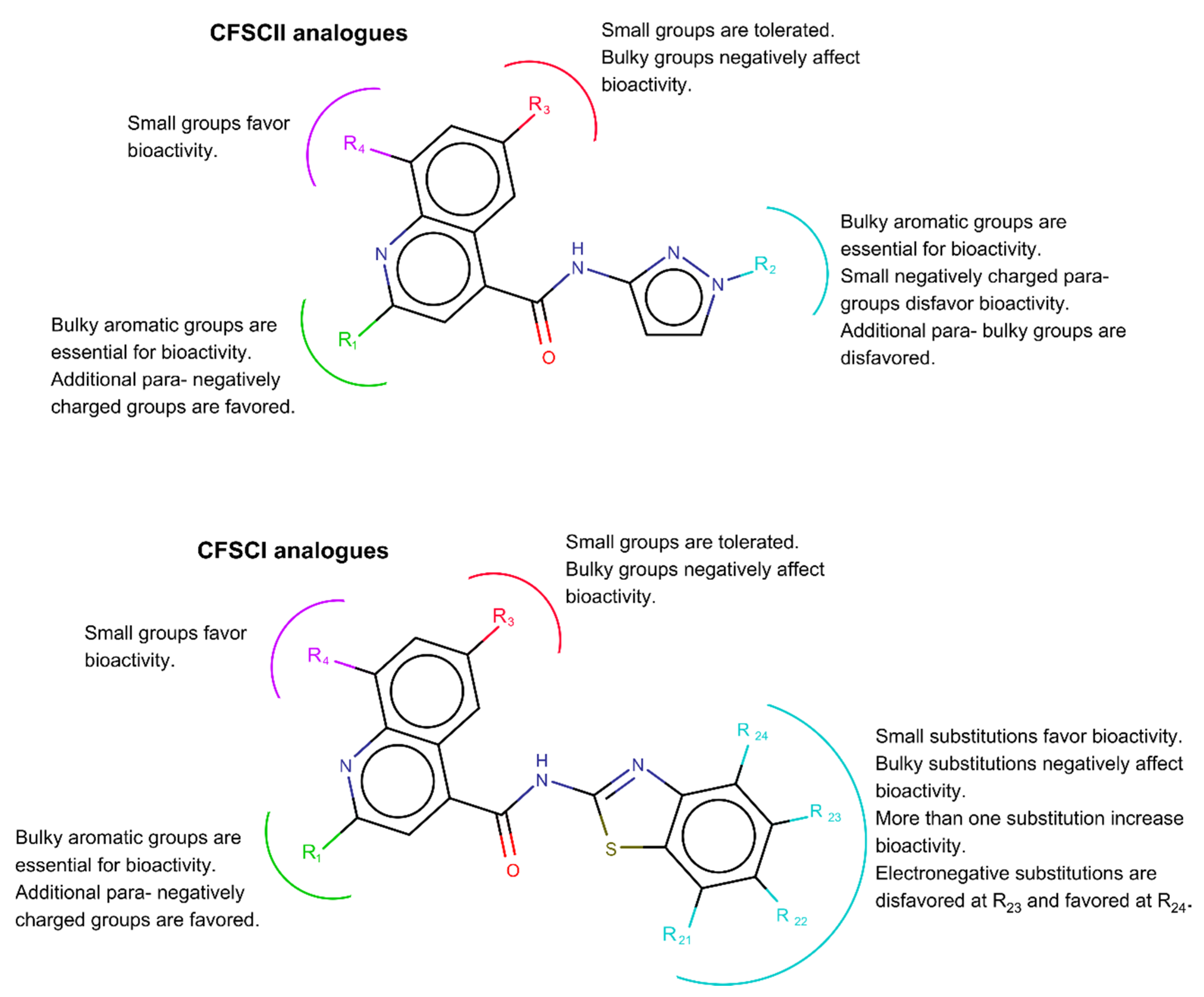
3.2. 3D-QSAR Analysis of All Tested Analogues of CSFC Lead Molecules
3.3. Isolation and Characterization of Drug-Resistant Viruses
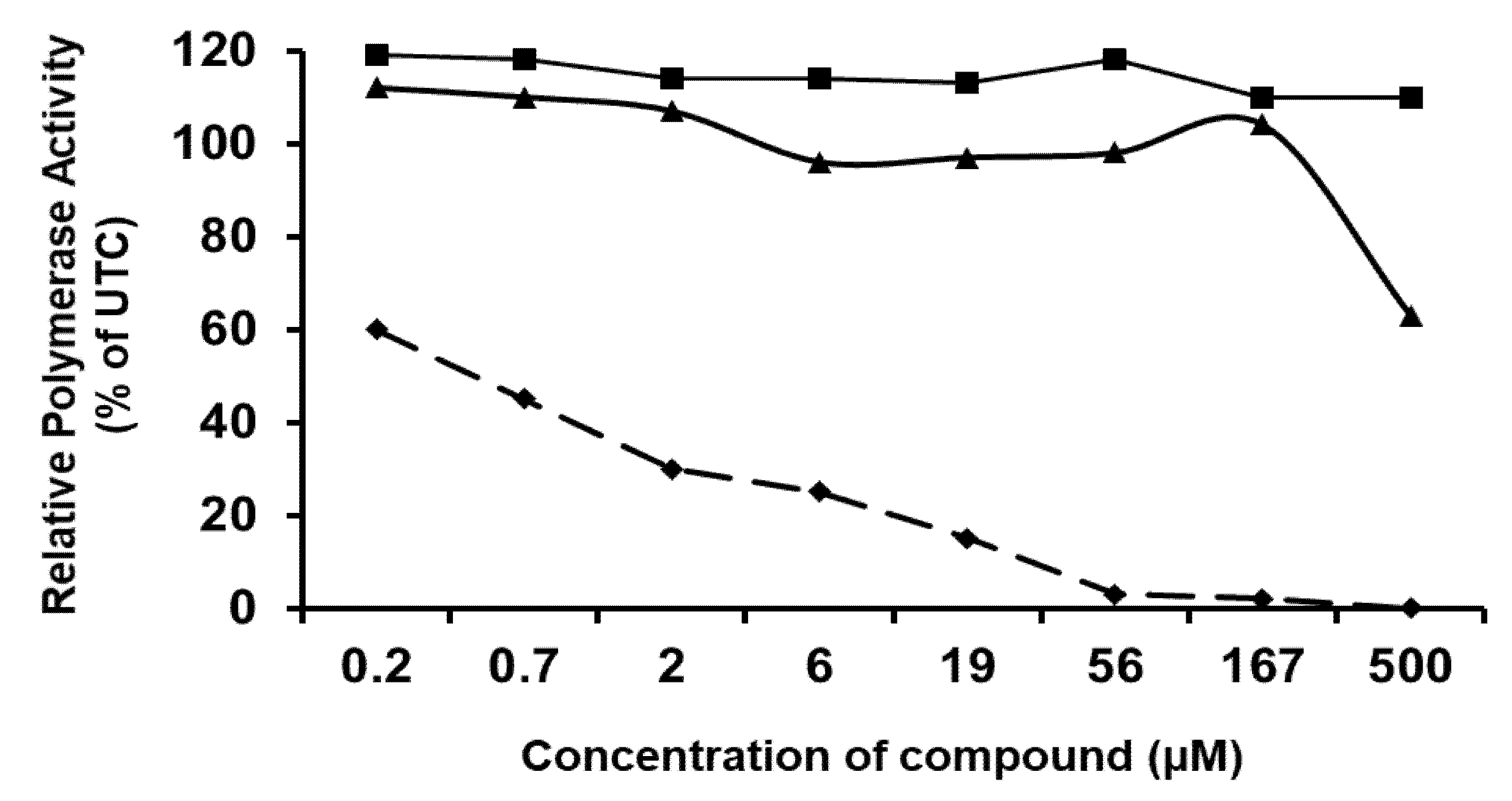
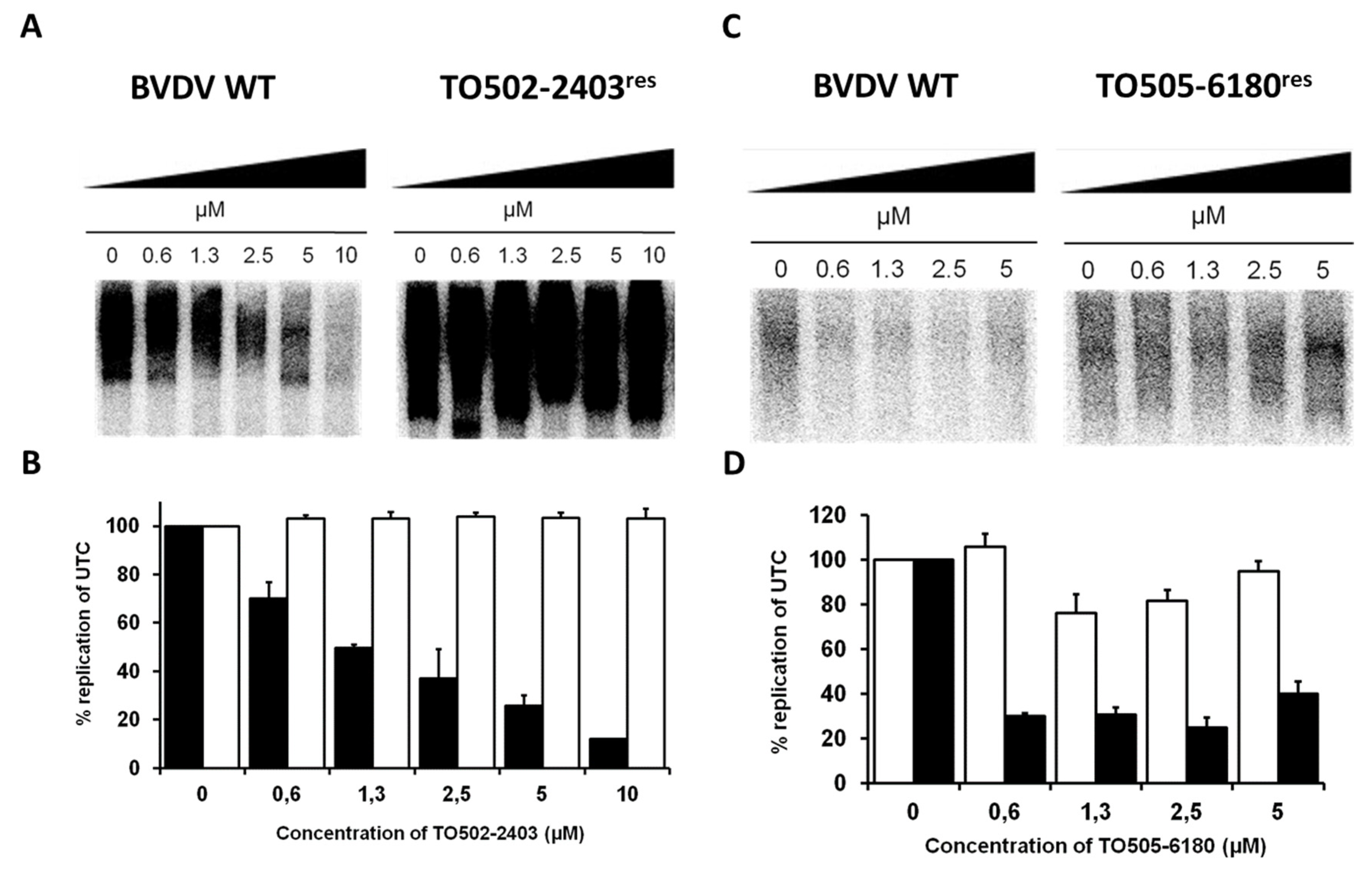
3.4. Effect of the CSFC Analogues on the BVDV RdRp and Replication Complexes
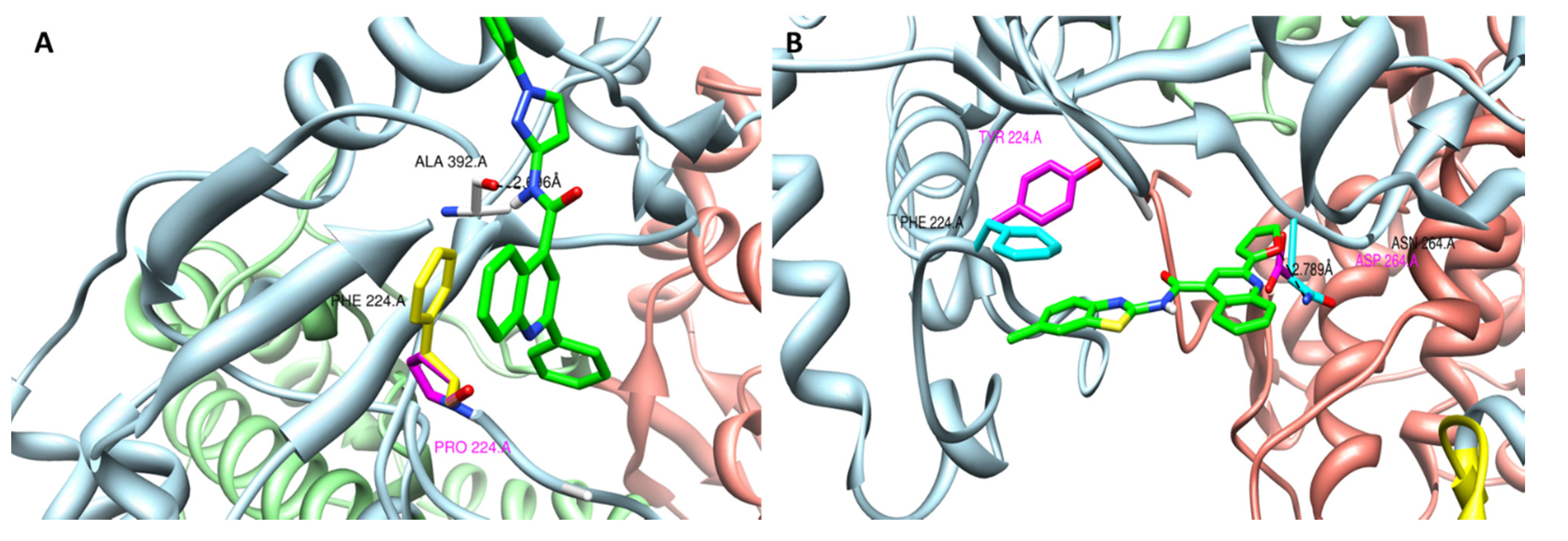
3.5. Computational Docking of CSFC Analogues in the BVDV RdRp Crystal Structure
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgements
Conflicts of Interest
References
- Hasler, B.; Howe, K.S.; Presi, P.; Stark, K.D. An economic model to evaluate the mitigation programme for bovine viral diarrhoea in Switzerland. Prev. Vet. Med. 2012, 106, 162–173. [Google Scholar] [CrossRef]
- Houe, H. Economic impact of, B.V.DV infection in dairies. Biologicals 2003, 31, 137–143. [Google Scholar] [CrossRef]
- Moennig, V.; Becher, P. Pestivirus control programs: How far have we come and where are we going? Anim. Health Res. Rev. 2015, 16, 83–87. [Google Scholar] [CrossRef] [PubMed]
- Colloff, A.; Watson, P.; Duff, J.P.; Scholes, S. Haemorrhagic disease in cattle with genotype 1 bovine viral diarrhoea virus infection. Vet. Rec. 2012, 171, 530. [Google Scholar] [CrossRef] [PubMed]
- Friedgut, O.; Rotenberg, D.; Brenner, J.; Yehuda, S.; Paz, R.; Alpert, N.; Ram, A.; Yadin, H.; Grummer, B. Description of the first acute bovine diarrhea virus-2 outbreak in Israel. Vet. J. 2011, 189, 108–110. [Google Scholar] [CrossRef] [PubMed]
- Hessman, B.E.; Sjeklocha, D.B.; Fulton, R.W.; Ridpath, J.F.; Johnson, B.J.; McElroy, D.R. Acute bovine viral diarrhea associated with extensive mucosal lesions, high morbidity, and mortality in a commercial feedlot. J. Vet. Diagn. Investig. 2012, 24, 397–404. [Google Scholar] [CrossRef]
