Computational Study of Natural Compounds for the Clearance of Amyloid-Βeta: A Potential Therapeutic Management Strategy for Alzheimer’s Disease
Abstract
1. Introduction
2. Results and Discussion
2.1. Protein-Protein Interaction Study
2.2. Comparison of Vnc, Ajm, and Eme With Positive Control Curcumin
3. Materials and Methods
3.1. Preparation of Receptor-Protein Structures
3.2. Preparation of Ligand Structure
3.3. Molecular Interaction Study
3.4. Protein-Protein Interaction Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AD | Alzheimer’s disease |
| Aβ | Amyloid-beta |
| RAGE | Receptor for advanced glycation end products |
| BBB | Blood-brain barrier |
| Vnc | Vincamine |
| Ajm | Ajmalicine |
| Eme | Emetine |
References
- Alzheimer’s Association. Alzheimer’s disease facts and figures. Alzheimers Dement. 2015, 11, 332–384. [Google Scholar]
- McGeer, P.L.; McGeer, E.G. The amyloid cascade-inflammatory hypothesis of Alzheimer disease: Implications for therapy. Acta Neuropathol. 2013, 126, 479–497. [Google Scholar] [CrossRef]
- Glabe, C.G. Structural Classification of Toxic Amyloid Oligomers. J. Boil. Chem. 2008, 283, 29639–29643. [Google Scholar] [CrossRef]
- Kayed, R.; Sokolov, Y.; Edmonds, B.; McIntire, T.M.; Milton, S.C.; Hall, J.E.; Glabe, C.G. Permeabilization of Lipid Bilayers Is a Common Conformation-dependent Activity of Soluble Amyloid Oligomers in Protein Misfolding Diseases. J. Boil. Chem. 2004, 279, 46363–46366. [Google Scholar] [CrossRef]
- Quist, A.; Doudevski, I.; Lin, H.; Azimova, R.; Ng, D.; Frangione, B.; Kagan, B.; Ghiso, J.; Lal, R. Amyloid ion channels: A common structural link for the protein-misfolding disease. Proc. Natl. Acad. Sci. USA 2005, 102, 10427–10432. [Google Scholar] [CrossRef]
- Deane, R.J. Is RAGE still a therapeutic target for Alzheimer’s disease? Future Med. Chem. 2012, 4, 915–925. [Google Scholar] [CrossRef]
