Effect of Red and Blue Light on Anthocyanin Accumulation and Differential Gene Expression in Strawberry (Fragaria × ananassa)
Abstract
1. Introduction
2. Results
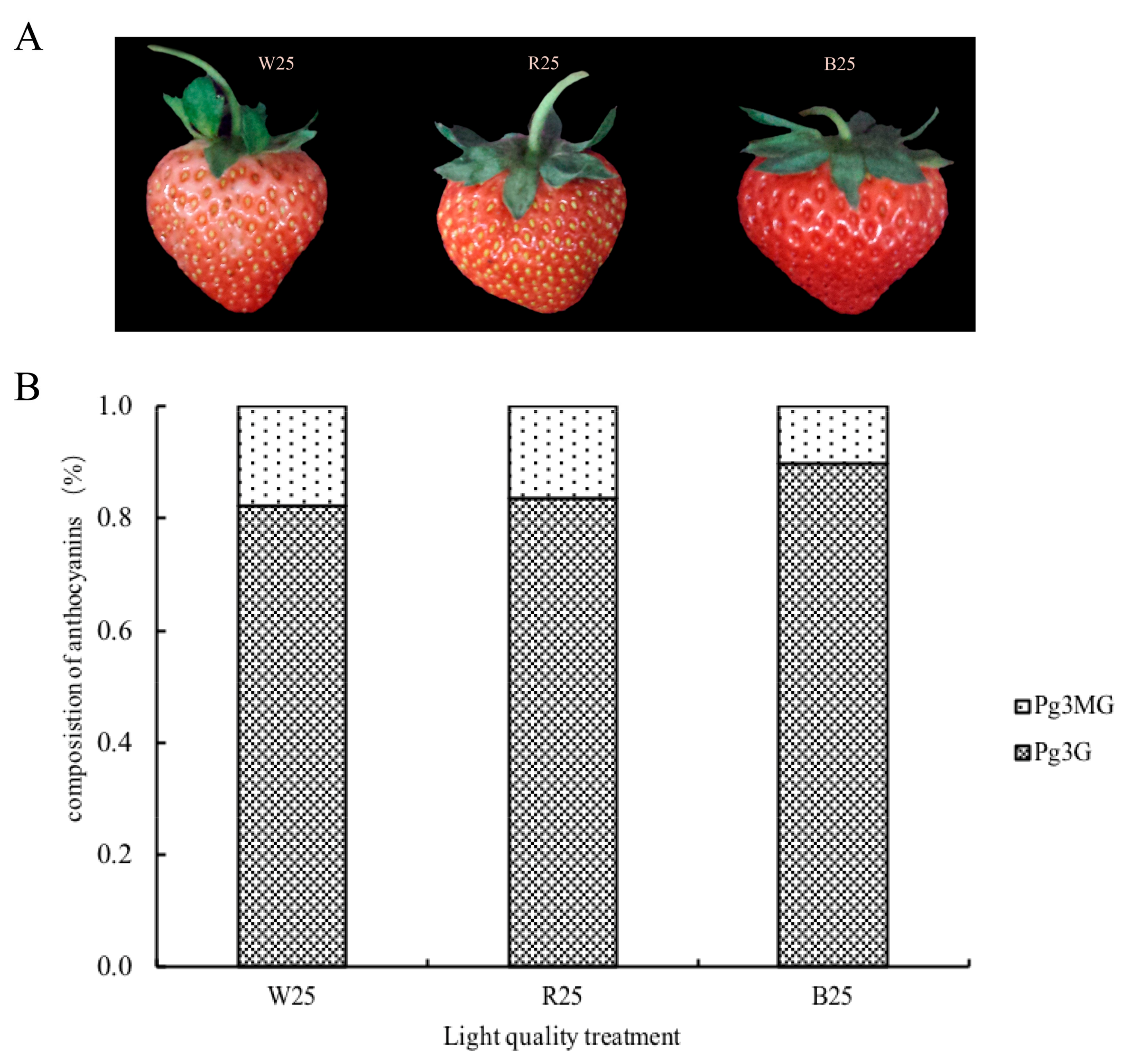
2.1. Anthocyanin Accumulation in Response to Different Light Quality
2.2. Transcriptome Sequencing and Gene Mapping
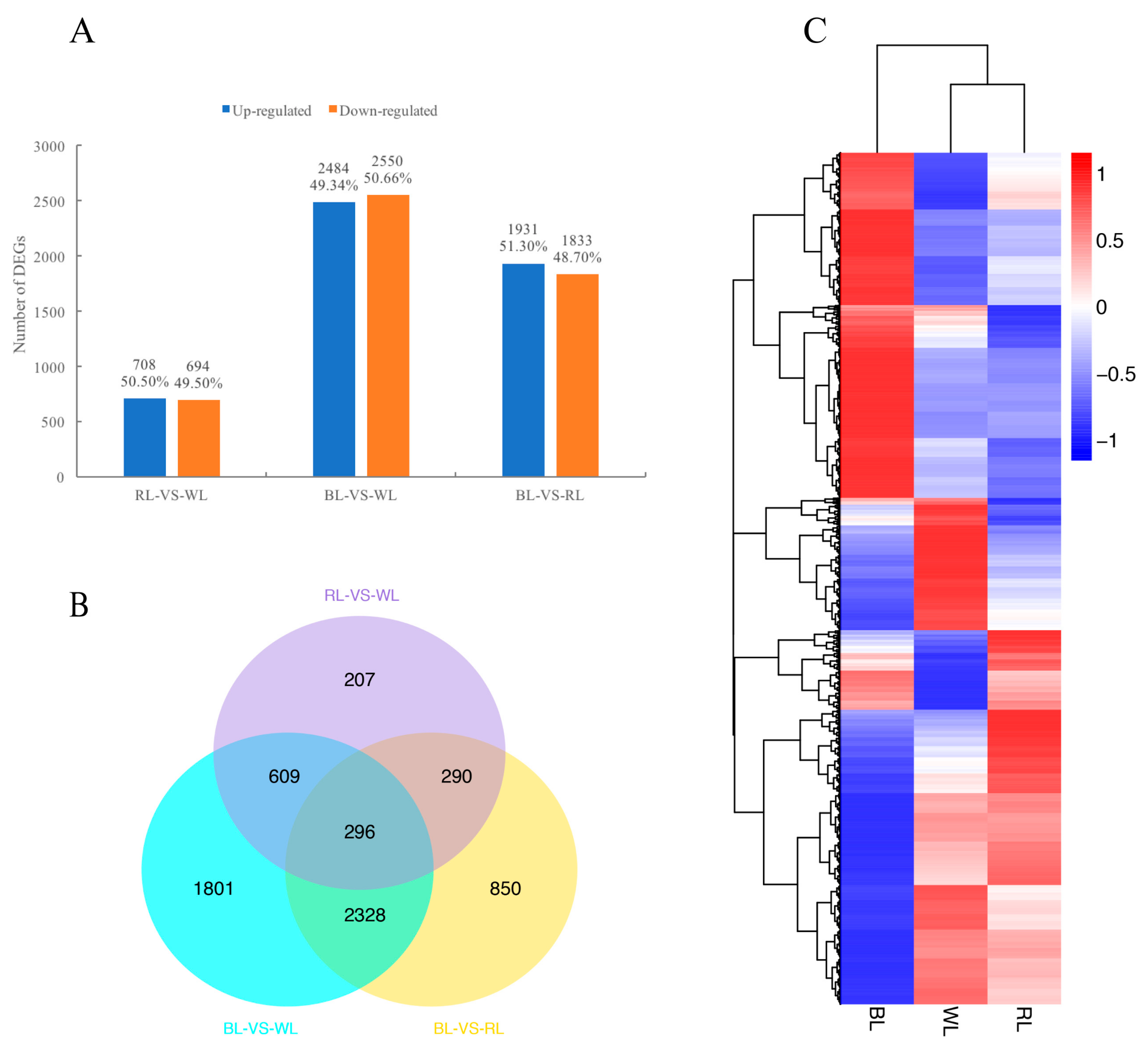
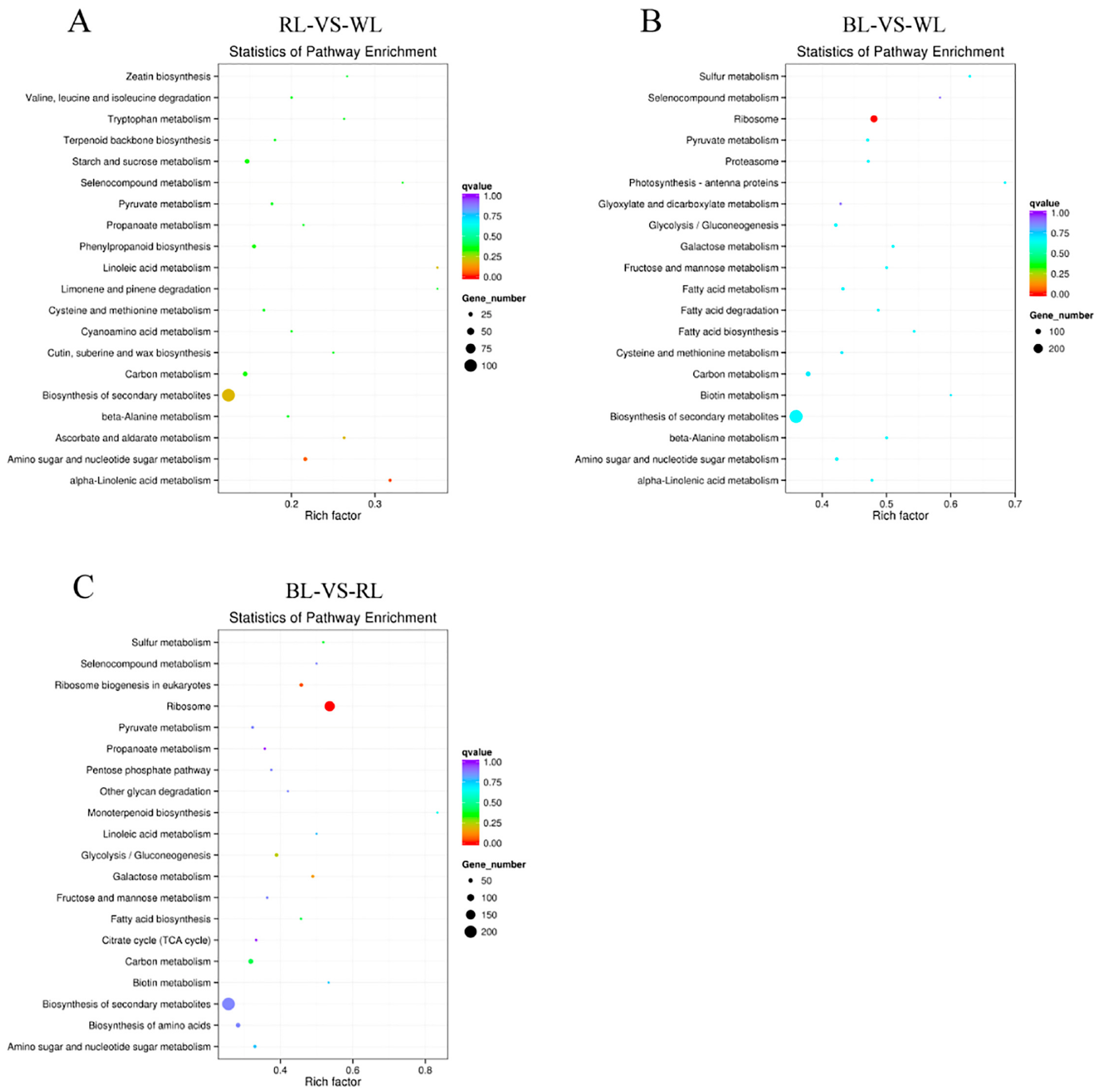
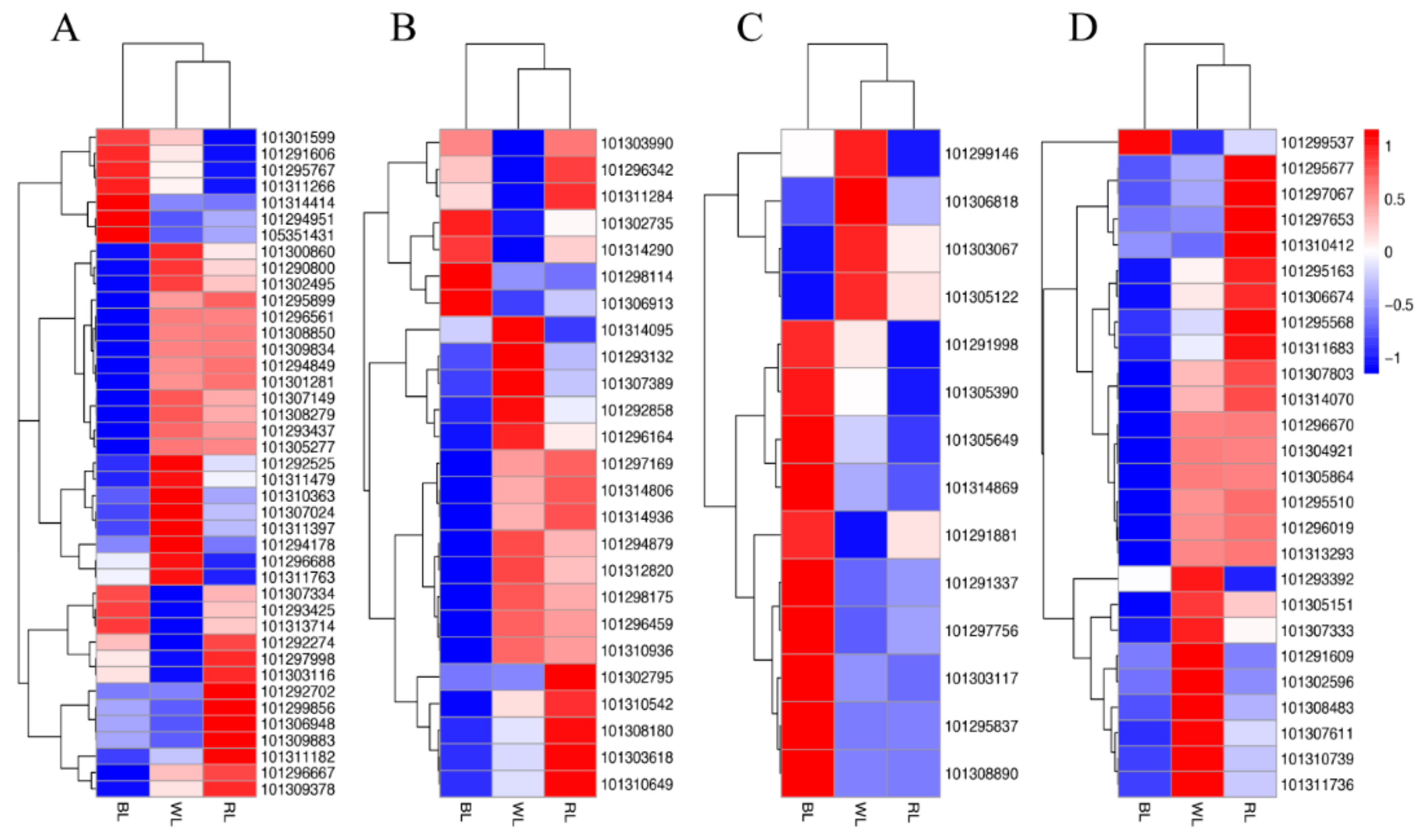
2.3. Global Analysis of Differential Gene Expression and Function Annotation
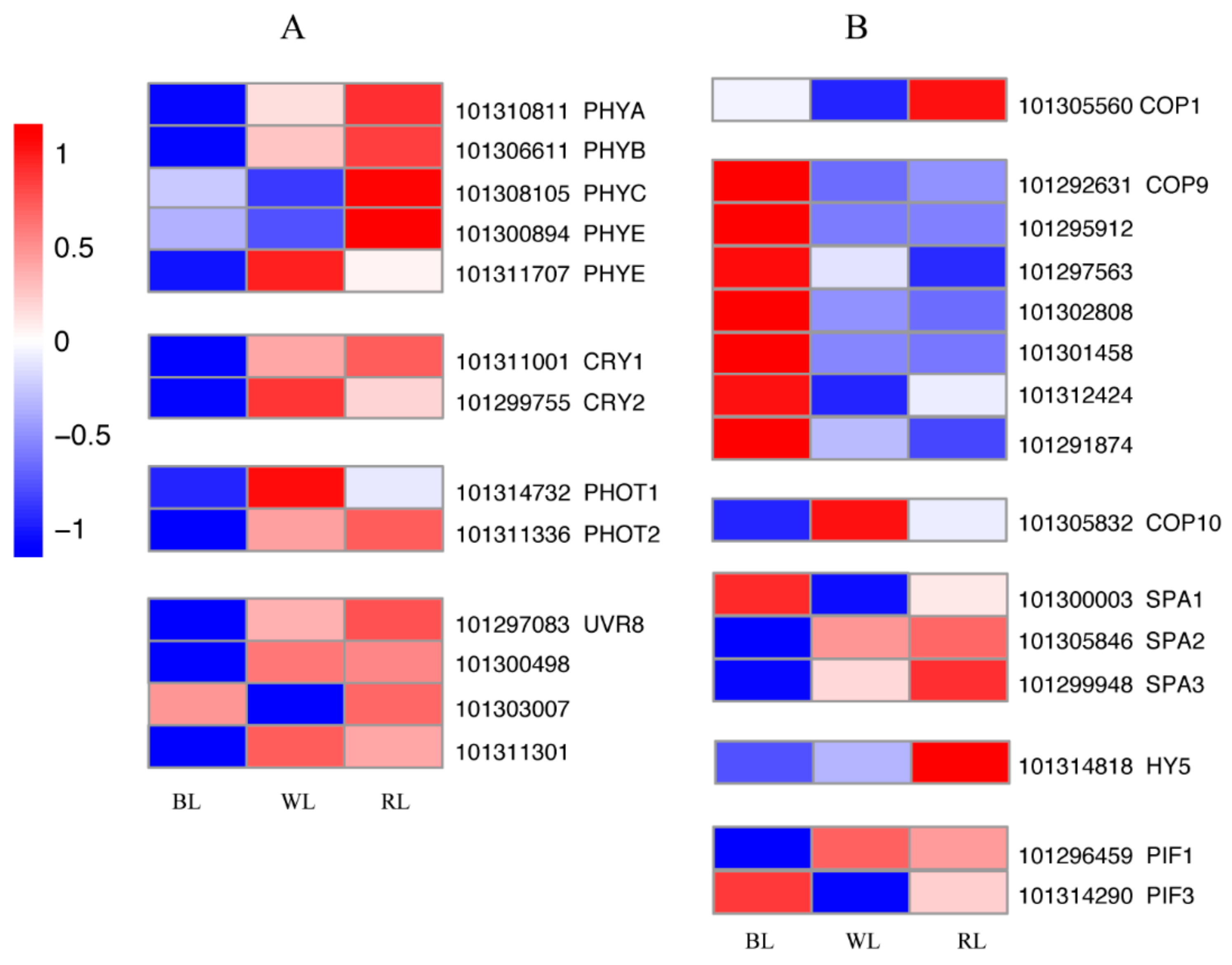
2.4. Light Signal Perception and Transduction Components
2.5. Transcription Factors (TFs)
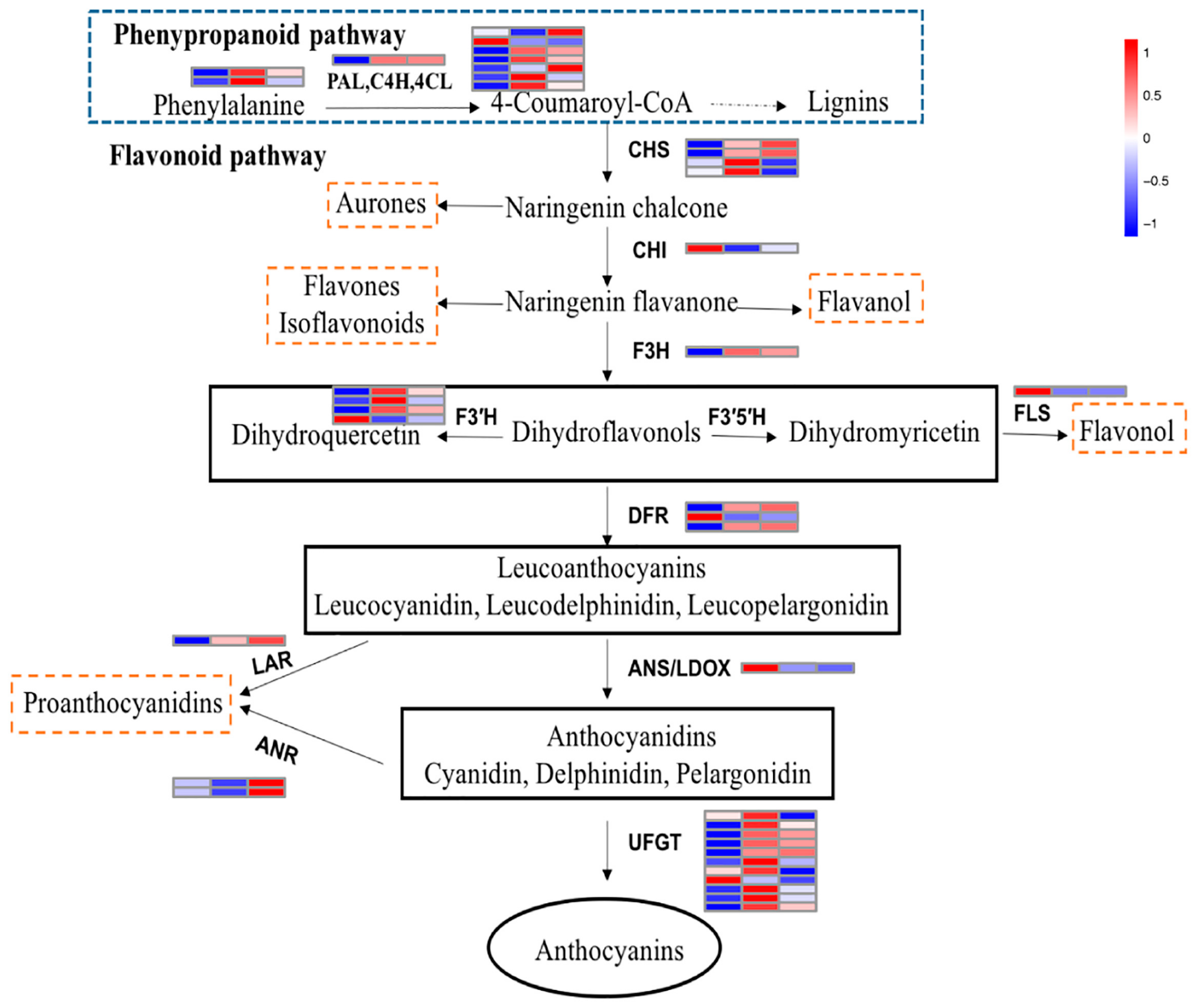
2.6. Structural Genes Involved in the Anthocyanin Pathway
2.7. Validation of Digital Expression Profiles by qRT-PCR
3. Discussion
4. Materials and Methods
4.1. Plant Materials
4.2. Anthocyanin and Proanthocyanidin Determination
4.3. RNA Extraction, cDNA Library Preparation and Illumina Sequencing
4.4. RNA-Seq Data Preprocessing Reads Mapping
4.5. Differentially-Expressed Gene (DEG) Identification and Functional Annotation
4.6. Quantitative Real-Time PCR (qRT-PCR) Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Conflicts of Interest
References