- Tautz, N.; Tews, B.A.; Meyers, G. The Molecular Biology of Pestiviruses. Adv. Virus Res. 2015, 93, 47–160. [Google Scholar] [CrossRef]
- Postel, A.; Moennig, V.; Becher, P. Classical Swine Fever in Europe—The current Situation. Berl. Munch. Tierarztl. 2013, 126, 468–475. [Google Scholar] [CrossRef]
- Koenig, P.; Lange, E.; Reimann, I.; Beer, M. CP7_E2alf: A safe and efficient marker vaccine strain for oral immunisation of wild boar against Classical swine fever virus (CSFV). Vaccine 2007, 25, 3391–3399. [Google Scholar] [CrossRef]
- Von Rosen, T.; Rangelova, D.; Nielsen, J.; Rasmussen, T.B.; Uttenthal, A. DIVA vaccine properties of the live chimeric pestivirus strain, C.P.7_E2gif. Vet. Microbiol. 2014, 170, 224–231. [Google Scholar] [CrossRef]
- Stringfellow, D.A.; Riddell, K.P.; Givens, M.D.; Galik, P.K.; Sullivan, E.; Dykstra, C.C.; Robl, J.; Kasinathan, P. Bovine viral diarrhea virus (BVDV) in cell lines used for somatic cell cloning. Theriogenology 2005, 63, 1004–1013. [Google Scholar] [CrossRef] [PubMed]
- Durantel, D.; Carrouee-Durantel, S.; Branza-Nichita, N.; Dwek, R.A.; Zitzmann, N. Effects of interferon, ribavirin, and iminosugar derivatives on cells persistently infected with noncytopathic bovine viral diarrhea virus. Antimicrob. Agents Chemother. 2004, 48, 497–504. [Google Scholar] [CrossRef] [PubMed]
- Givens, M.D.; Stringfellow, D.A.; Dykstra, C.C.; Riddell, K.P.; Galik, P.K.; Sullivan, E.; Robl, J.; Kasinathan, P.; Kumar, A.; Boykin, D. Prevention and elimination of bovine viral diarrhea virus infections in fetal fibroblast cells. Antivir. Res. 2004, 64, 113–118. [Google Scholar] [CrossRef]
- Finkielsztein, L.M.; Moltrasio, G.Y.; Caputto, M.E.; Castro, E.F.; Cavallaro, L.V.; Moglioni, A.G. What is Known About the Antiviral Agents Active Against Bovine Viral Diarrhea Virus (BVDV)? Curr. Med. Chem. 2010, 17, 2933–2955. [Google Scholar] [CrossRef] [PubMed]