- Yan, S.D.; Chen, X.; Fu, J.; Chen, M.; Zhu, H.; Roher, A.; Slattery, T.; Zhao, L.; Nagashima, M.; Morser, J.; et al. RAGE and amyloid-beta peptide neurotoxicity in Alzheimer’s disease. Nature 1996, 382, 685–691. [Google Scholar] [CrossRef]
- Du Yan, S.; Zhu, H.; Zhu, A.; Golabek, A.; Du, H.; Roher, A.; Yu, J.; Soto, C.; Schmidt, A.M.; Stern, D.; et al. Receptor-dependent cell stress and amyloid accumulation in systemic amyloidosis. Nat. Med. 2000, 6, 643–651. [Google Scholar] [CrossRef]
- Paresce, D.M.; Ghosh, R.N.; Maxfield, F.R. Microglial cells internalize aggregates of the Alzheimer’s disease amyloid beta-protein via a scavenger receptor. Neuron 1996, 17, 553–565. [Google Scholar] [CrossRef]
- Wang, H.Y.; Lee, D.H.; D’Andrea, M.R.; Peterson, P.A.; Shank, R.P.; Reitz, A.B. Aβ (1–42) binds to α-7 nicotinic acetylcholine receptor with high affinity. J. Biol. Chem. 2000, 275, 5626–5632. [Google Scholar] [CrossRef]
- Yaar, M.; Zhai, S.; Pilch, P.F.; Doyle, S.M.; Eisenhauer, P.B.; Fine, R.E.; Gilchrest, B.A. Binding of beta-amyloid to the p75 neurotrophin receptor induces apoptosis. J. Clin. Investig. 1997, 100, 2333–2340. [Google Scholar] [CrossRef]
- Zhang, L.; Fang, Y.; Xu, Y.; Lian, Y.; Xie, N.; Wu, T.; Zhang, H.; Sun, L.; Zhang, R.; Wang, Z. Curcumin Improves Amyloid β-Peptide (1-42) Induced Spatial Memory Deficits through BDNF-ERK Signaling Pathway. PLoS ONE 2015, 10, e0131525. [Google Scholar] [CrossRef]
- Stefani, M.; Rigacci, S. Protein Folding and Aggregation into Amyloid: The Interference by Natural Phenolic Compounds. Int. J. Mol. Sci. 2013, 14, 12411–12457. [Google Scholar] [CrossRef]
- Bratkovic, I.H.; Gaspersic, J.; Smid, L.M.; Bresjanac, M.; Jerala, R. Curcumin binds to the α-helical intermediate and to the amyloid form of prion protein—A new mechanism for the inhibition of PrPSc accumulation. J. Neurochem. 2008, 104, 1553–1564. [Google Scholar] [CrossRef]
