Multi-Level Comparison of Machine Learning Classifiers and Their Performance Metrics
Abstract
1. Introduction
2. Results and Discussion
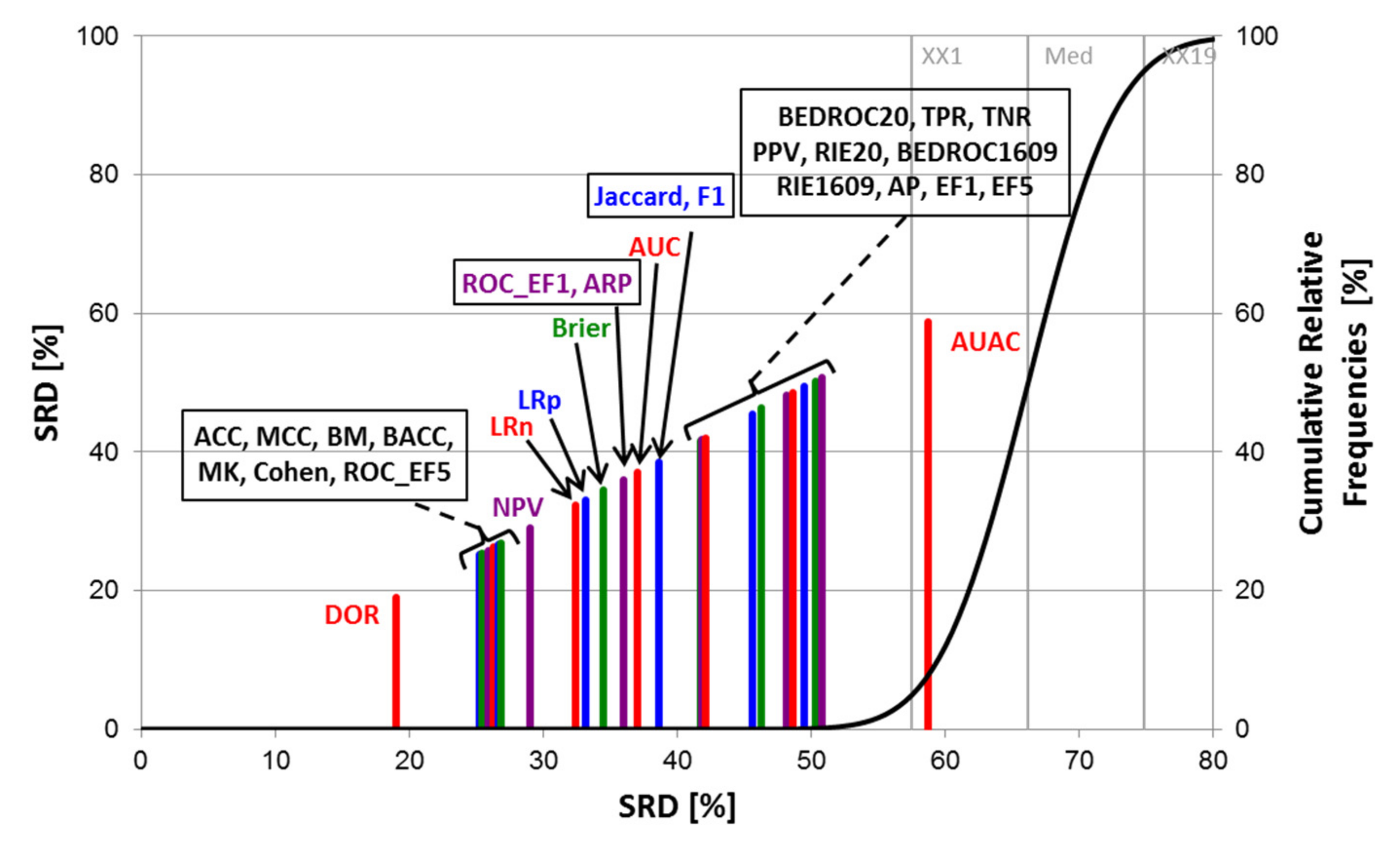
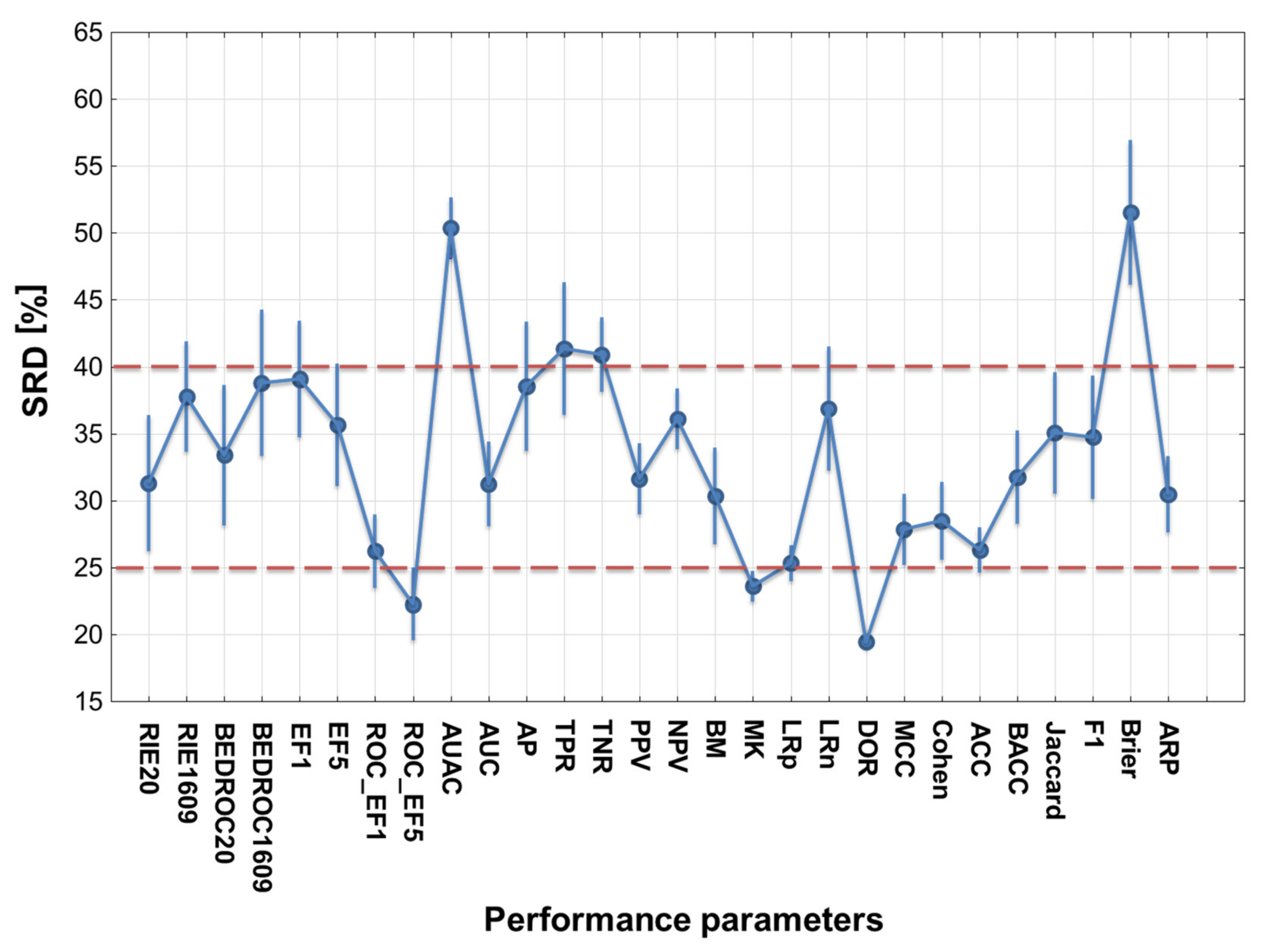
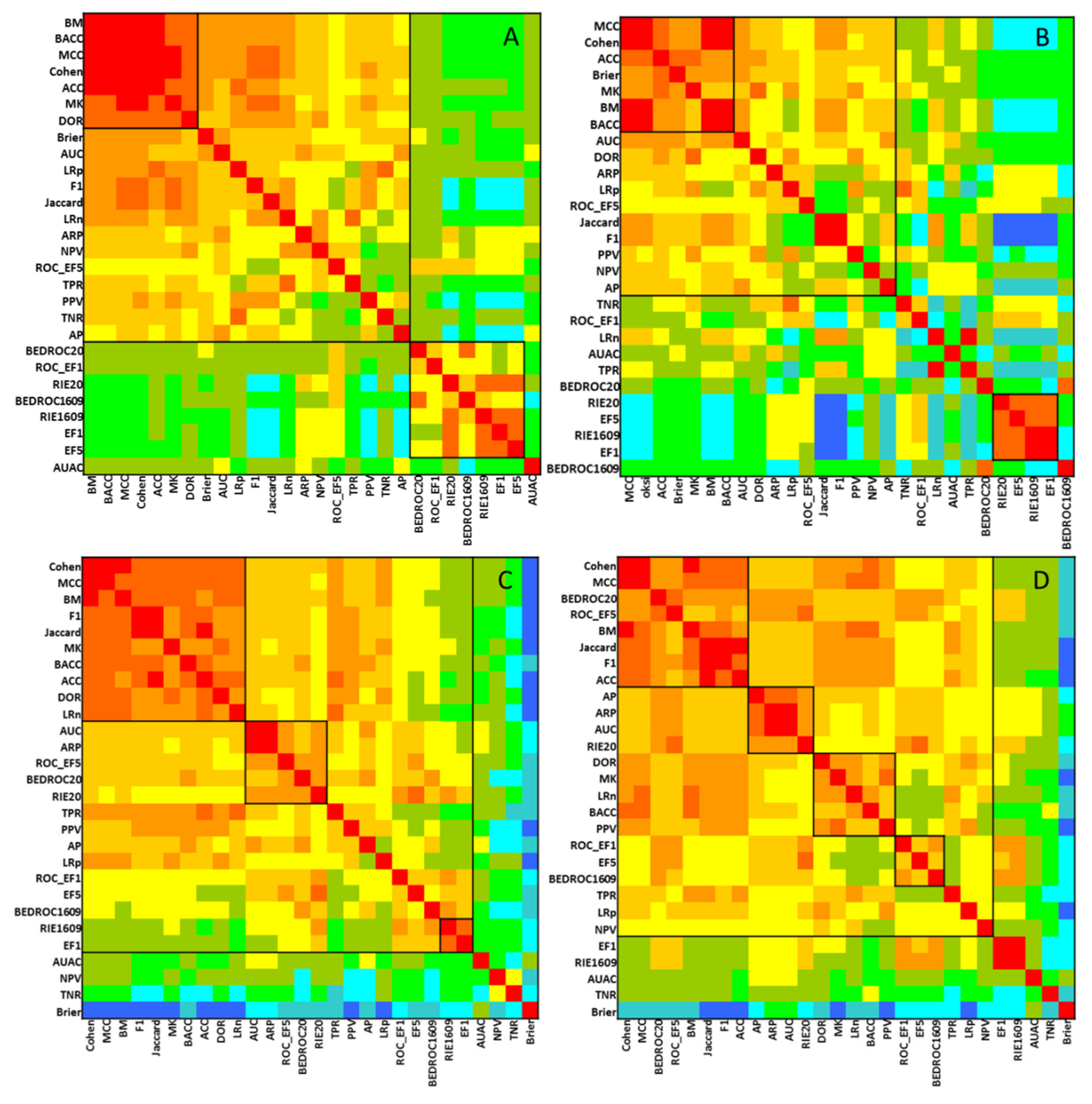
2.1. Statistical Evaluation of Performance Parameters
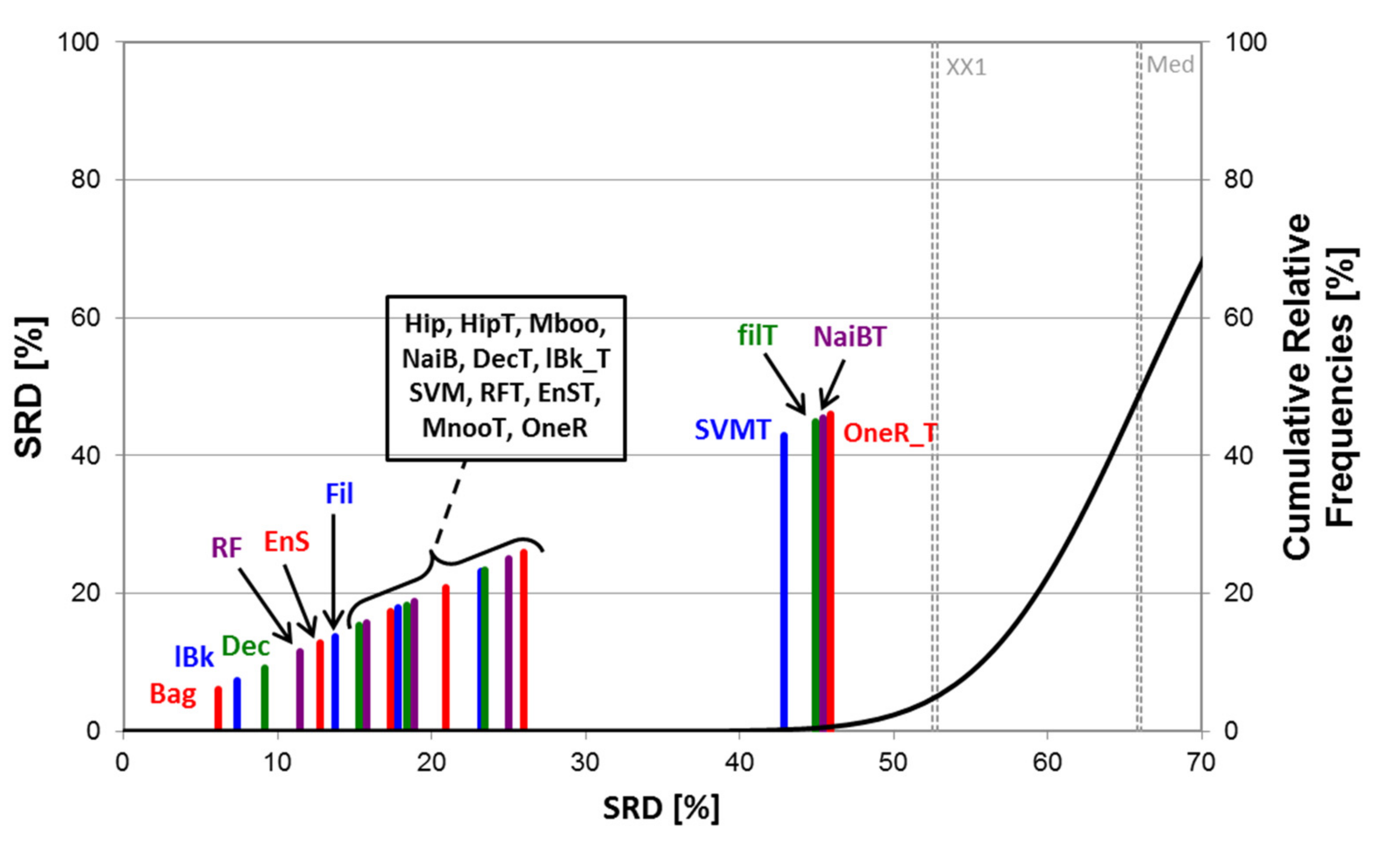
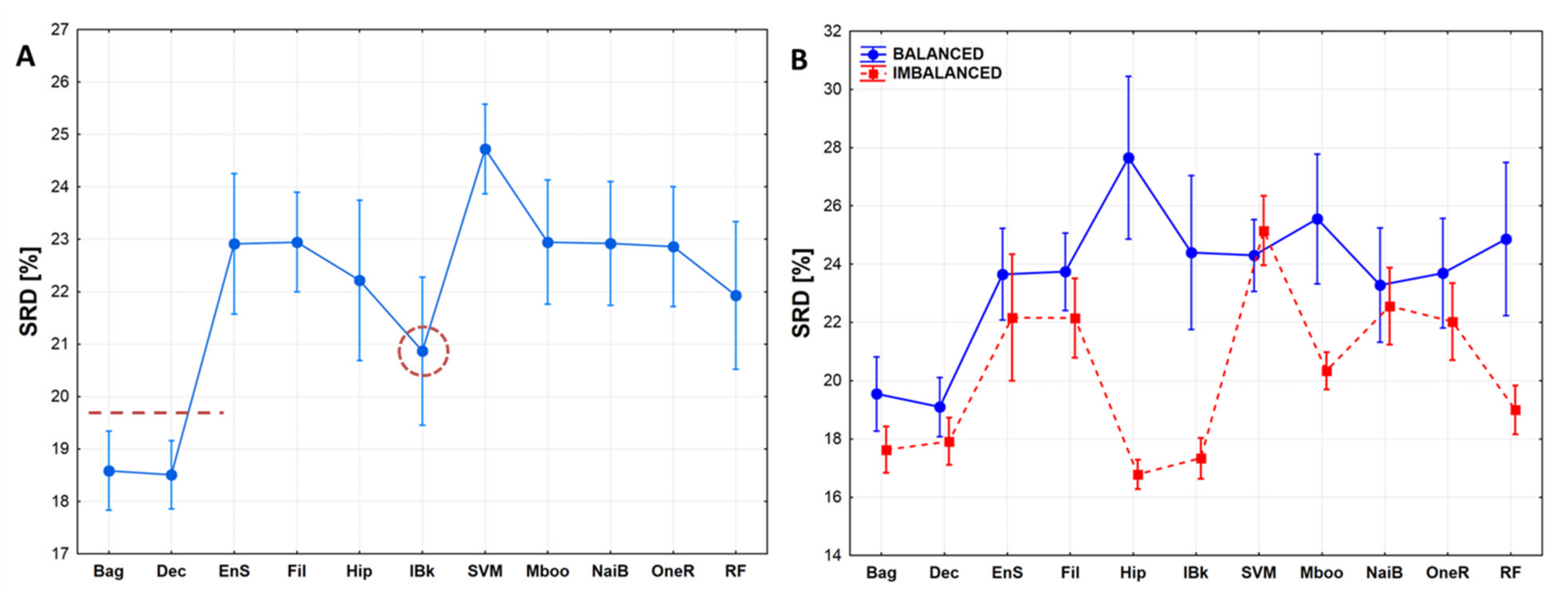
2.2. Statistical Evaluation of Machine Learning Models
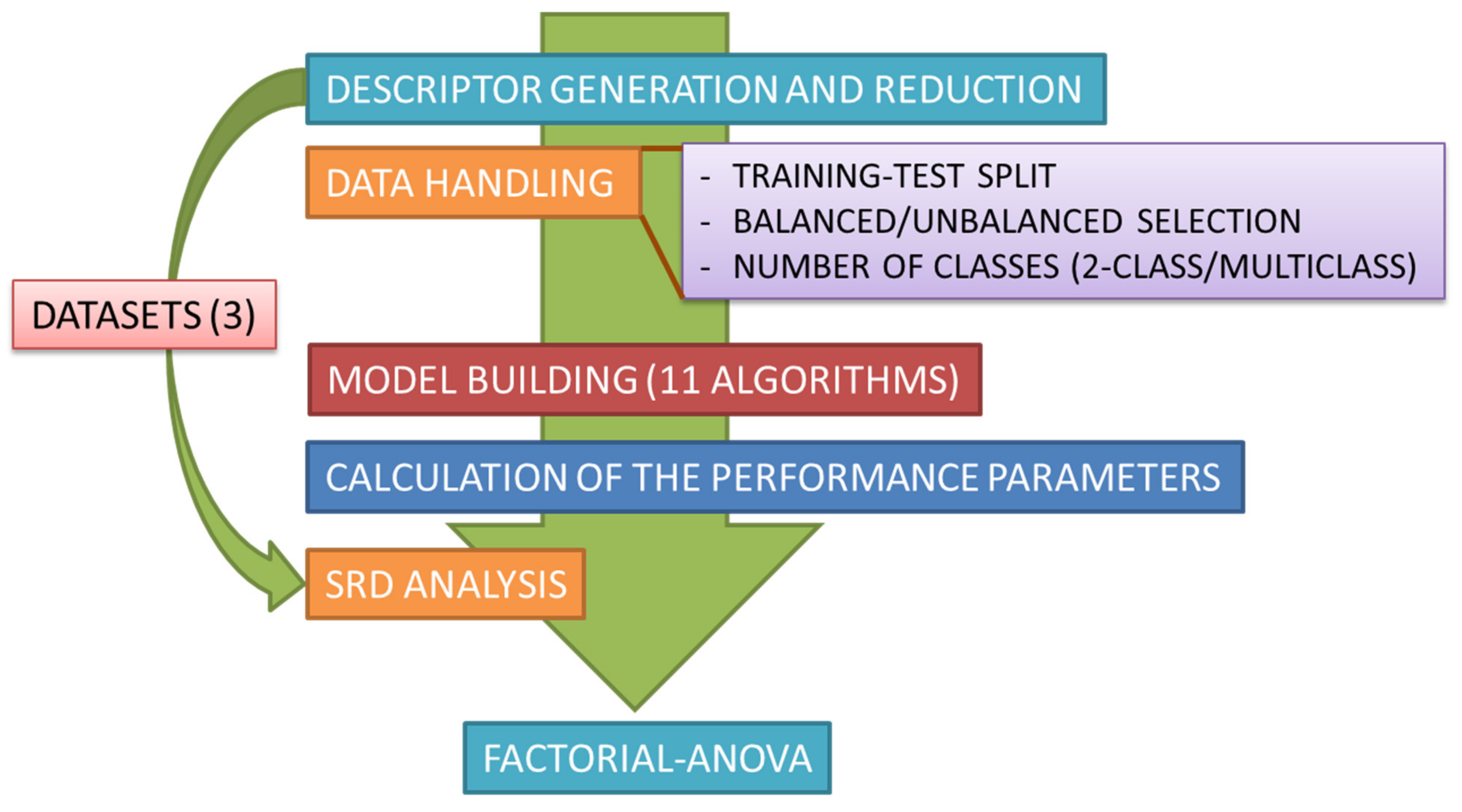
3. Methods
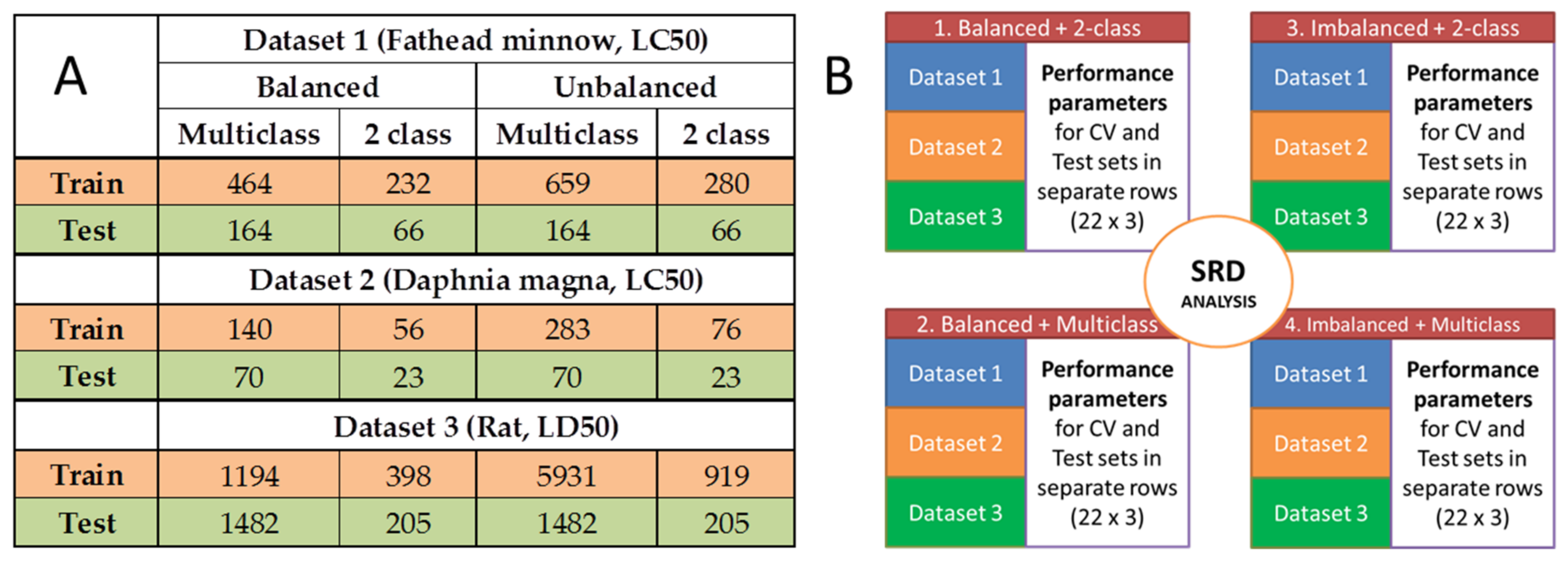
3.1. Datasets
3.2. Machine Learning Algorithms
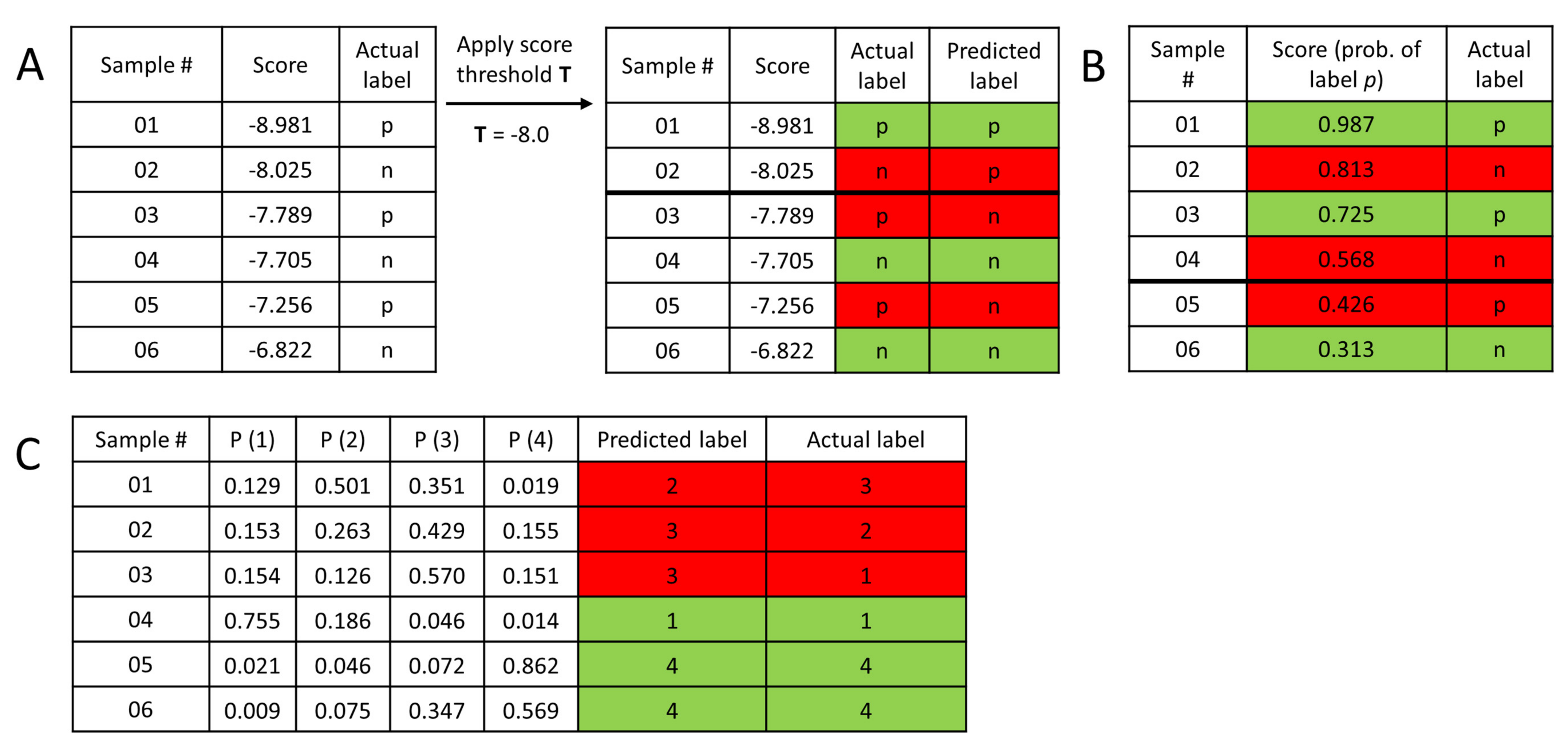