- Branza-Nichita, N.; Durantel, D.; Carrouee-Durantel, S.; Dwek, R.A.; Zitzmann, N. Antiviral effect of N-butyldeoxynojirimycin against bovine viral diarrhea virus correlates with misfolding of E2 envelope proteins and impairment of their association into E1-E2 heterodimers. J. Virol. 2001, 75, 3527–3536. [Google Scholar] [CrossRef] [PubMed]
- Bukhtiyarova, M.; Rizzo, C.J.; Kettner, C.A.; Korant, B.D.; Scarnati, H.T.; King, R.W. Inhibition of the bovine viral diarrhoea virus, NS.3 serine protease by a boron-modified peptidyl mimetic of its natural substrate. Antivir. Chem. Chemoth. 2001, 12, 367–373. [Google Scholar] [CrossRef]
- Baginski, S.G.; Pevear, D.C.; Seipel, M.; Sun, S.C.C.; Benetatos, C.A.; Chunduru, S.K.; Rice, C.M.; Collett, M.S. Mechanism of action of a pestivirus antiviral compound. Proc. Natl. Acad. Sci. USA 2000, 97, 7981–7986. [Google Scholar] [CrossRef]
- King, R.W.; Scarnati, H.T.; Priestley, E.S.; De Lucca, I.; Bansal, A.; Williams, J.K. Selection of a thiazole urea-resistant variant of bovine viral diarrhoea virus that maps to the, RNA-dependent, RNA polymerase. Antivir. Chem. Chemother. 2002, 13, 315–323. [Google Scholar] [CrossRef]
- Sun, J.H.; Lemm, J.A.; O’Boyle, D.R., 2nd.; Racela, J.; Colonno, R.; Gao, M. Specific inhibition of bovine viral diarrhea virus replicase. J. Virol. 2003, 77, 6753–6760. [Google Scholar] [CrossRef]
- Paeshuyse, J.; Leyssen, P.; Mabery, E.; Boddeker, N.; Vrancken, R.; Froeyen, M.; Ansari, I.H.; Dutartre, H.; Rozenski, J.; Gil, L.H.V.G.; et al. A novel, highly selective inhibitor of pestivirus replication that targets the viral, RNA-dependent, RNA polymerase. J. Virol. 2006, 80, 149–160. [Google Scholar] [CrossRef]
