Click Chemistry as a Tool for Cell Engineering and Drug Delivery
Abstract
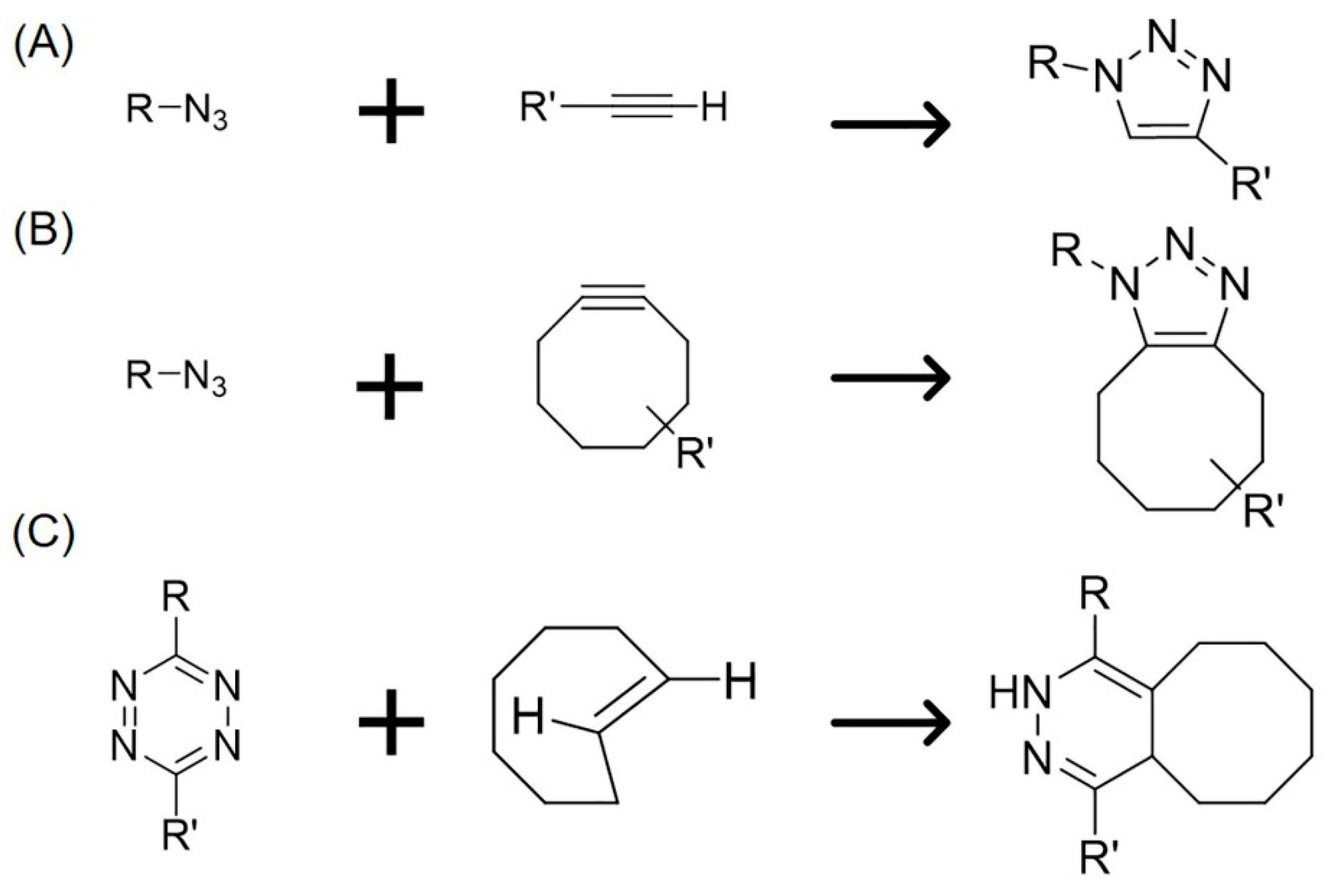
1. Introduction
2. Click Chemistry as a Tool for Cell Engineering in Cell Transplantation
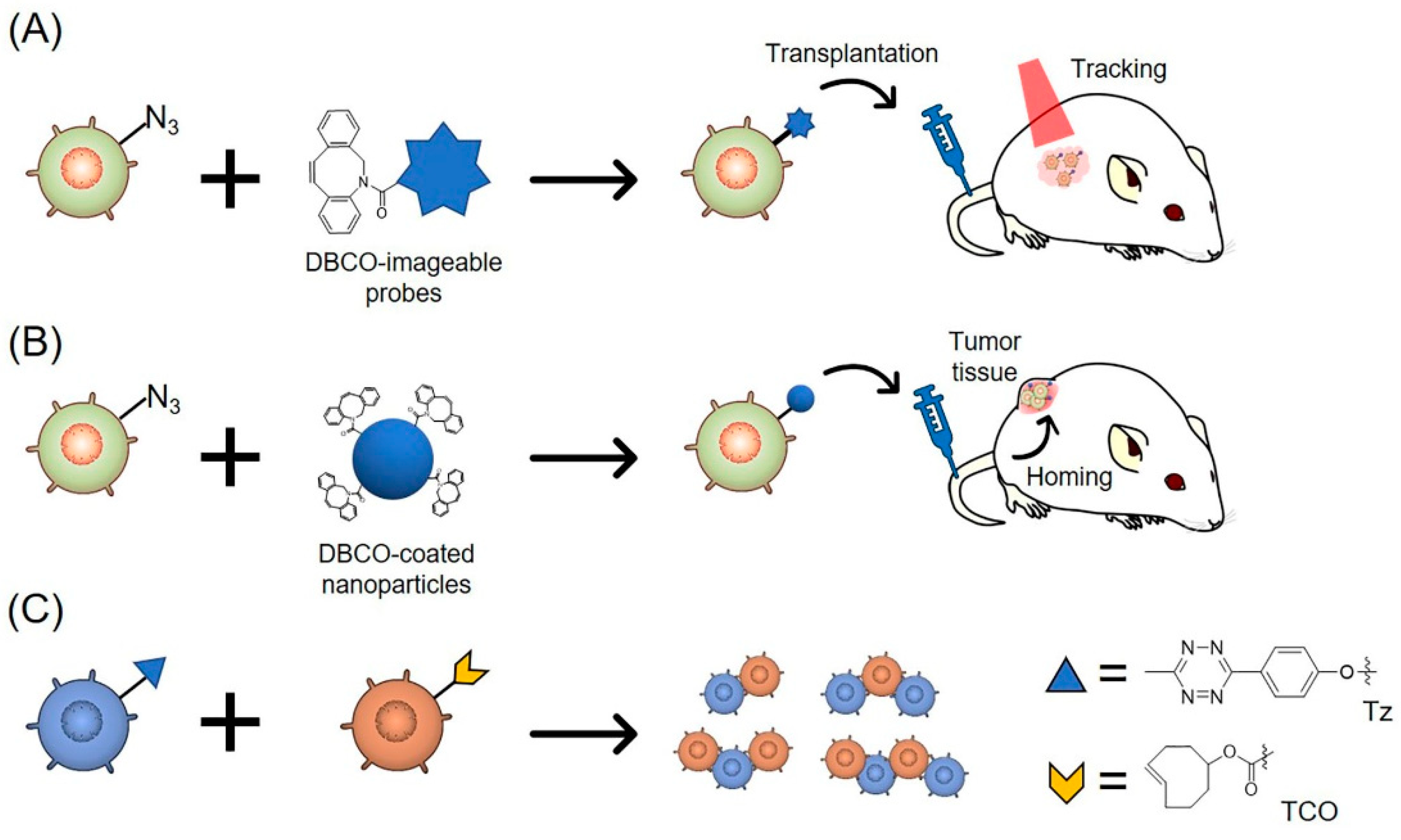
2.1. Click Chemistry as a Tracking Tool for Transplanted Cells
2.2. Click Chemistry as a Tool for Cell-Based Drug Delivery
2.3. Applications of Click Chemistry for the Formation of Cell Complexes
3. Applications of Click Chemistry for Drug Delivery in the Diagnosis and Treatment of Diseases
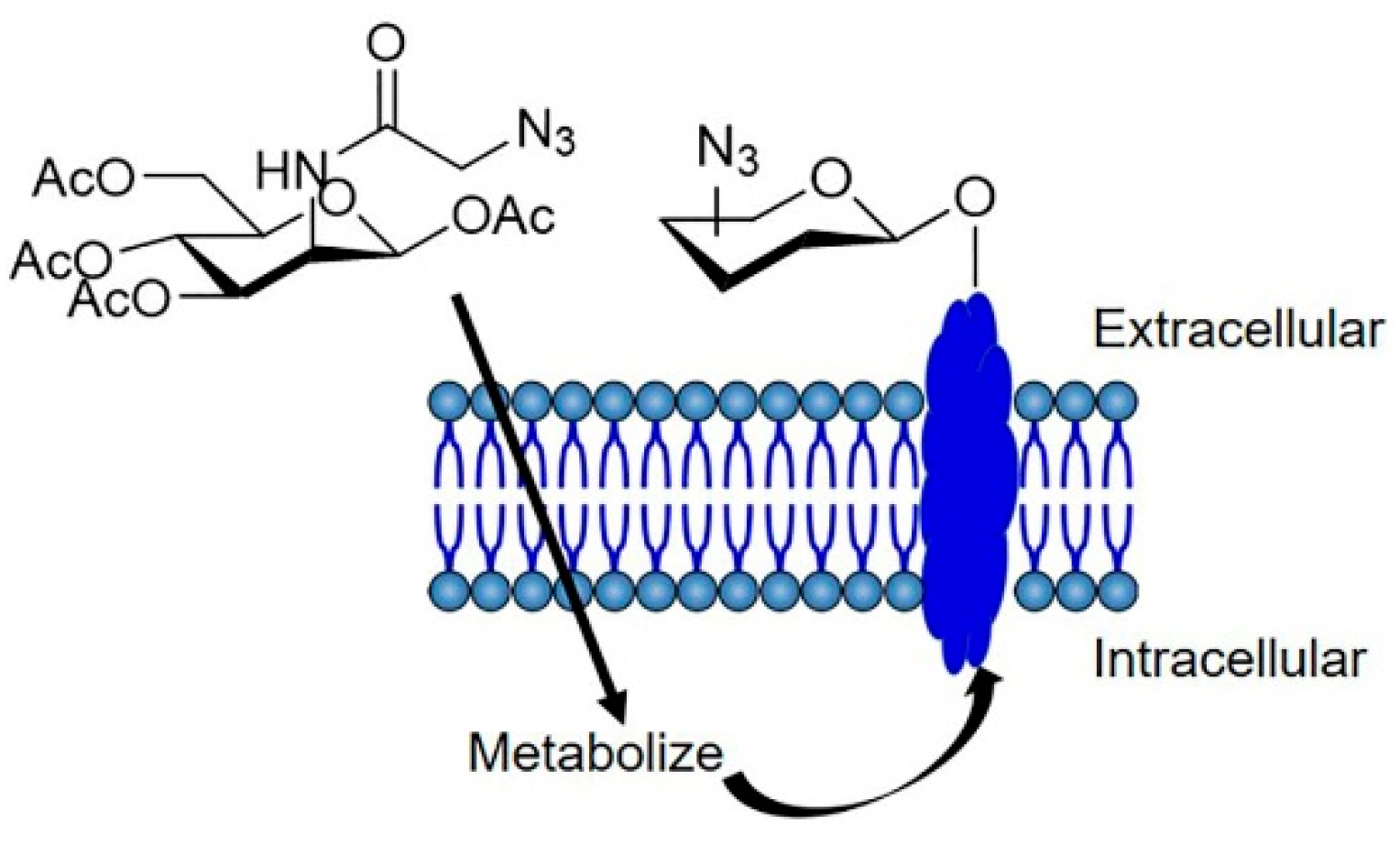
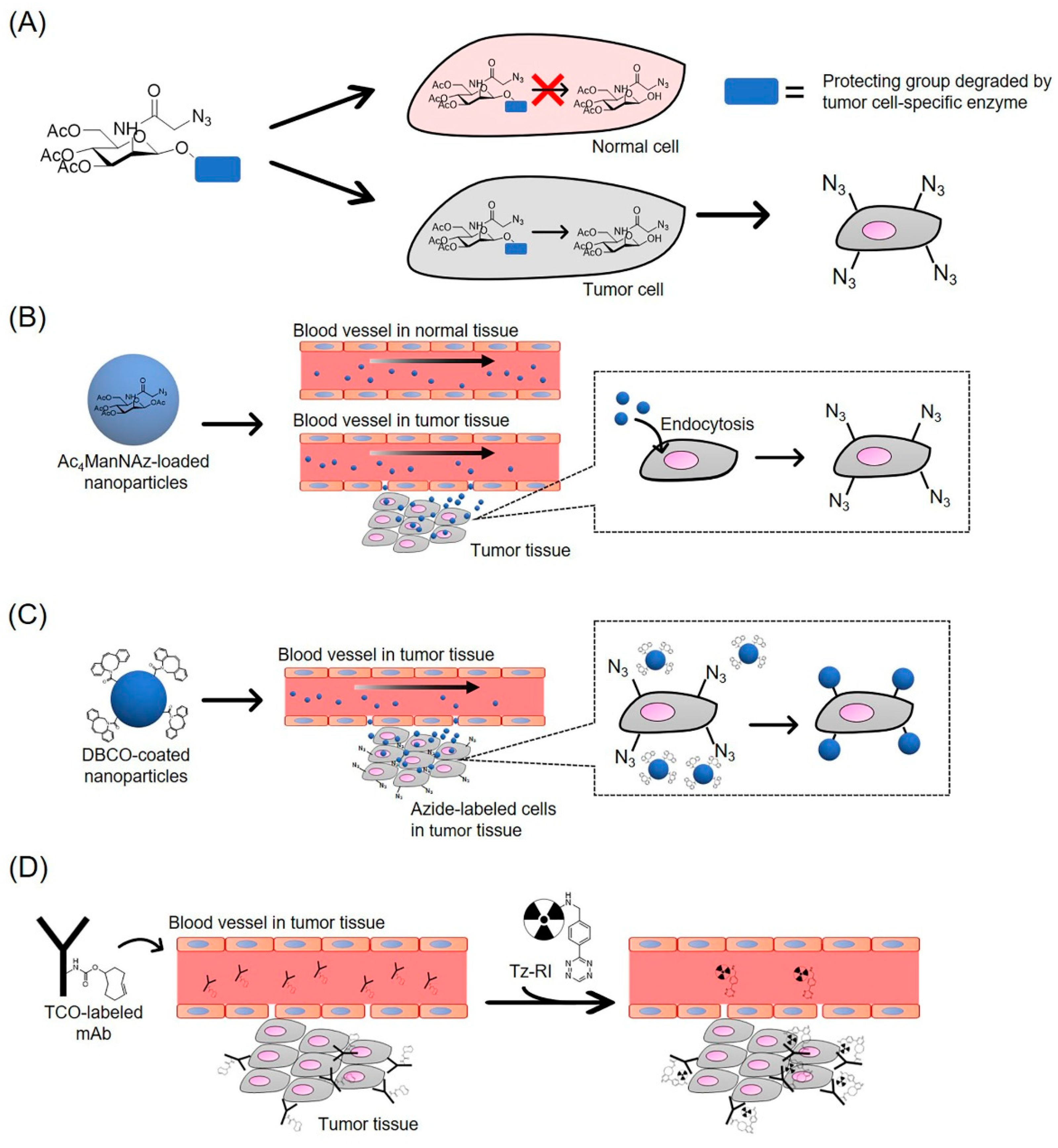
3.1. Tumor-Specific Labeling with Azide Groups
3.2. Tumor-Targeting Delivery by Click Chemistry for Cancer Therapy
3.3. Application of iEDDA Reaction for Tumor-Specific Delivery
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| A33-TCO | trans-Cyclooctene-modified humanized anti-a differentiation antigen (A33) monoclonal antibody |
| Ac4GalNAz | N-Azidoacetylgalactosamine tetraacylated |
| Ac3ManNAz | 1,3,4-Tri-O-acetyl-N-azidoacetylmannosamine |
| Ac4ManNAz | N-Azidoacetylmannosamine tetraacylated |
| ADCs | Monoclonal antibody-drug conjugates |
| ADIBO | Azadibenzocyclooctyne |
| ASCs | Adipose-derived mesenchymal stem cells |
| BCN | Bicyclo[6.1.0]nonyne |
| BCN-Ce6-CNPs | Glycol chitosan nanoparticle conjugated with bicyclo[6.1.0]nonyne and chlorine e6 |
| BCN-CNPs | Bicyclo[6.1.0]nonyne modified imageable glycol chitosan nanoparticle |
| CC49 | Anti-tumor associated glycoprotein 72 monoclonal anti body |
| CC49-TCO-DOX | CC49 conjugate with doxorubicin (DOX) via trans-cyclooctene linker |
| CEA | Carcinoembryonic antigen |
| CT | Computed tomography |
| CuAAC | Copper (I)-catalyzed azide-alkyne 1,3-dipolar cycloaddition |
| DBCO | Dibenzyl cyclooctyne- |
| DBCO-650 | Dibenzyl cyclooctyne-SETA 650 |
| DBCO-liposome | Dibenzyl cyclooctyne conjugated PEGylated liposome |
| DBCO-NPs | Dibenzyl cyclooctyne-modified poly (lactide-co-glycolide) nanoparticles |
| DBCO-VC-Dox | Dibenzyl cyclooctyne-valine-cysteine-doxorubicin conjugate |
| DCL-AAM | Histone deacetylase/cathepsin l-responsive acetylated azidomannose |
| DIFO | Difluorinated cyclooctyne |
| DOTA | 1,4,7,10-Tetraazacyclododecane- N,N’,N”,N’”-tetraacetic acid |
| DOX | Doxorubicin |
| EGFR | Epidermal growth factor receptor |
| EPR | Enhanced permeability and retention |
| HER-2 | Human epidermal growth factor receptor 2 |
| iEDDA | Inverse electron demand Diels-Alder |
| mAb | Monoclonal antibodies |
| MR | Magnetic resonance |
| MSCs | Mesenchymal stem cells |
| Nano-MPs | Nano-sized metabolic precursors |
| NIR | Near-infrared |
| NOTA | 1,4,7-Triazacyclononane-N,N′,N”-triacetic acid |
| OCT | Cyclooctyne |
| PEG | Poly(ethylene glycol) |
| PET | Positron emission tomography |
| PLGA | Poly (lactide-co-glycolide) |
| PSA | Prostate-specific antigen |
| RI | Radio isotope |
| RR-S-Ac3ManNAz | Arg-Arg-p-aminobenzyloxycarbonyl-1,3,4-tri-O-acetyl-N-azidoacetylmannosamine |
| SPAAC | Strain-promoted [3 + 2] azide-alkyne cycloaddition |
| SPECT | Single-photon emission computed tomography |
| TAG72 | Tumor associated glycoprotein 72 |
| TCO | trans-Cyclooctene |
| TCO-anti-VEGFR2 | trans-Cyclooctene-conjugated anti-vascular endothelial growth factor receptor 2 |
| Tz | Tetrazines |
References
- Kolb, H.C.; Finn, M.G.; Sharpless, K.B. Click chemistry: Diverse chemical function from a few good reactions. Angew. Chem. Int. Ed. Engl. 2001, 40, 2004–2021. [Google Scholar] [CrossRef]
- Yang, M.; Li, J.; Chen, P.R. Transition metal-mediated bioorthogonal protein chemistry in living cells. Chem. Soc. Rev. 2014, 43, 6511–6526. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Zhang, Z. Development and applications of the copper-catalyzed azide-alkyne cycloaddition (CuAAC) as a bioorthogonal reaction. Molecules 2016, 21, 1393. [Google Scholar] [CrossRef]
- Agard, N.J.; Prescher, J.A.; Bertozzi, C.R. A strain-promoted [3 + 2] azide-alkyne cycloaddition for covalent modification of biomolecules in living systems. J. Am. Chem. Soc. 2004, 126, 15046–15047. [Google Scholar] [CrossRef] [PubMed]
- Sletten, E.M.; Bertozzi, C.R. From mechanism to mouse: A tale of two bioorthogonal reactions. Acc. Chem. Res. 2011, 44, 666–676. [Google Scholar] [CrossRef] [PubMed]
- Debets, M.F.; van Berkel, S.S.; Schoffelen, S.; Rutjes, F.P.; van Hest, J.C.; van Delft, F. Aza-dibenzocyclooctynes for fast and efficient enzyme PEGylation via copper-free [3 + 2] cycloaddition. Chem. Commun. 2010, 46, 97–99. [Google Scholar] [CrossRef] [PubMed]
- Kuzmin, A.; Poloukhtine, A.; Wolfert, M.A.; Popik, V.V. Surface functionalization using catalyst-free azide-alkyne cycloaddition. Bioconjug. Chem. 2010, 21, 2076–2085. [Google Scholar] [CrossRef] [PubMed]