3.3. Performance Metrics
3.4. Statistical Evaluation
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Vamathevan, J.; Clark, D.; Czodrowski, P.; Dunham, I.; Ferran, E.; Lee, G.; Li, B.; Madabhushi, A.; Shah, P.; Spitzer, M.; et al. Applications of machine learning in drug discovery and development. Nat. Rev. Drug Discov. 2019, 18, 463–477. [Google Scholar] [CrossRef]
- Chen, H.; Engkvist, O.; Wang, Y.; Olivecrona, M.; Blaschke, T. The rise of deep learning in drug discovery. Drug Discov. Today 2018, 23, 1241–1250. [Google Scholar] [CrossRef] [PubMed]
- Rudin, C. Stop explaining black box machine learning models for high stakes decisions and use interpretable models instead. Nat. Mach. Intell. 2019, 1, 206–215. [Google Scholar] [CrossRef]
- Berrar, D. Performance Measures for Binary Classification. Encycl. Bioinform. Comput. Biol. 2019, 546–560. [Google Scholar]
- Héberger, K. Sum of ranking differences compares methods or models fairly. TrAC Trends Anal. Chem. 2010, 29, 101–109. [Google Scholar] [CrossRef]
- Rácz, A.; Bajusz, D.; Héberger, K. Consistency of QSAR models: Correct split of training and test sets, ranking of models and performance parameters. SAR QSAR Environ. Res. 2015, 26, 683–700. [Google Scholar] [CrossRef]