- Paeshuyse, J.; Chezal, J.M.; Froeyen, M.; Leyssen, P.; Dutartre, H.; Vrancken, R.; Canard, B.; Letellier, C.; Li, T.; Mittendorfer, H.; et al. The imidazopyrrolopyridine analogue, AG110 is a novel, highly selective inhibitor of pestiviruses that targets the viral, RNA-dependent, RNA polymerase at a hot spot for inhibition of viral replication. J. Virol. 2007, 81, 11046–11053. [Google Scholar] [CrossRef] [PubMed]
- Paeshuyse, J.; Letellier, C.; Froeyen, M.; Dutartre, H.; Vrancken, R.; Canard, B.; De Clercq, E.; Gueiffier, A.; Teulade, J.C.; Herdewijn, P.; et al. A pyrazolotriazolopyrimidinamine inhibitor of bovine viral diarrhea virus replication that targets the viral, RNA-dependent, RNA polymerase. Antivir. Res. 2009, 82, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Newcomer, B.W.; Marley, M.S.; Ridpath, J.F.; Neill, J.D.; Boykin, D.W.; Kumar, A.; Givens, M.D. Efficacy of an antiviral compound to inhibit replication of multiple pestivirus species. Antivir. Res. 2012, 96, 127–129. [Google Scholar] [CrossRef] [PubMed]
- Castro, E.F.; Fabian, L.E.; Caputto, M.E.; Gagey, D.; Finkielsztein, L.M.; Moltrasio, G.Y.; Moglioni, A.G.; Campos, R.H.; Cavallaro, L.V. Inhibition of bovine viral diarrhea virus, RNA synthesis by thiosemicarbazone derived from 5,6-dimethoxy-1-indanone. J. Virol. 2011, 85, 5436–5445. [Google Scholar] [CrossRef] [PubMed]
- Finkielsztein, L.M.; Castro, E.F.; Fabian, L.E.; Moltrasio, G.Y.; Campos, R.H.; Cavallaro, L.V.; Moglioni, A.G. New 1-indanone thiosemicarbazone derivatives active against, B.V.DV. Eur. J. Med. Chem. 2008, 43, 1767–1773. [Google Scholar] [CrossRef] [PubMed]
- Tonelli, M.; Boido, V.; La Colla, P.; Loddo, R.; Posocco, P.; Paneni, M.S.; Fermeglia, M.; Pricl, S. Pharmacophore modeling, resistant mutant isolation, docking, and, MM.-PBSA analysis: Combined experimental/computer-assisted approaches to identify new inhibitors of the bovine viral diarrhea virus (BVDV). Bioorg. Med. Chem. 2010, 18, 2304–2316. [Google Scholar] [CrossRef] [PubMed]