- He, J.; Giusti, M.M. Anthocyanins: Natural colorants with health-promoting properties. Annu. Rev. Food Sci. Technol. 2010, 1, 163–187. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Butelli, E.; Martin, C. Engineering anthocyanin biosynthesis in plants. Curr. Opin. Plant. Biol. 2014, 19, 81–90. [Google Scholar] [CrossRef] [PubMed]
- Kong, J.M.; Chia, L.S.; Goh, N.K.; Chia, T.F.; Brouillard, R. Analysis and biological activities of anthocyanins. Phytochemistry 2003, 64, 923–933. [Google Scholar] [CrossRef]
- Prior, R.L.; Wu, X. Anthocyanins: Structural characteristics that result in unique metabolic patterns and biological activities. Free Radical Res. 2006, 40, 1014–1028. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, Y.; Ohmiya, A. Seeing is believing: Engineering anthocyanin and carotenoid biosynthetic pathways. Curr. Opin. Biotechnol. 2008, 19, 190–197. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Gou, J.; Jia, Z.; Yang, L.; Sun, Y.; Xiao, X.; Song, F.; Luo, K. Molecular cloning and characterization of two genes encoding dihydroflavonol-4-reductase from populus trichocarpa. PLoS ONE 2012, 7, e30364. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Dubos, C.; Lepiniec, L. Transcriptional control of flavonoid biosynthesis by MYB–bHLH–WDR complexes. Trends Plant. Sci. 2015, 20, 176–185. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Grain, D.; Bobet, S.; Le Gourrierec, J.; Thévenin, J.; Kelemen, Z.; Lepiniec, L.; Dubos, C. Complexity and robustness of the flavonoid transcriptional regulatory network revealed by comprehensive analyses of MYB–bHLH–WDR complexes and their targets in Arabidopsis seed. New Phytol. 2014, 202, 132–144. [Google Scholar] [CrossRef] [PubMed]
- Stracke, R.; Werber, M.; Weisshaar, B. The R2R3-MYB gene family in Arabidopsis thaliana. Curr. Opin. Plant. Biol. 2001, 4, 447–456. [Google Scholar] [CrossRef]
- Albert, N.W.; Lewis, D.H.; Zhang, H.; Schwinn, K.E.; Jameson, P.E.; Davies, K.M. Members of an R2R3-MYB transcription factor family in petunia are developmentally and environmentally regulated to control complex floral and vegetative pigmentation patterning. Plant. J. 2011, 65, 771–784. [Google Scholar] [CrossRef] [PubMed]
- Medina-Puche, L.; Cumplido-Laso, G.; Amil-Ruiz, F.; Hoffmann, T.; Ring, L.; Rodríguez-Franco, A.; Caballero, J.L.; Schwab, W.; Muñoz-Blanco, J.; Blanco-Portales, R. MYB10 plays a major role in the regulation of flavonoid/phenylpropanoid metabolism during ripening of Fragaria × ananassa fruits. J. Exp. Bot. 2014, 65, 401–417. [Google Scholar] [CrossRef] [PubMed]
