Comparative Proteomic Analysis of Rana chensinensis Oviduct
Abstract
1. Introduction
2. Results
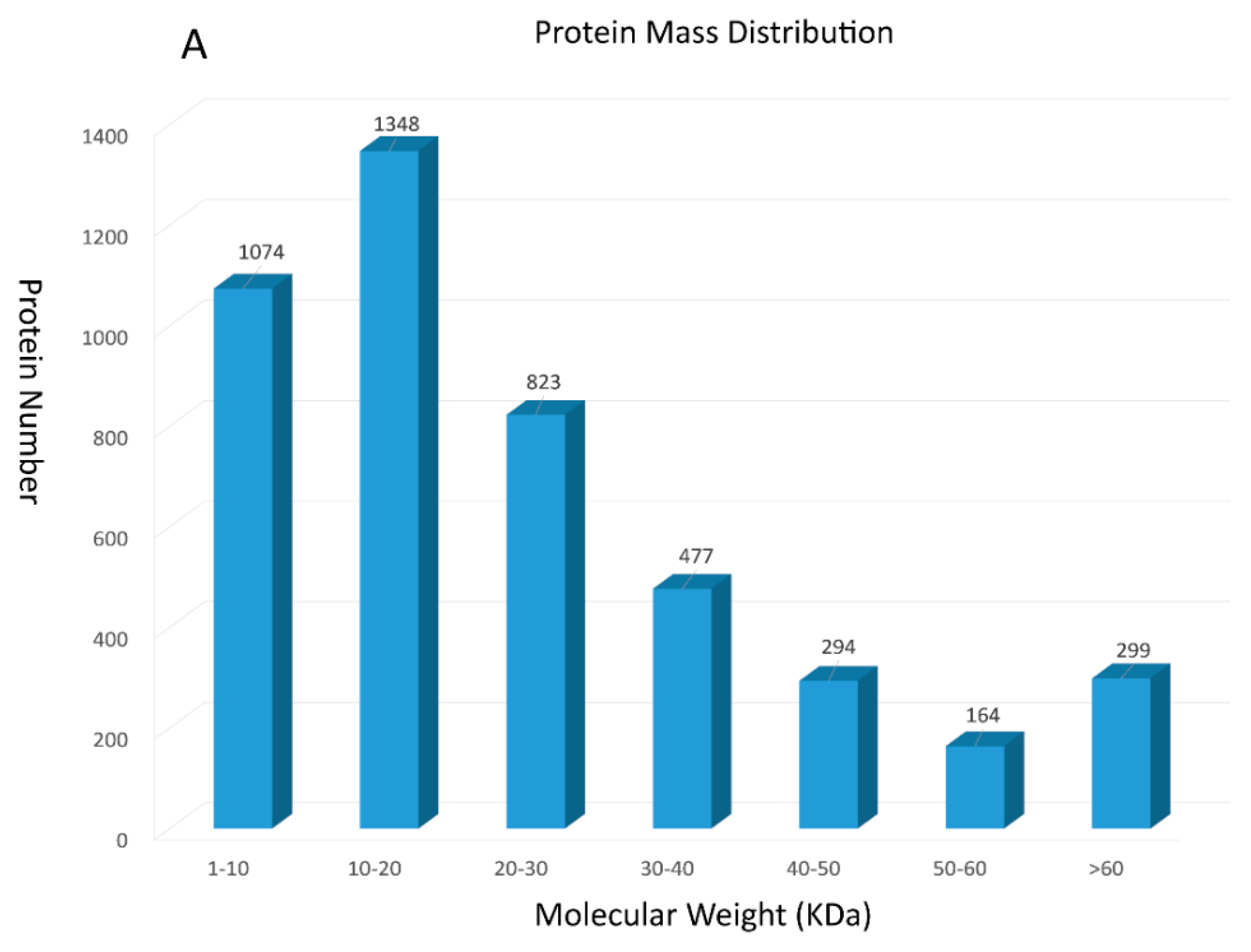
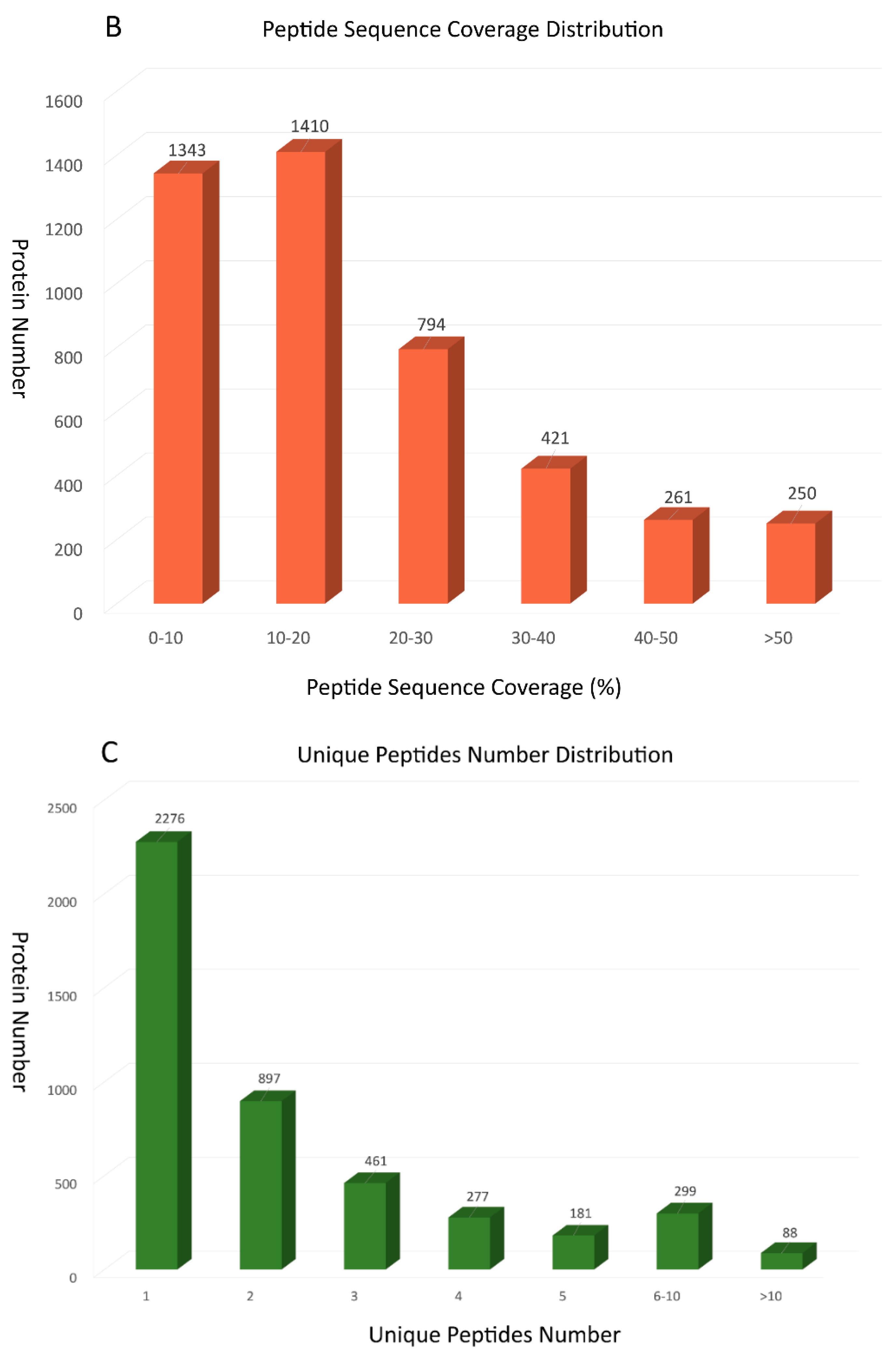
2.1. Protein Identified through Itraq Technology
2.2. Differentially Expressed Proteins
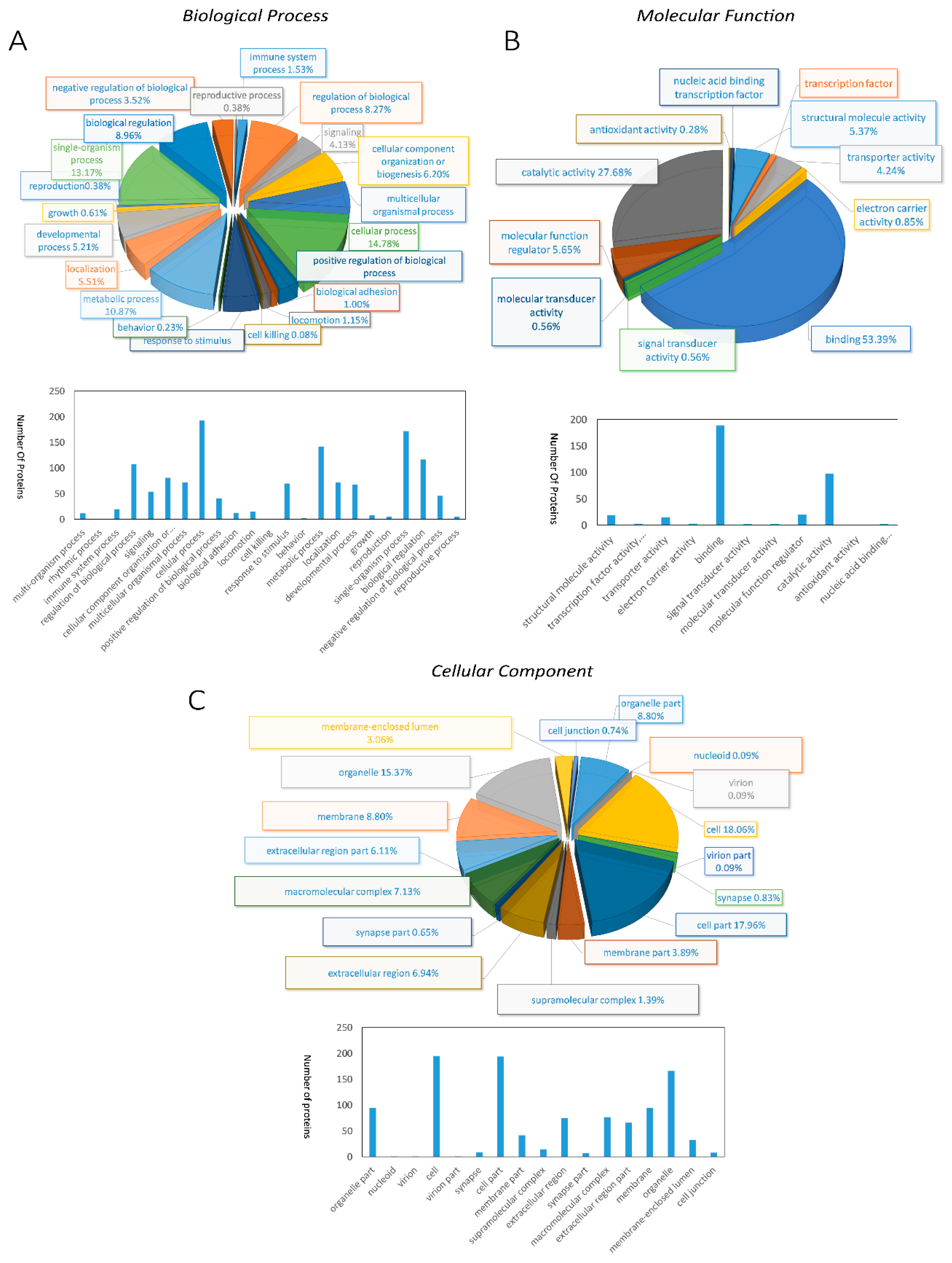
2.3. Gene Ontology Annotation and Classification Profiles
2.4. KEGG Pathway Annotation and Reasonable Enrichment Analysis
3. Discussion
3.1. Changes in Proteins Associated with Focal Adhesion and Extracellular Martix
3.2. Enzymes Involved in Energy Metabolism Pathway
3.3. Enzymes Changed in Pyruvate Metabolism Pathway
3.4. Down-Regulation of Lipid Metabolism during Breeding Period
3.5. Expression of 5-Aminolevulinate Synthase (ALAS2) in Heme Biosynthesis
4. Materials and Methods
4.1. Animals and Treatment Procedure
4.2. Protein Extraction, Digestion, and iTRAQ Labeling
4.3. Peptide Fractionation with Strong Cation Exchange (SCX) Chromatography
4.4. LC-MS/MS Analysis
4.5. Protein Identification and Data Analysis
4.6. Bioinformatics Analysis of Differentially Expressed Proteins
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Liu, Y.; Weng, J.; Huang, S.; Shen, Y.; Sheng, X.; Han, Y.; Xu, M.; Weng, Q. Immunoreactivities of PPARγ2, leptin and leptin receptor in oviduct of Chinese brown frog during breeding period and pre-hibernation. Eur. J. Histochem. 2014, 58. [Google Scholar] [CrossRef] [PubMed]
- Gabler, C.; Killian, G.J.; Einspanier, R. Differential expression of extracellular matrix components in the bovine oviduct during the oestrous cycle. Reprod. Camb. Engl. 2001, 122, 121–130. [Google Scholar] [CrossRef]
- Mondéjar, I.; Acuña, O.; Izquierdo-Rico, M.; Coy, P.; Avilés, M. The Oviduct: Functional Genomic and Proteomic Approach: Transcriptome and Proteome of the Oviduct. Reprod. Domest. Anim. 2012, 47, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Weng, J.; Liu, Y.; Xu, Y.; Hu, R.; Zhang, H.; Sheng, X.; Watanabe, G.; Taya, K.; Weng, Q.; Xu, M. Expression of P450arom and Estrogen Receptor Alpha in the Oviduct of Chinese Brown Frog (Rana dybowskii) during Prehibernation. Int. J. Endocrinol. 2015, 2015, 1–9. [Google Scholar] [CrossRef] [PubMed]
- The Pharmacopoeia Commission of PRC. Pharmacopoeia of the People’s Republic of China; The Pharmacopoeia Commission of PRC: Beijing, China, 2000; pp. 255–256. [Google Scholar]
