Quinolones: Mechanism, Lethality and Their Contributions to Antibiotic Resistance
Abstract
1. Introduction
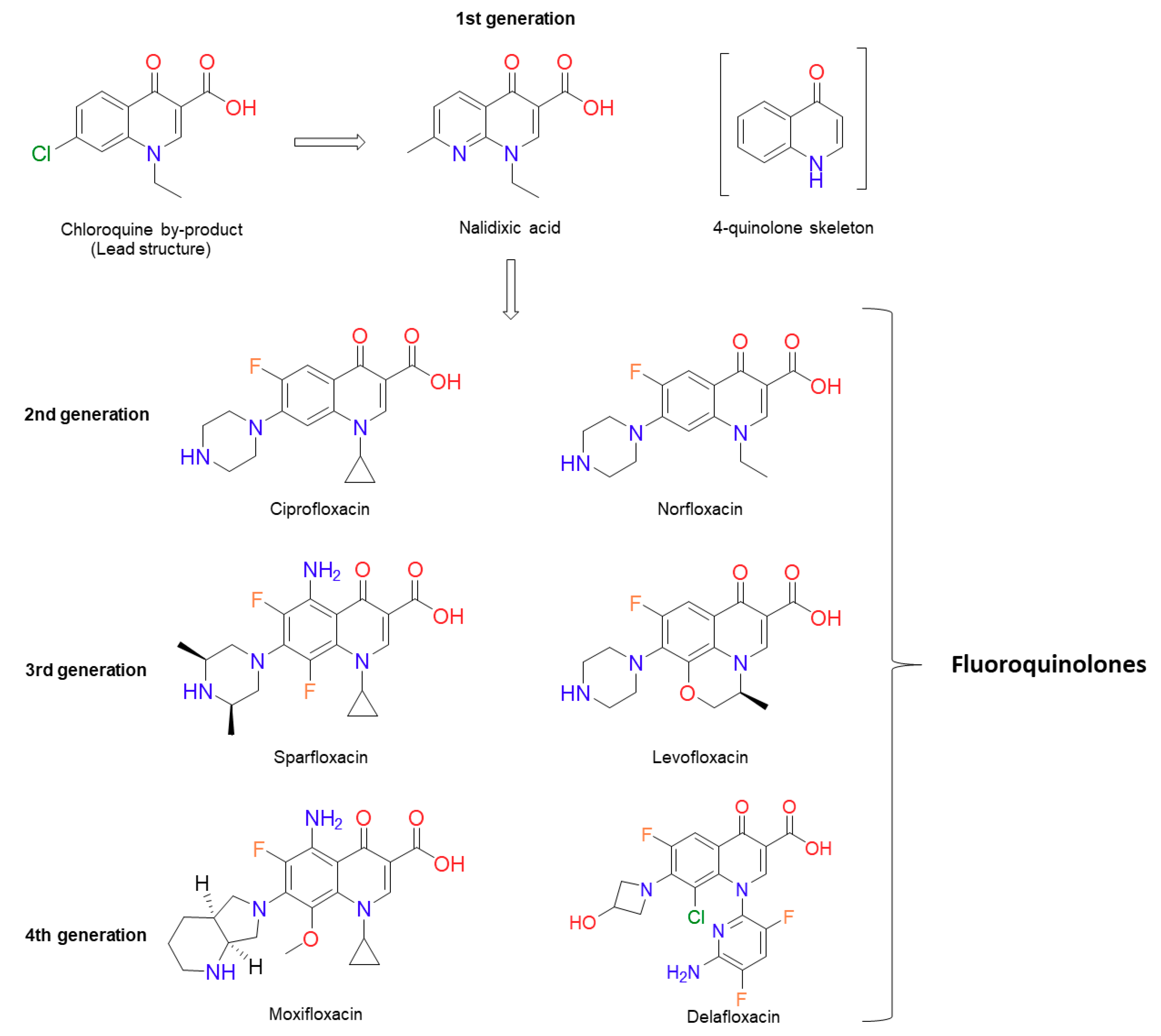
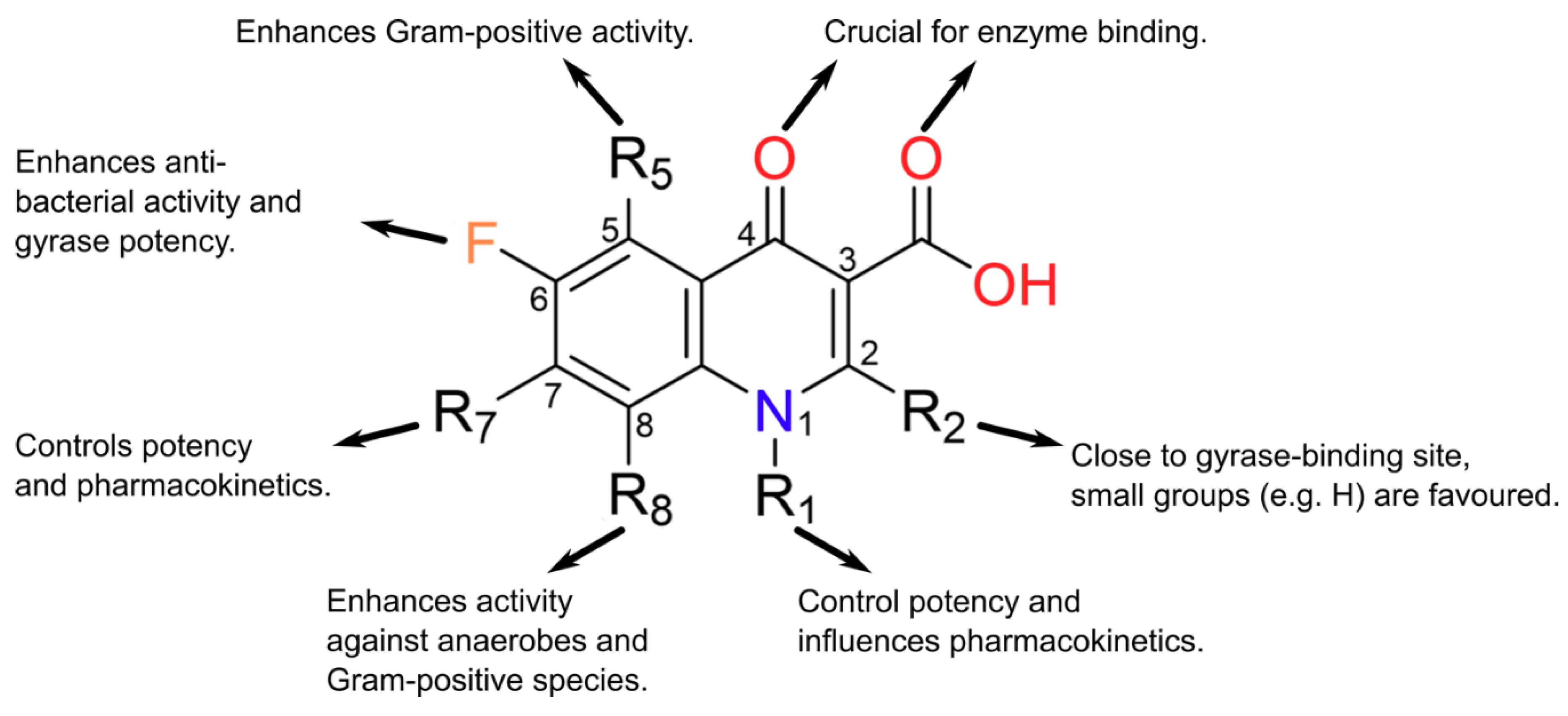
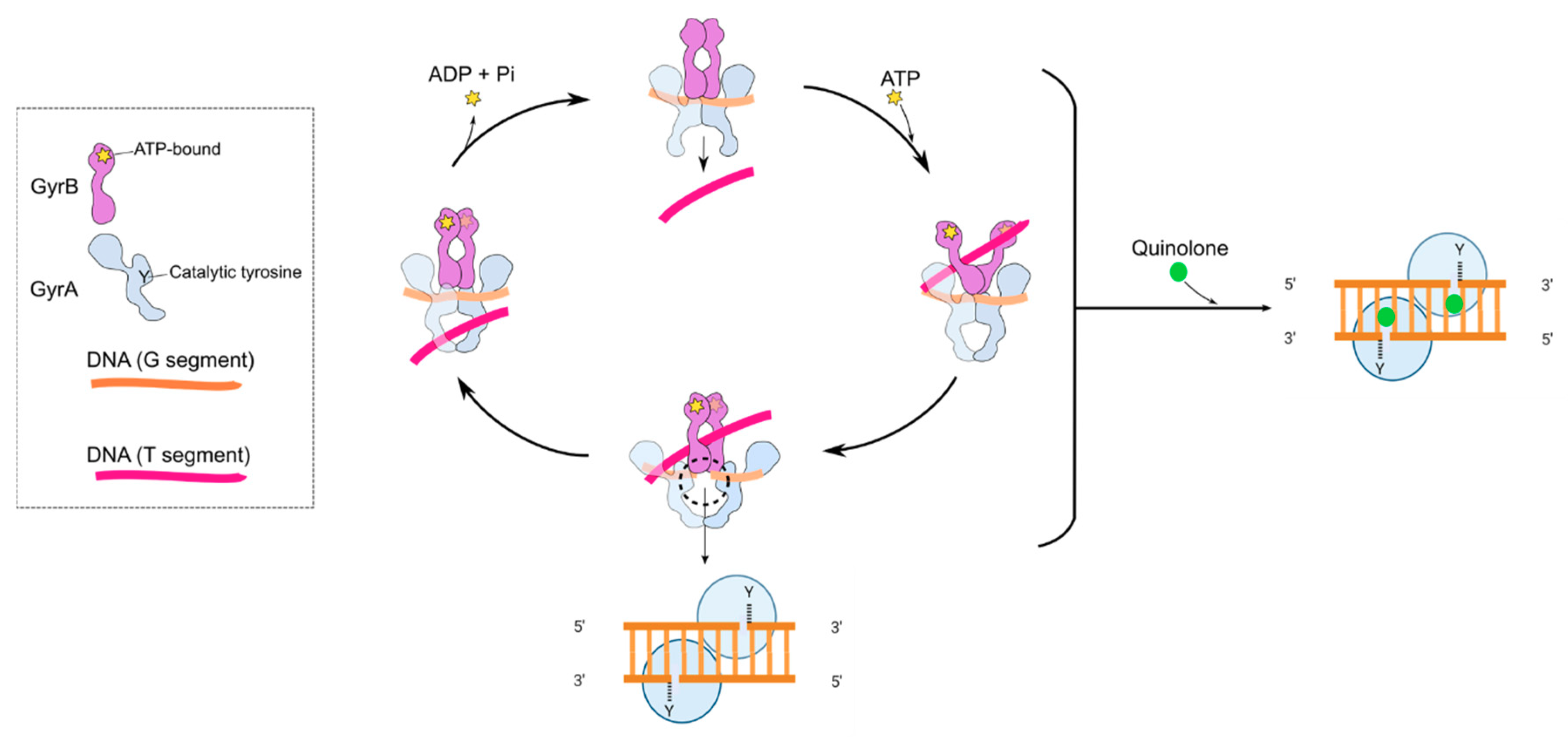
2. Background on Quinolones
3. Quinolone Lethality
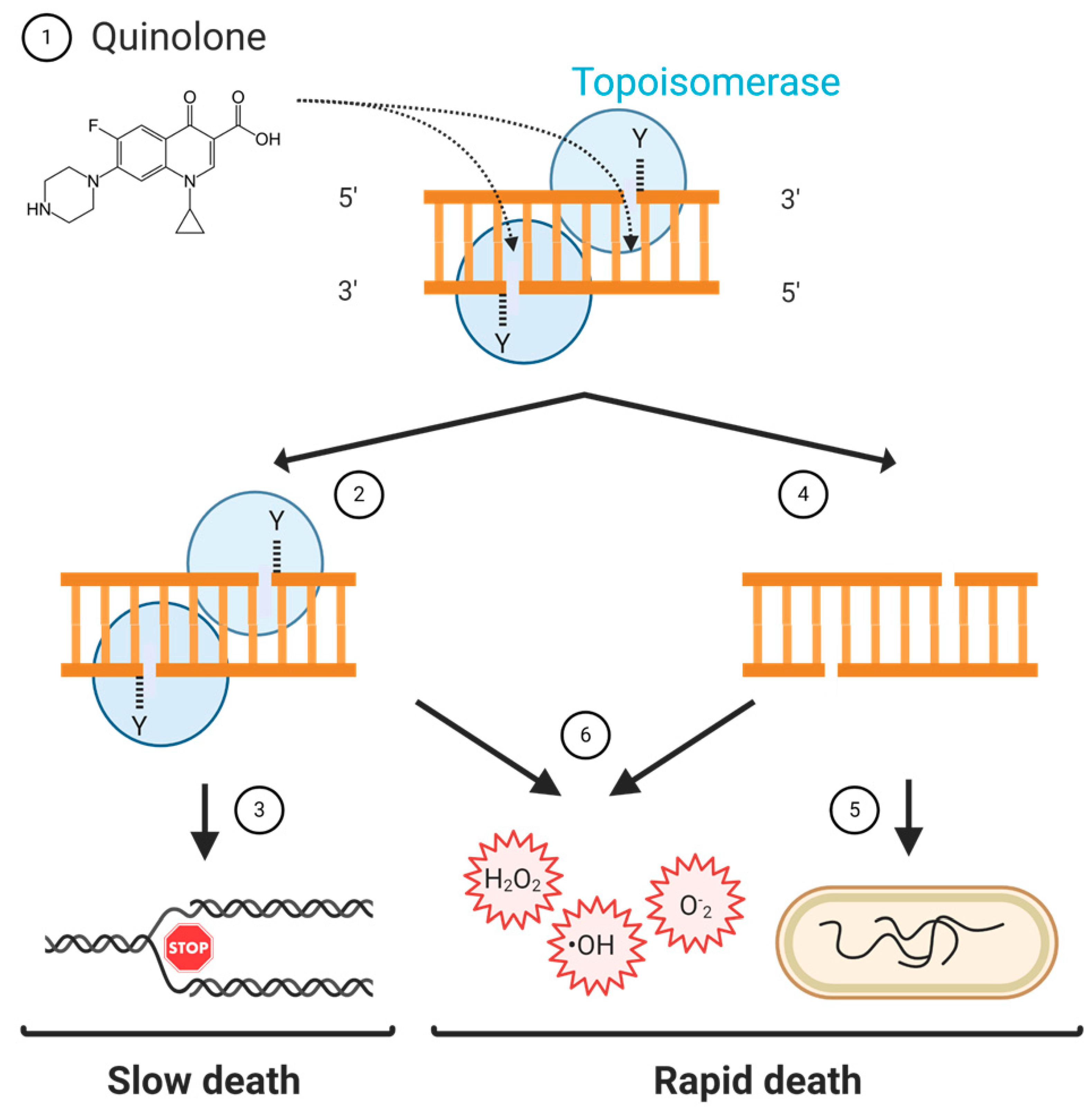
3.1. Slow Death
3.1.1. Block of Replication and Transcription
3.1.2. Inhibition of DNA and RNA Synthesis
3.2. Rapid Death
3.2.1. Processing of the Quinolone-Poisoned Gyrase
3.2.2. Nuclease and Protease Activity
3.2.3. Gyrase Subunit Dissociation
3.2.4. Chromosome Fragmentation
3.2.5. Reactive Oxygen Species (ROS) Formation
3.3. Repair of Quinolone-Induced DNA Damage
4. Resistance to Quinolones
4.1. Mutations in DNA Gyrase and Topo IV
4.2. Plasmid-Mediated Quinolone Resistance (PMQR)
4.3. Altered Drug Transport
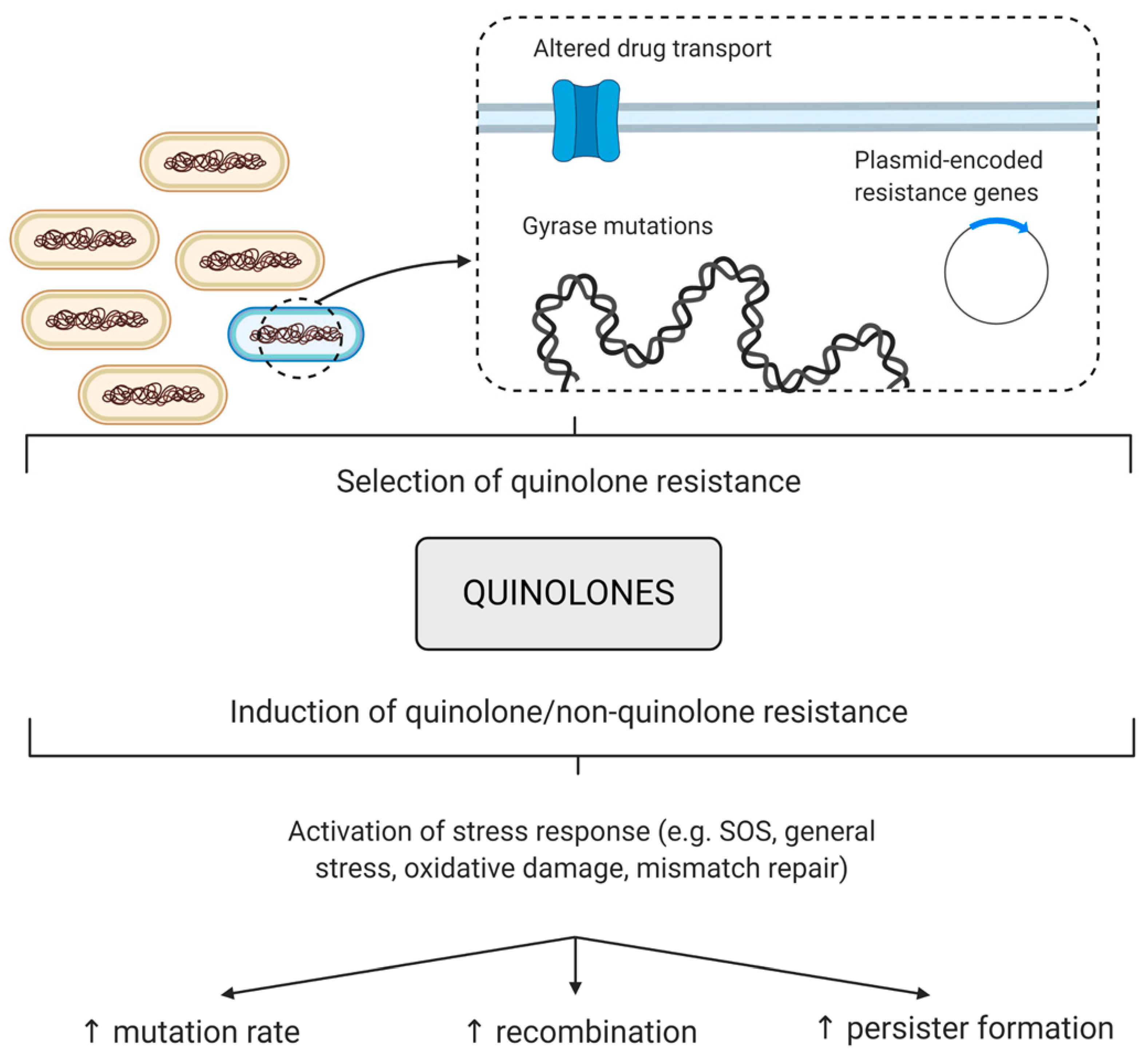
5. Quinolone-Induced AMR
5.1. Treatment with Quinolones Increases Resistance to Non-Quinolone Antibiotics (QIAR)
5.2. Sublethal FQ Treatment Increases Mutation Rate
5.3. Sublethal FQ Treatment Stimulates Recombination
5.4. Sublethal FQ Stimulates the Formation of Persisters
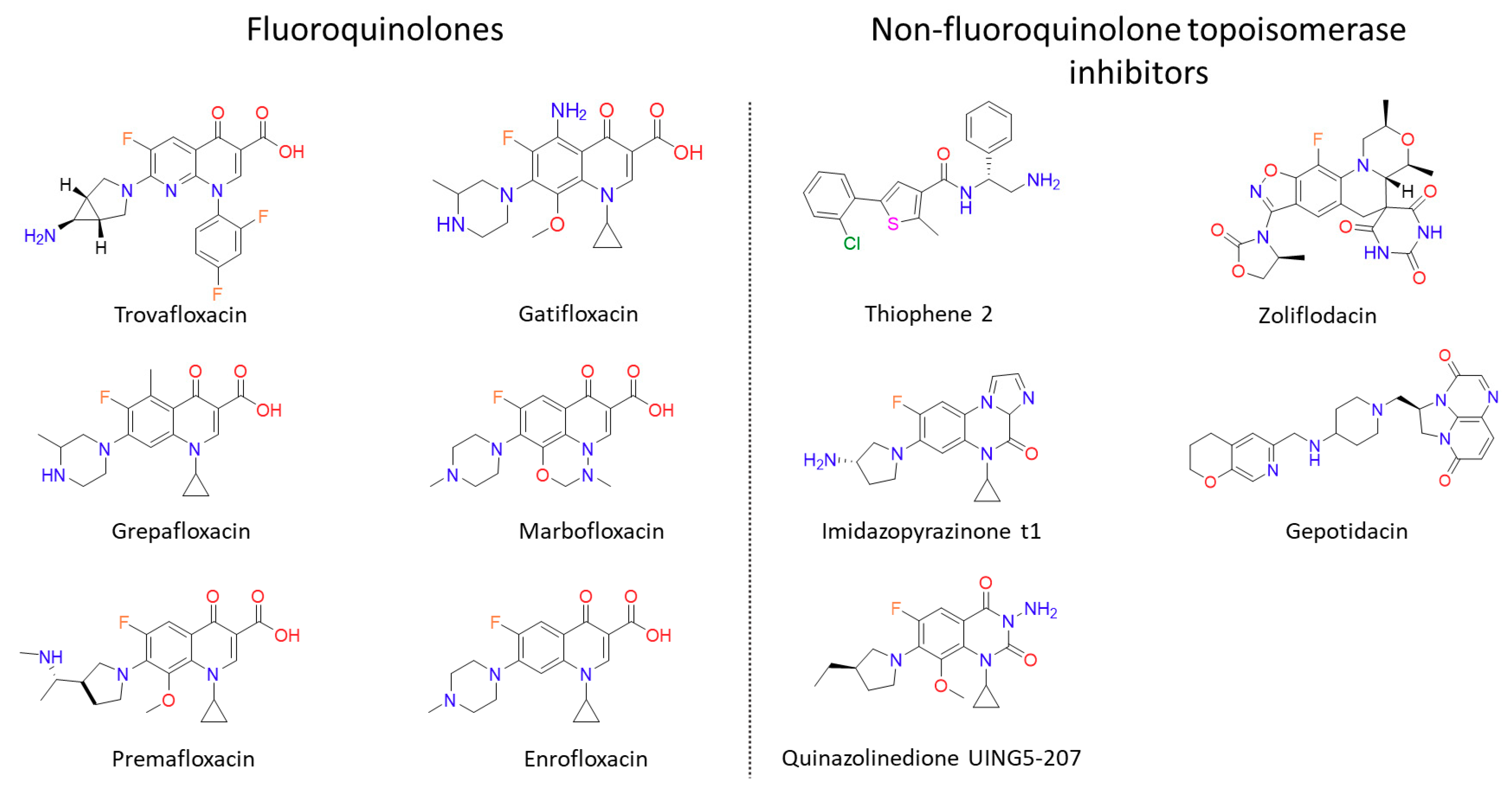
6. Future Prospects
Funding
Acknowledgments
Conflicts of Interest
References
- Lesher, G.Y.; Froelich, E.J.; Gruett, M.D.; Bailey, J.H.; Brundage, R.P. 1,8-Naphthyridone derivatives. A new class of chemotherapeutic agents. J. Med. Pharm. Chem. 1962, 91, 1063–1065. [Google Scholar] [CrossRef]
- Bisacchi, G.S. Origins of the quinolone class of antibacterials: An expanded “Discovery Story”. J. Med. Chem. 2015, 58, 4874–4882. [Google Scholar] [CrossRef]
- Gillespie, S.H. The role of moxifloxacin in tuberculosis therapy. ERR 2016, 25, 19–28. [Google Scholar] [CrossRef]
- Oliphant, C.M.; Green, G.M. Quinolones: A comprehensive review. Am. Fam. Physician 2002, 65, 455–464. [Google Scholar]
- Aldred, K.J.; Kerns, R.J.; Osheroff, N. Mechanism of quinolone action and resistance. Biochemistry 2014, 53, 1565–1574. [Google Scholar] [CrossRef]
- EMEA/H/A-31/1452. Quinolone-and Fluoroquinolone-Containing Medicinal Products; European Commision: Amsterdam, The Netherlands, 2018. [Google Scholar]
- Correia, S.; Poeta, P.; Hébraud, M.; Capelo, J.L.; Igrejas, G. Mechanisms of quinolone action and resistance: Where do we stand? J. Med. Microbiol. 2017, 66, 551–559. [Google Scholar] [CrossRef]
- Emmerson, A.M.; Jones, A.M. The quinolones: Decades of development and use. J. Antimicrob. Chemother. 2003, 51, 13–20. [Google Scholar] [CrossRef]
- Appelbaum, P.C.; Hunter, P.A. The fluoroquinolone antibacterials: Past, present and future perspectives. Int. J. Antimicrob. Agents 2000, 16, 5–15. [Google Scholar] [CrossRef]
- Collin, F.; Karkare, S.; Maxwell, A. Exploiting bacterial DNA gyrase as a drug target: Current state and perspectives. Appl. Microbiol. Biotechnol. 2011, 92, 479–497. [Google Scholar] [CrossRef]
- Andriole, V.T. The quinolones: Past, present, and future. Clin. Infect. Dis. 2005, 41, S113–S119. [Google Scholar] [CrossRef]
- Drlica, K.; Zhao, X.; Malik, M.; Hiasa, H.; Mustaev, A.; Kerns, R. Fluoroquinolone Resistance. In Bacterial Resistance to Antibiotics—From Molecules to Man; Bonev, B.B., Brown, N.M., Eds.; John Wiley & Sons: Hoboken, NJ, USA, 2019; pp. 125–161. [Google Scholar] [CrossRef]
- Ball, P. Quinolone generations: Natural history or natural selection? J. Antimicrob. Chemother. 2000, 46, 17–24. [Google Scholar] [CrossRef]
- Rohlfing, S.R.; Gerster, J.R.; Kvam, D.C. Bioevaluation of the antibacterial flumequine for urinary tract use. Antimicrob. Agents Chemother. 1976, 10, 20–24. [Google Scholar] [CrossRef]
- Ito, A.; Hirai, K.; Inoue, M.; Koga, H.; Suzue, S.; Irikura, T.; Mitsuhashi, S. In vitro antibacterial activity of AM-715, a new nalidixic acid analog. Antimicrob. Agents Chemother. 1980, 17, 103–108. [Google Scholar] [CrossRef]
- Wolfson, J.S.; Hooper, D.C. Norfloxacin: A new targeted fluoroquinolone antimicrobial agent. Ann. Intern. Med. 1988, 108, 238–251. [Google Scholar] [CrossRef]
- Wolfson, J.S.; Hooper, D.C. Fluoroquinolone antimicrobial agents. Clin. Microbiol. Rev. 1989, 2, 378–424. [Google Scholar] [CrossRef]
- Van Bambeke, F.; Michot, J.M.; Van Eldere, J.; Tulkens, P.M. Quinolones in 2005: An update. Clin. Microbiol. Infect. 2005, 11, 256–280. [Google Scholar] [CrossRef]
- Adhami, Z.N.; Wise, R.; Weston, D.; Crump, B. The pharmacokinetics and tissue penetration of norfloxacin. J. Antimicrob. Chemother. 1984, 13, 87–92. [Google Scholar] [CrossRef]
- Aznar, J.; Caballero, M.C.; Lozano, M.C.; de Miguel, C.; Palomares, J.C.; Perea, E.J. Activities of new quinoline derivatives against genital pathogens. Antimicrob. Agents Chemother. 1985, 27, 76–78. [Google Scholar] [CrossRef]
- Holmes, B.; Brogden, R.N.; Richards, D.M. Norfloxacin. Drugs 1985, 30, 482–513. [Google Scholar] [CrossRef]
- Hooper, D.C.; Wolfson, J.S.; Souza, K.S.; Tung, C.; McHugh, G.L.; Swartz, M.N. Genetic and biochemical characterization of norfloxacin resistance in Escherichia coli. Antimicrob. Agents Chemother. 1986, 29, 639–644. [Google Scholar] [CrossRef]
- Norrby, S.R.; Jonsson, M. Antibacterial activity of norfloxacin. Antimicrob. Agents Chemother. 1983, 23, 15–18. [Google Scholar] [CrossRef]
- Schacht, P.; Chyský, V.; Gruenwaldt, G.; Hullmann, R.; Weuta, H.; Arcieri, G.; Griffith, E.; O’Brien, B.; Branolte, J.; Bruck, H.; et al. Worldwide clinical data on efficacy and safety of ciprofloxacin. Infection 1988, 16, S29–S43. [Google Scholar] [CrossRef]
- Segev, S.; Yaniv, I.; Haverstock, D.; Reinhart, H. Safety of long-term therapy with ciprofloxacin: Data analysis of controlled clinical trials and review. Clin. Infect. Dis. 1999, 28, 299–308. [Google Scholar] [CrossRef]
- World Health Organization. WHO Model List of Essential Medicines, 20th List (March 2017, Amended August 2017); World Health Organization: Geneva, Switzerland, 2017. [Google Scholar]
- Mitscher, L.A. Bacterial topoisomerase inhibitors: Quinolone and pyridone antibacterial agents. Chem. Rev. 2005, 105, 559–592. [Google Scholar] [CrossRef]