- Jewett, J.C.; Sletten, E.M.; Bertozzi, C.R. Rapid Cu-free click chemistry with readily synthesized biarylazacyclooctynones. J. Am. Chem. Soc. 2010, 132, 3688–3690. [Google Scholar] [CrossRef] [PubMed]
- Dommerholt, J.; Schmidt, S.; Temming, R.; Hendriks, L.J.; Rutjes, F.P.; van Hest, J.C.; Lefeber, D.J.; Friedl, P.; van Delft, F.L. Readily accessible bicyclononynes for bioorthogonal labeling and three-dimensional imaging of living cells. Angew. Chem. Int. Ed. Engl. 2010, 49, 9422–9425. [Google Scholar] [CrossRef]
- Ning, X.; Guo, J.; Wolfert, M.A.; Boons, G.J. Visualizing metabolically labeled glycoconjugates of living cells by copper-free and fast Huisgen cycloadditions. Angew. Chem. Int. Ed. Engl. 2008, 47, 2253–2255. [Google Scholar] [CrossRef]
- Baskin, J.M.; Prescher, J.A.; Laughlin, S.T.; Agard, N.J.; Chang, P.V.; Miller, I.A.; Lo, A.; Codelli, J.A.; Bertozzi, C.R. Copper-free click chemistry for dynamic in vivo imaging. Proc. Natl. Acad. Sci. USA 2007, 104, 16793–16797. [Google Scholar] [CrossRef]
- Dadfar, S.M.M.; Sekula-Neuner, S.; Bog, U.; Trouillet, V.; Hirtz, M. Site-specific surface functionalization via microchannel cantilever spotting (µCS): Comparison between azide-alkyne and thiol-alkyne click chemistry reactions. Small 2018, 14, e1800131. [Google Scholar] [CrossRef] [PubMed]
- Blackman, M.L.; Royzen, M.; Fox, J.M. Tetrazine ligation: Fast bioconjugation based on inverse-electron-demand Diels-Alder reactivity. J. Am. Chem. Soc. 2008, 130, 13518–13519. [Google Scholar] [CrossRef] [PubMed]
- Devaraj, N.K.; Weissleder, R.; Hilderbrand, S.A. Tetrazine-based cycloadditions: Application to pretargeted live cell imaging. Bioconjug. Chem. 2008, 19, 2297–2299. [Google Scholar] [CrossRef] [PubMed]
- Patterson, D.M.; Nazarova, L.A.; Xie, B.; Kamber, D.N.; Prescher, J.A. Functionalized cyclopropenes as bioorthogonal chemical reporters. J. Am. Chem. Soc. 2012, 134, 18638–18643. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Šečkutė, J.; Cole, C.M.; Devaraj, N.K. Live-cell imaging of cyclopropene tags with fluorogenic tetrazine cycloadditions. Angew. Chem. Int. Ed. Engl. 2012, 51, 7476–7479. [Google Scholar] [CrossRef] [PubMed]
- Engelsma, S.B.; Willems, L.I.; van Paaschen, C.E.; van Kasteren, S.; van der Marel, G.A.; Overkleeft, H.S.; Filippov, D.V. Acylazetine as a dienophile in bioorthogonal inverse electron-demand Diels-Alder ligation. Org. Lett. 2014, 16, 2744–2747. [Google Scholar] [CrossRef]
- Eising, S.; Lelivelt, F.; Bonger, K.M. Vinylboronic acids as fast reacting, synthetically accessible, and stable bioorthogonal reactants in the carboni-lindsey reaction. Angew. Chem. Int. Ed. Engl. 2016, 55, 12243–12247. [Google Scholar] [CrossRef]
- Kiick, K.L.; Saxon, E.; Tirrell, D.A.; Bertozzi, C.R. Incorporation of azides into recombinant proteins for chemoselective modification by the Staudinger ligation. Proc. Natl. Acad. Sci. USA 2002, 99, 19–24. [Google Scholar] [CrossRef]
- Saxon, E.; Bertozzi, C.R. Cell surface engineering by a modified Staudinger reaction. Science 2000, 287, 2007–2010. [Google Scholar] [CrossRef]
- Agard, N.J.; Bertozzi, C.R. Chemical approaches to perturb, profile, and perceive glycans. Acc. Chem. Res. 2009, 42, 788–797. [Google Scholar] [CrossRef] [PubMed]
- Ricks, T.J.; Cassilly, C.D.; Carr, A.J.; Alves, D.S.; Alam, S.; Tscherch, K.; Yokley, T.W.; Workman, C.E.; Morrell-Falvey, J.L.; Barrera, F.N.; et al. Labeling of phosphatidylinositol lipid products in cells through metabolic engineering by using a clickable myo-inositol probe. Chembiochem 2018. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, B.L.; Guo, Z.; Bernardes, G.J.L. Inverse electron demand Diels-Alder reactions in chemical biology. Chem. Soc. Rev. 2017, 46, 4895–4950. [Google Scholar] [CrossRef] [PubMed]
- Späte, A.K.; Bußkamp, H.; Niederwieser, A.; Schart, V.F.; Marx, A.; Wittmann, V. Rapid labeling of metabolically engineered cell-surface glycoconjugates with a carbamate-linked yclopropane reporter. Bioconjug. Chem. 2014, 25, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Du, J.; Meledeo, M.A.; Wang, Z.; Khanna, H.S.; Paruchuri, V.D.; Yarema, K.J. Metabolic glycoengineering: sialic acid and beyond. Glycobiology 2009, 19, 1382–1401. [Google Scholar] [CrossRef]
- Laughlin, S.T.; Bertozzi, C.R. In vivo imaging of Caenorhabditis elegans glycans. ACS Chem. Biol. 2009, 4, 1068–1072. [Google Scholar] [CrossRef] [PubMed]
- Han, S.S.; Lee, D.E.; Shim, H.E.; Lee, S.; Jung, T.; Oh, J.H.; Lee, H.A.; Moon, S.H.; Jeon, J.; Yoon, S.; et al. Physiological effects of Ac4ManNAz and optimization of metabolic labeling for cell tracking. Theranostics 2017, 7, 1164–1176. [Google Scholar] [CrossRef]
