Identification and Detection of Bioactive Peptides in Milk and Dairy Products: Remarks about Agro-Foods
Abstract
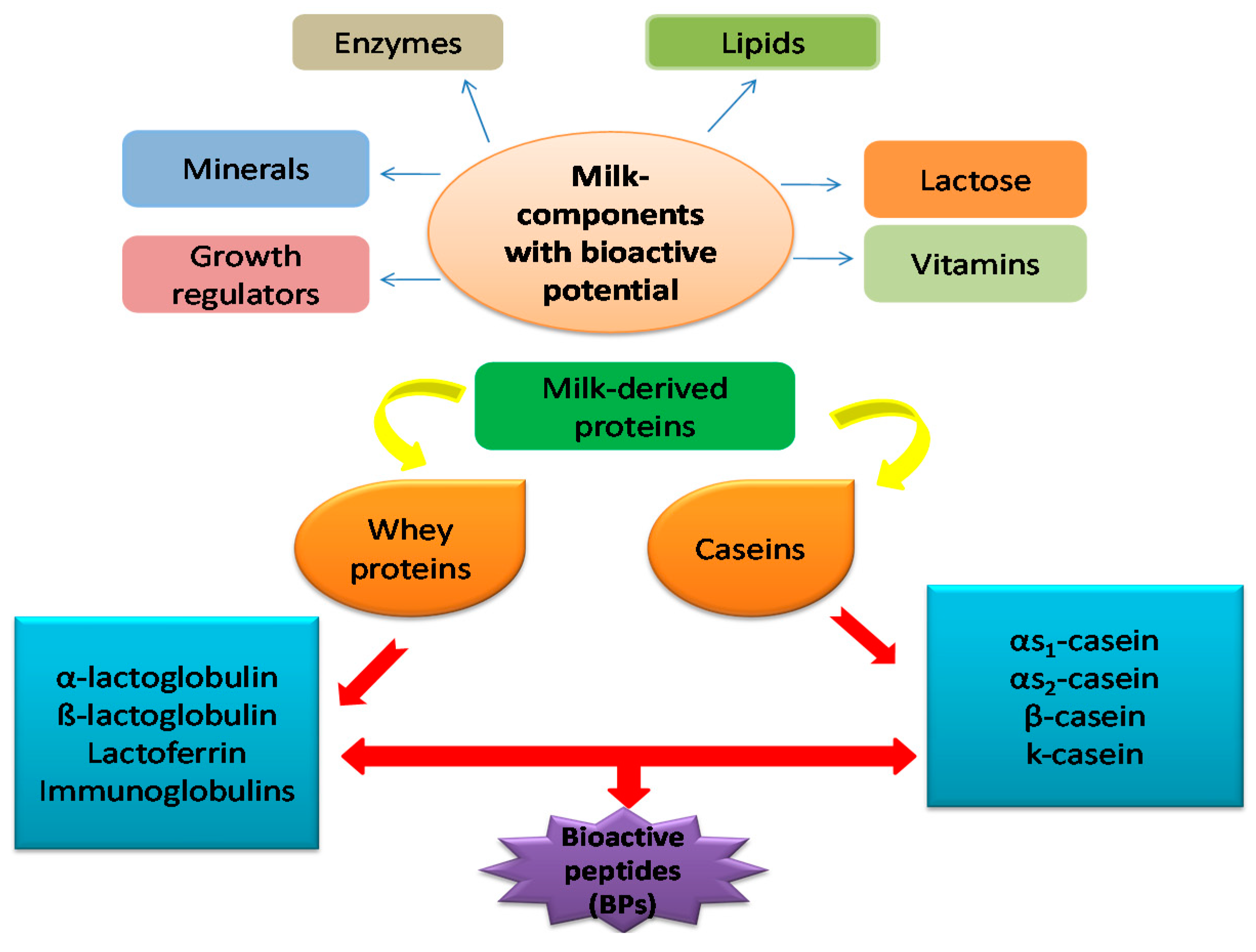
1. Introduction
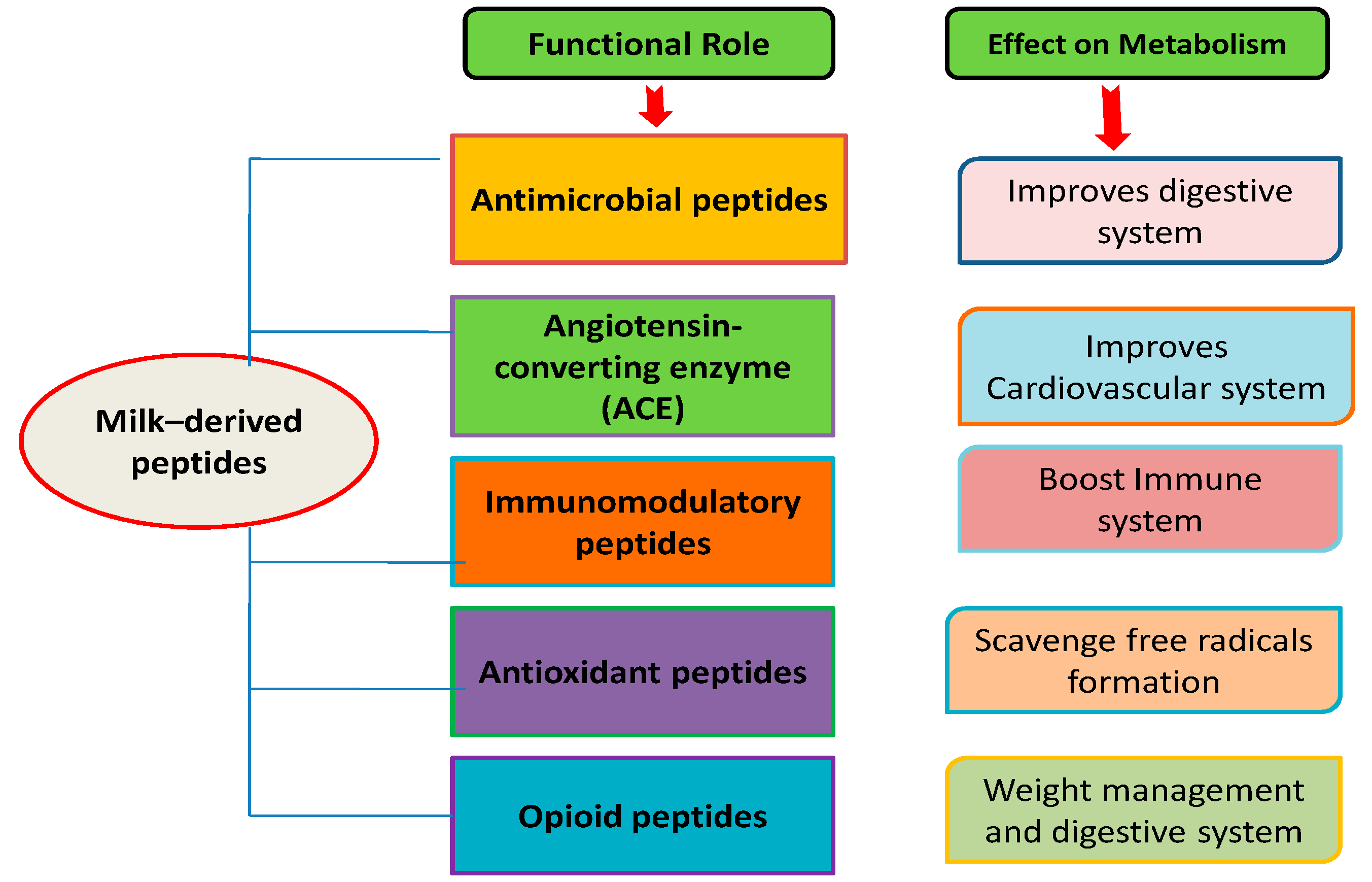
2. Primary Classes of Milk-Derived Bioactive Peptides
2.1. Antimicrobial Peptides
2.2. Angiotensin-Converting Enzyme (ACE)/Anti-Hypertensive Inhibitory Peptides
2.3. Immunomodulatory Peptides
2.4. Antioxidant Peptides
2.5. Opioid Peptides
3. Milk Fat Globule Membranes (MFGM): A Value-Added Product in Milk and Dairy Industry
4. Factors Affecting Milk Bioactive Peptides Composition and the Variations among Animal Genetics
5. Chemical Analysis of Individual Components of Milk and Dairy Products
5.1. Milk Components Analysis
5.1.1. Harmonized Methods
5.1.2. Chemical Analysis: Primary Methods
5.1.3. Electronic Analysis: Secondary Method
5.2. Dairy Product Composition Analysis
5.2.1. Sensory Analysis
5.2.2. Instrumental Analysis
6. Advanced Analytical Techniques of Milk and Dairy Products Analysis
6.1. Spectroscopic Detection
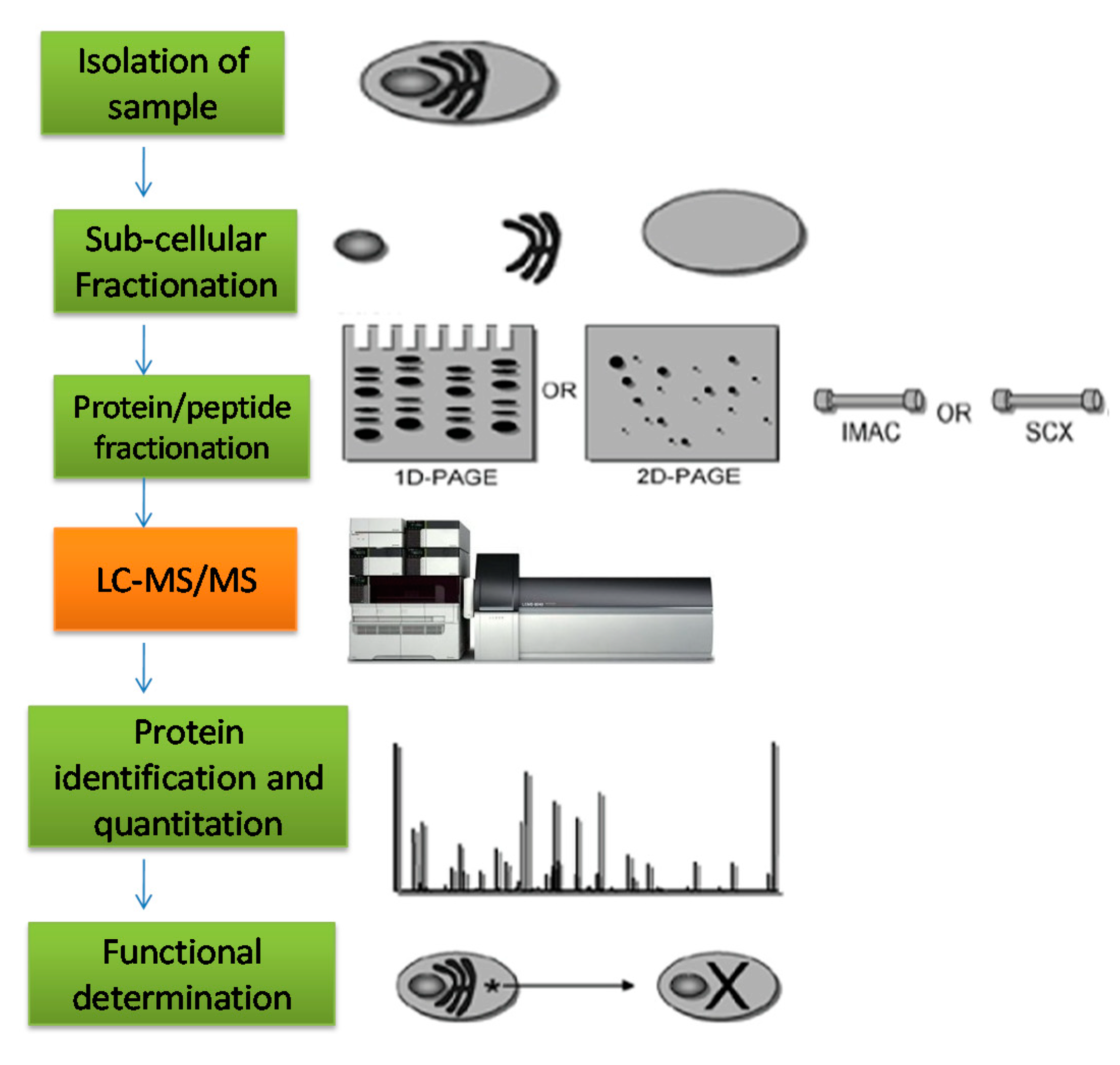
6.2. Peptidomics Profiling
6.3. Chromatography
6.4. Mass Spectrometry
6.5. Electrophoresis
6.6. Enzyme-Linked Immunosorbent Assay (ELISA)
6.7. Polymerase Chain Reaction (PCR) Based Detection
6.8. Role of Bioinformatics When Applied in Conjugation with Peptidomics
6.9. Endogenous Bioactive Peptides (BPs) Analysis
6.10. Coupling Different Techniques
7. Potential Roles and Applications for Milk and Dairy Industry Infant Formula Products
8. Knowledge Summary and Future Directions
9. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ACE | Angiotensin-converting enzyme |
| ADSA | American Dairy Science Association |
| AOACI | International Association of Official Analytical Chemists |
| API | Atmospheric pressure ionization |
| BPs | Bioactive peptides |
| CE | Capillary electrophoresis or |
| CMP | Caseino-macropeptide |
| DESI | Desorption electrospray ionization |
| ELISA | Enzyme-linked immunosorbent assay |
| ESI | Electrical pulverization |
| GC | Gas chromatography |
| IEC | Ion-exchange chromatography |
| Ig | Immunoglobulins |
| IR | Infrared |
| ISO | International Organization for Standardization |
| LC | Liquid chromatography |
| MALDI | Laser matrix-assisted desorption/ionization |
| MIR | Mid Infrared |
| MRM | Multiple reaction monitoring |
| MS | Mass spectrometry |
| NALDI | Nanostructure laser desorption/ionization |
| NIR | Near-infrared milk |
| PCR | Polymerase chain reaction |
| QDA | Quantitative descriptive analysis |
| SEC | Size-exclusion chromatography |
| USDA | United States Department of Agriculture |
References
- Bhattacharya, M.; Salcedo, J.; Robinson, R.C.; Henrick, B.M.; Barile, D. Peptidomic and glycomic profiling of commercial dairy products: Identification, quantification and potential bioactivities. NPJ Sci. Food 2019, 3, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, C.-C.; Hernández-Ledesma, B.; Fernández-Tomé, S.; Weinborn, V.; Barile, D.; de Moura Bell, J.M.L. Milk proteins, peptides, and oligosaccharides: Effects against the 21st century disorders. BioMed. Res. Int. 2015, 2015, 146840. [Google Scholar] [CrossRef] [PubMed]
- Kulinich, A.; Liu, L. Human milk oligosaccharides: The role in the fine-tuning of innate immune responses. Carbohydr. Res. 2016, 432, 62–70. [Google Scholar] [CrossRef]
- Smilowitz, J.T.; Lebrilla, C.B.; Mills, D.A.; German, J.B.; Freeman, S.L. Breast milk oligosaccharides: Structure-function relationships in the neonate. Annu. Rev. Nutr. 2014, 34, 143–169. [Google Scholar] [CrossRef] [PubMed]
- Andreas, N.J.; Kampmann, B.; Le-Doare, K.M. Human breast milk: A review on its composition and bioactivity. Early Hum. Dev. 2015, 91, 629–635. [Google Scholar] [CrossRef] [PubMed]
- Korhonen, H. Milk-derived bioactive peptides: From science to applications. J. Funct. Foods 2009, 1, 177–187. [Google Scholar] [CrossRef]
- Rai, M.; Da Silva, S.S. Nanotechnology for Bioenergy and Biofuel Production; Springer International Publishing: Berlin/Heidelberg, Germany, 2017. [Google Scholar] [CrossRef]
- Mohanty, D.P.; Mohapatra, S.; Misra, S.; Sahu, P.S. Milk derived bioactive peptides and their impact on human health–A review. Saudi J. Biol. Sci. 2016, 23, 577–583. [Google Scholar] [CrossRef]
- Gobbetti, M.; Stepaniak, L.; De Angelis, M.; Corsetti, A.; Di Cagno, R. Latent bioactive peptides in milk proteins: Proteolytic activation and significance in dairy processing. Crit. Rev. Food Sci. Nutr. 2002, 42, 223–239. [Google Scholar] [CrossRef]
- Marcone, S.; Belton, O.; Fitzgerald, D.J. Milk-derived bioactive peptides and their health promoting effects: A potential role in atherosclerosis. Br. J. Clin. Pharmacol. 2017, 83, 152–162. [Google Scholar] [CrossRef]
- Hernández-Ledesma, B.; Del Mar Contreras, M.; Recio, I. Antihypertensive peptides: Production, bioavailability and incorporation into foods. Adv. Colloid Interface Sci. 2011, 165, 23–35. [Google Scholar] [CrossRef]
- Lucey, J.A.; Otter, D.; Horne, D.S. A 100-year review: Progress on the chemistry of milk and its components. J. Dairy Sci. 2017, 100, 9916–9932. [Google Scholar] [CrossRef] [PubMed]
- Ricci-Cabello, I.; Olalla Herrera, M.; Artacho, R. Possible role of milk-derived bioactive peptides in the treatment and prevention of metabolic syndrome. Nutr. Rev. 2012, 70, 241–255. [Google Scholar] [CrossRef] [PubMed]
- Barbé, F.; Le Feunteun, S.; Rémond, D.; Ménard, O.; Jardin, J.; Henry, G.; Laroche, B.; Dupont, D. Tracking the in vivo release of bioactive peptides in the gut during digestion: Mass spectrometry peptidomic characterization of effluents collected in the gut of dairy matrix fed mini-pigs. Food Res. Int. 2014, 63, 147–156. [Google Scholar] [CrossRef]
- Jauhiainen, T.; Niittynen, L.; Orešič, M.; Järvenpää, S.; Hiltunen, T.P.; Rönnback, M.; Vapaatalo, H.; Korpela, R. Effects of long-term intake of lactotripeptides on cardiovascular risk factors in hypertensive subjects. Eur. J. Clin. Nutr. 2012, 66, 843–849. [Google Scholar] [CrossRef] [PubMed]
- Cicero, A.F.G.; Gerocarni, B.; Laghi, L.; Borghi, C. Blood pressure lowering effect of lactotripeptides assumed as functional foods: A meta-analysis of current available clinical trials. J. Hum. Hypertens. 2011, 25, 425–436. [Google Scholar] [CrossRef]
- Dupont, D.; Mandalari, G.; Mollé, D.; Jardin, J.; Rolet-Répécaud, O.; Duboz, G.; Léonil, J.; Mills, C.E.N.; Mackie, A.R. Food processing increases casein resistance to simulated infant digestion. Mol. Nutr. Food Res. 2010, 54, 1677–1689. [Google Scholar] [CrossRef]
- Kopf-Bolanz, K.A.; Schwander, F.; Gijs, M.; Vergeres, G.; Portmann, R.; Egger, L. Validation of an in vitro digestive system for studying macronutrient decomposition in humans. J. Nutr. 2012, 142, 245–250. [Google Scholar] [CrossRef]
- Picariello, G.; Ferranti, P.; Fierro, O.; Mamone, G.; Caira, S.; Di Luccia, A.; Monica, S.; Addeo, F. Peptides surviving the simulated gastrointestinal digestion of milk proteins: Biological and toxicological implications. J. Chromatogr. B 2010, 878, 295–308. [Google Scholar] [CrossRef]
- Lynch, J.M.; Barbano, D.M.; Healy, P.A.; Fleming, J.R. Effectiveness of temperature modification in decreasing the bias in milk fat test results between the Babcock and ether extraction methods. J. AOAC Int. 2003, 86, 768–774. [Google Scholar] [CrossRef]