- Ahuja, S.D.; Ashkin, D.; Avendano, M.; Banerjee, R.; Bauer, M.; Bayona, J.N.; Becerra, M.C.; Benedetti, A.; Burgos, M.; Centis, R.; et al. Multidrug resistant pulmonary tuberculosis treatment regimens and patient outcomes: An individual patient data meta-analysis of 9153 patients. PLoS Med. 2012, 9, e1001300. [Google Scholar] [CrossRef]
- Lucena, M.I.; Andrade, R.J.; Rodrigo, L.; Salmerón, J.; Alvarez, A.; Lopez-Garrido, M.J.; Camargo, R.; Alcantára, R. Trovafloxacin-induced acute hepatitis. Clin. Infect. Dis. 2000, 30, 400–401. [Google Scholar] [CrossRef]
- Mandell, L.; Tillotson, G. Safety of fluoroquinolones: An update. Can. J. Infect. Dis. 2002, 13, 54–61. [Google Scholar] [CrossRef]
- WHO. Critically Important Antimicrobials for Human Medicine, 5th ed.; World Health Organisation—WHO Advisory Group on Integrated Surveillance of Antimicrobial Resistance (AGISAR): Geneva, Switzerland, 2017; p. 48. [Google Scholar]
- Domagala, J.M. Structure-activity and structure-side-effect relationships for the quinolone antibacterials. J. Antimicrob. Chemother. 1994, 33, 685–706. [Google Scholar] [CrossRef]
- Gellert, M.; Mizuuchi, K.; O’Dea, M.H.; Itoh, T.; Tomizawa, J.-I. Nalidixic acid resistance: A second genetic character involved in DNA gyrase activity. Proc. Natl. Acad. Sci. USA 1977, 74, 4772–4776. [Google Scholar] [CrossRef]
- Sugino, A.; Peebles, C.L.; Kreuzer, K.N.; Cozzarelli, N.R. Mechanism of action of nalidixic acid: Purification of Escherichia coli nalA gene product and its relationship to DNA gyrase and a novel nicking-closing enzyme. Proc. Natl. Acad. Sci. USA 1977, 74, 4767–4771. [Google Scholar] [CrossRef]
- Bush, N.G.; Evans-Roberts, K.; Maxwell, A. DNA Topoisomerases. EcoSal Plus 2015, 6. [Google Scholar] [CrossRef]
- Bates, A.D.; Maxwell, A. DNA Topology, 2nd ed.; Oxford University Press: New York, NY, USA, 2005; p. 216. [Google Scholar]
- Gellert, M.; Mizuuchi, K.; O’Dea, M.H.; Nash, H.A. DNA gyrase: An enzyme that introduces superhelical turns into DNA. Proc. Natl. Acad. Sci. USA 1976, 73, 3872–3876. [Google Scholar] [CrossRef]
- Wang, J.C. DNA Topoisomerases. Annu. Rev. Biochem. 1996, 65, 635–692. [Google Scholar] [CrossRef]
- Berger, J.M. Type II DNA topoisomerases. Curr. Opin. Struct. Biol. 1998, 8, 26–32. [Google Scholar] [CrossRef]
- Dar, M.A.; Sharma, A.; Mondal, N.; Dhar, S.K. Molecular cloning of apicoplast-targeted Plasmodium falciparum DNA gyrase genes: Unique intrinsic ATPase activity and ATP-independent dimerization of PfGyrB subunit. Eukaryot. Cell 2007, 6, 398–412. [Google Scholar] [CrossRef]
- Wall, M.K.; Mitchenall, L.A.; Maxwell, A. Arabidopsis thaliana DNA gyrase is targeted to chloroplasts and mitochondria. Proc. Natl. Acad. Sci. USA 2004, 101, 7821–7826. [Google Scholar] [CrossRef]
- Corbett, K.D.; Berger, J.M. Structure, molecular mechanisms, and evolutionary relationships in DNA topoisomerases. Annu. Rev. Biophys. Biomol. Struct. 2004, 33, 95–118. [Google Scholar] [CrossRef]
- Roca, J.; Wang, J.C. The capture of a DNA double helix by an ATP-dependent protein clamp: A key step in DNA transport by type II DNA topoisomerases. Cell 1992, 71, 833–840. [Google Scholar] [CrossRef]
- Wang, J. Moving one DNA double helix through another by a type II DNA topoisomerase: The story of a simple molecular machine. Q. Rev. Biophys. 1998, 31, 107–144. [Google Scholar] [CrossRef]
- Drlica, K.; Zhao, X. DNA gyrase, topoisomerase IV, and the 4-quinolones. Microbiol. Mol. Biol. Rev. 1997, 61, 377–392. [Google Scholar] [CrossRef]
- Hooper, D.C.; Jacoby, G.A. Topoisomerase inhibitors: Fluoroquinolone mechanisms of action and resistance. Cold Spring Harb. Perspect. Med. 2016, 6, a025320. [Google Scholar] [CrossRef]
- Ferrero, L.; Cameron, B.; Manse, B.; Lagneaux, D.; Crouzet, J.; Famechon, A.; Blanche, F. Cloning and primary structure of Staphylococcus aureus DNA topoisomerase IV: A primary target of fluoroquinolones. Mol. Microbiol. 1994, 13, 641–653. [Google Scholar] [CrossRef]
- Khodursky, A.B.; Zechiedrich, E.L.; Cozzarelli, N.R. Topoisomerase IV is a target of quinolones in Escherichia coli. Proc. Natl. Acad. Sci. USA 1995, 92, 11801–11805. [Google Scholar] [CrossRef]
- Redgrave, L.S.; Sutton, S.B.; Webber, M.A.; Piddock, L.J. Fluoroquinolone resistance: Mechanisms, impact on bacteria, and role in evolutionary success. Trends Microbiol. 2014, 22, 438–445. [Google Scholar] [CrossRef]
- Cheng, G.; Hao, H.; Dai, M.; Liu, Z.; Yuan, Z. Antibacterial action of quinolones: From target to network. Eur. J. Med. Chem. 2013, 66, 555–562. [Google Scholar] [CrossRef]
- Drlica, K.; Hiasa, H.; Kerns, R.; Malik, M.; Mustaev, A.; Zhao, X. Quinolones: Action and resistance updated. Curr. Top. Med. Chem. 2009, 9, 981–998. [Google Scholar] [CrossRef]
- Mustaev, A.; Malik, M.; Zhao, X.; Kurepina, N.; Luan, G.; Oppegard, L.M.; Hiasa, H.; Marks, K.R.; Kerns, R.J.; Berger, J.M.; et al. Fluoroquinolone-gyrase-DNA complexes: Two modes of drug binding. J. Biol. Chem. 2014. [Google Scholar] [CrossRef]
- Bax, B.D.; Chan, P.F.; Eggleston, D.S.; Fosberry, A.; Gentry, D.R.; Gorrec, F.; Giordano, I.; Hann, M.M.; Hennessy, A.; Hibbs, M.; et al. Type IIA topoisomerase inhibition by a new class of antibacterial agents. Nature 2010, 466, 935–940. [Google Scholar] [CrossRef]
- Laponogov, I.; Sohi, M.K.; Veselkov, D.A.; Pan, X.-S.; Sawhney, R.; Thompson, A.W.; McAuley, K.E.; Fisher, L.M.; Sanderson, M.R. Structural insight into the quinolone–DNA cleavage complex of type IIA topoisomerases. Nat. Struct. Mol. Biol. 2009, 16, 667–669. [Google Scholar] [CrossRef]
- Wohlkonig, A.; Chan, P.F.; Fosberry, A.P.; Homes, P.; Huang, J.; Kranz, M.; Leydon, V.R.; Miles, T.J.; Pearson, N.D.; Perera, R.L.; et al. Structural basis of quinolone inhibition of type IIA topoisomerases and target-mediated resistance. Nat. Struct. Mol. Biol. 2010, 17, 1152–1153. [Google Scholar] [CrossRef]
- Tornaletti, S.; Pedrini, A.M. Studies on the interaction of 4-quinolones with DNA by DNA unwinding experiments. Biochim. Biophys. Acta 1988, 949, 279–287. [Google Scholar] [CrossRef]
- Tornaletti, S.; Pedrini, A.M. DNA unwinding induced by nalidixic acid binding to DNA. Biochem. Pharmacol. 1988, 37, 1881–1882. [Google Scholar] [CrossRef]
- Shen, L.L. A reply: “Do quinolones bind to DNA?”—Yes. Biochem. Pharmacol. 1989, 38, 2042–2044. [Google Scholar] [CrossRef]