- Chagné, D.; Lin-Wang, K.; Espley, R.V.; Volz, R.K.; How, N.M.; Rouse, S.; Brendolise, C.; Carlisle, C.M.; Kumar, S.; De Silva, N. An ancient duplication of apple myb transcription factors is responsible for novel red fruit-flesh phenotypes. Plant. Physiol. 2013, 161, 225–239. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Díaz, J.R.; Pérez-Díaz, J.; Madrid-Espinoza, J.; González-Villanueva, E.; Moreno, Y.; Ruiz-Lara, S. New member of the R2R3-MYB transcription factors family in grapevine suppresses the anthocyanin accumulation in the flowers of transgenic tobacco. Plant. Mol. Biol. 2016, 90, 63–76. [Google Scholar] [CrossRef] [PubMed]
- Lai, B.; Li, X.J.; Hu, B.; Qin, Y.H.; Huang, X.M.; Wang, H.C.; Hu, G.B. Lcmyb1 is a key determinant of differential anthocyanin accumulation among genotypes, tissues, developmental phases and aba and light stimuli in litchi chinensis. PLoS ONE 2014, 9, e86293. [Google Scholar] [CrossRef] [PubMed]
- Hichri, I.; Barrieu, F.; Bogs, J.; Kappel, C.; Delrot, S.; Lauvergeat, V. Recent advances in the transcriptional regulation of the flavonoid biosynthetic pathway. J. Exp. Bot. 2011, 62, 2465–2483. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, A.; Brown, M.; Hatlestad, G.; Akhavan, N.; Smith, T.; Hembd, A.; Moore, J.; Montes, D.; Mosley, T.; Resendez, J.; et al. TTG2 controls the developmental regulation of seed coat tannins in Arabidopsis by regulating vacuolar transport steps in the proanthocyanidin pathway. Dev. Biol. 2016, 419, 54–63. [Google Scholar] [CrossRef] [PubMed]
- Verweij, W.R.; Spelt, C.E.; Bliek, M.; de Vries, M.; Wit, N.; Faraco, M.; Koes, R.; Quattrocchio, F.M. Functionally similar wrky proteins regulate vacuolar acidification in petunia and hair development in Arabidopsis. Plant. Cell 2016, 28, 786–803. [Google Scholar] [CrossRef] [PubMed]
- Duan, S.; Wang, J.; Gao, C.; Jin, C.; Li, D.; Peng, D.; Du, G.; Li, Y.; Chen, M. Functional characterization of a heterologously expressed Brassica napus WRKY41-1 transcription factor in regulating anthocyanin biosynthesis in Arabidopsis thaliana. Plant. Sci. 2018, 268, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.N.; Li, W.C.; Wang, H.C.; Shi, S.Y.; Shu, B.; Liu, L.Q.; Wei, Y.Z.; Xie, J.H. Transcriptome profiling of light-regulated anthocyanin biosynthesis in the pericarp of litchi. Front. Plant. Sci. 2016, 7, 963. [Google Scholar] [CrossRef] [PubMed]
- Guan, L.; Dai, Z.; Wu, B.H.; Wu, J.; Merlin, I.; Hilbert, G.; Renaud, C.; Gomès, E.; Edwards, E.; Li, S.H. Anthocyanin biosynthesis is differentially regulated by light in the skin and flesh of white-fleshed and teinturier grape berries. Planta 2016, 243, 23–41. [Google Scholar] [CrossRef] [PubMed]