- Vernile, A.; Beresford, T.P.; Spano, G.; Massa, S.; Fox, P.F. Chemical studies of Pecorino Siciliano cheese throughout ripening. Milchwissenschaft 2007, 62, 280–284. [Google Scholar]
- Hayaloglu, A.A.; Cakmakci, S.; Brechany, E.Y.; Deegan, K.C.; McSweeney, P.L.H. Microbiology, biochemistry, and volatile composition of Tulum cheese ripened in goat’s skin or plastic bags. J. Dairy Sci. 2007, 90, 1102–1121. [Google Scholar] [CrossRef]
- Roncada, P.; Gaviraghi, A.; Liberatori, S.; Canas, B.; Bini, L.; Greppi, G.F. Identification of caseins in goat milk. Proteomics 2002, 2, 723–726. [Google Scholar] [CrossRef]
- Pohlentz, G.; Kölbl, S.; Peter-Katalinić, J. High sequence coverage by in-capillary proteolysis of native proteins and simultaneous analysis of the resulting peptides by nanoelectrospray ionization-mass spectrometry and tandem mass spectrometry. Proteomics 2005, 5, 1758–1763. [Google Scholar] [CrossRef] [PubMed]
- Cunsolo, V.; Galliano, F.; Muccilli, V.; Saletti, R.; Marletta, D.; Bordonaro, S.; Foti, S. Detection and characterization by high-performance liquid chromatography and mass spectrometry of a goat β-casein associated with a CSN2 null allele. Rapid Commun. Mass Spectrom. 2005, 19, 2943–2949. [Google Scholar] [CrossRef] [PubMed]
- Cunsolo, V.; Muccilli, V.; Saletti, R.; Marletta, D.; Foti, S. Detection and characterization by high-performance liquid chromatography and mass spectrometry of two truncated goat αs2-caseins. Rapid Commun. Mass Spectrom. 2006, 20, 1061–1070. [Google Scholar] [CrossRef]
- Moreno, F.J.; Recio, I.; Olano, A.; López-Fandiño, R. Heterogeneity of caprine κ-casein macropeptide. J. Dairy Res. 2001, 68, 197–208. [Google Scholar] [CrossRef] [PubMed]
- Moreno, F.J.; Recio, I.; Olano, A.; López-Fandiño, R. Chromatographic characterization of ovine κ-casein macropeptide. J. Dairy Res. 2000, 67, 349–359. [Google Scholar] [CrossRef]
- Mamone, G.; Caira, S.; Garro, G.; Nicolai, A.; Ferranti, P.; Picariello, G.; Malorni, A.; Chianese, L.; Addeo, F. Casein phosphoproteome: Identification of phosphoproteins by combined mass spectrometry and two-dimensional gel electrophoresis. Electrophoresis 2003, 24, 2824–2837. [Google Scholar] [CrossRef] [PubMed]
- Claverol, S.; Burlet-Schiltz, O.; Gairin, J.E.; Monsarrat, B. Characterization of protein variants and post-translational modifications: ESI-MSn analyses of intact proteins eluted from polyacrylamide gels. Mol. Cell. Proteom. 2003, 2, 483–493. [Google Scholar] [CrossRef]
- Panchaud, A.; Kussmann, M.; Affolter, M. Rapid enrichment of bioactive milk proteins and iterative, consolidated protein identification by multidimensional protein identification technology. Proteomics 2005, 5, 3836–3846. [Google Scholar] [CrossRef]
- Reinhardt, T.A.; Lippolis, J.D. Developmental changes in the milk fat globule membrane proteome during the transition from colostrum to milk. J. Dairy Sci. 2008, 91, 2307–2318. [Google Scholar] [CrossRef]
- Marvin, L.F.; Parisod, V.; Fay, L.B.; Guy, P.A. Characterization of lactosylated proteins of infant formula powders using two-dimensional gel electrophoresis and nanoelectrospray mass spectrometry. Electrophoresis 2002, 23, 2505–2512. [Google Scholar] [CrossRef]
- Mollé, D.; Morgan, F.; Bouhallab, S.; Léonil, J. Selective detection of lactolated peptides in hydrolysates by liquid chromatography/electrospray tandem mass spectrometry. Anal. Biochem. 1998, 259, 152–161. [Google Scholar] [CrossRef] [PubMed]
- Soeryapranata, E.; Powers, J.R.; Ünlü, G. Degradation of αs1-CN f1-23 by aminopeptidase N and endopeptidases E, O, O2, and O3 of Lactobacillus helveticus WSU19 under cheese ripening conditions. Int. Dairy J. 2008, 18, 178–186. [Google Scholar] [CrossRef]
- Soeryapranata, E.; Powers, J.R.; Ünlü, G. Cloning and characterization of debittering peptidases, PepE, PepO, PepO2, PepO3, and PepN, of Lactobacillus helveticus WSU19. Int. Dairy J. 2007, 17, 1096–1106. [Google Scholar] [CrossRef]
- Plath, A.; Krause, I.; Einspanier, R. Species identification in dairy products by three different DNA-based techniques. Z. für Leb. Und-forsch. A 1997, 205, 437–441. [Google Scholar] [CrossRef]
- Branciari, R.; NIJMAN, I.J.; Plas, M.E.; DI ANTONIO, E.; Lenstra, J.A. Species origin of milk in Italian mozzarella and Greek feta cheese. J. Food Prot. 2000, 63, 408–411. [Google Scholar] [CrossRef] [PubMed]
- Klotz, A.; Einspanier, R. ORIGINAL PAPERS-Development of a DNA-based screening method to detect cow milk in ewe, goat and buffalo milk and dairy products using PCR-LCR-EIA-technique. Milchwissenschaft 2001, 56, 67–69. [Google Scholar]
- Rea, S.; Chikuni, K.; Branciari, R.; Sangamayya, R.A.M.S.; Ranucci, D.; Avellini, P. Use of duplex polymerase chain reaction (duplex-PCR) technique to identify bovine and water buffalo milk used in making mozzarella cheese. J. Dairy Res. 2001, 68, 689–698. [Google Scholar] [CrossRef] [PubMed]
- Mafra, I.; Roxo, Á.; Ferreira, I.M.; Oliveira, M.B.P.P. A duplex polymerase chain reaction for the quantitative detection of cows’ milk in goats’ milk cheese. Int. Dairy J. 2007, 17, 1132–1138. [Google Scholar] [CrossRef]
- Lopparelli, R.M.; Cardazzo, B.; Balzan, S.; Giaccone, V.; Novelli, E. Real-time TaqMan polymerase chain reaction detection and quantification of cow DNA in pure water buffalo mozzarella cheese: Method validation and its application on commercial samples. J. Agric. Food Chem. 2007, 55, 3429–3434. [Google Scholar] [CrossRef]
- Levieux, D.; Venien, A. Rapid, sensitive two-site ELISA for detection of cows’ milk in goats’ or ewes’ milk using monoclonal antibodies. J. Dairy Res. 1994, 61, 91–99. [Google Scholar] [CrossRef]
- Richmond, H.D. Dairy Chemistry: A Practical Handbook for Dairy Chemists and Others Having Control of Dairies; Charles Griffin & Co Ltd.: London, UK, 1920. [Google Scholar]
- Muehlhoff, E.; Bennett, A.; McMahon, D. Milk and Dairy Products in Human Nutrition; Food and Agriculture Organization of the United Nations (FAO): Rome, Italy, 2013; ISBN 978-92-5-107863-1. [Google Scholar]
- Khan, I.T.; Nadeem, M.; Imran, M.; Ullah, R.; Ajmal, M.; Jaspal, M.H. Antioxidant properties of Milk and dairy products: A comprehensive review of the current knowledge. Lipids Health Dis. 2019, 18, 1–13. [Google Scholar] [CrossRef]
- Saxelin, M.; Korpela, R.; Mäyrä-Mäkinen, A. Introduction: Classifying functional dairy products. In Functional Dairy Products; Mattila-Sandholm, T., Saarela, M., Eds.; Woodhead Publishing Ltd. and CRC Press LLC: Cambridge, UK, 2003. [Google Scholar]
- Wang, Y.-C.; Yu, R.-C.; Chou, C.-C. Antioxidative activities of soymilk fermented with lactic acid bacteria and bifidobacteria. Food Microbiol. 2006, 23, 128–135. [Google Scholar] [CrossRef] [PubMed]
- Usta, B.; Yılmaz-Ersan, L. Antioxidant enzymes of milk and their biological effects. Ziraat Fakültesi Derg. Uludağ Üniversitesi 2013, 27, 123–130. [Google Scholar]
- Mohanty, D.P.; Tripathy, P.; Mohapatra, S.; Samantaray, D.P. Bioactive potential assessment of antibacterial peptide produced by Lactobacillus isolated from milk and milk products. Int. J. Curr. Microbiol. Appl. Sci 2014, 3, 72–80. [Google Scholar]