- Shen, Y.; Liu, Y.; Ma, J.; Ma, X.; Tian, Y.; Zhang, H.; Li, L.; Xu, M.; Weng, Q.; Watanabe, G.; et al. Immunoreactivity of c-kit receptor protein during the prehibernation period in the oviduct of the Chinese brown frog, Rana chensinensis. J. Vet. Med. Sci. 2012, 74, 209–213. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.; Yang, L.; Wang, C.; Ma, S.; Cui, L.; Huang, S.; Sheng, X.; Weng, Q.; Xu, M. Immunostimulatory Activity of Protein Hydrolysate from Oviductus Ranae on Macrophage In Vitro. Evid. Based Complement. Altern. Med. 2014, 2014, 1–11. [Google Scholar] [CrossRef]
- Wright, P.C.; Noirel, J.; Ow, S.-Y.; Fazeli, A. A review of current proteomics technologies with a survey on their widespread use in reproductive biology investigations. Theriogenology 2012, 77, 738–765. [Google Scholar] [CrossRef] [PubMed]
- Zieske, L.R. A perspective on the use of iTRAQ reagent technology for protein complex and profiling studies. J. Exp. Bot. 2006, 57, 1501–1508. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.W.; Wang, G.; Baek, S.J.; Shen, R.-F. Comparative Study of Three Proteomic Quantitative Methods, DIGE, cICAT, and iTRAQ, Using 2D Gel- or LC−MALDI TOF/TOF. J. Proteome Res. 2006, 5, 651–658. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Mao, X.; Huang, J.; Ding, Y.; Wu, J.; Dong, S.; Kong, L.; Gao, G.; Li, C.-Y.; Wei, L. KOBAS 2.0: A web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 2011, 39, W316–W322. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Mao, X.; Cai, T.; Luo, J.; Wei, L. KOBAS server: A web-based platform for automated annotation and pathway identification. Nucleic Acids Res. 2006, 34, W720–W724. [Google Scholar] [CrossRef] [PubMed]
- Hynes, R.O.; Naba, A. Overview of the Matrisome—An Inventory of Extracellular Matrix Constituents and Functions. Cold Spring Harb. Perspect. Biol. 2012, 4, a004903. [Google Scholar] [CrossRef] [PubMed]
- Rozario, T.; DeSimone, D.W. The extracellular matrix in development and morphogenesis: A dynamic view. Dev. Biol. 2010, 341, 126–140. [Google Scholar] [CrossRef] [PubMed]
- Mouw, J.K.; Ou, G.; Weaver, V.M. Extracellular matrix assembly: A multiscale deconstruction. Nat. Rev. Mol. Cell Biol. 2014, 15, 771–785. [Google Scholar] [CrossRef] [PubMed]
- Kühn, K. Basement membrane (type IV) collagen. Matrix Biol. 1995, 14, 439–445. [Google Scholar] [CrossRef]
- Lamy, J.; Labas, V.; Harichaux, G.; Tsikis, G.; Mermillod, P.; Saint-Dizier, M. Regulation of the bovine oviductal fluid proteome. Reproduction 2016, 152, 629–644. [Google Scholar] [CrossRef] [PubMed]
- Kelemen-Valkony, I.; Kiss, M.; Csiha, J.; Kiss, A.; Bircher, U.; Szidonya, J.; Maróy, P.; Juhász, G.; Komonyi, O.; Csiszár, K.; et al. Drosophila basement membrane collagen col4a1 mutations cause severe myopathy. Matrix Biol. 2012, 31, 29–37. [Google Scholar] [CrossRef] [PubMed]