- Aldred, K.J.; Breland, E.J.; Vlckova, V.; Strub, M.P.; Neuman, K.C.; Kerns, R.J.; Osheroff, N. Role of the water-metal ion bridge in mediating interactions between quinolones and Escherichia coli topoisomerase IV. Biochemistry 2014, 53, 5558–5567. [Google Scholar] [CrossRef] [PubMed]
- Aldred, K.J.; McPherson, S.A.; Turnbough, C.L., Jr.; Kerns, R.J.; Osheroff, N. Topoisomerase IV-quinolone interactions are mediated through a water-metal ion bridge: Mechanistic basis of quinolone resistance. Nucleic Acids Res. 2013, 41, 4628–4639. [Google Scholar] [CrossRef] [PubMed]
- Hooper, D.C.; Jacoby, G.A. Mechanisms of drug resistance: Quinolone resistance. Ann. N. Y. Acad. Sci. 2015, 1354, 12–31. [Google Scholar] [CrossRef]
- Snyder, M.; Drlica, K. DNA gyrase on the bacterial chromosome: DNA cleavage induced by oxolinic acid. J. Mol. Biol. 1979, 131, 287–302. [Google Scholar] [CrossRef]
- Wentzell, L.M.; Maxwell, A. The complex of DNA gyrase and quinolone drugs on DNA forms a barrier to the T7 DNA polymerase replication complex. J. Mol. Biol. 2000, 304, 779–791. [Google Scholar] [CrossRef]
- Willmott, C.J.R.; Critchlow, S.E.; Eperon, I.C.; Maxwell, A. The complex of DNA gyrase and quinolone drugs with DNA forms a barrier to transcription by RNA polymerase. J. Mol. Biol. 1994, 242, 351–363. [Google Scholar] [CrossRef]
- Chen, C.-R.; Malik, M.; Snyder, M.; Drlica, K. DNA gyrase and topoisomerase IV on the bacterial chromosome: Quinolone-induced DNA cleavage. J. Mol. Biol. 1996, 258, 627–637. [Google Scholar] [CrossRef]
- Malik, M.; Zhao, X.; Drlica, K. Lethal fragmentation of bacterial chromosomes mediated by DNA gyrase and quinolones. Mol. Microbiol. 2006, 61, 810–825. [Google Scholar] [CrossRef] [PubMed]
- Carret, G.; Flandrois, J.P.; Lobry, J.R. Biphasic kinetics of bacterial killing by quinolones. J. Antimicrob. Chemother. 1991, 27, 319–327. [Google Scholar] [CrossRef] [PubMed]
- Drlica, K.; Malik, M.; Kerns, R.J.; Zhao, X. Quinolone-mediated bacterial death. Antimicrob. Agents Chemother. 2008, 52, 385–392. [Google Scholar] [CrossRef] [PubMed]
- Hong, Y.; Li, Q.; Gao, Q.; Xie, J.; Huang, H.; Drlica, K.; Zhao, X. Reactive oxygen species play a dominant role in all pathways of rapid quinolone-mediated killing. J. Antimicrob. Chemother. 2020, 75, 576–585. [Google Scholar] [CrossRef] [PubMed]
- Jeong, K.S.; Xie, Y.; Hiasa, H.; Khodursky, A.B. Analysis of pleiotropic transcriptional profiles: A case study of DNA gyrase inhibition. PLoS Genet. 2006, 2, e152. [Google Scholar] [CrossRef] [PubMed]
- Postow, L.; Crisona, N.J.; Peter, B.J.; Hardy, C.D.; Cozzarelli, N.R. Topological challenges to DNA replication: Conformations at the fork. Proc. Natl. Acad. Sci. USA 2001, 98, 8219–8226. [Google Scholar] [CrossRef]
- Schvartzman, J.B.; Stasiak, A. A topological view of the replicon. EMBO Rep. 2004, 5, 256–261. [Google Scholar] [CrossRef]
- Seol, Y.; Neuman, K.C. The dynamic interplay between DNA topoisomerases and DNA topology. Biophys. Rev. 2016, 8, 101–111. [Google Scholar] [CrossRef]
- Peter, B.J.; Ullsperger, C.; Hiasa, H.; Marians, K.J.; Cozzarelli, N.R. The structure of supercoiled intermediates in DNA replication. Cell 1998, 94, 819–827. [Google Scholar] [CrossRef]
- Sogo, J.M.; Stasiak, A.; Martínez-Robles, M.a.L.; Krimer, D.B.; Hernández, P.; Schvartzman, J.B. Formation of knots in partially replicated DNA molecules. J. Mol. Biol. 1999, 286, 637–643. [Google Scholar] [CrossRef]
- Cebrián, J.; Castán, A.; Martínez, V.; Kadomatsu-Hermosa, M.J.; Parra, C.; Fernández-Nestosa, M.J.; Schaerer, C.; Hernández, P.; Krimer, D.B.; Schvartzman, J.B. Direct evidence for the formation of precatenanes during DNA replication. J. Biol. Chem. 2015, 290, 13725–13735. [Google Scholar] [CrossRef] [PubMed]
- Sissi, C.; Palumbo, M. In front of and behind the replication fork: Bacterial type IIA topoisomerases. Cell. Mol. Life Sci. 2010, 67, 2001–2024. [Google Scholar] [CrossRef] [PubMed]
- Pohlhaus, J.R.; Kreuzer, K.N. Norfloxacin-induced DNA gyrase cleavage complexes block Escherichia coli replication forks, causing double-stranded breaks in vivo. Mol. Microbiol. 2005, 56, 1416–1429. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Malik, M.; Chan, N.; Drlica-Wagner, A.; Wang, J.-Y.; Li, X.; Drlica, K. Lethal action of quinolones against a temperature-sensitive dnaB replication mutant of Escherichia coli. Antimicrob. Agents Chemother. 2006, 50, 362–364. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, A.; Stokes, J.M.; Matic, I. Our evolving understanding of the mechanism of quinolones. Antibiotics 2018, 7, 32. [Google Scholar] [CrossRef] [PubMed]
- Shea, M.E.; Hiasa, H. The RuvAB branch migration complex can displace topoisomerase IV·quinolone·DNA ternary complexes. J. Biol. Chem. 2003, 278, 48485–48490. [Google Scholar] [CrossRef] [PubMed]
- Hsiang, Y.-H.; Lihou, M.G.; Liu, L.F. Arrest of replication forks by drug-stabilized topoisomerase I-DNA cleavable complexes as a mechanism of cell killing by camptothecin. Cancer Res. 1989, 49, 5077–5082. [Google Scholar]
- Tsao, Y.P.; Russo, A.; Nyamuswa, G.; Silber, R.; Liu, L.F. Interaction between replication forks and topoisomerase I-DNA cleavable complexes: Studies in a cell-free SV40 DNA replication system. Cancer Res. 1993, 53, 5908–5914. [Google Scholar]
- Hsiang, Y.H.; Hertzberg, R.; Hecht, S.; Liu, L.F. Camptothecin induces protein-linked DNA breaks via mammalian DNA topoisomerase I. J. Biol. Chem. 1985, 260, 14873–14878. [Google Scholar]
- Strumberg, D.; Pilon, A.A.; Smith, M.; Hickey, R.; Malkas, L.; Pommier, Y. Conversion of topoisomerase I cleavage complexes on the leading strand of ribosomal DNA into 5’-phosphorylated DNA double-strand breaks by replication runoff. Mol. Cell. Biol. 2000, 20, 3977–3987. [Google Scholar] [CrossRef]
- Cortés, F.; Piñero, J.; Ortiz, T. Importance of replication fork progression for the induction of chromosome damage and SCE by inhibitors of DNA topoisomerases. Mutat. Res. Lett. 1993, 303, 71–76. [Google Scholar] [CrossRef]
- Chow, R.T.; Dougherty, T.J.; Fraimow, H.S.; Bellin, E.Y.; Miller, M.H. Association between early inhibition of DNA synthesis and the MICs and MBCs of carboxyquinolone antimicrobial agents for wild-type and mutant [gyrA nfxB(ompF) acrA] Escherichia coli K-12. Antimicrob. Agents Chemother. 1988, 32, 1113–1118. [Google Scholar] [CrossRef]