- Lahov, E.; Regelson, W. Antibacterial and immunostimulating casein-derived substances from milk: Casecidin, isracidin peptides. Food Chem. Toxicol. 1996, 34, 131–145. [Google Scholar] [CrossRef]
- Zucht, H.-D.; Raida, M.; Adermann, K.; Mägert, H.-J.; Forssmann, W.-G. Casocidin-I: A casein-αs2 derived peptide exhibits antibacterial activity. FEBS Lett. 1995, 372, 185–188. [Google Scholar] [CrossRef]
- Recio, I.; Visser, S. Two ion-exchange chromatographic methods for the isolation of antibacterial peptides from lactoferrin: In situ enzymatic hydrolysis on an ion-exchange membrane. J. Chromatogr. A 1999, 831, 191–201. [Google Scholar] [CrossRef]
- van der Kraan, M.I.A.; Nazmi, K.; Teeken, A.; Groenink, J.; van’t Hof, W.; Veerman, E.C.I.; Bolscher, J.G.M.; Amerongen, A.V.N. Lactoferrampin, an antimicrobial peptide of bovine lactoferrin, exerts its candidacidal activity by a cluster of positively charged residues at the C-terminus in combination with a helix-facilitating N-terminal part. Biol. Chem. 2005, 386, 137–142. [Google Scholar] [CrossRef]
- Farrell, H.M., Jr.; Jimenez-Flores, R.; Bleck, G.T.; Brown, E.M.; Butler, J.E.; Creamer, L.K.; Hicks, C.L.; Hollar, C.M.; Ng-Kwai-Hang, K.F.; Swaisgood, H.E. Nomenclature of the proteins of cows’ milk—Sixth revision. J. Dairy Sci. 2004, 87, 1641–1674. [Google Scholar] [CrossRef]
- Manso, M.A.; Lopez-Fandino, R. κ-casein macropeptides from cheese whey: Physicochemical, biological, nutritional, and technological features for possible uses. Food Rev. Int. 2004, 20, 329–355. [Google Scholar] [CrossRef]
- Bellamy, W.; Wakabayashi, H.; Takase, M.; Kawase, K.; Shimamura, S.; Tomita, M. Killing of Candida albicans by lactoferricin B, a potent antimicrobial peptide derived from the N-terminal region of bovine lactoferrin. Med. Microbiol. Immunol. 1993, 182, 97–105. [Google Scholar] [CrossRef] [PubMed]
- Farnaud, S.; Evans, R.W. Lactoferrin—a multifunctional protein with antimicrobial properties. Mol. Immunol. 2003, 40, 395–405. [Google Scholar] [CrossRef]
- Pan, Y.; Rowney, M.; Guo, P.; Hobman, P. Biological properties of lactoferrin: An overview. Aust. J. Dairy Technol. 2007, 62, 31. [Google Scholar]
- Korhonen, H.; Pihlanto, A. Technological options for the production of health-promoting proteins and peptides derived from milk and colostrum. Curr. Pharm. Des. 2007, 13, 829–843. [Google Scholar] [CrossRef]
- FitzGerald, R.J.; Murray, B.A.; Walsh, D.J. Hypotensive peptides from milk proteins. J. Nutr. 2004, 134, 980S–988S. [Google Scholar] [CrossRef]
- Vercruysse, L.; Van Camp, J.; Smagghe, G. ACE inhibitory peptides derived from enzymatic hydrolysates of animal muscle protein: A review. J. Agric. Food Chem. 2005, 53, 8106–8115. [Google Scholar] [CrossRef]
- FitzGerald, R.J.; Meisel, H. Milk protein-derived peptide inhibitors of angiotensin-I-converting enzyme. Br. J. Nutr. 2000, 84, 33–37. [Google Scholar] [CrossRef]
- Nakamura, Y.; Yamamoto, N.; Sakai, K.; Takano, T. Antihypertensive effect of sour milk and peptides isolated from it that are inhibitors to angiotensin I-converting enzyme. J. Dairy Sci. 1995, 78, 1253–1257. [Google Scholar] [CrossRef]
- Aihara, K.; Ishii, H.; Yoshida, M. Casein-derived tripeptide, Val-Pro-Pro (VPP), modulates monocyte adhesion to vascular endothelium. J. Atheroscler. Thromb. 2009, 16, 594–603. [Google Scholar] [CrossRef]
- Seppo, L.; Jauhiainen, T.; Poussa, T.; Korpela, R. A fermented milk high in bioactive peptides has a blood pressure–lowering effect in hypertensive subjects. Am. J. Clin. Nutr. 2003, 77, 326–330. [Google Scholar] [CrossRef]
- Murakami, M.; Tonouchi, H.; Takahashi, R.; Kitazawa, H.; Kawai, Y.; Negishi, H.; Saito, T. Structural analysis of a new anti-hypertensive peptide (β-lactosin B) isolated from a commercial whey product. J. Dairy Sci. 2004, 87, 1967–1974. [Google Scholar] [CrossRef]
- Saito, T.; Nakamura, T.; Kitazawa, H.; Kawai, Y.; Itoh, T. Isolation and structural analysis of antihypertensive peptides that exist naturally in Gouda cheese. J. Dairy Sci. 2000, 83, 1434–1440. [Google Scholar] [CrossRef]
- Bracquart, P.; Lorient, D. Effet des Acides Amines et Peptides Sur la Croissance de Streptococcus Thermophilus. iii: Peptides Comportant Glu, His et Met. Milchwissenschaft 1979, 34, 676–679. [Google Scholar]
- Maruyama, S.; Suzuki, H. A peptide inhibitor of angiotensin I converting enzyme in the tryptic hydrolysate of casein. Agric. Biol. Chem. 1982, 46, 1393–1394. [Google Scholar]
- Maruyama, S.; Nakagomi, K.; Tomizuka, N.; Suzuki, H. Angiotensin I-converting enzyme inhibitor derived from an enzymatic hydrolysate of casein. II. Isolation and bradykinin-potentiating activity on the uterus and the ileum of rats. Agric. Biol. Chem. 1985, 49, 1405–1409. [Google Scholar]
- Maruyama, S.; Mitachi, H.; Awaya, J.; Kurono, M.; Tomizuka, N.; Suzuki, H. Angiotensin I-converting enzyme inhibitory activity of the C-terminal hexapeptide of αs1-casein. Agric. Biol. Chem. 1987, 51, 2557–2561. [Google Scholar] [CrossRef]
- Perpetuo, E.A.; Juliano, L.; Lebrun, I. Biochemical and pharmacological aspects of two bradykinin-potentiating peptides obtained from tryptic hydrolysis of casein. J. Protein Chem. 2003, 22, 601–606. [Google Scholar] [CrossRef]
- Migliore-Samour, D.; Jolles, P. Casein, a prohormone with an immunomodulating role for the newborn? Experientia 1988, 44, 188–193. [Google Scholar] [CrossRef]
- Clare, D.A.; Catignani, G.L.; Swaisgood, H.E. Biodefense properties of milk: The role of antimicrobial proteins and peptides. Curr. Pharm. Des. 2003, 9, 1239–1255. [Google Scholar] [CrossRef] [PubMed]
- Gill, H.S.; Doull, F.; Rutherfurd, K.J.; Cross, M.L. Immunoregulatory peptides in bovine milk. Br. J. Nutr. 2000, 84, 111–117. [Google Scholar] [CrossRef] [PubMed]
- Meisel, H.; FitzGerald, R.J. Biofunctional peptides from milk proteins: Mineral binding and cytomodulatory effects. Curr. Pharm. Des. 2003, 9, 1289–1296. [Google Scholar] [PubMed]
- Monnai, M.; Horimoto, Y.; Otani, H. Immunomodificatory effect of dietary bovine kappa-caseinoglycopeptide on serum antibody levels and proliferative responses of lymphocytes in mice. Milchwissenschaft 1998, 53, 129–132. [Google Scholar]
- Abuja, P.M.; Albertini, R. Methods for monitoring oxidative stress, lipid peroxidation and oxidation resistance of lipoproteins. Clin. Chim. Acta 2001, 306, 1–17. [Google Scholar] [CrossRef]
- Halliwell, B. Lipid peroxidation, antioxidants and cardiovascular disease: How should we move forward? Cardiovasc. Res. 2000, 47, 410–418. [Google Scholar] [CrossRef]
- Halliwell, B.; Whiteman, M. Measuring reactive species and oxidative damage in vivo and in cell culture: How should you do it and what do the results mean? Br. J. Pharmacol. 2004, 142, 231–255. [Google Scholar] [CrossRef]
- Korhonen, H.; Pihlanto, A. Food-derived bioactive peptides-opportunities for designing future foods. Curr. Pharm. Des. 2003, 9, 1297–1308. [Google Scholar] [CrossRef]
- Cervato, G.; Cazzola, R.; Cestaro, B. Studies on the antioxidant activity of milk caseins. Int. J. Food Sci. Nutr. 1999, 50, 291–296. [Google Scholar]
- Wong, P.Y.Y.; Kitts, D.D. Chemistry of buttermilk solid antioxidant activity. J. Dairy Sci. 2003, 86, 1541–1547. [Google Scholar] [CrossRef]
- Suetsuna, K.; Ukeda, H.; Ochi, H. Isolation and characterization of free radical scavenging activities peptides derived from casein. J. Nutr. Biochem. 2000, 11, 128–131. [Google Scholar] [CrossRef]