- Héberger, K.; Rácz, A.; Bajusz, D. Which Performance Parameters Are Best Suited to Assess the Predictive Ability of Models? In Advances in QSAR Modeling; Roy, K., Ed.; Springer: Cham, Switzerland, 2017; pp. 89–104. [Google Scholar]
- Rácz, A.; Bajusz, D.; Héberger, K. Modelling methods and cross-validation variants in QSAR: A multi-level analysis. SAR QSAR Environ. Res. 2018, 29, 661–674. [Google Scholar] [CrossRef]
- Piir, G.; Kahn, I.; García-Sosa, A.T.; Sild, S.; Ahte, P.; Maran, U. Best Practices for QSAR Model Reporting: Physical and Chemical Properties, Ecotoxicity, Environmental Fate, Human Health, and Toxicokinetics Endpoints. Environ. Health Perspect. 2018, 126, 126001. [Google Scholar] [CrossRef]
- Andrić, F.; Bajusz, D.; Rácz, A.; Šegan, S.; Héberger, K. Multivariate assessment of lipophilicity scales—computational and reversed phase thin-layer chromatographic indices. J. Pharm. Biomed. Anal. 2016, 127, 81–93. [Google Scholar] [CrossRef]
- Toxicity Estimation Software Tool (TEST)—EPA. Available online: https://www.epa.gov/chemical-research/toxicity-estimation-software-tool-test (accessed on 10 July 2019).
- Globally Harmonized System of Classification and Labelling of Chemicals (GHS). Available online: https://pubchem.ncbi.nlm.nih.gov/ghs/ (accessed on 5 July 2019).
- Rácz, A.; Bajusz, D.; Héberger, K. Intercorrelation Limits in Molecular Descriptor Preselection for QSAR/QSPR. Mol. Inform. 2019, 28. [Google Scholar] [CrossRef]
- John, G.H.; Langley, P. Estimating Continuous Distributions in Bayesian Classifiers. In Proceedings of the UAI’95 Eleventh Conference on Uncertainty in Artificial Intelligence, Montréal, QC, Canada, 18–20 August 1995; pp. 338–345. [Google Scholar]
- Software Documentation. WEKA API—Filtered Classifier. Available online: http://weka.sourceforge.net/doc.stable/ (accessed on 17 July 2019).