- Piddock, L.J.; Walters, R.N.; Diver, J.M. Correlation of quinolone MIC and inhibition of DNA, RNA, and protein synthesis and induction of the SOS response in Escherichia coli. Antimicrob. Agents Chemother. 1990, 34, 2331–2336. [Google Scholar] [CrossRef] [PubMed]
- Deitz, W.H.; Cook, T.M.; Goss, W.A. Mechanism of action of nalidixic acid on Escherichia coli. 3. Conditions required for lethality. J. Bacteriol. 1966, 91, 768–773. [Google Scholar] [CrossRef] [PubMed]
- Malik, M.; Hussain, S.; Drlica, K. Effect of anaerobic growth on quinolone lethality with Escherichia coli. Antimicrob. Agents Chemother. 2007, 51, 28–34. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sun, Y.; Saha, S.; Wang, W.; Saha, L.K.; Huang, S.-Y.N.; Pommier, Y. Excision repair of topoisomerase DNA-protein crosslinks (TOP-DPC). DNA Repair 2020, 89, 102837. [Google Scholar] [CrossRef]
- Pommier, Y.; Huang, S.-y.N.; Gao, R.; Das, B.B.; Murai, J.; Marchand, C. Tyrosyl-DNA-phosphodiesterases (TDP1 and TDP2). DNA Repair 2014, 19, 114–129. [Google Scholar] [CrossRef]
- Cortes Ledesma, F.; El Khamisy, S.F.; Zuma, M.C.; Osborn, K.; Caldecott, K.W. A human 5′-tyrosyl DNA phosphodiesterase that repairs topoisomerase-mediated DNA damage. Nature 2009, 461, 674–678. [Google Scholar] [CrossRef]
- Pouliot, J.J.; Robertson, C.A.; Nash, H.A. Pathways for repair of topoisomerase I covalent complexes in Saccharomyces cerevisiae. Genes Cells 2001, 6, 677–687. [Google Scholar] [CrossRef]
- Pouliot, J.J.; Yao, K.C.; Robertson, C.A.; Nash, H.A. Yeast gene for a Tyr-DNA phosphodiesterase that repairs topoisomerase I complexes. Science 1999, 286, 552–555. [Google Scholar] [CrossRef]
- Aedo, S.; Tse-Dinh, Y.-C. SbcCD-mediated processing of covalent gyrase-DNA complex in Escherichia coli. Antimicrob. Agents Chemother. 2013, 57, 5116–5119. [Google Scholar] [CrossRef] [PubMed]
- Connelly, J.C.; de Leau, E.S.; Leach, D.R.F. Nucleolytic processing of a protein-bound DNA end by the E. coli SbcCD (MR) complex. DNA Repair 2003, 2, 795–807. [Google Scholar] [CrossRef]
- Tamayo, M.; Santiso, R.; Gosalvez, J.; Bou, G.; Fernández, J.L. Rapid assessment of the effect of ciprofloxacin on chromosomal DNA from Escherichia coli using an in situ DNA fragmentation assay. BMC Microbiol. 2009, 9, 69. [Google Scholar] [CrossRef] [PubMed]
- Andryukov, B.G.; Somova, L.M.; Timchenko, N.F. Molecular and genetic characteristics of cell death in prokaryotes. Mol. Gen. Microbiol. Virol. 2018, 33, 73–83. [Google Scholar] [CrossRef]
- Van Acker, H.; Coenye, T. The role of reactive oxygen species in antibiotic-mediated killing of bacteria. Trends Microbiol. 2017, 25, 456–466. [Google Scholar] [CrossRef]
- Whitaker, A.M.; Schaich, M.A.; Smith, M.R.; Flynn, T.S.; Freudenthal, B.D. Base excision repair of oxidative DNA damage: From mechanism to disease. Front. Biosci. (Landmark Ed.) 2017, 22, 1493–1522. [Google Scholar] [CrossRef]
- Imlay, J.A. Pathways of oxidative damage. Annu. Rev. Microbiol. 2003, 57, 395–418. [Google Scholar] [CrossRef]
- Dwyer, D.J.; Collins, J.J.; Walker, G.C. Unraveling the physiological complexities of antibiotic lethality. Annu. Rev. Pharmacol. Toxicol. 2015, 55, 313–332. [Google Scholar] [CrossRef]
- Hong, Y.; Zeng, J.; Wang, X.; Drlica, K.; Zhao, X. Post-stress bacterial cell death mediated by reactive oxygen species. Proc. Natl. Acad. Sci. USA 2019, 116, 10064–10071. [Google Scholar] [CrossRef]
- Dwyer, D.J.; Kohanski, M.A.; Hayete, B.; Collins, J.J. Gyrase inhibitors induce an oxidative damage cellular death pathway in Escherichia coli. Mol. Syst. Biol. 2007, 3, 91. [Google Scholar] [CrossRef]
- Liu, Y.; Imlay, J.A. Cell death from antibiotics without the involvement of reactive oxygen species. Science 2013, 339, 1210–1213. [Google Scholar] [CrossRef] [PubMed]
- Keren, I.; Wu, Y.; Inocencio, J.; Mulcahy, L.R.; Lewis, K. Killing by bactericidal antibiotics does not depend on reactive oxygen species. Science 2013, 339, 1213–1216. [Google Scholar] [CrossRef] [PubMed]
- Foti, J.J.; Devadoss, B.; Winkler, J.A.; Collins, J.J.; Walker, G.C. Oxidation of the guanine nucleotide pool underlies cell death by bactericidal antibiotics. Science 2012, 336, 315–319. [Google Scholar] [CrossRef] [PubMed]
- Belenky, P.; Ye, J.D.; Porter, C.B.M.; Cohen, N.R.; Lobritz, M.A.; Ferrante, T.; Jain, S.; Korry, B.J.; Schwarz, E.G.; Walker, G.C.; et al. Bactericidal antibiotics induce toxic metabolic perturbations that lead to cellular damage. Cell Rep. 2015, 13, 968–980. [Google Scholar] [CrossRef] [PubMed]
- Imlay, J.A. The molecular mechanisms and physiological consequences of oxidative stress: Lessons from a model bacterium. Nat. Rev. Microbiol. 2013, 11, 443–454. [Google Scholar] [CrossRef] [PubMed]
- Imlay, J.A. Diagnosing oxidative stress in bacteria: Not as easy as you might think. Curr. Opin. Microbiol. 2015, 24, 124–131. [Google Scholar] [CrossRef] [PubMed]
- Hong, Y.; Li, L.; Luan, G.; Drlica, K.; Zhao, X. Contribution of reactive oxygen species to thymineless death in Escherichia coli. Nat. Microbiol. 2017, 2, 1667–1675. [Google Scholar] [CrossRef]
- Dörr, T.; Lewis, K.; Vulić, M. SOS response induces persistence to fluoroquinolones in Escherichia coli. PLoS Genet. 2009, 5, e1000760. [Google Scholar] [CrossRef]
- Henderson, M.L.; Kreuzer, K.N. Functions that protect Escherichia coli from tightly bound DNA-protein complexes created by mutant EcoRII methyltransferase. PLoS ONE 2015, 10, e0128092. [Google Scholar] [CrossRef]
- McDaniel, L.S.; Rogers, L.H.; Hill, W.E. Survival of recombination-deficient mutants of Escherichia coli during incubation with nalidixic acid. J. Bacteriol. 1978, 134, 1195–1198. [Google Scholar] [CrossRef]
- Kreuzer, K.N. DNA damage responses in prokaryotes: Regulating gene expression, modulating growth patterns, and manipulating replication forks. Cold Spring Harb. Perspect. Biol. 2013, 5, a012674. [Google Scholar] [CrossRef] [PubMed]
- Pribis, J.P.; García-Villada, L.; Zhai, Y.; Lewin-Epstein, O.; Wang, A.Z.; Liu, J.; Xia, J.; Mei, Q.; Fitzgerald, D.M.; Bos, J.; et al. Gamblers: An antibiotic-induced evolvable cell subpopulation differentiated by reactive-oxygen-induced general stress response. Mol. Cell 2019, 74, 785–800.e7. [Google Scholar] [CrossRef] [PubMed]