- Musiu, S.; Purstinger, G.; Stallinger, S.; Vrancken, R.; Haegeman, A.; Koenen, F.; Leyssen, P.; Froeyen, M.; Neyts, J.; Paeshuyse, J. Substituted 2,6-bis(benzimidazol-2-yl)pyridines: A novel chemical class of pestivirus inhibitors that targets a hot spot for inhibition of pestivirus replication in the, RNA-dependent, RNA polymerase. Antivir. Res. 2014, 106, 71–79. [Google Scholar] [CrossRef]
- Asthana, S.; Shukla, S.; Vargiu, A.V.; Ceccarelli, M.; Ruggerone, P.; Paglietti, G.; Marongiu, M.E.; Blois, S.; Giliberti, G.; La Colla, P. Different molecular mechanisms of inhibition of bovine viral diarrhea virus and hepatitis C virus, RNA-dependent, RNA polymerases by a novel benzimidazole. Biochemistry 2013, 52, 3752–3764. [Google Scholar] [CrossRef]
- Giliberti, G.; Ibba, C.; Marongiu, E.; Loddo, R.; Tonelli, M.; Boido, V.; Laurini, E.; Posocco, P.; Fermeglia, M.; Pricl, S. Synergistic experimental/computational studies on arylazoenamine derivatives that target the bovine viral diarrhea virus, RNA-dependent, RNA polymerase. Bioorg. Med. Chem. 2010, 18, 6055–6068. [Google Scholar] [CrossRef]
- Musiu, S.; Leyssen, P.; Froeyen, M.; Chezal, J.M.; Neyts, J.; Paeshuyse, J. 3-(imidazo[1,2-a:5,4-b’]dipyridin-2-yl)aniline inhibits pestivirus replication by targeting a hot spot drug binding pocket in the, R.N.A-dependent, RNA polymerase. Antivir. Res. 2016, 129, 99–103. [Google Scholar] [CrossRef]
- Vrancken, R.; Paeshuyse, J.; Haegeman, A.; Puerstinger, G.; Froeyen, M.; Herdewijn, P.; Kerkhofs, P.; Neyts, J.; Koenen, F. Imidazo[4,5-c]pyridines inhibit the in vitro replication of the classical swine fever virus and target the viral polymerase. Antivir. Res. 2008, 77, 114–119. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, P.C.D.; Skillman, A.G.; Warren, G.L.; Ellingson, B.A.; Stahl, M.T. OMEGA [Internet]. Santa Fe, NM: OpenEye Scientific Software. Available online: http://www.eyesopen.com (accessed on 31 January 2020).