- Xie, R.; Dong, L.; Huang, R.; Hong, S.; Lei, R.; Chen, X. Targeted imaging and proteomic analysis of tumor-associated glycans in living animals. Angew. Chem. Int. Ed. Engl. 2014, 53, 14082–14086. [Google Scholar] [CrossRef]
- Yoon, H.I.; Yhee, J.Y.; Na, J.H.; Lee, S.; Lee, H.; Kang, S.W.; Chang, H.; Ryu, J.H.; Lee, S.; Kwon, I.C.; et al. Bioorthogonal copper free click chemistry for labeling and tracking of chondrocytes in vivo. Bioconjug. Chem. 2016, 27, 927–936. [Google Scholar] [CrossRef]
- Yoon, H.Y.; Koo, H.; Kim, K.; Kwon, I.C. Molecular imaging based on metabolic glycoengineering and bioorthogonal click chemistry. Biomaterials 2017, 132, 28–36. [Google Scholar] [CrossRef]
- Lee, S.Y.; Lee, S.; Lee, J.; Yhee, J.Y.; Yoon, H.I.; Park, S.J.; Koo, H.; Moon, S.H.; Lee, H.; Cho, Y.W.; et al. Non-invasive stem cell tracking in hindlimb ischemia animal model using bio-orthogonal copper-free click chemistry. Biochem. Biophys. Res. Commun. 2016, 479, 779–786. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Li, R.; Chen, Y.; Zhang, S.; Wang, W.; Li, F. Applying DNA rolling circle amplification in fluorescence imaging of cell surface glycans labeled by a metabolic method. Chem. Sci. 2016, 7, 6182–6189. [Google Scholar] [CrossRef] [PubMed]
- Layek, B.; Sadhukha, T.; Prabha, S. Glycoengineered mesenchymal stem cells as an enabling platform for two-step targeting of solid tumors. Biomaterials 2016, 88, 97–109. [Google Scholar] [CrossRef] [PubMed]
- Koo, H.; Choi, M.; Kim, E.; Hahn, S.K.; Weissleder, R.; Yun, S.H. Bioorthogonal click chemistry-based synthetic cell glue. Small 2015, 11, 6458–6466. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Yoon, H.I.; Na, J.H.; Jeon, S.; Lim, S.; Koo, H.; Han, S.S.; Kang, S.W.; Park, S.J.; Moon, S.H.; et al. In vivo stem cell tracking with imageable nanoparticles that bind bioorthogonal chemical receptors on the stem cell surface. Biomaterials 2017, 139, 12–29. [Google Scholar] [CrossRef] [PubMed]
- Kang, K.; Joo, S.; Choi, J.Y.; Geum, S.; Hong, S.P.; Lee, S.Y.; Kim, Y.H.; Kim, S.M.; Yoon, M.H.; Nam, Y.; et al. Tissue-based metabolic labeling of polysialic acids in living primary hippocampal neurons. Proc. Natl. Acad. Sci. USA 2015, 112, 241–248. [Google Scholar] [CrossRef]
- Lee, S.; Jung, S.; Koo, H.; Na, J.H.; Yoon, H.Y.; Shim, M.K.; Park, J.; Kim, J.H.; Lee, S.; Pomper, M.G.; et al. Nano-sized metabolic precursors for heterogeneous tumor-targeting strategy using bioorthogonal click chemistry in vivo. Biomaterials 2017, 148, 1–15. [Google Scholar] [CrossRef]
- Kang, S.W.; Lee, S.; Na, J.H.; Yoon, H.I.; Lee, D.E.; Koo, H.; Cho, Y.W.; Kim, S.H.; Jeong, S.Y.; Kwon, I.C.; et al. Cell labeling and tracking method without distorted signals by phagocytosis of macrophages. Theranostics 2014, 4, 420–431. [Google Scholar] [CrossRef]
- Prescher, J.A.; Dube, D.H.; Bertozzi, C.R. Chemical remodelling of cell surfaces in living animals. Nature 2004, 430, 873–877. [Google Scholar] [CrossRef]
- Koo, H.; Lee, S.; Na, J.H.; Kim, S.H.; Hahn, S.K.; Choi, K.; Kwon, I.C.; Jeong, S.Y.; Kim, K. Bioorthogonal copper-free click chemistry in vivo for tumor-targeted delivery of nanoparticles. Angew. Chem. Int. Ed. Engl. 2012, 51, 11836–11840. [Google Scholar] [CrossRef]
- Wang, H.; Wang, R.; Cai, K.; He, H.; Liu, Y.; Yen, J.; Wang, Z.; Xu, M.; Sun, Y.; Zhou, X.; et al. Selective in vivo metabolic cell-labeling-mediated cancer targeting. Nat. Chem. Biol. 2017, 13, 415–424. [Google Scholar] [CrossRef]
- Praveschotinunt, P.; Dorval Courchesne, N.M.; den Hartog, I.; Lu, C.; Kim, J.J.; Nguyen, P.Q.; Joshi, N.S. Tracking of engineered bacteria in vivo using nonstandard amino acid incorporation. ACS Synth. Biol. 2018, 7, 1640–1650. [Google Scholar] [CrossRef]
- Du, L.; Qin, H.; Ma, T.; Zhang, T.; Xing, D. In vivo imaging-guided photothermal/photoacoustic synergistic therapy with bioorthogonal metabolic glycoengineering-activated tumor targeting nanoparticles. ACS Nano 2017, 11, 8930–8943. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Li, J.; Xie, R.; Fan, X.; Liu, Y.; Zheng, S.; Ge, Y.; Chen, P.R. Bioorthogonal chemical activation of kinases in living systems. ACS Cent. Sci. 2016, 325–331. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wang, B.; Zhao, N.; Tian, Z.; Dai, Y.; Nie, Y.; Tian, J.; Wang, Z.; Chen, X. Improved tumor targeting and longer retention time of NIR fluorescent probes using bioorthogonal chemistry. Theranostics 2017, 7, 3794–3802. [Google Scholar] [CrossRef]
- Mao, F.; Tu, Q.; Wang, L.; Chu, F.; Li, X.; Li, H.S.; Xu, W. Mesenchymal stem cells and their therapeutic applications in inflammatory bowel disease. Oncotarget 2017, 8, 38008–38021. [Google Scholar] [CrossRef] [PubMed]