- Tzu, J.; Marinkovich, M.P. Bridging structure with function: Structural, regulatory, and developmental role of laminins. Int. J. Biochem. Cell Biol. 2008, 40, 199–214. [Google Scholar] [CrossRef] [PubMed]
- Koch, M.; Olson, P.F.; Albus, A.; Jin, W.; Hunter, D.D.; Brunken, W.J.; Burgeson, R.E.; Champliaud, M.-F. Characterization and Expression of the Laminin γ3 Chain: A Novel, Non-Basement Membrane–associated, Laminin Chain. J. Cell Biol. 1999, 145, 605–618. [Google Scholar] [CrossRef] [PubMed]
- Saotome, K.; Isomura, T.; Seki, T.; Nakamura, Y.; Nakamura, M. Structural changes in gonadal basement membranes during sex differentiation in the frog Rana rugosa. J. Exp. Zool. Part Ecol. Genet. Physiol. 2010, 313A, 369–380. [Google Scholar] [CrossRef] [PubMed]
- Madekurozwa, M.-C. Immunolocalization of Intermediate Filaments and Laminin in the Oviduct of the Immature and Mature Japanese Quail (Coturnix coturnix japonica). Anat. Histol. Embryol. 2014, 43, 210–220. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.S.; Alonso, J.L.; Ostuni, E.; Whitesides, G.M.; Ingber, D.E. Cell shape provides global control of focal adhesion assembly. Biochem. Biophys. Res. Commun. 2003, 307, 355–361. [Google Scholar] [CrossRef]
- Kapelnikov, A.; Zelinger, E.; Gottlieb, Y.; Rhrissorrakrai, K.; Gunsalus, K.C.; Heifetz, Y. Mating induces an immune response and developmental switch in the Drosophila oviduct. Proc. Natl. Acad. Sci. USA 2008, 105, 13912–13917. [Google Scholar] [CrossRef] [PubMed]
- Vigoreaux, J.O.; Saide, J.D.; Pardue, M.L. Structurally differentDrosophila striated muscles utilize distinct variants of Z-band-associated proteins. J. Muscle Res. Cell Motil. 1991, 12, 340–354. [Google Scholar] [CrossRef] [PubMed]
- Soundar, S.; Park, J.-H.; Huh, T.-L.; Colman, R.F. Evaluation by Mutagenesis of the Importance of 3 Arginines in α, β, and γ Subunits of Human NAD-dependent Isocitrate Dehydrogenase. J. Biol. Chem. 2003, 278, 52146–52153. [Google Scholar] [CrossRef] [PubMed]
- Bzymek, K.P.; Colman, R.F. Role of α-Asp 181, β-Asp 192, and γ-Asp 190 in the Distinctive Subunits of Human NAD-Specific Isocitrate Dehydrogenase. Biochemistry 2007, 46, 5391–5397. [Google Scholar] [CrossRef] [PubMed]
- Zeng, L.; Morinibu, A.; Kobayashi, M.; Zhu, Y.; Wang, X.; Goto, Y.; Yeom, C.J.; Zhao, T.; Hirota, K.; Shinomiya, K.; et al. Aberrant IDH3α expression promotes malignant tumor growth by inducing HIF-1-mediated metabolic reprogramming and angiogenesis. Oncogene 2015, 34, 4758–4766. [Google Scholar] [CrossRef] [PubMed]