- Storvik, K.A.M.; Foster, P.L. RpoS, the stress response sigma factor, plays a dual role in the regulation of Escherichia coli’s error-prone DNA polymerase IV. J. Bacteriol. 2010, 192, 3639–3644. [Google Scholar] [CrossRef] [PubMed]
- Davies, J.; Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. Rev. 2010, 74, 417–433. [Google Scholar] [CrossRef] [PubMed]
- Fair, R.J.; Tor, Y. Antibiotics and bacterial resistance in the 21st century. Perspect. Med. Chem. 2014, 6, 25–64. [Google Scholar] [CrossRef]
- Phillips, I.; Casewell, M.; Cox, T.; De Groot, B.; Friis, C.; Jones, R.; Nightingale, C.; Preston, R.; Waddell, J. Does the use of antibiotics in food animals pose a risk to human health? A critical review of published data. J. Antimicrob. Chemother. 2004, 53, 28–52. [Google Scholar] [CrossRef]
- Livermore, D.M.; Hope, R.; Reynolds, R.; Blackburn, R.; Johnson, A.P.; Woodford, N. Declining cephalosporin and fluoroquinolone non-susceptibility among bloodstream Enterobacteriaceae from the UK: Links to prescribing change? J. Antimicrob. Chemother. 2013, 68, 2667–2674. [Google Scholar] [CrossRef]
- Spellberg, B.; Doi, Y. The rise of fluoroquinolone-resistant Escherichia coli in the community: Scarier than we thought. J. Infect. Dis. 2015, 212, 1853–1855. [Google Scholar] [CrossRef]
- Dalhoff, A. Global fluoroquinolone resistance epidemiology and implictions for clinical use. Interdiscip. Perspect. Infect. Dis. 2012, 2012, 37. [Google Scholar] [CrossRef]
- Kim, E.S.; Hooper, D.C. Clinical importance and epidemiology of quinolone resistance. Infect. Chemother. 2014, 46, 226–238. [Google Scholar] [CrossRef]
- Yoshida, H.; Bogaki, M.; Nakamura, M.; Nakamura, S. Quinolone resistance-determining region in the DNA gyrase gyrA gene of Escherichia coli. Antimicrob. Agents Chemother. 1990, 34, 1271–1272. [Google Scholar] [CrossRef] [PubMed]
- Avalos, E.; Catanzaro, D.; Catanzaro, A.; Ganiats, T.; Brodine, S.; Alcaraz, J.; Rodwell, T. Frequency and geographic distribution of gyrA and gyrB mutations associated with fluoroquinolone resistance in clinical Mycobacterium tuberculosis isolates: A systematic review. PLoS ONE 2015, 10, e0120470. [Google Scholar] [CrossRef] [PubMed]
- Gensberg, K.; Jin, Y.F.; Piddock, L.J. A novel gyrB mutation in a fluoroquinolone-resistant clinical isolate of Salmonella Typhimurium. FEMS Microbiol. Lett. 1995, 132, 57–60. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, H.; Bogaki, M.; Nakamura, M.; Yamanaka, L.M.; Nakamura, S. Quinolone resistance-determining region in the DNA gyrase gyrB gene of Escherichia coli. Antimicrob. Agents Chemother. 1991, 35, 1647–1650. [Google Scholar] [CrossRef]
- Morais Cabral, J.H.; Jackson, A.P.; Smith, C.V.; Shikotra, N.; Maxwell, A.; Liddington, R.C. Crystal structure of the breakage-reunion domain of DNA gyrase. Nature 1997, 388, 903–906. [Google Scholar] [CrossRef] [PubMed]
- Friedman, S.M.; Lu, T.; Drlica, K. Mutation in the DNA gyrase A gene of Escherichia coli that expands the quinolone resistance-retermining region. Antimicrob. Agents Chemother. 2001, 45, 2378–2380. [Google Scholar] [CrossRef]
- Ince, D.; Hooper, D.C. Quinolone resistance due to reduced target enzyme expression. J. Bacteriol. 2003, 185, 6883–6892. [Google Scholar] [CrossRef]
- Michel-Briand, Y.; Uccelli, V.; Laporte, J.-M.; Plesiat, P. Elimination of plasmids from Enterobacteriaceae by 4-quinolone derivatives. J. Antimicrob. Chemother. 1986, 18, 667–674. [Google Scholar] [CrossRef]
- Weisser, J.; Wiedemann, B. Elimination of plasmids by enoxacin and ofloxacin at near inhibitory concentrations. J. Antimicrob. Chemother. 1986, 18, 575–583. [Google Scholar] [CrossRef]
- Weisser, J.; Wiedemann, B. Inhibition of R-plasmid transfer in Escherichia coli by 4-quinolones. Antimicrob. Agents Chemother. 1987, 31, 531–534. [Google Scholar] [CrossRef]
- Martínez-Martínez, L.; Pascual, A.; Jacoby, G.A. Quinolone resistance from a transferable plasmid. Lancet 1998, 351, 797–799. [Google Scholar] [CrossRef]
- Vetting, M.W.; Hegde, S.S.; Fajardo, J.E.; Fiser, A.; Roderick, S.L.; Takiff, H.E.; Blanchard, J.S. Pentapeptide repeat proteins. Biochem. 2006, 45, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Jacoby, G.A.; Chow, N.; Waites, K.B. Prevalence of plasmid-mediated quinolone resistance. Antimicrob. Agents Chemother. 2003, 47, 559–562. [Google Scholar] [CrossRef] [PubMed]
- Montero, C.; Mateu, G.; Rodriguez, R.; Takiff, H. Intrinsic resistance of Mycobacterium smegmatis to fluoroquinolones may be influenced by new pentapeptide protein MfpA. Antimicrob. Agents Chemother. 2001, 45, 3387. [Google Scholar] [CrossRef]
- Collin, F.; Maxwell, A. The microbial toxin microcin B17: Prospects for the development of new antibacterial agents. J. Mol. Biol. 2019, 431, 3400–3426. [Google Scholar] [CrossRef]
- Hegde, S.S.; Vetting, M.W.; Roderick, S.L.; Mitchenall, L.A.; Maxwell, A.; Takiff, H.E.; Blanchard, J.S. A fluoroquinolone resistance protein from Mycobacterium tuberculosis that mimics DNA. Science 2005, 308, 1480–1483. [Google Scholar] [CrossRef]
- Xiong, X.; Bromley, E.H.C.; Oelschlaeger, P.; Woolfson, D.N.; Spencer, J. Structural insights into quinolone antibiotic resistance mediated by pentapeptide repeat proteins: Conserved surface loops direct the activity of a Qnr protein from a gram-negative bacterium. Nucleic Acids Res. 2011, 39, 3917–3927. [Google Scholar] [CrossRef]
- Garrido, M.C.; Herrero, M.; Kolter, R.; Moreno, F. The export of the DNA replication inhibitor Microcin B17 provides immunity for the host cell. EMBO J. 1988, 7, 1853–1862. [Google Scholar] [CrossRef]
- Jacoby, G.A.; Corcoran, M.A.; Hooper, D.C. Protective effect of Qnr on agents other than quinolones that target DNA gyrase. Antimicrob. Agents Chemother. 2015, 59, 6689–6695. [Google Scholar] [CrossRef]
- Robicsek, A.; Strahilevitz, J.; Jacoby, G.A.; Macielag, M.; Abbanat, D.; Hye Park, C.; Bush, K.; Hooper, D.C. Fluoroquinolone-modifying enzyme: A new adaptation of a common aminoglycoside acetyltransferase. Nat. Med. 2006, 12, 83–88. [Google Scholar] [CrossRef]
- Heisig, P.; Tschorny, R. Characterization of fluoroquinolone-resistant mutants of Escherichia coli selected in vitro. Antimicrob. Agents Chemother. 1994, 38, 1284–1291. [Google Scholar] [CrossRef] [PubMed]