- Galipeau, J.; Sensébé, L. Mesenchymal stromal cells: Clinical challenges and therapeutic opportunities. Cell Stem Cell 2018, 22, 824–833. [Google Scholar] [CrossRef]
- Lee, R.H.; Pulin, A.A.; Seo, M.J.; Kota, D.J.; Ylostalo, J.; Larson, B.L.; Semprun-Prieto, L.; Delafontaine, P.; Prockop, D.J. Intravenous hMSCs improve myocardial infarction in mice because cells embolized in lung are activated to secrete the anti-inflammatory protein TSG-6. Cell Stem Cell 2009, 5, 54–63. [Google Scholar] [CrossRef] [PubMed]
- Kerkelä, E.; Hakkarainen, T.; Mäkelä, T.; Raki, M.; Kambur, O.; Kilpinen, L.; Nikkilä, J.; Lehtonen, S.; Ritamo, I.; Pernu, R. Transient proteolytic modification of mesenchymal stromal cells increases lung clearance rate and targeting to injured tissue. Stem Cells Transl. Med. 2013, 2, 510–520. [Google Scholar] [CrossRef] [PubMed]
- Kircher, M.F.; Gambhir, S.S.; Grimm, J. Noninvasive cell-tracking methods. Nat. Rev. Clin. Oncol. 2011, 8, 677–688. [Google Scholar] [CrossRef]
- Takayama, Y.; Kusamori, K.; Hayashi, M.; Tanabe, N.; Matsuura, S.; Tsujimura, M.; Katsumi, H.; Sakane, T.; Nishikawa, M.; Yamamoto, A. Long-term drug modification to the surface of mesenchymal stem cells by the avidin-biotin complex method. Sci. Rep. 2017, 7, 16953. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, R.; Ghosh, M.; Chowdhury, A.S.; Ray, N. Cell tracking in microscopic video using matching and linking of bipartite graphs. Comput. Methods Programs Biomed. 2013, 112, 422–431. [Google Scholar] [CrossRef] [PubMed]
- Yukawa, H.; Watanabe, M.; Kaji, N.; Okamoto, Y.; Tokeshi, M.; Miyamoto, Y.; Noguchi, H.; Baba, Y.; Hayashi, S. Monitoring transplanted adipose tissue-derived stem cells combined with heparin in the liver by fluorescence imaging using quantum dots. Biomaterials 2012, 33, 2177–2186. [Google Scholar] [CrossRef] [PubMed]
- Gu, E.; Chen, W.Y.; Gu, J.; Burridge, P.; Wu, J.C. Molecular imaging of stem cells: Tracking survival, biodistribution, tumorigenicity, and immunogenicity. Theranostics 2012, 2, 335–345. [Google Scholar] [CrossRef]
- Hakamata, Y.; Murakami, T.; Kobayashi, E. "Firefly rats" as an organ/cellular source for long-term in vivo bioluminescent imaging. Transplantation 2006, 81, 1179–1184. [Google Scholar] [CrossRef]
- Li, Z.; Suzuki, Y.; Huang, M.; Cao, F.; Xie, X.; Connolly, A.J.; Yang, P.C.; Wu, J.C. Comparison of reporter gene and iron particle labeling for tracking fate of human embryonic stem cells and differentiated endothelial cells in living subjects. Stem Cells 2008, 26, 864–873. [Google Scholar] [CrossRef]
- Patrick, P.S.; Hammersley, J.; Loizou, L.; Kettunen, M.; Rodrigues, T.B.; Hu, D.E.; Tee, S.S.; Hesketh, R.; Lyons, S.K.; Soloviev, D.; et al. Dual-modality gene reporter for in vivo imaging. Proc. Natl. Acad. Sci. USA 2014, 111, 415–420. [Google Scholar] [CrossRef] [PubMed]
- Iwano, S.; Sugiyama, M.; Hama, H.; Watakabe, A.; Hasegawa, N.; Kuchimaru, T.; Tanaka, K.Z.; Takahashi, M.; Ishida, Y.; Hata, J.; et al. Single-cell bioluminescence imaging of deep tissue in freely moving animals. Science 2018, 359, 935–939. [Google Scholar] [CrossRef] [PubMed]
- Higuchi, Y.; Wu, C.; Chang, K.L.; Irie, K.; Kawakami, S.; Yamashita, F.; Hashida, M. Polyamidoamine dendrimer-conjugated quantum dots for efficient labeling of primary cultured mesenchymal stem cells. Biomaterials 2011, 32, 6676–6682. [Google Scholar] [CrossRef] [PubMed]
- Anderson, S.A.; Glod, J.; Arbab, A.S.; Noel, M.; Ashari, P.; Fine, H.A.; Frank, J.A. Noninvasive MR imaging of magnetically labeled stem cells to directly identify neovasculature in a glioma model. Blood 2005, 105, 420–425. [Google Scholar] [CrossRef]
- Rodriguez-Porcel, M. In vivo imaging and monitoring of transplanted stem cells: Clinical applications. Curr. Cardiol. Rep. 2010, 12, 51–58. [Google Scholar] [CrossRef]
- Kusamori, K.; Takayama, Y.; Nishikawa, M. Stable surface modification of mesenchymal stem cells using the avidin-biotin complex technique. Curr. Protoc. Stem Cell Biol. 2018, 47, e66. [Google Scholar] [CrossRef] [PubMed]
- Ogihara, Y.; Yukawa, H.; Kameyama, T.; Nishi, H.; Onoshima, D.; Ishikawa, T.; Torimoto, T.; Baba, Y. Labeling and in vivo visualization of transplanted adipose tissue-derived stem cells with safe cadmium-free aqueous ZnS coating of ZnS-AgInS2 nanoparticles. Sci. Rep. 2017, 7, 40047. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Park, G.; Hyun, H.; Lee, J.H.; Ashitate, Y.; Choi, J.; Hong, G.H.; Owens, E.A.; Henary, M.; Choi, H.S. Near-infrared lipophilic fluorophores for tracking tissue growth. Biomed. Mater. 2013, 8, 014110. [Google Scholar] [CrossRef]