- Soundar, S.; O’Hagan, M.; Fomulu, K.S.; Colman, R.F. Identification of Mn2+-binding Aspartates from α, β, and γ Subunits of Human NAD-dependent Isocitrate Dehydrogenase. J. Biol. Chem. 2006, 281, 21073–21081. [Google Scholar] [CrossRef] [PubMed]
- Dumollard, R.; Campbell, K.; Halet, G.; Carroll, J.; Swann, K. Regulation of cytosolic and mitochondrial ATP levels in mouse eggs and zygotes. Dev. Biol. 2008, 316, 431–440. [Google Scholar] [CrossRef] [PubMed]
- Soleilhavoup, C.; Riou, C.; Tsikis, G.; Labas, V.; Harichaux, G.; Kohnke, P.; Reynaud, K.; de Graaf, S.P.; Gerard, N.; Druart, X. Proteomes of the Female Genital Tract During the Oestrous Cycle. Mol. Cell. Proteom. 2016, 15, 93–108. [Google Scholar] [CrossRef] [PubMed]
- Hugentobler, S.A.; Humpherson, P.G.; Leese, H.J.; Sreenan, J.M.; Morris, D.G. Energy substrates in bovine oviduct and uterine fluid and blood plasma during the oestrous cycle. Mol. Reprod. Dev. 2008, 75, 496–503. [Google Scholar] [CrossRef] [PubMed]
- Clarke, C.F.; Williams, W.; Teruya, J.H. Ubiquinone biosynthesis in Saccharomyces cerevisiae. Isolation and sequence of COQ3, the 3,4-dihydroxy-5-hexaprenylbenzoate methyltransferase gene. J. Biol. Chem. 1991, 266, 16636–16644. [Google Scholar] [PubMed]
- Dumollard, R.; Ward, Z.; Carroll, J.; Duchen, M.R. Regulation of redox metabolism in the mouse oocyte and embryo. Dev. Camb. Engl. 2007, 134, 455–465. [Google Scholar] [CrossRef] [PubMed]
- Ben-Meir, A.; Burstein, E.; Borrego-Alvarez, A.; Chong, J.; Wong, E.; Yavorska, T.; Naranian, T.; Chi, M.; Wang, Y.; Bentov, Y.; et al. Coenzyme Q10 restores oocyte mitochondrial function and fertility during reproductive aging. Aging Cell 2015, 14, 887–895. [Google Scholar] [CrossRef] [PubMed]
- Wyss, M.; Kaddurah-Daouk, R. Creatine and Creatinine Metabolism. Physiol. Rev. 2000, 80, 1107–1213. [Google Scholar] [CrossRef] [PubMed]
- Ellery, S.J.; LaRosa, D.A.; Kett, M.M.; Della Gatta, P.A.; Snow, R.J.; Walker, D.W.; Dickinson, H. Maternal creatine homeostasis is altered during gestation in the spiny mouse: Is this a metabolic adaptation to pregnancy? BMC Pregnancy Childbirth 2015, 15. [Google Scholar] [CrossRef] [PubMed]