- Everett, M.J.; Jin, Y.F.; Ricci, V.; Piddock, L.J. Contributions of individual mechanisms to fluoroquinolone resistance in 36 Escherichia coli strains isolated from humans and animals. Antimicrob. Agents Chemother. 1996, 40, 2380–2386. [Google Scholar] [CrossRef] [PubMed]
- Bolla, J.-M.; Alibert-Franco, S.; Handzlik, J.; Chevalier, J.; Mahamoud, A.; Boyer, G.; Kieć-Kononowicz, K.; Pagès, J.-M. Strategies for bypassing the membrane barrier in multidrug resistant Gram-negative bacteria. FEBS Lett. 2011, 585, 1682–1690. [Google Scholar] [CrossRef] [PubMed]
- Fernández, L.; Hancock, R.E.W. Adaptive and mutational resistance: Role of porins and efflux pumps in drug resistance. Clin. Microbiol. Rev. 2012, 25, 661–681. [Google Scholar] [CrossRef]
- Alekshun, M.N.; Levy, S.B. Regulation of chromosomally mediated multiple antibiotic resistance: The mar regulon. Antimicrob. Agents Chemother. 1997, 41, 2067–2075. [Google Scholar] [CrossRef]
- Alekshun, M.N.; Levy, S.B. Molecular Mechanisms of Antibacterial Multidrug Resistance. Cell 2007, 128, 1037–1050. [Google Scholar] [CrossRef]
- Davin-Regli, A.; Bolla, J.-M.; James, C.E.; Lavigne, J.-P.; Chevalier, J.; Garnotel, E.; Molitor, A.; Pages, J.-M. Membrane permeability and regulation of drug “influx and efflux” in enterobacterial pathogens. Curr. Drug Targets 2008, 9, 750–759. [Google Scholar] [CrossRef]
- Van Bambeke, F.; Balzi, E.; Tulkens, P.M. Antibiotic efflux pumps. Biochem. Pharmacol. 2000, 60, 457–470. [Google Scholar] [CrossRef]
- Papkou, A.; Hedge, J.; Kapel, N.; Young, B.; MacLean, R.C. Efflux pump activity potentiates the evolution of antibiotic resistance across Staphylococcus aureus isolates. Nat. Commun. 2020, 11, 3970. [Google Scholar] [CrossRef]
- Schmalstieg, A.M.; Srivastava, S.; Belkaya, S.; Deshpande, D.; Meek, C.; Leff, R.; van Oers, N.S.C.; Gumbo, T. The antibiotic resistance arrow of time: Efflux pump induction is a general first step in the evolution of mycobacterial drug resistance. Antimicrob. Agents Chemother. 2012, 56, 4806–4815. [Google Scholar] [CrossRef]
- Hernández, A.; Sánchez, M.B.; Martínez, J.L. Quinolone resistance: Much more than predicted. Front. Microbiol. 2011, 2, 22. [Google Scholar] [CrossRef] [PubMed]
- Piddock, L.J.V.; Hall, M.C.; Walters, R.N. Phenotypic characterization of quinolone-resistant mutants of Enterobacteriaceae selected from wild type, gyrA type and multiplyresistant (marA) type strains. J. Antimicrob. Chemother. 1991, 28, 185–198. [Google Scholar] [CrossRef] [PubMed]
- Strahilevitz, J.; Jacoby, G.A.; Hooper, D.C.; Robicsek, A. Plasmid-mediated quinolone resistance: A multifaceted threat. Clin. Microbiol. Rev. 2009, 22, 664–689. [Google Scholar] [CrossRef] [PubMed]
- Costa, S.S.; Viveiros, M.; Amaral, L.; Couto, I. Multidrug efflux pumps in Staphylococcus aureus: An update. Open Microbiol J. 2013, 7, 59–71. [Google Scholar] [CrossRef] [PubMed]
- Goldman, J.D.; White, D.G.; Levy, S.B. Multiple antibiotic resistance (mar) locus protects Escherichia coli from rapid cell killing by fluoroquinolones. Antimicrob. Agents Chemother. 1996, 40, 1266–1269. [Google Scholar] [CrossRef]
- Kern, W.V.; Oethinger, M.; Jellen-Ritter, A.S.; Levy, S.B. Non-target gene mutations in the development of fluoroquinolone resistance in Escherichia coli. Antimicrob. Agents Chemother. 2000, 44, 814–820. [Google Scholar] [CrossRef]
- Maneewannakul, K.; Levy, S.B. Identification for mar mutants among quinolone-resistant clinical isolates of Escherichia coli. Antimicrob. Agents Chemother. 1996, 40, 1695–1698. [Google Scholar] [CrossRef]
- Oethinger, M.; Podglajen, I.; Kern, W.V.; Levy, S.B. Overexpression of the marA or soxS regulatory gene in clinical topoisomerase mutants of Escherichia coli. Antimicrob. Agents Chemother. 1998, 42, 2089–2094. [Google Scholar] [CrossRef]
- Jurado, S.; Medina, A.; de la Fuente, R.; Ruiz-Santa-Quiteria, J.A.; Orden, J.A. Resistance to non-quinolone antimicrobials in commensal Escherichia coli isolates from chickens treated orally with enrofloxacin. Jpn. J. Vet. Res. 2015, 63, 195–200. [Google Scholar]
- Kohanski, M.A.; DePristo, M.A.; Collins, J.J. Sublethal antibiotic treatment leads to multidrug resistance via radical-induced mutagenesis. Mol. Cell 2010, 37, 311–320. [Google Scholar] [CrossRef]
- Pereira, R.V.; Siler, J.D.; Ng, J.C.; Davis, M.A.; Grohn, Y.T.; Warnick, L.D. Effect of on-farm use of antimicrobial drugs on resistance in fecal Escherichia coli of preweaned dairy calves. J. Dairy Sci. 2014, 97, 7644–7654. [Google Scholar] [CrossRef] [PubMed]
- Chuppava, B.; Keller, B.; El-Wahab, A.A.; Meißner, J.; Kietzmann, M.; Visscher, C. Resistance of Escherichia coli in turkeys after therapeutic or environmental exposition with enrofloxacin depending on flooring. Int. J. Environ. Res. Public Health 2018, 15, 1993. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Hao, H.; Dai, M.; Zhang, H.; Ning, J.; Cheng, G.; Shabbir, M.A.B.; Sajid, A.; Yuan, Z. Resistance and virulence mechanisms of Escherichia coli selected by Enrofloxacin in Chicken. Antimicrob. Agents Chemother. 2019, 63, e01824-18. [Google Scholar] [CrossRef] [PubMed]
- Tattevin, P.; Basuino, L.; Chambers, H.F. Subinhibitory fluoroquinolone exposure selects for reduced beta-lactam susceptibility in methicillin-resistant Staphylococcus aureus and alterations in the SOS-mediated response. Res. Microbiol. 2009, 160, 187–192. [Google Scholar] [CrossRef] [PubMed]
- Lopez, E.; Blazquez, J. Effect of subinhibitory concentrations of antibiotics on intrachromosomal homologous recombination in Escherichia coli. Antimicrob. Agents Chemother. 2009, 53, 3411–3415. [Google Scholar] [CrossRef] [PubMed]
- Lopez, E.; Elez, M.; Matic, I.; Blazquez, J. Antibiotic-mediated recombination: Ciprofloxacin stimulates SOS-independent recombination of divergent sequences in Escherichia coli. Mol. Microbiol. 2007, 64, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Cirz, R.T.; Jones, M.B.; Gingles, N.A.; Minogue, T.D.; Jarrahi, B.; Peterson, S.N.; Romesberg, F.E. Complete and SOS-mediated response of Staphylococcus aureus to the antibiotic ciprofloxacin. J. Bacteriol. 2007, 189, 531–539. [Google Scholar] [CrossRef]