- Kidd, S.; Spaeth, E.; Dembinski, J.L.; Dietrich, M.; Watson, K.; Klopp, A.; Battula, V.L.; Weil, M.; Andreeff, M.; Marini, F.C. Direct evidence of mesenchymal stem cell tropism for tumor and wounding microenvironments using in vivo bioluminescent imaging. Stem Cells 2009, 27, 2614–2623. [Google Scholar] [CrossRef] [PubMed]
- Nitzsche, F.; Müller, C.; Lukomska, B.; Jolkkonen, J.; Deten, A.; Boltze, J. Concise review: MSC adhesion cascade-insights into homing and transendothelial migration. Stem Cells 2017, 35, 1446–1460. [Google Scholar] [CrossRef] [PubMed]
- Studeny, M.; Marini, F.C.; Champlin, R.E.; Zompetta, C.; Fidler, I.J.; Andreeff, M. Bone marrow-derived mesenchymal stem cells as vehicles for interferon-beta delivery into tumors. Cancer Res. 2002, 62, 3603–3608. [Google Scholar]
- Studeny, M.; Marini, F.C.; Dembinski, J.L.; Zompetta, C.; Cabreira-Hansen, M.; Bekele, B.N.; Champlin, R.E.; Andreeff, M. Mesenchymal stem cells: Potential precursors for tumor stroma and targeted-delivery vehicles for anticancer agents. J. Natl. Cancer Inst. 2004, 96, 1593–1603. [Google Scholar] [CrossRef]
- Sadhukha, T.; O’Brien, T.D.; Prabha, S. Nano-engineered mesenchymal stem cells as targeted therapeutic carriers. J. Control. Release. 2014, 196, 243–251. [Google Scholar] [CrossRef]
- Zhao, Y.; Tang, S.; Guo, J.; Alahdal, M.; Cao, S.; Yang, Z.; Zhang, F.; Shen, Y.; Sun, M.; Mo, R.; et al. Targeted delivery of doxorubicin by nano-loaded mesenchymal stem cells for lung melanoma metastases therapy. Sci. Rep. 2017, 7, 44758. [Google Scholar] [CrossRef]
- Huang, W.C.; Lu, I.L.; Chiang, W.H.; Lin, Y.W.; Tsai, Y.C.; Chen, H.H.; Chang, C.W.; Chiang, C.S.; Chiu, H.C. Tumortropic adipose-derived stem cells carrying smart nanotherapeutics for targeted delivery and dual-modality therapy of orthotopic glioblastoma. J. Control. Release. 2017, 254, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Nold, P.; Hartmann, R.; Feliu, N.; Kantner, K.; Gamal, M.; Pelaz, B.; Hühn, J.; Sun, X.; Jungebluth, P.; Del Pino, P.; et al. Optimizing conditions for labeling of mesenchymal stromal cells (MSCs) with gold nanoparticles: A prerequisite for in vivo tracking of MSCs. J. Nanobiotechnology 2017, 15, 24. [Google Scholar] [CrossRef] [PubMed]
- Agnihotri, S.A.; Mallikarjuna, N.N.; Aminabhavi, T.M. Recent advances on chitosan-based micro- and nanoparticles in drug delivery. J. Control. Release. 2004, 100, 5–28. [Google Scholar] [CrossRef] [PubMed]
- Chang, P.V.; Prescher, J.A.; Sletten, E.M.; Baskin, J.M.; Miller, I.A.; Agard, N.J.; Lo, A.; Bertozzi, C.R. Copper-free click chemistry in living animals. Proc. Natl. Acad. Sci. USA 2010, 107, 1821–1826. [Google Scholar] [CrossRef] [PubMed]
- Jain, V.; Jain, S.; Mahajan, S.C. Nanomedicines based drug delivery systems for anti-cancer targeting and treatment. Curr. Drug Deliv. 2015, 12, 177–191. [Google Scholar] [CrossRef] [PubMed]
- England, C.G.; Hernandez, R.; Eddine, S.B.; Cai, W. Molecular imaging of pancreatic cancer with antibodies. Mol. Pharm. 2016, 13, 8–24. [Google Scholar] [CrossRef]
- Carter, P. Improving the efficacy of antibody-based cancer therapies. Nat. Rev. Cancer 2001, 1, 118–129. [Google Scholar] [CrossRef]
- Guo, J.; Luan, X.; Cong, Z.; Sun, Y.; Wang, L.; McKenna, S.L.; Cahill, M.R.; O’Driscoll, C.M. The potential for clinical translation of antibody-targeted nanoparticles in the treatment of acute myeloid leukaemia. J. Control. Release. 2018, 286, 154–166. [Google Scholar] [CrossRef]
- Matsumura, Y.; Maeda, H. A new concept for macromolecular therapeutics in cancer chemotherapy: Mechanism of tumoritropic accumulation of proteins and the antitumor agent smancs. Cancer Res. 1986, 46, 6387–6392. [Google Scholar]
- Yasunaga, M.; Manabe, S.; Tsuji, A.; Furuta, M.; Ogata, K.; Koga, Y.; Saga, T.; Matsumura, Y. Development of antibody-drug conjugates using DDS and molecular imaging. Bioengineering 2017, 4, 78. [Google Scholar] [CrossRef]
- Loganzo, F.; Sung, M.; Gerber, H.P. Mechanisms of resistance to antibody-drug conjugates. Mol. Cancer Ther. 2016, 15, 2825–2834. [Google Scholar] [CrossRef] [PubMed]
- Rong, J.; Han, J.; Dong, L.; Tan, Y.; Yang, H.; Feng, L.; Wang, Q.W.; Meng, R.; Zhao, J.; Wang, S.Q.; et al. Glycan imaging in intact rat hearts and glycoproteomic analysis reveal the upregulation of sialylation during cardiac hypertrophy. J. Am. Chem. Soc. 2014, 136, 17468–17476. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Koo, H.; Na, J.H.; Han, S.; Min, H.S.; Lee, S.J.; Kim, S.H.; Yun, S.H.; Jeong, S.Y.; Kwon, I.C.; et al. Chemical tumor-targeting of nanoparticles based on metabolic glycoengineering and click chemistry. ACS Nano 2014, 8, 2048–2063. [Google Scholar] [CrossRef] [PubMed]