- Ginger, M.L.; Ngazoa, E.S.; Pereira, C.A.; Pullen, T.J.; Kabiri, M.; Becker, K.; Gull, K.; Steverding, D. Intracellular Positioning of Isoforms Explains an Unusually Large Adenylate Kinase Gene Family in the Parasite Trypanosoma brucei. J. Biol. Chem. 2005, 280, 11781–11789. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Gonzalez, A.; Kourembanas, S.; Wyatt, T.A.; Mitsialis, S.A. Mutation of Murine Adenylate Kinase 7 Underlies a Primary Ciliary Dyskinesia Phenotype. Am. J. Respir. Cell Mol. Biol. 2009, 40, 305–313. [Google Scholar] [CrossRef] [PubMed]
- Acuña, O.S.; Avilés, M.; López-Úbeda, R.; Guillén-Martínez, A.; Soriano-Úbeda, C.; Torrecillas, A.; Coy, P.; Izquierdo-Rico, M.J. Differential gene expression in porcine oviduct during the oestrous cycle. Reprod. Fertil. Dev. 2017, 29, 2387. [Google Scholar] [CrossRef] [PubMed]
- Warren, D.E. The role of mineralized tissue in the buffering of lactic acid during anoxia and exercise in the leopard frog Rana pipiens. J. Exp. Biol. 2005, 208, 1117–1124. [Google Scholar] [CrossRef] [PubMed]
- Donohoe, P.H.; Boutilier, R.G. The use of extracellular lactate as an oxidative substrate in the oxygen-limited frog. Respir. Physiol. 1999, 116, 171–179. [Google Scholar] [CrossRef]
- Abboud, J.; Storey, K.B. Novel control of lactate dehydrogenase from the freeze tolerant wood frog: Role of posttranslational modifications. PeerJ 2013, 1, e12. [Google Scholar] [CrossRef] [PubMed]
- Kiss, A.J.; Muir, T.J.; Lee, R.E., Jr.; Costanzo, J.P. Seasonal Variation in the Hepatoproteome of the Dehydration- and Freeze-Tolerant Wood Frog, Rana sylvatica. Int. J. Mol. Sci. 2011, 12, 8406–8414. [Google Scholar] [CrossRef] [PubMed]
- Costanzo, J.P.; do Amaral, M.C.F.; Rosendale, A.J.; Lee, R.E. Hibernation physiology, freezing adaptation and extreme freeze tolerance in a northern population of the wood frog. J. Exp. Biol. 2013, 216, 3461–3473. [Google Scholar] [CrossRef] [PubMed]
- Kumashiro, N.; Beddow, S.A.; Vatner, D.F.; Majumdar, S.K.; Cantley, J.L.; Guebre-Egziabher, F.; Fat, I.; Guigni, B.; Jurczak, M.J.; Birkenfeld, A.L.; et al. Targeting Pyruvate Carboxylase Reduces Gluconeogenesis and Adiposity and Improves Insulin Resistance. Diabetes 2013, 62, 2183–2194. [Google Scholar] [CrossRef] [PubMed]
- Ikeda, Y.; Yamamoto, J.; Okamura, M.; Fujino, T.; Takahashi, S.; Takeuchi, K.; Osborne, T.F.; Yamamoto, T.T.; Ito, S.; Sakai, J. Transcriptional Regulation of the Murine Acetyl-CoA Synthetase 1 Gene through Multiple Clustered Binding Sites for Sterol Regulatory Element-binding Proteins and a Single Neighboring Site for Sp1. J. Biol. Chem. 2001, 276, 34259–34269. [Google Scholar] [CrossRef] [PubMed]
- Luong, A.; Hannah, V.C.; Brown, M.S.; Goldstein, J.L. Molecular Characterization of Human Acetyl-CoA Synthetase, an Enzyme Regulated by Sterol Regulatory Element-binding Proteins. J. Biol. Chem. 2000, 275, 26458–26466. [Google Scholar] [CrossRef] [PubMed]