- Rival, S.G.; Boeriu, C.G.; Wichers, H.J. Caseins and casein hydrolysates. 2. Antioxidative properties and relevance to lipoxygenase inhibition. J. Agric. Food Chem. 2001, 49, 295–302. [Google Scholar] [CrossRef] [PubMed]
- Okada, Y.; Okada, M. Scavenging effect of water soluble proteins in broad beans on free radicals and active oxygen species. J. Agric. Food Chem. 1998, 46, 401–406. [Google Scholar] [CrossRef] [PubMed]
- Lindmark-Månsson, H.; Åkesson, B. Antioxidative factors in milk. Br. J. Nutr. 2000, 84, 103–110. [Google Scholar] [CrossRef] [PubMed]
- Brantl, V. Novel opioid peptides derived from human β-casein: Human β-casomorphins. Eur. J. Pharmacol. 1984, 106, 213–214. [Google Scholar] [CrossRef]
- Molina, P.E.; Abumrad, N.N. Metabolic effects of opiates and opioid peptides. Adv. Neuroimmunol. 1994, 4, 105–116. [Google Scholar] [CrossRef]
- Dziuba, J.; Minkiewicz, P.; Nałecz, D.; Iwaniak, A. Database of biologically active peptide sequences. Food/Nahrung 1999, 43, 190–195. [Google Scholar] [CrossRef]
- Calvo, C.; Cesselin, F.; Gelman, M.; Glowinski, J. Identification of an opioid peptide secreted by rat embryonic mixed brain cells as a promoter of macrophage migration. Eur. J. Neurosci. 2000, 12, 2676–2684. [Google Scholar] [CrossRef]
- Sturner, R.A.; Chang, K.J. Opioid peptide content in infant formulas. Pediatr. Res. 1988, 23, 4–10. [Google Scholar]
- Tomé, D.; Debabbi, H. Physiological effects of milk protein components. Int. Dairy J. 1998, 8, 383–392. [Google Scholar] [CrossRef]
- Meisel, H.; Fitzgerald, R.J. Opioid peptides encrypted in intact milk protein sequences. Br. J. Nutr. 2000, 84, 27–31. [Google Scholar] [CrossRef] [PubMed]
- Meisel, H. Overview on milk protein-derived peptides. Int. Dairy J. 1998, 8, 363–373. [Google Scholar] [CrossRef]
- Dong, X.M.; Revol, J.-F.; Gray, D.G. Effect of microcrystallite preparation conditions on the formation of colloid crystals of cellulose. Cellulose 1998, 5, 19–32. [Google Scholar] [CrossRef]
- Daniel, H.; Vohwinkel, M.; Rehner, G. Effect of casein and β-casomorphins on gastrointestinal motility in rats. J. Nutr. 1990, 120, 252–257. [Google Scholar] [CrossRef]
- Matthies, H.; Stark, H.; Hartrodt, B.; Ruethrich, H.-L.; Spieler, H.-T.; Barth, A.; Neubert, K. Derivatives of β-casomorphins with high analgesic potency. Peptides 1984, 5, 463–470. [Google Scholar] [CrossRef]
- Meisel, H.; Schlimme, E. Milk proteins: Precursors of bioactive peptides. Trends Food Sci. Technol. 1990, 1, 41–43. [Google Scholar] [CrossRef]
- Tokas, J.; Punia, H. Milk Fat Globule Membrane (MFGM): An Ingredient of Dairy Products as Nutraceutical. Trends Pept. Protein Sci. 2019, 4, 1–7. [Google Scholar]
- Saeland, E.; de Jong, M.A.W.P.; Nabatov, A.A.; Kalay, H.; Geijtenbeek, T.B.H.; van Kooyk, Y. MUC1 in human milk blocks transmission of human immunodeficiency virus from dendritic cells to T cells. Mol. Immunol. 2009, 46, 2309–2316. [Google Scholar] [CrossRef]
- Matsumoto, M.; Hara, K.; Kimata, H.; Benno, Y.; Shimamoto, C. Exfoliation of Helicobacter pylori from gastric mucin by glycopolypeptides from buttermilk. J. Dairy Sci. 2005, 88, 49–54. [Google Scholar] [CrossRef]
- Martin, H.M.; Hancock, J.T.; Salisbury, V.; Harrison, R. Role of xanthine oxidoreductase as an antimicrobial agent. Infect. Immun. 2004, 72, 4933–4939. [Google Scholar] [CrossRef] [PubMed]
- Phan, T.T.Q.; Asaduzzaman, M.; Le, T.T.; Fredrick, E.; Van der Meeren, P.; Dewettinck, K. Composition and emulsifying properties of a milk fat globule membrane enriched material. Int. Dairy J. 2013, 29, 99–106. [Google Scholar] [CrossRef]
- El-Loly, M. Composition, properties and nutritional aspects of milk fat globule membrane-a review. Polish J. Food Nutr. Sci. 2011, 61, 7–32. [Google Scholar] [CrossRef]
- Cavaletto, M.; Giuffrida, M.G.; Conti, A. Milk fat globule membrane components—A proteomic approach. Adv. Exp. Med. Biol. 2008, 606, 129–141. [Google Scholar]
- Kvistgaard, A.S.; Pallesen, L.T.; Arias, C.F.; Lopez, S.; Petersen, T.E.; Heegaard, C.W.; Rasmussen, J.T. Inhibitory effects of human and bovine milk constituents on rotavirus infections. J. Dairy Sci. 2004, 87, 4088–4096. [Google Scholar] [CrossRef]
- Gallier, S.; Gragson, D.; Jiménez-Flores, R.; Everett, D.W. β-Casein–phospholipid monolayers as model systems to understand lipid–protein interactions in the milk fat globule membrane. Int. Dairy J. 2012, 22, 58–65. [Google Scholar] [CrossRef]
- Dewettinck, K.; Rombaut, R.; Thienpont, N.; Le, T.T.; Messens, K.; Van Camp, J. Nutritional and technological aspects of milk fat globule membrane material. Int. Dairy J. 2008, 18, 436–457. [Google Scholar] [CrossRef]
- Albenzio, M.; Caroprese, M.; Santillo, A.; Marino, R.; Taibi, L.; Sevi, A. Effects of somatic cell count and stage of lactation on the plasmin activity and cheese-making properties of ewe milk. J. Dairy Sci. 2004, 87, 533–542. [Google Scholar] [CrossRef]
- Albenzio, M.; Caroprese, M.; Santillo, A.; Marino, R.; Muscio, A.; Sevi, A. Proteolytic patterns and plasmin activity in ewes’ milk as affected by somatic cell count and stage of lactation. J. Dairy Res. 2005, 72, 86–92. [Google Scholar] [CrossRef]
- Albenzio, M.; Santillo, A.; d’Angelo, F.; Sevi, A. Focusing on casein gene cluster and protein profile in Garganica goat milk. J. Dairy Res. 2009, 76, 83. [Google Scholar] [CrossRef]
- Kelly, A.L.; O’Flaherty, F.; Fox, P.F. Indigenous proteolytic enzymes in milk: A brief overview of the present state of knowledge. Int. Dairy J. 2006, 16, 563–572. [Google Scholar] [CrossRef]
- Yamamoto, N.; Akino, A.; Takano, T. Antihypertensive effect of the peptides derived from casein by an extracellular proteinase from Lactobacillus helveticus CP790. J. Dairy Sci. 1994, 77, 917–922. [Google Scholar] [CrossRef]
- Recio, I.; Quirós, A.; Hernández-Ledesma, B.; Gómez-Ruiz, J.A.; Miguel, M.; Amigo, L.; López-Expósito, I.; Ramos, M.; Aleixandre, A. Bioactive peptides identified in enzyme hydrolysates from milk caseins and procedure for their obtention. European Patent PCT/ES 2,006,070,079, 8 June 2006. [Google Scholar]
- Quirós, A.; Ramos, M.; Muguerza, B.; Delgado, M.A.; Miguel, M.; Aleixandre, A.; Recio, I. Identification of novel antihypertensive peptides in milk fermented with Enterococcus faecalis. Int. Dairy J. 2007, 17, 33–41. [Google Scholar] [CrossRef]
- Vargas-Bello-Pérez, E.; Márquez-Hernández, R.I.; Hernández-Castellano, L.E. Bioactive peptides from milk: Animal determinants and their implications in human health. J. Dairy Res. 2019, 86, 136–144. [Google Scholar] [CrossRef]
- Walker, G.P.; Dunshea, F.R.; Doyle, P.T. Effects of nutrition and management on the production and composition of milk fat and protein: A review. Aust. J. Agric. Res. 2004, 55, 1009–1028. [Google Scholar] [CrossRef]
- Sok, M.; Ouellet, D.R.; Firkins, J.L.; Pellerin, D.; Lapierre, H. Amino acid composition of rumen bacteria and protozoa in cattle. J. Dairy Sci. 2017, 100, 5241–5249. [Google Scholar] [CrossRef]
- Tacoma, R.; Fields, J.; Ebenstein, D.B.; Lam, Y.-W.; Greenwood, S.L. Characterization of the bovine milk proteome in early-lactation Holstein and Jersey breeds of dairy cows. J. Proteom. 2016, 130, 200–210. [Google Scholar] [CrossRef] [PubMed]