- Cirz, R.T.; O’Neill, B.M.; Hammond, J.A.; Head, S.R.; Romesberg, F.E. Defining the Pseudomonas aeruginosa SOS response and its role in the global response to the antibiotic ciprofloxacin. J. Bacteriol. 2006, 188, 7101–7110. [Google Scholar] [CrossRef]
- Gillespie, S.H.; Basu, S.; Dickens, A.L.; O’Sullivan, D.M.; McHugh, T.D. Effect of subinhibitory concentrations of ciprofloxacin on Mycobacterium fortuitum mutation rates. J. Antimicrob. Chemother. 2005, 56, 344–348. [Google Scholar] [CrossRef]
- O’Sullivan, D.M.; Hinds, J.; Butcher, P.D.; Gillespie, S.H.; McHugh, T.D. Mycobacterium tuberculosis DNA repair in response to subinhibitory concentrations of ciprofloxacin. J. Antimicrob. Chemother. 2008, 62, 1199–1202. [Google Scholar] [CrossRef]
- Marshall, B.M.; Levy, S.B. Food animals and antimicrobials: Impacts on human health. Clin. Microbiol. Rev. 2011, 24, 718–733. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.J.; Park, N.H.; Mechesso, A.F.; Lee, K.J.; Park, S.C. The phenotypic and molecular resistance induced by a single-exposure to sub-mutant prevention concentration of marbofloxacin in Salmonella Typhimurium isolates from swine. Vet. Microbiol. 2017, 207, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Pers, C.; Søgaard, P.; Pallesen, L. Selection of multiple resistance in Salmonella enteritidis during treatment with ciprofloxacin. Scand. J. Infect. Dis. 1996, 28, 529–531. [Google Scholar] [CrossRef] [PubMed]
- Cebrian, L.; RodrÍGuez, J.C.; Escribano, I.; Royo, G. Effect of exposure to fluoroquinolones and beta-lactams on the in vitro activity of other groups of antibiotics in Salmonella spp. APMIS 2006, 114, 523–528. [Google Scholar] [CrossRef]
- Ching, C.; Zaman, M.H. Development and selection of low-level multi-drug resistance over an extended range of sub-inhibitory ciprofloxacin concentrations in Escherichia coli. Sci. Rep. 2020, 10, 8754. [Google Scholar] [CrossRef]
- Piddock, L.J.V.; Wise, R. Induction of the SOS response in Escherichia coli by 4-quinolone antimicrobial agents. FEMS Microbiol. Lett. 1987, 41, 289–294. [Google Scholar] [CrossRef][Green Version]
- Lewin, C.S.; Howard, B.M.A.; Ratcliff, N.T.; Smith, J.T. 4-Quinolones and the SOS response. J. Med. Microbiol. 1989, 29, 139–144. [Google Scholar] [CrossRef]
- Ysern, P.; Clerch, B.; Castańo, M.; Gibert, I.; Barbé, J.; Llagostera, M. Induction of SOS genes in Escherichia coli and mutagenesis in Salmonella typhimurium by fluoroquinolones. Mutagenesis 1990, 5, 63–66. [Google Scholar] [CrossRef]
- Cirz, R.T.; Chin, J.K.; Andes, D.R.; de Crecy-Lagard, V.; Craig, W.A.; Romesberg, F.E. Inhibition of mutation and combating the evolution of antibiotic resistance. PLoS Biol. 2005, 3, e176. [Google Scholar] [CrossRef]
- Cirz, R.T.; Romesberg, F.E. Induction and inhibition of ciprofloxacin resistance-conferring mutations in hypermutator bacteria. Antimicrob. Agents Chemother. 2006, 50, 220–225. [Google Scholar] [CrossRef]
- Thi, T.D.; López, E.; Rodríguez-Rojas, A.; Rodríguez-Beltrán, J.; Couce, A.; Guelfo, J.R.; Castañeda-García, A.; Blázquez, J. Effect of recA inactivation on mutagenesis of Escherichia coli exposed to sublethal concentrations of antimicrobials. J. Antimicrob. Chemother. 2011, 66, 531–538. [Google Scholar] [CrossRef] [PubMed]
- Blazquez, J.; Rodriguez-Beltran, J.; Matic, I. Antibiotic-induced genetic variation: How it arises and how it can be prevented. Annu. Rev. Microbiol. 2018, 72, 209–230. [Google Scholar] [CrossRef] [PubMed]
- Song, L.Y.; Goff, M.; Davidian, C.; Mao, Z.; London, M.; Lam, K.; Yung, M.; Miller, J.H. Mutational consequences of ciprofloxacin in Escherichia coli. Antimicrob. Agents Chemother. 2016, 60, 6165–6172. [Google Scholar] [CrossRef] [PubMed]
- Long, H.; Miller, S.F.; Strauss, C.; Zhao, C.; Cheng, L.; Ye, Z.; Griffin, K.; Te, R.; Lee, H.; Chen, C.-C.; et al. Antibiotic treatment enhances the genome-wide mutation rate of target cells. Proc. Natl. Acad. Sci. USA 2016, 113, E2498–E2505. [Google Scholar] [CrossRef] [PubMed]
- Bearson, B.L.; Brunelle, B.W. Fluoroquinolone induction of phage-mediated gene transfer in multidrug-resistant Salmonella. Int. J. Antimicrob. Agents 2015, 46, 201–204. [Google Scholar] [CrossRef]
- Barrett, T.C.; Mok, W.W.K.; Murawski, A.M.; Brynildsen, M.P. Enhanced antibiotic resistance development from fluoroquinolone persisters after a single exposure to antibiotic. Nat. Commun. 2019, 10, 1177. [Google Scholar] [CrossRef]
- Goneau, L.W.; Yeoh, N.S.; MacDonald, K.W.; Cadieux, P.A.; Burton, J.P.; Razvi, H.; Reid, G. Selective target inactivation rather than global metabolic dormancy causes antibiotic tolerance in uropathogens. Antimicrob. Agents Chemother. 2014, 58, 2089–2097. [Google Scholar] [CrossRef]
- Theodore, A.; Lewis, K.; Vulić, M. Tolerance of Escherichia coli to fluoroquinolone antibiotics depends on specific components of the SOS response pathway. Genetics 2013, 195, 1265–1276. [Google Scholar] [CrossRef]
- Gutierrez, A.; Jain, S.; Bhargava, P.; Hamblin, M.; Lobritz, M.A.; Collins, J.J. Understanding and sensitizing density-dependent persistence to quinolone antibiotics. Mol. Cell 2017, 68, 1147–1154.e3. [Google Scholar] [CrossRef]
- Fisher, R.A.; Gollan, B.; Helaine, S. Persistent bacterial infections and persister cells. Nat. Rev. Microbiol. 2017, 15, 453–464. [Google Scholar] [CrossRef]
- Torres-Barceló, C.; Kojadinovic, M.; Moxon, R.; MacLean, R.C. The SOS response increases bacterial fitness, but not evolvability, under a sublethal dose of antibiotic. Proc. R. Soc. B 2015, 282, 20150885. [Google Scholar] [CrossRef] [PubMed]
- Almalki, Z.S.; Yue, X.; Xia, Y.; Wigle, P.R.; Guo, J.J. Utilization, Spending, and Price Trends for Quinolones in the US Medicaid Programs: 25 Years’ Experience 1991–2015. Pharmacoecon. Open 2017, 1, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Durkin, M.J.; Keller, M.; Butler, A.M.; Kwon, J.H.; Dubberke, E.R.; Miller, A.C.; Polgreen, P.M.; Olsen, M.A. An assessment of inappropriate antibiotic use and guideline adherence for uncomplicated urinary tract infections. Open Forum Infect. Dis. 2018, 5. [Google Scholar] [CrossRef] [PubMed]
- Richards, G.A.; Brink, A.J.; Feldman, C. Rational use of the fluoroquinolones. S. Afr. Med. J. S. Afr. Med. Tydskr. 2019, 109, 378–381. [Google Scholar] [CrossRef] [PubMed]