- Donkor, J.; Sariahmetoglu, M.; Dewald, J.; Brindley, D.N.; Reue, K. Three Mammalian Lipins Act as Phosphatidate Phosphatases with Distinct Tissue Expression Patterns. J. Biol. Chem. 2006, 282, 3450–3457. [Google Scholar] [CrossRef] [PubMed]
- Phan, J.; Péterfy, M.; Reue, K. Lipin Expression Preceding Peroxisome Proliferator-activated Receptor-γ Is Critical for Adipogenesis in Vivo and in Vitro. J. Biol. Chem. 2004, 279, 29558–29564. [Google Scholar] [CrossRef] [PubMed]
- Ramanadham, S.; Ali, T.; Ashley, J.W.; Bone, R.N.; Hancock, W.D.; Lei, X. Calcium-independent phospholipases A 2 and their roles in biological processes and diseases. J. Lipid Res. 2015, 56, 1643–1668. [Google Scholar] [CrossRef] [PubMed]
- Barbour, S.E.; Kapur, A.; Deal, C.L. Regulation of phosphatidylcholine homeostasis by calcium-independent phospholipase A2. Biochim. Biophys. Acta 1999, 1439, 77–88. [Google Scholar] [CrossRef]
- Song, Y.; Wilkins, P.; Hu, W.; Murthy, K.S.; Chen, J.; Lee, Z.; Oyesanya, R.; Wu, J.; Barbour, S.E.; Fang, X. Inhibition of calcium-independent phospholipase A2 suppresses proliferation and tumorigenicity of ovarian carcinoma cells. Biochem. J. 2007, 406, 427–436. [Google Scholar] [CrossRef] [PubMed]
- Chang, T.Y.; Chang, C.C.Y.; Cheng, D. Acyl-coenzyme A:cholesterol acyltransferase. Annu. Rev. Biochem. 1997, 66, 613–638. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-J.; Bian, Y.; Luo, J.; Lu, M.; Xiong, Y.; Guo, S.-Y.; Yin, H.-Y.; Lin, X.; Li, Q.; Chang, C.C.Y.; et al. Cholesterol and fatty acids regulate cysteine ubiquitylation of ACAT2 through competitive oxidation. Nat. Cell Biol. 2017, 19, 808–819. [Google Scholar] [CrossRef] [PubMed]
- Boden, G. Obesity, insulin resistance and free fatty acids. Curr. Opin. Endocrinol. Diabetes Obes. 2011, 18, 139–143. [Google Scholar] [CrossRef] [PubMed]
- Cable, E.E.; Miller, T.G.; Isom, H.C. Regulation of Heme Metabolism in Rat Hepatocytes and Hepatocyte Cell Lines: δ-Aminolevulinic Acid Synthase and Heme Oxygenase Are Regulated by Different Heme-Dependent Mechanisms. Arch. Biochem. Biophys. 2000, 384, 280–295. [Google Scholar] [CrossRef] [PubMed]
- Wingert, R.A.; Galloway, J.L.; Barut, B.; Foott, H.; Fraenkel, P.; Axe, J.L.; Weber, G.J.; Dooley, K.; Davidson, A.J.; Schmidt, B.; et al. Deficiency of glutaredoxin 5 reveals Fe–S clusters are required for vertebrate haem synthesis. Nature 2005, 436, 1035–1039. [Google Scholar] [CrossRef] [PubMed]