- Bernabucci, U.; Basiricò, L.; Morera, P.; Dipasquale, D.; Vitali, A.; Cappelli, F.P.; Calamari, L. Effect of summer season on milk protein fractions in Holstein cows. J. Dairy Sci. 2015, 98, 1815–1827. [Google Scholar] [CrossRef] [PubMed]
- Gandini, G.; Turri, F.; Rizzi, R.; Crotta, M.; Minozzi, G.; Pizzi, F. Economic evaluation of genetic improvement in local breeds: The case of the Verzaschese goat. Ital. J. Anim. Sci. 2017, 16, 199–207. [Google Scholar] [CrossRef]
- Roman, S.; Sánchez-Siles, L.M.; Siegrist, M. The importance of food naturalness for consumers: Results of a systematic review. Trends food Sci. Technol. 2017, 67, 44–57. [Google Scholar] [CrossRef]
- ISO/IEC 17025-General Requirements for the Competence of Testing and Calibration Laboratories; International Organization for Standardization: Geneva, Switzerland, 1999.
- International, A. Guidelines for collaborative study procedures to validate characteristics of a method of analysis. J. AOAC Int. 1995, 78, 143A–160A. [Google Scholar]
- Ellison, S.; Williams, A.; da Silva, R.B.; Bremser, W.; Brzyski, A.; Fodor, P.; Kaarls, R.; Kaus, R.; Magnuson, B.; di Meane, E.A.; et al. Quantifying Uncertainty in Analytical Measurements; EURACHEM: Caparica, Portugal, 2000. [Google Scholar]
- Ellison, S.L.R.; Williams, A. Disseminating traceability in chemical measurement: Principles of a new EURACHEM/CITAC guide. Accredit. Qual. Assur. 2003, 8, 483–485. [Google Scholar] [CrossRef]
- Lynch, J.M.; Barbano, D.M.; Healy, P.A.; Richard Fleming, J. Performance evaluation of the Babcock and ether extraction methods: 1989 through 1992. J. AOAC Int. 1994, 77, 976–981. [Google Scholar] [CrossRef]
- Lynch, J.M.; Barbano, D.M.; Healy, P.A.; Fleming, J.R. Performance evaluation of direct forced-air total solids and Kjeldahl total nitrogen methods: 1990 through 1995. J. AOAC Int. 1997, 80, 1038–1043. [Google Scholar] [CrossRef] [PubMed]
- Barbano, D.M.; Lynch, J.M. Crude and protein nitrogen bases for protein measurement and their impact on current testing accuracy. J. Dairy Sci. 1992, 75, 3210–3217. [Google Scholar] [CrossRef]
- Barbano, D.M.; Clark, J.L.; Dunham, C.E. Comparison of Babcock and ether extraction methods for determination of fat content of milk: Collaborative study. J. Assoc. Off. Anal. Chem. 1988, 71, 898–914. [Google Scholar] [CrossRef] [PubMed]
- Barbano, D.M.; Clark, J.L.; Dunham, C.E.; Flemin, R.J. Kjeldahl method for determination of total nitrogen content of milk: Collaborative study. J. Assoc. Off. Anal. Chem. 1990, 73, 849–859. [Google Scholar] [CrossRef]
- Barbano, D.M.; Lynch, J.M.; Fleming, J.R. Direct and indirect determination of true protein content of milk by Kjeldahl analysis: Collaborative study. J. Assoc. Off. Anal. Chem. 1991, 74, 281–288. [Google Scholar] [CrossRef]
- Lynch, J.M.; Barbano, D.M.; Fleming, J.R. Indirect and direct determination of the casein content of milk by Kjeldahl nitrogen analysis: Collaborative study. J. AOAC Int. 1998, 81, 763–774. [Google Scholar] [CrossRef]
- Lynch, J.M.; Barbano, D.M. Kjeldahl nitrogen analysis as a reference method for protein determination in dairy products. J. AOAC Int. 1999, 82, 1389–1398. [Google Scholar] [CrossRef]
- Clark, J.L.; Barbano, D.M.; Dunham, C.E. Comparison of two methods for determination of total solids content of milk: Collaborative study. J. Assoc. Off. Anal. Chem. 1989, 72, 712–718. [Google Scholar] [CrossRef]
- Agnet, Y. Fourier transform infrared spectrometry. Int. Dairy Fed. Bull. 1998, 332, 58–68. [Google Scholar]
- McKenna, D. 4. Standardization in cheese plants. In Practical Guide for Control of Cheese Yield; International Dairy Federation: Brussels, Belgium, 2000; pp. 28–35. [Google Scholar]
- McBride, R.L.; Hall, C. Cheese grading versus consumer acceptability: An inevitable discrepancy. Aust. J. Dairy Technol. 1979, 34, 66. [Google Scholar]
- Lawless, H.T.; Heymann, H. Sensory Evaluation of Food. Principles and Practices; Springer: New York, NY, USA, 1999. [Google Scholar]
- Drake, M.A.; McIngvale, S.C.; Gerard, P.D.; Cadwallader, K.R.; Civille, G.V. Development of a descriptive language for Cheddar cheese. J. Food Sci. 2001, 66, 1422–1427. [Google Scholar] [CrossRef]
- Acree, T.E.; Barnard, J.; Cunningham, D.G. A procedure for the sensory analysis of gas chromatographic effluents. Food Chem. 1984, 14, 273–286. [Google Scholar] [CrossRef]
- Karagül-Yüceer, Y.; Drake, M.A.; Cadwallader, K.R. Aroma-active components of liquid cheddar whey. J. Food Sci. 2003, 68, 1215–1219. [Google Scholar] [CrossRef]
- Carpino, S.; Mallia, S.; La Terra, S.; Melilli, C.; Licitra, G.; Acree, T.E.; Barbano, D.M.; Van Soest, P.J. Composition and aroma compounds of Ragusano cheese: Native pasture and total mixed rations. J. Dairy Sci. 2004, 87, 816–830. [Google Scholar] [CrossRef]
- Xue, H.; Hu, W.; Son, H.; Han, Y.; Yang, Z. Indirect ELISA for detection and quantification of bovine milk in goat milk. Food Sci. 2010, 24. [Google Scholar]
- Stănciuc, N.; Râpeanu, G. Identification of adulterated sheep and goat cheeses marketed in Romania by immunocromatographic assay. Food Agric. Immunol. 2010, 21, 157–164. [Google Scholar] [CrossRef]
- Dallas, D.C.; Guerrero, A.; Khaldi, N.; Borghese, R.; Bhandari, A.; Underwood, M.A.; Lebrilla, C.B.; German, J.B.; Barile, D. A peptidomic analysis of human milk digestion in the infant stomach reveals protein-specific degradation patterns. J. Nutr. 2014, 144, 815–820. [Google Scholar] [CrossRef]
- Mamone, G.; Picariello, G.; Caira, S.; Addeo, F.; Ferranti, P. Analysis of food proteins and peptides by mass spectrometry-based techniques. J. Chromatogr. A 2009, 1216, 7130–7142. [Google Scholar] [CrossRef]
- Dallas, D.C.; Guerrero, A.; Parker, E.A.; Robinson, R.C.; Gan, J.; German, J.B.; Barile, D.; Lebrilla, C.B. Current peptidomics: Applications, purification, identification, quantification, and functional analysis. Proteomics 2015, 15, 1026–1038. [Google Scholar] [CrossRef] [PubMed]
- Panchaud, A.; Affolter, M.; Kussmann, M. Mass spectrometry for nutritional peptidomics: How to analyze food bioactives and their health effects. J. Proteom. 2012, 75, 3546–3559. [Google Scholar] [CrossRef] [PubMed]
- Lahrichi, S.L.; Affolter, M.; Zolezzi, I.S.; Panchaud, A. Food peptidomics: Large scale analysis of small bioactive peptides—A pilot study. J. Proteom. 2013, 88, 83–91. [Google Scholar] [CrossRef]
- Coutte, F.; Lecouturier, D.; Firdaous, L.; Kapel, R.; Bazinet, L.; Cabassud, C.; Dhulster, P. Recent trends in membrane bioreactors. In Current Developments in Biotechnology and Bioengineering; Elsevier: Amsterdam, The Netherlands, 2017; pp. 279–311. [Google Scholar]
- Wu, S.; Qi, W.; Li, T.; Lu, D.; Su, R.; He, Z. Simultaneous production of multi-functional peptides by pancreatic hydrolysis of bovine casein in an enzymatic membrane reactor via combinational chromatography. Food Chem. 2013, 141, 2944–2951. [Google Scholar] [CrossRef] [PubMed]
- Hayes, M.; Ross, R.P.; Fitzgerald, G.F.; Hill, C.; Stanton, C. Casein-derived antimicrobial peptides generated by Lactobacillus acidophilus DPC6026. Appl. Environ. Microbiol. 2006, 72, 2260–2264. [Google Scholar] [CrossRef]