- Shim, M.K.; Yoon, H.Y.; Ryu, J.H.; Koo, H.; Lee, S.; Park, J.H.; Kim, J.H.; Lee, S.; Pomper, M.G.; Kwon, I.C.; et al. Cathepsin B-specific metabolic precursor for in vivo tumor-specific fluorescence imaging. Angew. Chem. Int. Ed. Engl. 2016, 55, 14698–14703. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Gauthier, M.; Kelly, J.R.; Miller, R.J.; Xu, M.; O’Brien, W.D., Jr.; Cheng, J. Targeted ultrasound-assisted cancer-selective chemical labeling and subsequent cancer imaging using click chemistry. Angew. Chem. Int. Ed. Engl. 2016, 55, 5452–5456. [Google Scholar] [CrossRef]
- Lee, S.B.; Kim, H.L.; Jeong, H.J.; Lim, S.T.; Sohn, M.H.; Kim, D.W. Mesoporous silica nanoparticle pretargeting for PET imaging based on a rapid bioorthogonal reaction in a living body. Angew. Chem. Int. Ed. Engl. 2013, 52, 10549–10552. [Google Scholar] [CrossRef]
- Neves, A.A.; Stöckmann, H.; Wainman, Y.A.; Kuo, J.C.; Fawcett, S.; Leeper, F.J.; Brindle, K.M. Imaging cell surface glycosylation in vivo using "double click" chemistry. Bioconjug. Chem. 2013, 24, 934–941. [Google Scholar] [CrossRef] [PubMed]
- Rossin, R.; Verkerk, P.R.; van den Bosch, S.M.; Vulders, R.C.; Verel, I.; Lub, J.; Robillard, M.S. In vivo chemistry for pretargeted tumor imaging in live mice. Angew. Chem. Int. Ed. Engl. 2010, 49, 3375–3378. [Google Scholar] [CrossRef]
- Zeglis, B.M.; Sevak, K.K.; Reiner, T.; Mohindra, P.; Carlin, S.D.; Zanzonico, P.; Weissleder, R.; Lewis, J.S. A pretargeted PET imaging strategy based on bioorthogonal Diels-Alder click chemistry. J. Nucl. Med. 2013, 54, 1389–1396. [Google Scholar] [CrossRef] [PubMed]
- Keinänen, O.; Fung, K.; Pourat, J.; Jallinoja, V.; Vivier, D.; Pillarsetty, N.K.; Airaksinen, A.J.; Lewis, J.S.; Zeglis, B.M.; Sarparanta, M. Pretargeting of internalizing trastuzumab and cetuximab with a 18F-tetrazine tracer in xenograft models. EJNMMI Res. 2017, 7, 95. [Google Scholar] [CrossRef]
- Devaraj, N.K.; Thurber, G.M.; Keliher, E.J.; Marinelli, B.; Weissleder, R. Reactive polymer enables efficient in vivo bioorthogonal chemistry. Proc. Natl. Acad. Sci. USA 2012, 10, 4762–4767. [Google Scholar] [CrossRef] [PubMed]
- Rossin, R.; Läppchen, T.; van den Bosch, S.M.; Laforest, R.; Robillard, M.S. Diels-Alder reaction for tumor pretargeting: In vivo chemistry can boost tumor radiation dose compared with directly labeled antibody. J. Nucl. Med. 2013, 54, 1989–1995. [Google Scholar] [CrossRef] [PubMed]
- Rossin, R.; van den Bosch, S.M.; Ten Hoeve, W.; Carvelli, M.; Versteegen, R.M.; Lub, J.; Robillard, M.S. Highly reactive trans-cyclooctene tags with improved stability for Diels-Alder chemistry in living systems. Bioconjug. Chem. 2013, 24, 1210–1217. [Google Scholar] [CrossRef] [PubMed]
- Rossin, R.; van Duijnhoven, S.M.; Läppchen, T.; van den Bosch, S.M.; Robillard, M.S. Trans-cyclooctene tag with improved properties for tumor pretargeting with the Diels-Alder reaction. Mol. Pharm. 2014, 11, 3090–3096. [Google Scholar] [CrossRef]
- Nichols, B.; Qin, Z.; Yang, J.; Vera, D.R.; Devaraj, N.K. 68Ga chelating bioorthogonal tetrazine polymers for the multistep labeling of cancer biomarkers. Chem. Commun. 2014, 50, 5215–5217. [Google Scholar] [CrossRef] [PubMed]
- Zlitni, A.; Janzen, N.; Foster, F.S.; Valliant, J.F. Catching bubbles: Targeting ultrasound microbubbles using bioorthogonal inverse-electron-demand Diels-Alder reactions. Angew. Chem. Int. Ed. Engl. 2014, 53, 6459–6463. [Google Scholar] [CrossRef] [PubMed]
- Houghton, J.L.; Zeglis, B.M.; Abdel-Atti, D.; Sawada, R.; Scholz, W.W.; Lewis, J.S. Pretargeted immuno-PET of pancreatic cancer: Overcoming circulating antigen and internalized antibody to reduce radiation doses. J. Nucl. Med. 2016, 57, 453–459. [Google Scholar] [CrossRef]
- Houghton, J.L.; Membreno, R.; Abdel-Atti, D.; Cunanan, K.M.; Carlin, S.; Scholz, W.W.; Zanzonico, P.B.; Lewis, J.S.; Zeglis, B.M. Establishment of the in vivo efficacy of pretargeted radioimmunotherapy utilizing inverse electron demand Diels-Alder click chemistry. Mol. Cancer Ther. 2017, 16, 124–133. [Google Scholar] [CrossRef]
- Rondon, A.; Ty, N.; Bequignat, J.B.; Quintana, M.; Briat, A.; Witkowski, T.; Bouchon, B.; Boucheix, C.; Miot-Noirault, E.; Pouget, J.P.; et al. Antibody PEGylation in bioorthogonal pretargeting with trans-cyclooctene/tetrazine cycloaddition: In vitro and in vivo evaluation in colorectal cancer models. Sci. Rep. 2017, 7, 14918. [Google Scholar] [CrossRef]
- Emmetiere, F.; Irwin, C.; Viola-Villegas, N.T.; Longo, V.; Cheal, S.M.; Zanzonico, P.; Pillarsetty, N.; Weber, W.A.; Lewis, J.S.; Reiner, T. 18F-labeled-bioorthogonal liposomes for in vivo targeting. Bioconjug. Chem. 2013, 24, 1784–1789. [Google Scholar] [CrossRef]
- Rossin, R.; van Duijnhoven, S.M.; Ten Hoeve, W.; Janssen, H.M.; Kleijn, L.H.; Hoeben, F.J.; Versteegen, R.M.; Robillard, M.S. Triggered drug release from an antibody-drug conjugate using fast “click-to-release” chemistry in mice. Bioconjug. Chem. 2016, 27, 1697–1706. [Google Scholar] [CrossRef] [PubMed]