- Yanagisawa, D.; Taguchi, H.; Yamamoto, A.; Shirai, N.; Hirao, K.; Tooyama, I. Curcuminoid Binds to Amyloid-β1-42 Oligomer and Fibril. J. Alzheimer’s Dis. 2011, 24, 33–42. [Google Scholar] [CrossRef]
- Yang, F.; Lim, G.P.; Begum, A.N.; Ubeda, O.J.; Simmons, M.R.; Ambegaokar, S.S.; Chen, P.P.; Kayed, R.; Glabe, C.G.; Frautschy, S.A.; et al. Curcumin inhibits formation of amyloid β oligomers and fibrils, binds plaques, and reduces amyloid in vivo. J. Biol. Chem. 2005, 280, 5892–5901. [Google Scholar] [CrossRef]
- Pandey, N.; Strider, J.; Nolan, W.C.; Yan, S.X.; Galvin, J.E. Curcumin inhibits aggregation of α-synuclein. Acta Neuropathol. 2008, 115, 479–489. [Google Scholar] [CrossRef]
- Na Zhao, L.; Chiu, S.-W.; Benoit, J.; Chew, L.Y.; Mu, Y. The Effect of Curcumin on the Stability of Aβ Dimers. J. Phys. Chem. B 2012, 116, 7428–7435. [Google Scholar] [CrossRef]
- Balasubramanian, K. Molecular orbital basis for yellow curry spice curcumin’s prevention of Alzheimer’s disease. J. Agric. Food Chem. 2006, 54, 3512–3520. [Google Scholar] [CrossRef]
- Hardy, J.; Selkoe, D.J. The amyloid hypothesis of Alzheimer’s disease: Progress and problems on the road to therapeutics. Science 2002, 297, 353–356. [Google Scholar] [CrossRef]
- Ngo, S.T.; Li, M.S. Top-leads from natural products for the treatment of Alzheimer’s disease: Docking and molecular dynamics study. Mol. Simul. 2013, 39, 279–291. [Google Scholar] [CrossRef]
- Williams, P.; Sorribas, A.; Howes, M.J.R. Natural products as a source of Alzheimer’s drug leads. Nat. Prod. Rep. 2011, 28, 48–77. [Google Scholar] [CrossRef]
- Howes, M.-J.R.; Houghton, P.J. Plants used in Chinese and Indian traditional medicine for improvement of memory and cognitive function. Pharm. Biochem. Behav. 2003, 75, 513–527. [Google Scholar] [CrossRef]