- Tosco, P.; Balle, T.; Shiri, F. Open3DALIGN: An open-source software aimed at unsupervised ligand alignment. J. Comput. Aided Mol. Des. 2011, 25, 777–783. [Google Scholar] [CrossRef] [PubMed]
- Ponder, J.W.; Richards, F.M. An Efficient Newton-Like Method for Molecular Mechanics Energy Minimization of Large Molecules. J. Comput. Chem. 1987, 8, 1016–1024. [Google Scholar] [CrossRef]
- Cruz-Monteagudo, M.; Medina-Franco, J.L.; Perera-Sardina, Y.; Borges, F.; Tejera, E.; Paz, Y.M.C.; Pérez-Castillo, Y.; Sánchez-Rodríguez, A.; Contreras-Posada, Z.; Cordeiro, M.N. Probing the Hypothesis of, S.A.R Continuity Restoration by the Removal of Activity Cliffs Generators in, Q.S.AR. Curr. Pharm. Des. 2016, 22, 5043–5056. [Google Scholar] [CrossRef] [PubMed]
- Cruz-Monteagudo, M.; Medina-Francos, J.L.; Perez-Castillo, Y.; Nicolotti, O.; Cordeiro, M.N.; Borges, F. Activity cliffs in drug discovery: Dr Jekyll or Mr Hyde? Drug Discov. Today 2014, 19, 1069–1080. [Google Scholar] [CrossRef]
- Sud, M. MayaChemTools: An Open Source Package for Computational Drug Discovery. J. Chem. Inf. Model. 2016, 56, 2292–2297. [Google Scholar] [CrossRef]
- Tosco, P.; Balle, T. Open3DQSAR: A new open-source software aimed at high-throughput chemometric analysis of molecular interaction fields. J. Mol. Model. 2011, 17, 201–208. [Google Scholar] [CrossRef]
- Zhong, W.D.; Gutshall, L.L.; Del Vecchio, A.M. Identification and characterization of an, RNA-dependent, RNA polymerase activity within the nonstructural protein 5B region of bovine viral diarrhea virus. J. Virol. 1998, 72, 9365–9369. [Google Scholar] [CrossRef]