- Han, Z.; Luo, X.G.; Tian, H.; Men, Y.; Wang, N.; Jiang, Y.; Zhang, T.C. Separation of angiotensin-converting enzyme inhibitors from fermented milk prepared with Lactobacillus casei HZ1. Adv. Mater. Res. 2012, 345, 161–167. [Google Scholar] [CrossRef]
- Jin, Y.; Yu, Y.; Qi, Y.; Wang, F.; Yan, J.; Zou, H. Peptide profiling and the bioactivity character of yogurt in the simulated gastrointestinal digestion. J. Proteom. 2016, 141, 24–46. [Google Scholar] [CrossRef]
- Li, C.; Kwok, L.-Y.; Mi, Z.; Bala, J.; Xue, J.; Yang, J.; Ma, Y.; Zhang, H.; Chen, Y. Characterization of the angiotensin-converting enzyme inhibitory activity of fermented milks produced with Lactobacillus casei. J. Dairy Sci. 2017, 100, 9495–9507. [Google Scholar] [CrossRef]
- Dupont, D. Peptidomic as a tool for assessing protein digestion. Curr. Opin. Food Sci. 2017, 16, 53–58. [Google Scholar] [CrossRef]
- Giacometti, J.; Buretić-Tomljanović, A. Peptidomics as a tool for characterizing bioactive milk peptides. Food Chem. 2017, 230, 91–98. [Google Scholar] [CrossRef]
- Combes, C.; Paterson, E.; Amadò, R. Isolation and identification of low-molecular-weight peptides from Emmentaler cheese. J. Food Sci. 2002, 67, 553–559. [Google Scholar] [CrossRef]
- Toelstede, S.; Hofmann, T. Sensomics mapping and identification of the key bitter metabolites in Gouda cheese. J. Agric. Food Chem. 2008, 56, 2795–2804. [Google Scholar] [CrossRef] [PubMed]
- Aparna, G.; Bimlesh, M.; Rajesh, K.; Sangwan, R.B. Identification of antioxidant peptides in cheddar cheese made with adjunct culture Lactobacillus casei ssp. casei 300. Milchwissenschaft 2010, 65, 396–399. [Google Scholar]
- Sforza, S.; Cavatorta, V.; Lambertini, F.; Galaverna, G.; Dossena, A.; Marchelli, R. Cheese peptidomics: A detailed study on the evolution of the oligopeptide fraction in Parmigiano-Reggiano cheese from curd to 24 months of aging. J. Dairy Sci. 2012, 95, 3514–3526. [Google Scholar] [CrossRef]
- Miclo, L.; Roux, E.; Genay, M.; Brusseaux, E.; Poirson, C.; Jameh, N.; Perrin, C.; Dary, A. Variability of hydrolysis of β-, αs1-, and αs2-caseins by 10 strains of Streptococcus thermophilus and resulting bioactive peptides. J. Agric. Food Chem. 2012, 60, 554–565. [Google Scholar] [CrossRef]
- Rauh, V.M.; Johansen, L.B.; Ipsen, R.; Paulsson, M.; Larsen, L.B.; Hammershøj, M. Plasmin activity in UHT milk: Relationship between proteolysis, age gelation, and bitterness. J. Agric. Food Chem. 2014, 62, 6852–6860. [Google Scholar] [CrossRef]
- Jensen, S.; Jansson, T.; Eggers, N.; Clausen, M.R.; Larsen, L.B.; Jensen, H.B.; Ray, C.; Sundgren, A.; Andersen, H.J.; Bertram, H.C. Storage-induced changes in the sensory characteristics and volatiles of conventional and lactose-hydrolyzed UHT processed milk. Eur. Food Res. Technol. 2015, 240, 1247–1257. [Google Scholar] [CrossRef]
- Nielsen, S.D.; Jansson, T.; Le, T.T.; Jensen, S.; Eggers, N.; Rauh, V.; Sundekilde, U.K.; Sørensen, J.; Andersen, H.J.; Bertram, H.C. Correlation between sensory properties and peptides derived from hydrolysed-lactose UHT milk during storage. Int. Dairy J. 2017, 68, 23–31. [Google Scholar] [CrossRef]
- Dallas, D.C.; Guerrero, A.; Parker, E.A.; Garay, L.A.; Bhandari, A.; Lebrilla, C.B.; Barile, D.; German, J.B. Peptidomic profile of milk of Holstein cows at peak lactation. J. Agric. Food Chem. 2014, 62, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Nongonierma, A.B.; FitzGerald, R.J. Strategies for the discovery and identification of food protein-derived biologically active peptides. Trends Food Sci. Technol. 2017, 69, 289–305. [Google Scholar] [CrossRef]
- Zigoneanu, I.G.; Sims, C.E.; Allbritton, N.L. Separation of peptide fragments of a protein kinase C substrate fused to a β-hairpin by capillary electrophoresis. Anal. Bioanal. Chem. 2015, 407, 8999–9008. [Google Scholar] [CrossRef]
- Kim, H.-O.; Li-Chan, E.C.Y. Application of Fourier transform Raman spectroscopy for prediction of bitterness of peptides. Appl. Spectrosc. 2006, 60, 1297–1306. [Google Scholar] [CrossRef] [PubMed]
- Li-Chan, E.C.Y. Vibrational spectroscopy applied to the study of milk proteins. Lait 2007, 87, 443–458. [Google Scholar] [CrossRef]
- Andersen, C.M.; Mortensen, G. Fluorescence spectroscopy: A rapid tool for analyzing dairy products. J. Agric. Food Chem. 2008, 56, 720–729. [Google Scholar] [CrossRef]
- Roufik, S.; Gauthier, S.F.; Dufour, É.; Turgeon, S.L. Interactions between bovine β-lactoglobulin A and various bioactive peptides as studied by front-face fluorescence spectroscopy. J. Agric. Food Chem. 2006, 54, 4962–4969. [Google Scholar] [CrossRef]
- Barbosa, C.M.S.; Morais, H.A.; Delvivo, F.M.; Mansur, H.S.; De Oliveira, M.C.; Silvestre, M.P.C. Papain hydrolysates of casein: Molecular weight profile and encapsulation in lipospheres. J. Sci. Food Agric. 2004, 84, 1891–1900. [Google Scholar] [CrossRef]
- Guerrero, A.; Dallas, D.C.; Contreras, S.; Bhandari, A.; Cánovas, A.; Islas-Trejo, A.; Medrano, J.F.; Parker, E.A.; Wang, M.; Hettinga, K. Peptidomic analysis of healthy and subclinically mastitic bovine milk. Int. Dairy J. 2015, 46, 46–52. [Google Scholar] [CrossRef]
- Mansor, R.; Mullen, W.; Albalat, A.; Zerefos, P.; Mischak, H.; Barrett, D.C.; Biggs, A.; Eckersall, P.D. A peptidomic approach to biomarker discovery for bovine mastitis. J. Proteom. 2013, 85, 89–98. [Google Scholar] [CrossRef]
- Meltretter, J.; Schmidt, A.; Humeny, A.; Becker, C.-M.; Pischetsrieder, M. Analysis of the peptide profile of milk and its changes during thermal treatment and storage. J. Agric. Food Chem. 2008, 56, 2899–2906. [Google Scholar] [CrossRef]
- Bouglé, D.; Bouhallab, S. Dietary bioactive peptides: Human studies. Crit. Rev. Food Sci. Nutr. 2017, 57, 335–343. [Google Scholar] [CrossRef]
- Yamamoto, N. Antihypertensive peptides derived from food proteins. Pept. Sci. 1997, 43, 129–134. [Google Scholar] [CrossRef]
- Agyei, D.; Ongkudon, C.M.; Wei, C.Y.; Chan, A.S.; Danquah, M.K. Bioprocess challenges to the isolation and purification of bioactive peptides. Food Bioprod. Process. 2016, 98, 244–256. [Google Scholar] [CrossRef]
- LeBlanc, J.G.; Matar, C.; Valdez, J.C.; LeBlanc, J.; Perdigon, G. Immunomodulating effects of peptidic fractions issued from milk fermented with Lactobacillus helveticus. J. Dairy Sci. 2002, 85, 2733–2742. [Google Scholar] [CrossRef]
- Georgalaki, M.; Zoumpopoulou, G.; Mavrogonatou, E.; Van Driessche, G.; Alexandraki, V.; Anastasiou, R.; Papadelli, M.; Kazou, M.; Manolopoulou, E.; Kletsas, D. Evaluation of the antihypertensive angiotensin-converting enzyme inhibitory (ACE-I) activity and other probiotic properties of lactic acid bacteria isolated from traditional Greek dairy products. Int. Dairy J. 2017, 75, 10–21. [Google Scholar] [CrossRef]
- Khalaf, R.; Baur, D.; Pfister, D. Optimization of reversed-phase chromatography methods for peptide analytics. J. Chromatogr. A 2015, 1425, 198–203. [Google Scholar] [CrossRef]
- Underberg, W.J.M.; Hoitink, M.A.; Reubsaet, J.L.E.; Waterval, J.C.M. Separation and detection techniques for peptides and proteins in stability research and bioanalysis. J. Chromatogr. B Biomed. Sci. Appl. 2000, 742, 401–409. [Google Scholar] [CrossRef]
- Van Eeckhaut, A.; Mangelings, D. Toward greener analytical techniques for the absolute quantification of peptides in pharmaceutical and biological samples. J. Pharm. Biomed. Anal. 2015, 113, 181–188. [Google Scholar] [CrossRef] [PubMed]