- Alam, A.; Shaikh, S.; Ahmad, S.; Ansari, M.; Shakil, S.; Rizvi, S.M.D.; Shakil, S.; Imran, M.; Haneef, M.; Abuzenadah, A.; et al. Molecular Interaction of Human Brain Acetylcholinesterase with a Natural Inhibitor Huperzine-B: An Enzoinformatics Approach. CNS Neurol. Disord. Drug Targets 2014, 13, 487–490. [Google Scholar] [CrossRef]
- Mackic, J.B.; Bading, J.; Ghiso, J.; Walker, L.; Wisniewski, T.; Frangione, B.; Zlokovic, B.V. Circulating amyloid-βpeptide crosses the blood-brain barrier in aged monkeys and contributes to Alzheimer’s disease lesions. Vascul. Pharmacol. 2002, 38, 303–313. [Google Scholar] [CrossRef]
- Martel, C.L.; Mackic, J.B.; McComb, J.G.; Ghiso, J.; Zlokovic, B.V. Blood-brain barrier uptake of the 40 and 42 amino acid sequences of circulating Alzheimer’s amyloid βin guinea pigs. Neurosci. Lett. 1996, 206, 157–160. [Google Scholar] [CrossRef]
- Deane, R.; Du Yan, S.; Submamaryan, R.K.; LaRue, B.; Jovanovic, S.; Hogg, E.; Welch, D.; Manness, L.; Lin, C.; Yu, J.; et al. RAGE mediates amyloid-beta peptide transport across the blood-brain barrier and accumulation in brain. Nat. Med. 2003, 9, 907–913. [Google Scholar] [CrossRef]
- Awasthi, M.; Singh, S.; Pandey, V.P.; Dwivedi, U.N. Alzheimer’s disease: An overview of amyloid beta dependent pathogenesis and its therapeutic implications along with in silico approaches emphasizing the role of natural products. J. Neurol. Sci. 2016, 361, 256–271. [Google Scholar] [CrossRef]
- Kalim, M.; Khan, A.; Baig, M.H.; Arif, J.M.; Lohani, M.; Jamal, F. Efficacy of natural inhibitors against pkc: An in silico approach to combat cancer. Recent Res. Sci. Technol. 2011, 3, 90–93. [Google Scholar]
- Wang, J.; Wang, W.; Kollman, P.A.; Case, D.A. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graph. Model. 2006, 25, 247–260. [Google Scholar] [CrossRef]