- Choi, K.H.; Groarke, J.M.; Young, D.C.; Kuhn, R.J.; Smith, J.L.; Pevear, D.C.; Rossmann, M.G. The structure of the, RNA-dependent, RNA polymerase from bovine viral diarrhea virus establishes the role of, G.T.P in de novo initiation. Proc. Natl. Acad. Sci. USA 2004, 101, 4425–4430. [Google Scholar] [CrossRef]
- Morris, G.M.; Goodsell, D.S.; Halliday, R.S.; Huey, R.; Hart, W.E.; Belew, R.K.; Olson, A.J. Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J. Comput. Chem. 1998, 19, 1639–1662. [Google Scholar] [CrossRef]
- Oh, J.W.; Ito, T.; Lai, M.M.C. A recombinant hepatitis C virus, RNA-dependent, RNA polymerase capable of copying the full-length viral, RNA. J. Virol. 1999, 73, 7694–7702. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.H.; Gallei, A.; Becher, P.; Rossmann, M.G. The structure of bovine viral diarrhea virus, RNA-dependent, RNA polymerase and its amino-terminal domain. Structure 2006, 14, 1107–1113. [Google Scholar] [CrossRef] [PubMed]
- Uttenthal, A.; Grondahl, C.; Hoyer, M.J.; Houe, H.; van Maanen, C.; Rasmussen, T.B.; Larsen, L.E. Persistent, B.V.DV infection in mousedeer infects calves—Do we know the reservoirs for, BVDV? Prev. Vet. Med. 2005, 72, 87–91. [Google Scholar] [CrossRef] [PubMed]
- Newcomer, B.W.; Neill, J.D.; Marley, M.S.; Ridpath, J.F.; Givens, M.D. Mutations induced in the, NS5B gene of bovine viral diarrhea virus by antiviral treatment convey resistance to the compound. Virus Res. 2013, 174, 95–100. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds are available from the authors. |
| Virus Strain and Mutations | EC50 (µM) of the Indicated Inhibitor a | ||||||
|---|---|---|---|---|---|---|---|
| LZ37 | BPIP | AG110 | BBP | TO502-2403 | TO505-6180 | 2′-C-Met-Cyt | |
| LZ37res F224P | 60 (9) 303x | >100 >1429x | |||||
| BPIPres F224S I261M | 78 (7) 391x | >100 >1429x | |||||
| AG110res A221K,R E291G | 3.7 (0.7) 19x | >100 >1429x | |||||
| BBPres I261M | 5 (1) 25x | >100 >1429x | |||||