- Davio, S.R.; Kienle, K.M.; Collins, B.E. Interdomain interactions in the chimeric protein toxin sCD4 (178)-PE40: A differential scanning calorimetry (DSC) study. Pharm. Res. 1995, 12, 642–648. [Google Scholar] [CrossRef]
- Reubsaet, J.L.E.; Beijnen, J.H.; Bult, A.; Hop, E.; Vermaas, R.; Kellekule, Y.; Kettenes-van den Bosch, J.J.; Underberg, W.J.M. Structural identification of the degradation products of the antitumor peptide antagonist [Arg6, D-Trp7, 9, MePhe8] substance P (6–11). Anal. Chem. 1995, 67, 4431–4436. [Google Scholar] [CrossRef]
- Sánchez-Rivera, L.; Martínez-Maqueda, D.; Cruz-Huerta, E.; Miralles, B.; Recio, I. Peptidomics for discovery, bioavailability and monitoring of dairy bioactive peptides. Food Res. Int. 2014, 63, 170–181. [Google Scholar] [CrossRef]
- Schrader, M.; Schulz-Knappe, P.; Fricker, L.D. Historical perspective of peptidomics. EuPA Open Proteom. 2014, 3, 171–182. [Google Scholar] [CrossRef]
- Foltz, M.; Meynen, E.E.; Bianco, V.; van Platerink, C.; Koning, T.M.M.G.; Kloek, J. Angiotensin converting enzyme inhibitory peptides from a lactotripeptide-enriched milk beverage are absorbed intact into the circulation. J. Nutr. 2007, 137, 953–958. [Google Scholar] [CrossRef] [PubMed]
- Kaiser, S.; Martin, M.; Lunow, D.; Rudolph, S.; Mertten, S.; Möckel, U.; Deußen, A.; Henle, T. Tryptophan-containing dipeptides are bioavailable and inhibit plasma human angiotensin-converting enzyme in vivo. Int. Dairy J. 2016, 52, 107–114. [Google Scholar] [CrossRef]
- Morifuji, M.; Ishizaka, M.; Baba, S.; Fukuda, K.; Matsumoto, H.; Koga, J.; Kanegae, M.; Higuchi, M. Comparison of different sources and degrees of hydrolysis of dietary protein: Effect on plasma amino acids, dipeptides, and insulin responses in human subjects. J. Agric. Food Chem. 2010, 58, 8788–8797. [Google Scholar] [CrossRef]
- Gillette, M.A.; Carr, S.A. Quantitative analysis of peptides and proteins in biomedicine by targeted mass spectrometry. Nat. Methods 2013, 10, 28–34. [Google Scholar] [CrossRef]
- Hopfgartner, G.; Varesio, E. New approaches for quantitative analysis in biological fluids using mass spectrometric detection. TrAC Trends Anal. Chem. 2005, 24, 583–589. [Google Scholar] [CrossRef]
- Han, X.; Aslanian, A.; Yates III, J.R. Mass spectrometry for proteomics. Curr. Opin. Chem. Biol. 2008, 12, 483–490. [Google Scholar] [CrossRef]
- John, H.; Walden, M.; Schäfer, S.; Genz, S.; Forssmann, W.-G. Analytical procedures for quantification of peptides in pharmaceutical research by liquid chromatography–mass spectrometry. Anal. Bioanal. Chem. 2004, 378, 883–897. [Google Scholar] [CrossRef]
- del Mar Contreras, M.; Lpez-Expsito, I.; Hernndez-Ledesma, B.; Ramos, M.; Recio, I. Application of mass spectrometry to the characterization and quantification of food-derived bioactive peptides. J. AOAC Int. 2008, 91, 981–994. [Google Scholar] [CrossRef]
- Lesur, A.; Varesio, E.; Domon, B.; Hopfgartner, G. Peptides quantification by liquid chromatography with matrix-assisted laser desorption/ionization and selected reaction monitoring detection. J. Proteome Res. 2012, 11, 4972–4982. [Google Scholar] [CrossRef] [PubMed]
- O’Keeffe, M.B.; FitzGerald, R.J. Identification of short peptide sequences in complex milk protein hydrolysates. Food Chem. 2015, 184, 140–146. [Google Scholar] [CrossRef] [PubMed]
- Kütt, M.L.; Malbe, M.; Stagsted, J. Nanostructure-assisted laser desorption/ionization (NALDI) for analysis of peptides in milk and colostrum. Agron. Res. 2011, 9, 415–420. [Google Scholar]
- Le Maux, S.; Nongonierma, A.B.; FitzGerald, R.J. Improved short peptide identification using HILIC–MS/MS: Retention time prediction model based on the impact of amino acid position in the peptide sequence. Food Chem. 2015, 173, 847–854. [Google Scholar] [CrossRef] [PubMed]
- Mayer, H.K. Milk species identification in cheese varieties using electrophoretic, chromatographic and PCR techniques. Int. Dairy J. 2005, 15, 595–604. [Google Scholar] [CrossRef]
- Cartoni, G.; Coccioli, F.; Jasionowska, R.; Masci, M. Determination of cows’ milk in goats’ milk and cheese by capillary electrophoresis of the whey protein fractions. J. Chromatogr. A 1999, 846, 135–141. [Google Scholar] [CrossRef]
- Molina, E.; Martín-Álvarez, P.J.; Ramos, M. Analysis of cows’, ewes’ and goats’ milk mixtures by capillary electrophoresis: Quantification by multivariate regression analysis. Int. Dairy J. 1999, 9, 99–105. [Google Scholar] [CrossRef]
- Bottero, M.T.; Civera, T.; Anastasio, A.; Turi, R.M.; Rosati, S. Identification of cow’s milk in “buffalo” cheese by duplex polymerase chain reaction. J. Food Prot. 2002, 65, 362–366. [Google Scholar] [CrossRef] [PubMed]
- Popelka, P.; Popelka, P.; Horská, D.; Golian, J.; Marcinčák, S. Detection of sheep milk and cheeses adulteration using enzyme immunoanalysis (ELISA). Slov. Vet. J. 2002, 27, 36–37. [Google Scholar]
- Haza, A.I.; Morales, P.; Martín, R.; García, T.; Anguita, G.; Sanz, B.; Hernández, P.E. Detection and quantification of goat’s cheese in ewe’s cheese using a monoclonal antibody and two ELISA formats. J. Sci. Food Agric. 1999, 79, 1043–1047. [Google Scholar] [CrossRef]
- Zeleňáková, L.; Golian, J.; Zajác, P. Application of ELISA tests for the detection of goat milk in sheep milk. Milchwissenschaft 2008, 63, 137–141. [Google Scholar]
- Costa, N.; Ravasco, F.; Miranda, R.; Duthoit, M.; Roseiro, L.B. Evaluation of a commercial ELISA method for the quantitative detection of goat and cow milk in ewe milk and cheese. Small Rumin. Res. 2008, 79, 73–79. [Google Scholar] [CrossRef]
- Zarranz, M.J.; Izco, J.M. Validation parameters of an ELISA test to detect presence of milk from other species. Milchwissenschaft 2007, 62, 159–163. [Google Scholar]
- Taylor, S.L.; Nordlee, J.A.; Niemann, L.M.; Lambrecht, D.M. Allergen immunoassays—Considerations for use of naturally incurred standards. Anal. Bioanal. Chem. 2009, 395, 83–92. [Google Scholar] [CrossRef]
- Feligini, M.; Bonizzi, I.; Curik, V.C.; Parma, P.; Greppi, G.F.; Enne, G. Detection of adulteration in Italian mozzarella cheese using mitochondrial DNA templates as biomarkers. Food Technol. Biotechnol. 2005, 43, 91–95. [Google Scholar]
- López-Calleja, I.; Alonso, I.G.; Fajardo, V.; Rodríguez, M.A.; Hernández, P.E.; García, T.; Martín, R. PCR detection of cows’ milk in water buffalo milk and mozzarella cheese. Int. Dairy J. 2005, 15, 1122–1129. [Google Scholar] [CrossRef]
- Lopez-Calleja, I.; Gonzalez, I.; Fajardo, V.; Martin, I.; Hernandez, P.E.; Garcia, T.; Martin, R. Application of polymerase chain reaction to detect adulteration of sheep’s milk with goats’ milk. J. Dairy Sci. 2005, 88, 3115–3120. [Google Scholar] [CrossRef]
- Mafra, I.; Ferreira, I.M.; Faria, M.A.; Oliveira, B.P.P. A novel approach to the quantification of bovine milk in ovine cheeses using a duplex polymerase chain reaction method. J. Agric. Food Chem. 2004, 52, 4943–4947. [Google Scholar] [CrossRef]
- Mašková, E.; Paulíčková, I. PCR-based detection of cow’s milk in goat and sheep cheeses marketed in the Czech Republic. Czech. J. Food Sci. 2006, 24, 127–132. [Google Scholar] [CrossRef]
- Bobková, A.; Židek, R.; Flimelová, E.; Bobko, M.; Fiková, M. Application of PCR method for milk adulteration and milk products identification. Potravinárstvo 2009, 1, 1–3. [Google Scholar]