- Rossin, R.; Versteegen, R.M.; Wu, J.; Khasanov, A.; Wessels, H.J.; Steenbergen, E.J.; Ten Hoeve, W.; Janssen, H.M.; van Onzen, A.H.A.M.; Hudson, P.J.; et al. Chemically triggered drug release from an antibody-drug conjugate leads to potent antitumour activity in mice. Nat. Commun. 2018, 9, 1484. [Google Scholar] [CrossRef] [PubMed]
- Marabelle, A.; Kohrt, H.; Caux, C.; Levy, R. Intratumoral immunization: A new paradigm for cancer therapy. Clin. Cancer Res. 2014, 20, 1747–1756. [Google Scholar] [CrossRef] [PubMed]
- Chang, P.V.; Dube, D.H.; Sletten, E.M.; Bertozzi, C.R. A strategy for the selective imaging of glycans using caged metabolic precursors. J. Am. Chem. Soc. 2010, 132, 9516–9518. [Google Scholar] [CrossRef] [PubMed]
- Denmeade, S.R.; Nagy, A.; Gao, J.; Lilja, H.; Schally, A.V.; Isaacs, J.T. Enzymatic activation of a doxorubicin-peptide prodrug by prostate-specific antigen. Cancer Res. 1998, 58, 2537–2540. [Google Scholar] [PubMed]
- Aggarwal, N.; Sloane, B.F. Cathepsin B: Multiple roles in cancer. Proteom. Clin. Appl. 2014, 8, 427–437. [Google Scholar] [CrossRef]
- Xie, R.; Hong, S.; Feng, L.; Rong, J.; Chen, X. Cell-selective metabolic glycan labeling based on ligand-targeted liposomes. J. Am. Chem. Soc. 2012, 134, 9914–9917. [Google Scholar] [CrossRef] [PubMed]
- Low, P.S.; Henne, W.A.; Doorneweerd, D.D. Discovery and development of folic-acid-based receptor targeting for imaging and therapy of cancer and inflammatory diseases. Acc. Chem. Res. 2008, 41, 120–129. [Google Scholar] [CrossRef]
- Paszko, E.; Ehrhardt, C.; Senge, M.O.; Kelleher, D.P.; Reynolds, J.V. Nanodrug applications in photodynamic therapy. Photodiagnosis Photodyn. Ther. 2011, 8, 14–29. [Google Scholar] [CrossRef]
- Kamimura, M.; Omoto, A.; Chiu, H.-C.; Soga, K. Enhanced red upconversion emission of NaYF4: Yb3+, Er3+, Mn2+ nanoparticles for near-infrared-induced photodynamic therapy and fluorescence imaging. Chem. Lett. 2017, 46, 1076–1078. [Google Scholar] [CrossRef]
| Compound | Non-Toxic Concentration | Incubation Time | Cell Type | Reference (Year) |
|---|---|---|---|---|
| Ac4ManNAz | 5 μM | 3 days | B16 | [32] (2016) |
| 10 μM | 3 days | A549 | [27] (2017) | |
| 20 μM | 3 days | MSC (human) | [33] (2016) | |
| 50 μM | 3 days | NIH3T3 | [34] (2015) | |
| 50 μM | 1 day | ASC (human) | [35] (2017) | |
| 3 days | Jurkat T lymphocyte | [34] (2015) | ||
| 3 days | Chondrocyte (rabbit) | [29] (2016) | ||
| 7 days | MSC (human) | [33] (2016) | ||
| Ac3ManNAz | <5 μM | 2 days | Primary hippocampal neurons (rat) | [36] (2015) |
| 100 μM | 2 days | U87 | [37] (2017) | |
| BCN-CNP-Cy5 | 500 μg/mL | 1 day | ASC (human) | [35] (2017) |
| DBCO-650 | 50 μM | 1 h | Chondrocyte (rabbit) | [29] (2016) |
| DBCO-Cy5 | 20 μM | 1 h | ASC (human) | [31] (2016) |
| 100 μM | 48 h | A549 | [38] (2014) | |
| TCO-DBCO | 100 μM | 30 min | NIH3T3 | [34] (2015) |
| A549 | ||||
| Jurkat T lymphocyte | ||||
| Tz-DBCO | 100 μM | 30 min | NIH3T3 | [34] (2015) |
| A549 | ||||
| Jurkat T lymphocyte |
| Compound | Non-Toxic Dose | Administration Route | Animal Type | Reference |
|---|---|---|---|---|
| Ac4ManNAz | 300 mg/kg/day daily for 7 days | ip | Mouse | [39] (2004) |
| 10 mg/kg/day daily for 3 days | it | Mouse | [40] (2012) | |
| 40 mg/kg/day daily for 3 days | iv | Mouse | [41] (2017) | |
| 10 nmol | Oral | Mouse | [42] (2018) | |
| Ac4ManNAz-LP | 40 mg/kg/day of Ac4ManNAz daily for 4 days | iv | Mouse | [43] (2017) |
| BCN-Lipo | 10 mg/kg | iv | Mouse | [37] (2017) |
| DBCO-Cy5 | 5 nmol | iv | Mouse | [38] (2014) |
| DBCO-Cy5.5 | 5 μg (4.25 nmol) | iv | Mouse | [33] (2016) |
| DBCO-lipo | 10 mg/kg | iv | Mouse | [40] (2012) |
| DBCO modified polymeric nanoparticles | 100 μg/body | iv | Mouse | [33] (2016) |
| DBCO-ZnPc-LP | 2.5 mg/kg | iv | Mouse | [43] (2017) |
| 3,6-dimethyl-1,2,4,5-Tz | 6 mg/kg | iv | Mouse | [44] (2016) |
| Tz-Cy5.5 | 8 nmol | iv | Mouse | [45] (2017) |
| GEBP11-TCO | 4 nmol | iv | Mouse | [45] (2017) |
| Reaction Type | Azide Sugar | Probe | Imaging | Cell Type | Transplantation Route | Type of Animals | Reference |
|---|---|---|---|---|---|---|---|
| SPAAC | Ac4ManNAz | DBCO-Cy5 (administration after cell transplantation) | FL | A549 (human) | Injection into liver | Nude mice | [38] (2014) |
| SPAAC | Ac4ManNAz | DBCO-650 | FL | Chondrocyte (rabbit) | sc | Nude mice | [29] (2016) |
| SPAAC | Ac4ManNAz | DBCO-Cy5.5 | FL | MSC (human) | iv and ip | Nude mice | [33] (2016) |
| SPAAC | Ac4ManNAz | DBCO-Cy5 | FL | ASC (human) | Injection into hindlimb | Nude mice | [31] (2016) |
| SPAAC | Ac4ManNAz | BCN-CNPs | FL, MR, CT | MSC (human) | sc | Nude mice | [35] (2017) |