- Wang, J.; Wolf, R.M.; Caldwell, J.W.; Kollman, P.A.; Case, D.A. Development and testing of a general amber force field. J. Comput. Chem. 2004, 25, 1157–1174. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera?A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Gasteiger, J.; Marsili, M. A new model for calculating atomic charges in molecules. Tetrahedron Lett. 1978, 19, 3181–3184. [Google Scholar] [CrossRef]
- Morris, G.M.; Goodsell, D.S.; Halliday, R.S.; Huey, R.; Hart, W.E.; Belew, R.K.; Olson, A.J. Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J. Comput. Chem. 1998, 19, 1639–1662. [Google Scholar] [CrossRef]
- Rarey, M.; Kramer, B.; Lengauer, T.; Klebe, G. A fast flexible docking method using an incremental construction algorithm. J. Mol. Biol. 1996, 26, 470–489. [Google Scholar] [CrossRef]
- Goodsell, D.S.; Morris, G.M.; Olson, A.J. Automated docking of flexible ligands: Applications of AutoDock. J. Mol. Recognit. 1996, 9, 1–5. [Google Scholar] [CrossRef]
- Ahmad, S.S.; Akhtar, S.; Danish Rizvi, S.M.; Kamal, M.A.; Sayeed, U.; Khan, M.K.A.; Siddiqui, M.H.; Arif, J.M. Screening and Elucidation of Selected Natural Compounds for Anti-Alzheimer’s Potential Targeting BACE-1 Enzyme: A Case Computational Study. Curr. Comput. Aided Drug Des. 2017, 13, 311–318. [Google Scholar] [CrossRef]
- Chen, R.; Li, L.; Weng, Z. ZDOCK: An initial-stage protein-docking algorithm. Proteins Struct. Funct. Bioinform. 2003, 52, 80–87. [Google Scholar] [CrossRef]
- Hwang, H.; Vreven, T.; Pierce, B.G.; Hung, J.-H.; Weng, Z. Performance of ZDOCK and ZRANK in CAPRI Rounds 13–19. Proteins Struct. Funct. Bioinform. 2010, 78, 3104–3110. [Google Scholar] [CrossRef]
- Chowdhury, R.; Rasheed, M.; Keidel, D.; Moussalem, M.; Olson, A.; Sanner, M.; Bajaj, C. Protein-Protein Docking with F2Dock 2.0 and GB-Rerank. PLoS ONE 2013, 8, e51307. [Google Scholar] [CrossRef][Green Version]