- Iwaniak, A.; Minkiewicz, P.; Darewicz, M.; Protasiewicz, M.; Mogut, D. Chemometrics and cheminformatics in the analysis of biologically active peptides from food sources. J. Funct. Foods 2015, 16, 334–351. [Google Scholar] [CrossRef]
- Nongonierma, A.B.; FitzGerald, R.J. Structure activity relationship modelling of milk protein-derived peptides with dipeptidyl peptidase IV (DPP-IV) inhibitory activity. Peptides 2016, 79, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Majumder, K.; Wu, J. Angiotensin I converting enzyme inhibitory peptides from simulated in vitro gastrointestinal digestion of cooked eggs. J. Agric. Food Chem. 2009, 57, 471–477. [Google Scholar] [CrossRef] [PubMed]
- Sagardia, I.; Iloro, I.; Elortza, F.; Bald, C. Quantitative structure–activity relationship based screening of bioactive peptides identified in ripened cheese. Int. Dairy J. 2013, 33, 184–190. [Google Scholar] [CrossRef]
- Sauer, S.; Luge, T. Nutriproteomics: Facts, concepts, and perspectives. Proteomics 2015, 15, 997–1013. [Google Scholar] [CrossRef] [PubMed]
- Català-Clariana, S.; Benavente, F.; Giménez, E.; Barbosa, J.; Sanz-Nebot, V. Identification of bioactive peptides in hypoallergenic infant milk formulas by capillary electrophoresis–mass spectrometry. Anal. Chim. Acta 2010, 683, 119–125. [Google Scholar] [CrossRef]
- Herrero, M.; Ibañez, E.; Cifuentes, A. Capillary electrophoresis-electrospray-mass spectrometry in peptide analysis and peptidomics. Electrophoresis 2008, 29, 2148–2160. [Google Scholar] [CrossRef]
- Fournier, M.L.; Gilmore, J.M.; Martin-Brown, S.A.; Washburn, M.P. Multidimensional separations-based shotgun proteomics. Chem. Rev. 2007, 107, 3654–3686. [Google Scholar] [CrossRef]
- Sommella, E.; Pepe, G.; Ventre, G.; Pagano, F.; Manfra, M.; Pierri, G.; Ismail, O.; Ciogli, A.; Campiglia, P. Evaluation of two sub-2 μm stationary phases, core–shell and totally porous monodisperse, in the second dimension of on-line comprehensive two dimensional liquid chromatography, a case study: Separation of milk peptides after expiration date. J. Chromatogr. A 2015, 1375, 54–61. [Google Scholar] [CrossRef]
- Guo, M.R.; Hendricks, G.M.; Kindstedt, P.S.; Flynn, A.; Fox, P.F. Nitrogen and mineral distribution in infant formulae. Int. Dairy J. 1996, 6, 963–979. [Google Scholar] [CrossRef]
- Fomon, M.D.; Samuel, J. Nutrition of Normal Infants; Mosby-Year Book, Inc.: Lowa, IA, USA, 1993; ISBN 1556642482. [Google Scholar]
- Gurr, M.I. Review of the progress of dairy science: Human and artificial milks for infant feeding. J. Dairy Res. 1981, 48, 519–554. [Google Scholar] [CrossRef]
- Nasirpour, A.; Scher, J.; Desobry, S. Baby foods: Formulations and interactions (a review). Crit. Rev. Food Sci. Nutr. 2006, 46, 665–681. [Google Scholar] [CrossRef] [PubMed]
- Darling, P.B.; Dunn, M.; Sarwar, G.; Brookes, S.; Ball, R.O.; Pencharz, P.B. Threonine kinetics in preterm infants fed their mothers’ milk or formula with various ratios of whey to casein. Am. J. Clin. Nutr. 1999, 69, 105–114. [Google Scholar] [CrossRef] [PubMed]
| Analytical Techniques | Chemometric Techniques | Application | References | |
|---|---|---|---|---|
| Electrophoretic Methods | Urea-PAGE | - | Evaluation of changes upon proteolysis in cheese | [21] |
| Urea-PAGE of casein | - | Determination of concentration of peptides and age-related differences | [22] | |
| Chromatographic and Spectrophotometric Techniques | ||||
| Mass Spectrometry | 2-DE; MALDI-TOF; ESI-IT | In-gel digestion | Polymorphism of goatαs1-casein | [23] |
| RP-HPLC < 1000 Da | Analysis of volatile compounds | Evaluation of changes upon proteolysis in cheese | [21] | |
| Sensory analysis | Differences among caseins peptides and sensory attributes | [22] | ||
| NanoESI-QTOF | In-capillary tryptic hydrolysis | Characterization of elephant milk proteins | [24] | |
| MALDI-TOF (reflectron), HPLC-ESI-IT | Tryptic digestion | Identification of truncated goat | [25] | |
| MALDI-TOF (reflectron), HPLC-ESI-IT | Tryptic digestion | Identification of truncated forms of goat αs2-CN A and E | [26] | |
| ESI-QqQ | Offline RP-HPLC | Degree of glycosylation and phosphorylation of ovine and caprine CMP | [27,28] | |
| 2-DE, immunoblotting, MALDI-TOF, ESI-QqQ | In-gel digestion immunoblotting | Phosphorylation and glycosylation of ovine caseins | [29] | |
| 1-DE and 2-DE, ESI-IT | Enzymatic digestion | Phosphorylation, glycosylation, and genetic variants of κ-casein | [30] | |
| 2-D LC-nanoESI-IT | Shotgun proteomics | Identification of minor human milk proteins | [31] | |
| 2-DE, HPLC-QTOF | In-gel digestion | Characterization of minor whey proteins | [32] | |
| HPLC-QTOF | In-gel digestion | Lactosylation of β-Lg, α-La, and αs2-CN in infant formula | [33] | |
| HPLC-ESI-QqQ | Tryptic digest in solution | Lactosylation of β-Lg | [34] | |
| MALDI-TOF | - | Degradation of αs1-CN f(1–23) by bacterial amino and endopeptidases | [35] | |
| MALDI-TOF (reflectron) | Specificity of peptidases from Lactobacillus helveticus | [36] | ||
| Polymerase Chain reaction (PCR) | PCR-RFLP | Bovine DNA in cheese | Ovine and caprine cheese samples | [37] |
| Bovine DNA in cheese | Commercial mozzarella and feta cheese samples | [38] | ||
| Bovine DNA in cheese | Experimental binary mixtures of bovine milk with ovine, caprine, and buffalo milk | [39] | ||
| Duplex PCR | Simultaneous detection of bovine and buffalo DNA in cheese and milk | Experimental mixtures of bovine and buffalo milk and commercial buffalo Mozzarella samples | [40] | |
| Commercial cheese samples | Simultaneous detection of bovine and caprine DNA in cheese | [41] | ||
| RT-PCR | Bovine DNA in cheese | Experimental and commercial mozzarella cheese | [42] | |
| ELISA | Identification of immunogens | Evaluation the fraction of goat’s milk for quantification and the existence of standard immunoglobulins (IgG) | [43] | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Punia, H.; Tokas, J.; Malik, A.; Sangwan, S.; Baloda, S.; Singh, N.; Singh, S.; Bhuker, A.; Singh, P.; Yashveer, S.; et al. Identification and Detection of Bioactive Peptides in Milk and Dairy Products: Remarks about Agro-Foods. Molecules 2020, 25, 3328. https://doi.org/10.3390/molecules25153328
Punia H, Tokas J, Malik A, Sangwan S, Baloda S, Singh N, Singh S, Bhuker A, Singh P, Yashveer S, et al. Identification and Detection of Bioactive Peptides in Milk and Dairy Products: Remarks about Agro-Foods. Molecules. 2020; 25(15):3328. https://doi.org/10.3390/molecules25153328
Chicago/Turabian StylePunia, Himani, Jayanti Tokas, Anurag Malik, Sonali Sangwan, Satpal Baloda, Nirmal Singh, Satpal Singh, Axay Bhuker, Pradeep Singh, Shikha Yashveer, and et al. 2020. "Identification and Detection of Bioactive Peptides in Milk and Dairy Products: Remarks about Agro-Foods" Molecules 25, no. 15: 3328. https://doi.org/10.3390/molecules25153328
APA StylePunia, H., Tokas, J., Malik, A., Sangwan, S., Baloda, S., Singh, N., Singh, S., Bhuker, A., Singh, P., Yashveer, S., Agarwal, S., & Mor, V. S. (2020). Identification and Detection of Bioactive Peptides in Milk and Dairy Products: Remarks about Agro-Foods. Molecules, 25(15), 3328. https://doi.org/10.3390/molecules25153328