- Vimolmangkang, S.; Zheng, D.; Han, Y.; Khan, M.A.; Soria-Guerra, R.E.; Korban, S.S. Transcriptome analysis of the exocarp of apple fruit identifies light-induced genes involved in red color pigmentation. Gene 2014, 534, 78–87. [Google Scholar] [CrossRef] [PubMed]
- Yu, B.; Zhang, D.; Huang, C.; Qian, M.; Zheng, X.; Teng, Y.; Su, J.; Shu, Q. Isolation of anthocyanin biosynthetic genes in red chinese sand pear (Pyrus pyrifolia nakai) and their expression as affected by organ/tissue, cultivar, bagging and fruit side. Sci. Hortic. 2012, 136, 29–37. [Google Scholar] [CrossRef]
- Reyes, L.F.; Miller, J.C.; Cisneros-Zevallos, L. Environmental conditions influence the content and yield of anthocyanins and total phenolics in purple-and red-flesh potatoes during tuber development. Am. J. Potato Res. 2004, 81, 187–193. [Google Scholar] [CrossRef]
- Carvalho, I.S.; Cavaco, T.; Carvalho, L.M.; Duque, P. Effect of photoperiod on flavonoid pathway activity in sweet potato (Ipomoea batatas (L.) lam.) leaves. Food Chem. 2010, 118, 384–390. [Google Scholar] [CrossRef]