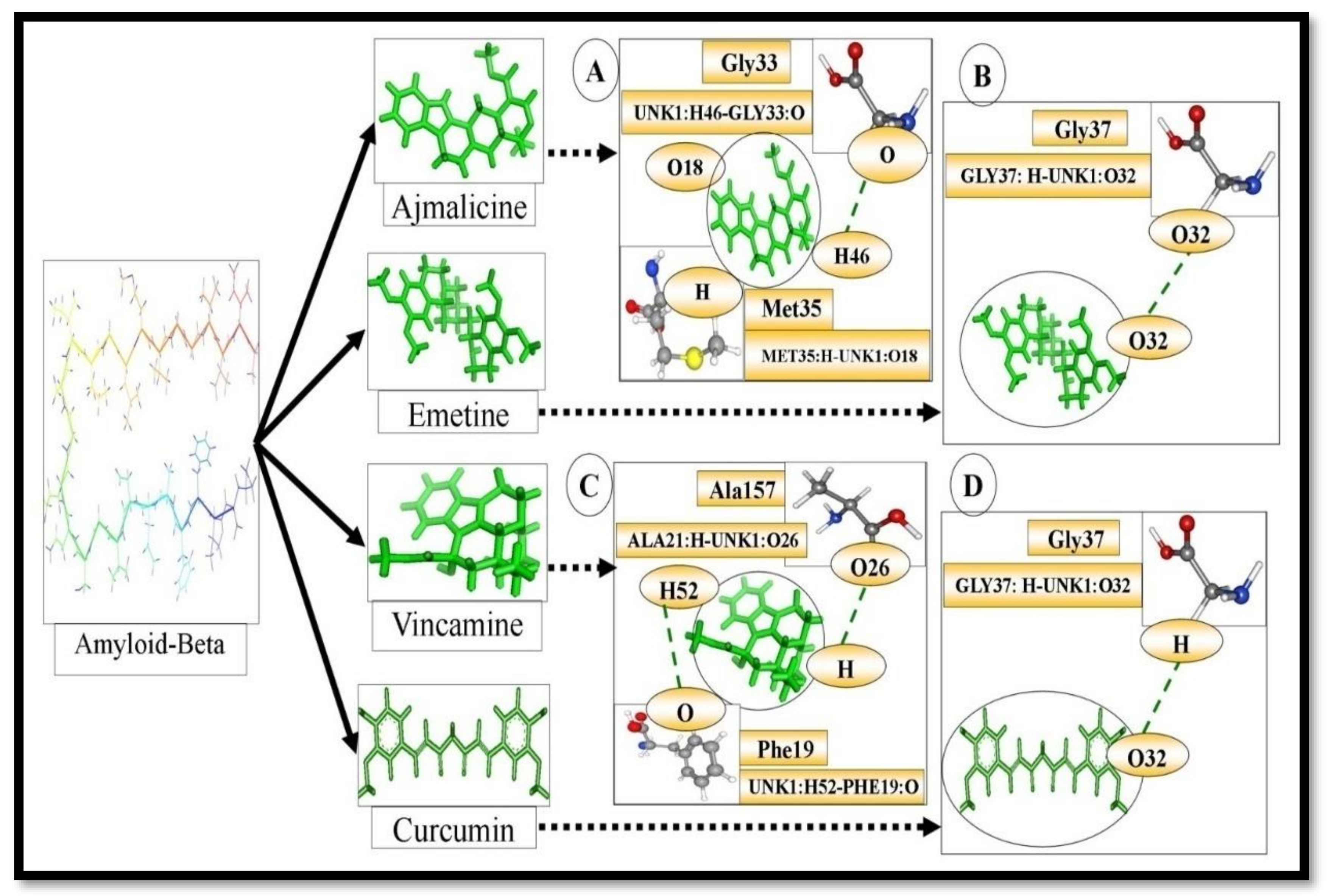
| S.N | Target | Ligands Name | Interaction Amino Acid | H-bond Distance (Å) | H-bond |
|---|---|---|---|---|---|
| 1. | Β-Amyloid- | Vincamine | Phe19, Phe20, Ala21, Asp23, Ile32, Gly33, Leu34, Met35, and Val36 | 1.87477 2.11324 | ALA21:H-UNK1: O26 UNK1: H52-PHE19:O |
| 2. | Ajmalicine | Phe19, Phe20, Ala21, Gly22, Asp23, Ile32, Gly33,Leu34, Met35, and Val36 | 1.81314 1.87937 | MET35:H-UNK1: O18 UNK1: H46-GLY33:O | |
| 3. | Emetine | Ala21, Gl22, Asp23, Gly33, Leu34, Met35, Val36, and Gly37 | 2.41182 | GLY37:H-UNK1: O32 | |
| 4. | Curcumin | Ala21, Glu22, Asp23, Val24, Gly25, Leu34, Met35, Val36, and Gly37. | 2.13938 | UNK1: H43-GLY37:O |
| S. No | Target | Ligand Name | Binding Energy (Kcal/mol) | Inhibition Constant (Ki) | vdw+hb+ Desolvation Energy (Kcal/mol) | Internal Molecular Energy (Kcal/mol) | Electrostatic Energy (Kcal/mol) |
|---|---|---|---|---|---|---|---|
| 1. | β-Amyloid | Vincamine | −5.45 | 249.96 µM | −5.33 | −6.11 | −0.78 |
| 2. | Ajmalicine | −6.66 | 13.23 µM | −6.27 | −7.25 | −0.98 | |
| 3. | Emetine | −6.99 | 7.5 µM | −8.2 | −9.08 | −0.88 | |
| 4. | Curcumin | −3.61 | 136.2 µM | −5.56 | −5.59 | −0.03 |
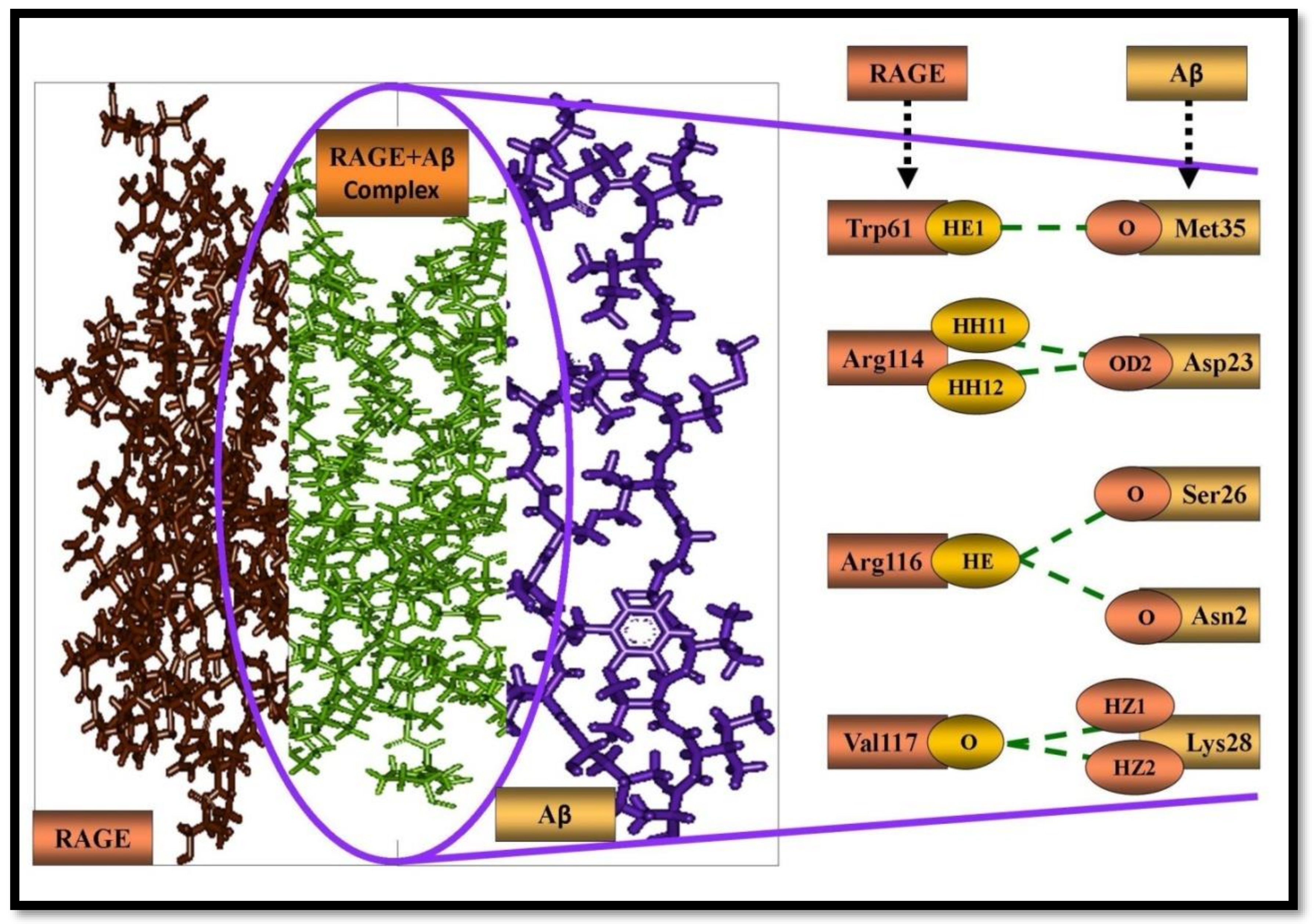
| S S.No | Amino Acid Residues of Aβ | Amino Acid Residues of RAGE | Donor Atom | Acceptor Atom | H-Bonds | Distance of H-Bond (Å) |
|---|---|---|---|---|---|---|
| 1. | Met35 | Trp61 | TRP61:HE1 | MET35:O | TRP61:HE1 - MET35:O | 2.31028 |
| 2. | Asp23 | Arg114 | ARG114:HH11 | ASP23:OD2 | ARG114:HH11 -ASP23:OD2 | 1.40911 |
| 3. | Asp23 | Arg114 | ARG114:HH12 | ASP23:OD2 | ARG114:HH12 -ASP23:OD2 | 2.02749 |
| 4. | Ser26 | Arg116 | ARG116:HE | SER26:O | ARG116:HE - SER26:O | 2.48357 |
| 5. | Asn27 | Arg116 | ARG116: HE | ASN27:O | ARG116: HE - ASN27:O | 2.04783 |