- Smith, S.J.; Cox, T.M. Translational control of erythroid delta-aminolevulinate synthase in immature human erythroid cells by heme. Cell. Mol. Biol. (Noisy-Le-Grand) 1997, 43, 103–114. [Google Scholar]
- Wiśniewski, J.R.; Zougman, A.; Nagaraj, N.; Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 2009, 6, 359–362. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.-L.; Zhou, Y.-H.; Chen, J.-Q.; Huang, Q.-Y.; Han, Q.; Liu, B.; Cheng, G.-D.; Li, Y.-H. Quantitative proteomic analysis of sub-MIC erythromycin inhibiting biofilm formation of S. suis in vitro. J. Proteom. 2015, 116, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Li, Y.; Yao, B.; Sun, M.; Wang, Z.; Zhao, Y. Transcriptome Sequencing and de novo Analysis for Oviductus Ranae of Rana chensinensis Using Illumina RNA-Seq Technology. J. Genet. Genom. 2013, 40, 137–140. [Google Scholar] [CrossRef] [PubMed]
- Koenig, T.; Menze, B.H.; Kirchner, M.; Monigatti, F.; Parker, K.C.; Patterson, T.; Steen, J.J.; Hamprecht, F.A.; Steen, H. Robust Prediction of the MASCOT Score for an Improved Quality Assessment in Mass Spectrometric Proteomics. J. Proteome Res. 2008, 7, 3708–3717. [Google Scholar] [CrossRef] [PubMed]
- Dong, M.; Gu, J.; Zhang, L.; Chen, P.; Liu, T.; Deng, J.; Lu, H.; Han, L.; Zhao, B. Comparative proteomics analysis of superior and inferior spikelets in hybrid rice during grain filling and response of inferior spikelets to drought stress using isobaric tags for relative and absolute quantification. J. Proteom. 2014, 109, 382–399. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.; Han, X.; Geng, C.; Zhao, Y.; Zhang, Z.; Qiu, F. Comparative proteomic analysis revealing the complex network associated with waterlogging stress in maize (Zea mays L.) seedling root cells. Proteomics 2015, 15, 135–147. [Google Scholar] [CrossRef] [PubMed]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene Ontology: Tool for the unification of biology. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Gotz, S.; Garcia-Gomez, J.M.; Terol, J.; Williams, T.D.; Nagaraj, S.H.; Nueda, M.J.; Robles, M.; Talon, M.; Dopazo, J.; Conesa, A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008, 36, 3420–3435. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of Rana chensinensis oviduct are available from the authors. |
| Pathway Terms 1 | DEPs with KEGG Annotation 2 | DEPs with KEGG Enrichment 3 |
|---|---|---|
| Pathways in cancer | 9 | 0 |
| RNA transport | 8 | 4 |
| Spliceosome | 8 | 0 |
| Focal adhesion | 7 | 4 |
| PI3K–Akt signaling | 7 | 0 |