- Steindal, A.L.H.; Mølmann, J.; Bengtsson, G.B.; Johansen, T.J. Influence of day length and temperature on the content of health-related compounds in broccoli (Brassica oleracea L. Var. Italica). J. Agric. Food Chem. 2013, 61, 10779–10786. [Google Scholar] [CrossRef] [PubMed]
- Zoratti, L.; Karppinen, K.; Luengo Escobar, A.; Häggman, H.; Jaakola, L. Light-controlled flavonoid biosynthesis in fruits. Front. Plant. Sci. 2014, 5, 534. [Google Scholar] [CrossRef] [PubMed]
- Lau, O.S.; Deng, X.W. The photomorphogenic repressors COP1 and DET1: 20 years later. Trends Plant. Sci. 2012, 17, 584–593. [Google Scholar] [CrossRef] [PubMed]
- Stracke, R.; Favory, J.J.; Gruber, H.; Bartelniewoehner, L.; Bartels, S.; Binkert, M.; Funk, M.; Weisshaar, B.; Ulm, R. The Arabidopsis bzip transcription factor HY5 regulates expression of the PFG1/MYB12 gene in response to light and ultraviolet-b radiation. Plant. Cell. Environ. 2010, 33, 88–103. [Google Scholar] [PubMed]
- Shin, D.H.; Choi, M.; Kim, K.; Bang, G.; Cho, M.; Choi, S.B.; Choi, G.; Park, Y.I. Hy5 regulates anthocyanin biosynthesis by inducing the transcriptional activation of the MYB75/PAP1 transcription factor in Arabidopsis. FEBS Lett. 2013, 587, 1543–1547. [Google Scholar] [CrossRef] [PubMed]