| 6. | Lys28 | Val117 | LYS28:HZ1 | VAL117:O | LYS28:HZ1 - VAL117:O | 2.15927 |
| 7. | Lys28 | Val117 | LYS28:HZ2 | VAL117:O | LYS28:HZ2 - AVAL117:O | 1.10912 |
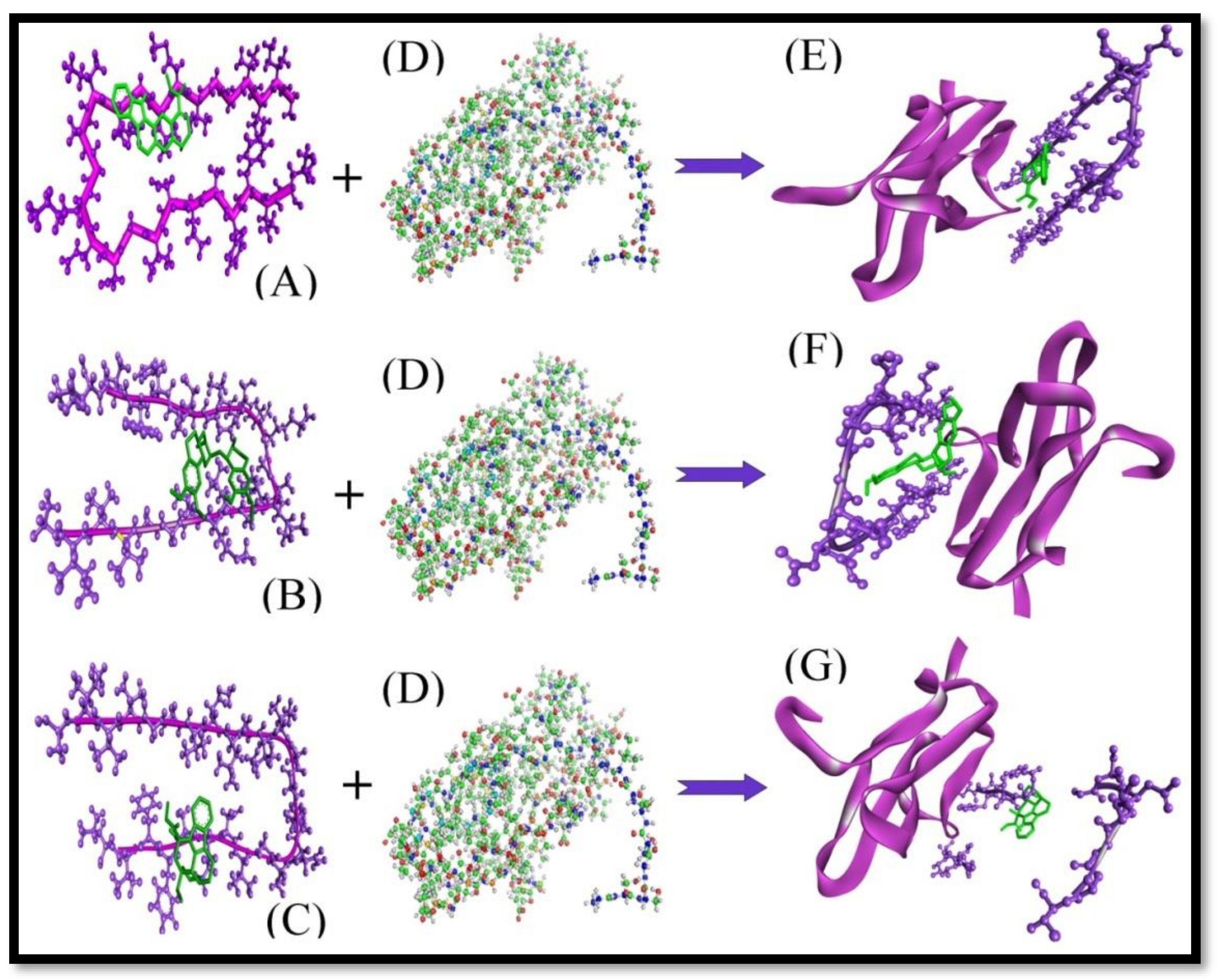
| S.No. | Protein-Protein interaction | Z-dock Score |
|---|---|---|
| 1. | Aβ+RAGE | 1269.55 |
| Complex-Protein Interaction | Z-dock Score | |
| 2. | (Aβ+Ajm)+ RAGE | 911.83 |
| 3. | (Aβ+Eme)+ RAGE | 940.69 |
| 4. | (Aβ+Vnc)+ RAGE | 907.98 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ahmad, S.S.; Khan, H.; Danish Rizvi, S.M.; Ansari, S.A.; Ullah, R.; Rastrelli, L.; Mahmood, H.M.; Siddiqui, M.H. Computational Study of Natural Compounds for the Clearance of Amyloid-Βeta: A Potential Therapeutic Management Strategy for Alzheimer’s Disease. Molecules 2019, 24, 3233. https://doi.org/10.3390/molecules24183233
Ahmad SS, Khan H, Danish Rizvi SM, Ansari SA, Ullah R, Rastrelli L, Mahmood HM, Siddiqui MH. Computational Study of Natural Compounds for the Clearance of Amyloid-Βeta: A Potential Therapeutic Management Strategy for Alzheimer’s Disease. Molecules. 2019; 24(18):3233. https://doi.org/10.3390/molecules24183233
Chicago/Turabian StyleAhmad, Syed Sayeed, Haroon Khan, Syed Mohd. Danish Rizvi, Siddique Akber Ansari, Riaz Ullah, Luca Rastrelli, Hafiz Majid Mahmood, and Mohd. Haris Siddiqui. 2019. "Computational Study of Natural Compounds for the Clearance of Amyloid-Βeta: A Potential Therapeutic Management Strategy for Alzheimer’s Disease" Molecules 24, no. 18: 3233. https://doi.org/10.3390/molecules24183233
APA StyleAhmad, S. S., Khan, H., Danish Rizvi, S. M., Ansari, S. A., Ullah, R., Rastrelli, L., Mahmood, H. M., & Siddiqui, M. H. (2019). Computational Study of Natural Compounds for the Clearance of Amyloid-Βeta: A Potential Therapeutic Management Strategy for Alzheimer’s Disease. Molecules, 24(18), 3233. https://doi.org/10.3390/molecules24183233