- Aha, D.W.; Kibler, D.; Albert, M.K. Instance-based learning algorithms. Mach. Learn. 1991, 6, 37–66. [Google Scholar] [CrossRef]
- Smusz, S.; Kurczab, R.; Bojarski, A.J. A multidimensional analysis of machine learning methods performance in the classification of bioactive compounds. Chemom. Intell. Lab. Syst. 2013, 128, 89–100. [Google Scholar] [CrossRef]
- Webb, G.I. MultiBoosting: A Technique for Combining Boosting and Wagging. Mach. Learn. 2000, 40, 159–196. [Google Scholar] [CrossRef]
- Chang, C.C.; Lin, C.J. LIBSVM—A lbrary for support vector machines. ACM Trans. Intell. Syst. Technol. (TIST) 2011, 2, 27. [Google Scholar]
- Holte, R.C. Very Simple Classification Rules Perform Well on Most Commonly Used Datasets. Mach. Learn. 1993, 91, 63–90. [Google Scholar] [CrossRef]
- Breiman, L.E.O. Bagging Predictors. Mach. Learn. 1996, 140, 123–140. [Google Scholar] [CrossRef]
- Melville, P.; Mooney, R.J. Constructing Diverse Classifier Ensembles using Artificial Training Examples. In Proceedings of the IJCAI-2003, Acapulco, Mexico, 9–15 August 2003; pp. 505–510. [Google Scholar]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Matthews, B.W. Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochim. Biophys. Acta Protein Struct. 1975, 405, 442–451. [Google Scholar] [CrossRef]
- Cramer, H. Mathematical Methods of Statistics; Princeton University Press: Princeton, NJ, USA, 1946; ISBN 0-691-08004-6. [Google Scholar]
- Powers, D.M.W. Evaluation: From Precision, Recall and F-Factor to ROC, Informedness, Markedness & Correlation. J. Mach. Learn. Technol. 2011, 2, 37–63. [Google Scholar]
- Nicholls, A. Confidence limits, error bars and method comparison in molecular modeling. Part 1: The calculation of confidence intervals. J. Comput. Aided. Mol. Des. 2014, 28, 887–918. [Google Scholar] [CrossRef]
- Czodrowski, P. Count on kappa. J. Comput. Aided. Mol. Des. 2014, 28, 1049–1055. [Google Scholar] [CrossRef] [PubMed]
- Sheridan, R.P.; Singh, S.B.; Fluder, E.M.; Kearsley, S.K. Protocols for Bridging the Peptide to Nonpeptide Gap in Topological Similarity Searches. J. Chem. Inf. Comput. Sci. 2001, 41, 1395–1406. [Google Scholar] [CrossRef] [PubMed]
- Fawcett, T. An introduction to ROC analysis. Pattern Recognit. Lett. 2006, 27, 861–874. [Google Scholar] [CrossRef]
- Truchon, J.F.; Bayly, C.I. Evaluating Virtual Screening Methods: Good and Bad Metrics for the “Early Recognition” Problem. J. Chem. Inf. Model. 2007, 47, 488–508. [Google Scholar] [CrossRef] [PubMed]
- Kairys, V.; Fernandes, M.X.; Gilson, M.K. Screening Drug-Like Compounds by Docking to Homology Models: A Systematic Study. J. Chem. Inf. Model. 2006, 46, 365–379. [Google Scholar] [CrossRef] [PubMed]
- Kollár-Hunek, K.; Héberger, K. Method and model comparison by sum of ranking differences in cases of repeated observations (ties). Chemom. Intell. Lab. Syst. 2013, 127, 139–146. [Google Scholar] [CrossRef]
- Bajusz, D.; Rácz, A.; Héberger, K. Why is Tanimoto index an appropriate choice for fingerprint-based similarity calculations? J. Cheminform. 2015, 7, 20. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Not available. |

| Class 1 | Class 2 | Class 3 | Class 4 | Class 5 | Class 6 | |||
|---|---|---|---|---|---|---|---|---|
| Dataset 1 | Balanced | Training | 116 | 116 | 116 | 116 | ||
| Test | 29 | 50 | 48 | 37 | ||||
| Imbalanced | Training | 116 | 166 | 213 | 164 | |||
| Test | 29 | 50 | 48 | 37 | ||||
| Dataset 2 | Balanced | Training | 28 | 28 | 28 | 28 | ||
| Test | 8 | 8 | 24 | 15 | ||||
| Imbalanced | Training | 48 | 65 | 58 | 84 | |||
| Test | 8 | 8 | 24 | 15 | ||||
| Dataset 3 | Balanced | Training | 199 | 199 | 199 | 199 | 199 | 199 |
| Test | 58 | 132 | 267 | 587 | 291 | 14 | ||
| Imbalanced | Training | 199 | 557 | 1053 | 2325 | 1178 | 619 | |
| Test | 58 | 132 | 267 | 587 | 291 | 14 |
| Name (Abbreviation) | Class. Scheme | Details |
|---|---|---|
| Naïve Bayes (NaiB) | Bayes | This algorithm is based on the Bayes theorem and the assumption of the independence of all attributes. The samples are examined separately and the individual probability of belonging to a class is calculated for each particular class. Standard options were used in WEKA NäiveBayes node [14]. |
| FilteredClassifier (Fil) | Meta | The algorithm is running an arbitrary classifier on data that has been passed through an arbitrary filter. Attribute selection filter was used with CfsSubset Evaluation and the best first search method [15]. |
| lBk, k-nearest neighbour (lBk) | Lazy | One of the simplest algorithms, where the class membership is assigned based on the majority vote of the k-nearest neighbours of an instance. Euclidean distance was used as distance measure and k = 1 was the number of used neighbours [16]. |
| HyperPipe (Hip) | Misc | Fast and simple algorithm, which is working well with many attributes. The basic idea of the method is the construction of pipes with different pattern of attributes to each class. The samples are monitored and selected to each class based on the pipes and the corresponding class [17]. |
| MultiboostAB (Mboo) | Meta | This algorithm is the modified version of the AdaBoost technique with wagging. The idea of wagging is to assign random weights to the cases in each training set based on Poisson distribution. In this case Decision stump classifier was used. The number of iteration was 10 and the weight threshold was 100. The number of subcommittees was set to 3 [18]. |
| libSVM, library SVM (SVM) | Func.* | Support vector machine can define hyperplane(s) in a higher dimensional space to separate the classes of samples distinctly. The plane should have the maximum margin between data points. Support vectors (points) can maximize the margin of the classifier. Different kernel functions and optimization parameters can be used for the classification task with SVM [19]. In this case radial basis function (RBF) was used as the kernel. |
| oneR, based on 1-rule, (OneR) | Rule | This algorithm ranks the attributes based on the error rate (on the training set). The basic concept is connected to 1-rules algorithms, where the samples are classified based on a single attribute [20]. Numeric values are treated as continuous ones. In this case, bucket size was 6 (standard) for the discretizing procedure of the attributes. |