| Reaction Type | Tumor Labeling | Targeting Agent | Tumor Model | Purpose | Reference | |||
|---|---|---|---|---|---|---|---|---|
| Labeling Method | Labeling Agent | Administration Route | Targeting Agent | Administration Route | ||||
| SPAAC | MGE | Ac4ManNAz | it | DBCO-liposome | iv | A549 subcutaneous tumor | Imaging | [40] (2012) |
| SPAAC | MGE | RR-S-Ac3ManNAz | it | DBCO-Cy5.5 | iv | HT-29 subcutaneous tumor | Imaging | [84] (2016) |
| SPAAC | MGE | DCL-AAM | iv | DBCO-Cy5 | iv | LS174T subcutaneous tumor | Imaging | [41] (2017) |
| DBCO-VC-Dox | LS174T and MDA-MB-231 subcutaneous tumor and4T1 lung metastatic tumor | Therapy | ||||||
| SPAAC | MGE | Ac4ManNAz-loaded CNPs | iv | BCN-Ce6-CNPs | iv | A549 subcutaneous tumor | Imaging and therapy | [83] (2014) |
| SPAAC | MGE | Ac4ManNAz-liposome | iv | DBCO-ZnPc-liposome | iv | A549 subcutaneous tumor | Imaging andtherapy | [43] (2017) |
| SPAAC | MGE | Nano-MPs | iv | Cy5.5 labeled BCN-liposome | iv | U87 and MCF7 subcutaneous tumor | Imaging | [37] (2017) |
| SPAAC | MGE | Ac4ManNAz loaded microbubble | iv | DBCO-Cy5 | iv | 4T1 subcutaneous tumor | Imaging | [85] (2016) |
| SPAAC | Azide-labeling cells | Azide-labeling MSCs | it | NIR-dye containing DBCO-NPs | iv | A549-Luc subcutaneous tumor | Imaging | [33] (2016) |
| ip | Paclitaxel-loaded DBCO-NPs | ip | MA148 metastatic ovarian tumor | Therapy | ||||
| SPAAC | Nanoparticles | DBCO-PEG-MSNs | iv | 18F-azide | iv | U87 MG subcutaneous tumor | Imaging | [86] (2013) |
| SPAAC, Staudinger ligation and iEDDA | MGE | Ac4GalNAz | ip | Tz-DyLight | iv | LL2 subcutaneous tumor | Imaging | [87] (2013) |
| TMDIBO-TCO and PHOS-TCO | iv | |||||||
| iEDDA | Antigen-antibody reaction | CD49-TCO | iv | 111 In-Tz | iv | Colon carcinoma xenograft | Imaging | [88] (2010) |
| iEDDA | Antigen-antibody reaction | A33-TCO | iv | 64Cu-Tz-Bn-NOTA | iv | SW1222 subcutaneous tumor | Imaging | [89] (2013) |
| iEDDA | Antigen-antibody reaction | mAb-TCO | iv | 18F-Tz | iv | A431 and BT-474 subcutaneous tumor | Imaging | [90] (2017) |
| iEDDA | Antigen-antibody reaction | A33-TCO | iv | Polymer-modified Tz | iv | LS174 subcutaneous tumor | Imaging | [91] (2012) |
| iEDDA | Antigen-antibody reaction | CC49-TCO | iv | 177Lu-Tz | iv | LS174T subcutaneous tumor | Imaging | [92] (2013) |
| iEDDA | Antigen-antibody reaction | CC49-TCO | iv | 111In-Tz | iv | LS174T subcutaneous tumor | Imaging | [93] (2013) |
| iEDDA | Antigen-antibody reaction | TCO-oxymethylacetamide-tagged CC49 | iv | 111In-Tz | iv | LS174T subcutaneous tumor | Imaging | [94] (2014) |
| iEDDA | Antigen-antibody reaction | A33-TCO | iv | 68Ga-Tz-DTPA dextran | iv | LS174T subcutaneous tumor | Imaging | [95] (2014) |
| iEDDA | Antigen-antibody reaction | TCO-antiVEGFR2 | iv | MBTZ | iv | SKOV-3 subcutaneous tumor | Imaging | [96] (2014) |
| iEDDA | Antigen-antibody reaction | 5B1-TCO | iv | 64Cu-NOTA-PEG7-TCO | iv | Capan-2 and BxPC3 subcutaneous tumor | Imaging | [97] (2016) |
| iEDDA | Antigen-antibody reaction | 5B1-TCO | iv | 177Lu-DOTA-PEG7-Tz | iv | BxPC3 subcutaneous tumor | Therapy | [98] (2017) |
| 89Zr-Tz | Imaging | |||||||
| iEDDA | Antigen-antibody reaction | Ts29.2-PEG-TCO | iv | Tz-Cy5 | iv | HT29 subcutaneous tumor | Imaging | [99] (2017) |
| 35A7-PEG-TCO | iv | Tz-Cy5 | ip | A431-CEA-Luc peritoneal carcinomatosis tumor | ||||
| iEDDA | Membrane inserted peptide | pHLIP-Tz | it | 18F-TCO-liposome | iv | SKOV-3 subcutaneous tumor | Imaging | [100] (2013) |
| iEDDA | Aptamer (peptide) | GEBP11-TCO | iv | Cy5.5-Tz | iv | SGC-7901 subcutaneous tumor | Imaging | [45] (2017) |
| iEDDA | Antigen-antibody reaction | CC49-TCO-DOX | iv | Radio-labeled Tz | iv and ip | LS174T subcutaneous tumor | Therapy | [101] (2016) |
| iEDDA | Antigen-antibody reaction | TAG72-TCO-MMAE | iv | Tz-DOTA | iv | LS174T and OVCAR 3 subcutaneous tumor | Therapy | [102] (2018) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Takayama, Y.; Kusamori, K.; Nishikawa, M. Click Chemistry as a Tool for Cell Engineering and Drug Delivery. Molecules 2019, 24, 172. https://doi.org/10.3390/molecules24010172
Takayama Y, Kusamori K, Nishikawa M. Click Chemistry as a Tool for Cell Engineering and Drug Delivery. Molecules. 2019; 24(1):172. https://doi.org/10.3390/molecules24010172
Chicago/Turabian StyleTakayama, Yukiya, Kosuke Kusamori, and Makiya Nishikawa. 2019. "Click Chemistry as a Tool for Cell Engineering and Drug Delivery" Molecules 24, no. 1: 172. https://doi.org/10.3390/molecules24010172
APA StyleTakayama, Y., Kusamori, K., & Nishikawa, M. (2019). Click Chemistry as a Tool for Cell Engineering and Drug Delivery. Molecules, 24(1), 172. https://doi.org/10.3390/molecules24010172