| TO502-2403res F224P | >100 11x | 79 (5) 1580x | >100 63x | 60 (6) 200x | >100 >500x | >100 >1429x | 14 (2) |
| TO505-6180res F224Y N264D | >100 11x | 45 (3) 900x | 25 (4) 16x | >100 >333x | 32 (6) 160x | >100 >1429x | 2.5 ± 0.4 |
| Wild type (NADL) | 0.20 (0.06) | 0.07 (0.02) | |||||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Musiu, S.; Castillo, Y.P.; Muigg, A.; Pürstinger, G.; Leyssen, P.; Froeyen, M.; Neyts, J.; Paeshuyse, J. Quinolinecarboxamides Inhibit the Replication of the Bovine Viral Diarrhea Virus by Targeting a Hot Spot for the Inhibition of Pestivirus Replication in the RNA-Dependent RNA Polymerase. Molecules 2020, 25, 1283. https://doi.org/10.3390/molecules25061283
Musiu S, Castillo YP, Muigg A, Pürstinger G, Leyssen P, Froeyen M, Neyts J, Paeshuyse J. Quinolinecarboxamides Inhibit the Replication of the Bovine Viral Diarrhea Virus by Targeting a Hot Spot for the Inhibition of Pestivirus Replication in the RNA-Dependent RNA Polymerase. Molecules. 2020; 25(6):1283. https://doi.org/10.3390/molecules25061283
Chicago/Turabian StyleMusiu, Simone, Yunierkis Perez Castillo, Alexandra Muigg, Gerhard Pürstinger, Pieter Leyssen, Mathy Froeyen, Johan Neyts, and Jan Paeshuyse. 2020. "Quinolinecarboxamides Inhibit the Replication of the Bovine Viral Diarrhea Virus by Targeting a Hot Spot for the Inhibition of Pestivirus Replication in the RNA-Dependent RNA Polymerase" Molecules 25, no. 6: 1283. https://doi.org/10.3390/molecules25061283
APA StyleMusiu, S., Castillo, Y. P., Muigg, A., Pürstinger, G., Leyssen, P., Froeyen, M., Neyts, J., & Paeshuyse, J. (2020). Quinolinecarboxamides Inhibit the Replication of the Bovine Viral Diarrhea Virus by Targeting a Hot Spot for the Inhibition of Pestivirus Replication in the RNA-Dependent RNA Polymerase. Molecules, 25(6), 1283. https://doi.org/10.3390/molecules25061283