| Bagging (Bag) | Meta | The basic concept of bagging is the creation of different models based on the bootstrapped training sets. The average (or vote) of these multiple versions are used for the prediction of class memberships for each sample [21]. In this case the number of iterations for bagging was set to 10. |
| Ensemble Selection (EnS) | Meta | It combines several classifier algorithms in the ensemble selection. The average prediction of the models in the ensemble is applied for the class membership determination. The selection of the models is based on an error metric (in our case RMSE). Forward selection was used for the optimization process of the ensemble. Iterations (here, 100) are also carried out such as in the case of Bagging. |
| Decorate (Dec) | Meta | It is also an ensemble-type algorithm, where the ensembles are constructed directly with diverse hypotheses with the application of additional artificially-constructed training examples to the original one. The classifier is working on the union of the original training and the artificial data (diversity data). The new classifiers are added to the ensemble, if the training error is not increased [22]. Several iterations are carried out to make the prediction stronger. Here, we applied 10 iterations. |
| Random Forest (RF) | Trees | Random forest is a tree-based method, which can be used for classification and regression problems alike. The basic idea is that it builds many trees and each of them predicts a classification. The final classification is made by a voting of the sequences of trees. The trees are weak predictors, but together they produce an ensemble; with the vote of each tree, the method can make good predictions [23]. |
| Predicted + (PP) | Predicted − (PN) | |
|---|---|---|
| Actual + (P) | True positive (TP) | False negative (FN) |
| Actual − (N) | False positive (FP) | True negative (TN) |
| Name | Alternative Names | Formula | Complementary Metric | Complementary Metric Formula |
|---|---|---|---|---|
| True positive rate (TPR) | Sensitivity, recall, hit rate | False negative rate (FNR), miss rate | ||
| True negative rate (TNR) | Specificity, selectivity | False positive rate (FPR), fall-out | ||
| Positive predictive value (PPV) | precision | False discovery rate (FDR) | ||
| Negative predictive value (NPV) | False omission rate (FOR) |
| Name | Formula | Description |
|---|---|---|
| Accuracy (ACC), or Correct classification rate (CC) | Readily generalized to multiple classes. Complementary metric: misclassification rate (or zero-one loss, or Hamming loss). | |
| Balanced accuracy (BACC) | Alternative of accuracy for imbalanced datasets. Readily generalized to multiple (k) classes. : number of samples correctly predicted into class j : actual number of samples in class j | |
| F1 score (F1), or F measure | Harmonic mean of precision and recall | |
| Matthews correlation coefficient (MCC) [24], φ coefficient (Pearson) [25] | Readily generalized to multiple classes. : number of samples predicted into class j : actual number of samples in class j : total no. of correctly predicted samples n: total no. of samples | |
| Bookmaker informedness (BM), or Informedness [26] | ||
| Markedness (MK) [26] | ||
| Positive likelihood ratio (LR+) | ||
| Negative likelihood ratio (LR−) | ||
| Diagnostic odds ratio (DOR) | ||
| Enrichment factor (EF) | Ratio of true positives in the top x% of the predictions, divided by ratio of positives in the whole dataset. | |
| ROC enrichment (ROC_EF) [27] | Ratio of TPR and FPR at a fixed FPR value (x). Independent of dataset composition. | |
| Cohen’s kappa [28] | Readily generalized to multiple classes. baseline corresponds to the random agreement probability. : number of samples predicted into class j : actual number of samples in class j n: total no. of samples | |
| Jaccard score (J) | Jaccard-Tanimoto similarity between the sets of predicted and actual (true) labels for the complete set of samples. | |
| Brier score loss (B) | Readily generalized to multiple classes. is the predicted probability of sample i belonging to class j, while is the actual outcome (0 or 1). Requires predicted probability values for each class. The smaller the better. | |
| Robust initial enhancement (RIE) [29] | is the rank of positive sample i in the ordered list of samples and is a parameter that defines the exponential weight. The denominator corresponds to the average sum of the exponential when P positives are uniformly distributed in the ordered list containing n samples. |
| Name | Formula | Description |
|---|---|---|
| Area under the ROC curve (AUC) [30] | Area under the TPR-FPR curve | Probability that a randomly selected positive sample will be ranked before a randomly selected negative. |
| Area under the accumulation curve (AUAC) | Area under the TPR-score (or TPR-rank) curve | If the ranks are normalized, then 0 ≤ AUAC ≤ 1 Probability that a randomly selected positive will be ranked before a randomly selected sample from a uniform distribution. |
| Average precision (AP) | Area under the precision-recall (PPV-TPR) curve | |
| Boltzmann-enhanced discrimination of receiver operating characteristic (BEDROC) [31] | See the definition of RIE above, is a parameter that defines the exponential weight. 0 ≤ BEDROC ≤ 1 BEDROC is an analog of AUC that assigns an (exponentially) greater weight to high-ranked samples, thus tackling the “early recognition problem”. | |
| Average rank (position) of actives (positives) () [32] | is the rank of positive sample i in the total ranked list of samples. The smaller the better. |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rácz, A.; Bajusz, D.; Héberger, K. Multi-Level Comparison of Machine Learning Classifiers and Their Performance Metrics. Molecules 2019, 24, 2811. https://doi.org/10.3390/molecules24152811
Rácz A, Bajusz D, Héberger K. Multi-Level Comparison of Machine Learning Classifiers and Their Performance Metrics. Molecules. 2019; 24(15):2811. https://doi.org/10.3390/molecules24152811
Chicago/Turabian StyleRácz, Anita, Dávid Bajusz, and Károly Héberger. 2019. "Multi-Level Comparison of Machine Learning Classifiers and Their Performance Metrics" Molecules 24, no. 15: 2811. https://doi.org/10.3390/molecules24152811
APA StyleRácz, A., Bajusz, D., & Héberger, K. (2019). Multi-Level Comparison of Machine Learning Classifiers and Their Performance Metrics. Molecules, 24(15), 2811. https://doi.org/10.3390/molecules24152811