| Huntington’s disease | 7 | 0 |
| Alzheimer’s disease | 6 | 0 |
| Metabolic pathways | 24 | 11 |
| ECM–receptor interaction | 6 | 3 |
| Amoebiasis | 6 | 0 |
| Organism 1 | Number of Proteins 2 | Organism | Number of Proteins |
|---|---|---|---|
| Xenopus tropicalis | 163 | Salmo salar | 2 |
| Xenopus laevis | 61 | Bos taurus | 2 |
| Rana catesbeiana | 13 | Oryctolagus cuniculus | 1 |
| Taeniopygia guttata | 11 | Mus musculus | 1 |
| Gallus gallus | 8 | Ovis aries | 1 |
| Monodelphis domestica | 7 | Zonotrichia albicollis | 1 |
| Homo sapiens | 5 | Pan troglodytes | 1 |
| Rana esculenta | 4 | Pyrrhocoris apterus | 1 |
| Mus musculus | 4 | Canis familiaris | 1 |
| Ailuropoda melanoleuca | 4 | Oryzias latipes | 1 |
| Ornithorhynchus anatinus | 3 | Equus caballus | 1 |
| Danio rerio | 3 | Trepomonas agilis | 1 |
| Rattus norvegicus | 3 | Anoplopoma fimbria | 1 |
| Macaca mulatta | 2 | Helicoverpa zea | 1 |
| Bufo japonicus | 2 | Sus scrofa | 1 |
| Callithrix jacchus | 2 |
| Pathway Terms 1 | Input Number 2 | Background Number 3 | Gene Symbol of Input DEPs 4 | p-Value |
|---|---|---|---|---|
| Lysosome | 5 | 123 | acp2, lamp1, lgmn, hexb, LOC100496969 | 0.0049 |
| RNA transport | 4 | 140 | thoc6, eif3a, eif4a2, eif4b | 0.0292 |
| Glycosaminoglycan degradation | 2 | 18 | LOC100496969, hexb | 0.0292 |
| ECM–receptor interaction | 3 | 75 | lama2, lamb2, col4a1 | 0.0292 |
| Metabolic pathways | 11 | 1163 | idh3b, rpn2, coq3, pla2g6, lpin1, ak7, alas2, gamt, hexb, LOC100496969, acat2 | 0.0292 |
| Pyruvate metabolism | 3 | 43 | acss2.2.L, pc.1.L, ldha.S | 0.0338 |
| Focal adhesion | 4 | 191 | lama2, lamb2, col4a1, actn1 | 0.0451 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Su, H.; Zhang, H.; Wei, X.; Pan, D.; Jing, L.; Zhao, D.; Zhao, Y.; Qi, B. Comparative Proteomic Analysis of Rana chensinensis Oviduct. Molecules 2018, 23, 1384. https://doi.org/10.3390/molecules23061384
Su H, Zhang H, Wei X, Pan D, Jing L, Zhao D, Zhao Y, Qi B. Comparative Proteomic Analysis of Rana chensinensis Oviduct. Molecules. 2018; 23(6):1384. https://doi.org/10.3390/molecules23061384
Chicago/Turabian StyleSu, Hang, He Zhang, Xinghua Wei, Daian Pan, Li Jing, Daqing Zhao, Yu Zhao, and Bin Qi. 2018. "Comparative Proteomic Analysis of Rana chensinensis Oviduct" Molecules 23, no. 6: 1384. https://doi.org/10.3390/molecules23061384
APA StyleSu, H., Zhang, H., Wei, X., Pan, D., Jing, L., Zhao, D., Zhao, Y., & Qi, B. (2018). Comparative Proteomic Analysis of Rana chensinensis Oviduct. Molecules, 23(6), 1384. https://doi.org/10.3390/molecules23061384