- Ching, C.; Orubu, E.S.F.; Wirtz, V.J.; Zaman, M.H. Bacterial antibiotic resistance development and mutagenesis following exposure to subminimal inhibitory concentrations of fluoroquinolones in vitro: A systematic literature review protocol. BMJ Open 2019, 9, e030747. [Google Scholar] [CrossRef] [PubMed]
- Jorgensen, S.C.J.; Mercuro, N.J.; Davis, S.L.; Rybak, M.J. Delafloxacin: Place in therapy and review of microbiologic, clinical and pharmacologic properties. Infect. Dis. Ther 2018, 7, 197–217. [Google Scholar] [CrossRef]
- Chan, P.F.; Huang, J.; Bax, B.D.; Gwynn, M.N. Recent developments in inhibitors of bacterial type IIA topoisomerases. In Antibiotics: Targets, Mechanisms and Resistance; Gualerzi, C.O., Brandi, L., Fabbretti, A., Pon, C.L., Eds.; Wiley-VCH Verlag GmbH & Co: Weinheim, Germany, 2014; pp. 263–297. [Google Scholar]
- Candel, F.J.; Peñuelas, M. Delafloxacin: Design, development and potential place in therapy. Drug Design Dev. Ther. 2017, 11, 881–891. [Google Scholar] [CrossRef]
- Houssaye, S.; Gutmann, L.; Varon, E. Topoisomerase mutations associated with in vitro selection of resistance to moxifloxacin in Streptococcus pneumoniae. Antimicrob. Agents Chemother. 2002, 46, 2712–2715. [Google Scholar] [CrossRef]
- Li, X.; Zhao, X.; Drlica, K. Selection of Streptococcus pneumoniae mutants having reduced susceptibility to moxifloxacin and levofloxacin. Antimicrob. Agents Chemother. 2002, 46, 522–524. [Google Scholar] [CrossRef]
- Takei, M.; Fukuda, H.; Kishii, R.; Hosaka, M. Target preference of 15 duinolones against Staphylococcus aureus, based on antibacterial activities and target inhibition. Antimicrob. Agents Chemother. 2001, 45, 3544–3547. [Google Scholar] [CrossRef]
- Varon, E.; Janoir, C.; Kitzis, M.D.; Gutmann, L. ParC and GyrA may be interchangeable initial targets of some fluoroquinolones in Streptococcus pneumoniae. Antimicrob. Agents Chemother. 1999, 43, 302–306. [Google Scholar] [CrossRef] [PubMed]
- Kishii, R.; Takei, M.; Fukuda, H.; Hayashi, K.; Hosaka, M. Contribution of the 8-methoxy group to the activity of gatifloxacin against type II topoisomerases of Streptococcus pneumoniae. Antimicrob. Agents Chemother. 2003, 47, 77–81. [Google Scholar] [CrossRef]
- Chan, P.F.; Germe, T.; Bax, B.D.; Huang, J.; Thalji, R.K.; Bacqué, E.; Checchia, A.; Chen, D.; Cui, H.; Ding, X.; et al. Thiophene antibacterials that allosterically stabilize DNA-cleavage complexes with DNA gyrase. Proc. Natl. Acad. Sci. USA 2017, 114, E4492–E4500. [Google Scholar] [CrossRef]
- Drlica, K.; Mustaev, A.; Towle, T.R.; Luan, G.; Kerns, R.J.; Berger, J.M. Bypassing fluoroquinolone resistance with quinazolinediones: Studies of drug–gyrase–DNA complexes having implications for drug design. ACS Chem. Biol. 2014, 9, 2895–2904. [Google Scholar] [CrossRef] [PubMed]
- Germe, T.; Vörös, J.; Jeannot, F.; Taillier, T.; Stavenger, R.A.; Bacqué, E.; Maxwell, A.; Bax, B.D. A new class of antibacterials, the imidazopyrazinones, reveal structural transitions involved in DNA gyrase poisoning and mechanisms of resistance. Nucleic Acids Res. 2018, 46, 4114–4128. [Google Scholar] [CrossRef] [PubMed]
- Jeannot, F.; Taillier, T.; Despeyroux, P.; Renard, S.; Rey, A.; Mourez, M.; Poeverlein, C.; Khichane, I.; Perrin, M.-A.; Versluys, S.; et al. Imidazopyrazinones (IPYs): Non-quinolone bacterial topoisomerase inhibitors showing partial cross-resistance with quinolones. J. Med. Chem. 2018, 61, 3565–3581. [Google Scholar] [CrossRef]
- Askoura, M.; Mottawea, W.; Abujamel, T.; Taher, I. Efflux pump inhibitors (EPIs) as new antimicrobial agents against Pseudomonas aeruginosa. Libyan J. Med. 2011, 6. [Google Scholar] [CrossRef]
- Lomovskaya, O.; Warren, M.S.; Lee, A.; Galazzo, J.; Fronko, R.; Lee, M.; Blais, J.; Cho, D.; Chamberland, S.; Renau, T.; et al. Identification and characterization of inhibitors of multidrug resistance efflux pumps in Pseudomonas aeruginosa: Novel agents for combination therapy. Antimicrob. Agents Chemother. 2001, 45, 105–116. [Google Scholar] [CrossRef]
- Mo, C.Y.; Manning, S.A.; Roggiani, M.; Culyba, M.J.; Samuels, A.N.; Sniegowski, P.D.; Goulian, M.; Kohli, R.M. Systematically altering bacterial SOS activity under stress reveals therapeutic strategies for potentiating antibiotics. mSphere 2016, 1, e00163-16. [Google Scholar] [CrossRef]
- Klitgaard, R.N.; Jana, B.; Guardabassi, L.; Nielsen, K.L.; Løbner-Olesen, A. DNA damage repair and drug efflux as potential targets for reversing low or intermediate ciprofloxacin resistance in E. coli K-12. Front. Microbiol. 2018, 9, 1438. [Google Scholar] [CrossRef]
- Recacha, E.; Machuca, J.; Díaz de Alba, P.; Ramos-Güelfo, M.; Docobo-Pérez, F.; Rodriguez-Beltrán, J.; Blázquez, J.; Pascual, A.; Rodríguez-Martínez, J.M. Quinolone resistance reversion by targeting the SOS response. mBio 2017, 8, e00971-17. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Rosado, A.I.; Valencia, E.Y.; Rodríguez-Rojas, A.; Costas, C.; Galhardo, R.S.; Rodríguez-Beltrán, J.; Blázquez, J. N-acetylcysteine blocks SOS induction and mutagenesis produced by fluoroquinolones in Escherichia coli. J. Antimicrob. Chemother. 2019, 74, 2188–2196. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutionalaffiliations. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bush, N.G.; Diez-Santos, I.; Abbott, L.R.; Maxwell, A. Quinolones: Mechanism, Lethality and Their Contributions to Antibiotic Resistance. Molecules 2020, 25, 5662. https://doi.org/10.3390/molecules25235662
Bush NG, Diez-Santos I, Abbott LR, Maxwell A. Quinolones: Mechanism, Lethality and Their Contributions to Antibiotic Resistance. Molecules. 2020; 25(23):5662. https://doi.org/10.3390/molecules25235662
Chicago/Turabian StyleBush, Natassja G., Isabel Diez-Santos, Lauren R. Abbott, and Anthony Maxwell. 2020. "Quinolones: Mechanism, Lethality and Their Contributions to Antibiotic Resistance" Molecules 25, no. 23: 5662. https://doi.org/10.3390/molecules25235662
APA StyleBush, N. G., Diez-Santos, I., Abbott, L. R., & Maxwell, A. (2020). Quinolones: Mechanism, Lethality and Their Contributions to Antibiotic Resistance. Molecules, 25(23), 5662. https://doi.org/10.3390/molecules25235662