- Ang, L.H.; Chattopadhyay, S.; Wei, N.; Oyama, T.; Okada, K.; Batschauer, A.; Deng, X.W. Molecular interaction between COP1 and HY5 defines a regulatory switch for light control of Arabidopsis development. Mol. Cell. 1998, 1, 213–222. [Google Scholar] [CrossRef]
- Liu, Z.; Zhang, Y.; Wang, J.; Li, P.; Zhao, C.; Chen, Y.; Bi, Y. Phytochrome-interacting factors PIF4 and PIF5 negatively regulate anthocyanin biosynthesis under red light in arabidopsis seedlings. Plant. Sci. 2015, 238, 64–72. [Google Scholar] [CrossRef] [PubMed]
- Shin, J.; Park, E.; Choi, G. PIF3 regulates anthocyanin biosynthesis in an HY5-dependent manner with both factors directly binding anthocyanin biosynthetic gene promoters in Arabidopsis. Plant. J. 2007, 49, 981–994. [Google Scholar] [CrossRef] [PubMed]
- Azuma, A.; Udo, Y.; Sato, A.; Mitani, N.; Kono, A.; Ban, Y.; Yakushiji, H.; Koshita, Y.; Kobayashi, S. Haplotype composition at the color locus is a major genetic determinant of skin color variation in vitis× labruscana grapes. Theor. Appl. Genet. 2011, 122, 1427–1438. [Google Scholar] [CrossRef] [PubMed]
- Jaakola, L. New insights into the regulation of anthocyanin biosynthesis in fruits. Trends Plant. Sci 2013, 18, 477–483. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, H. Light-Controlled plant Cultivation System in Japan-Development of a Vegetable Factory Using Leds as a light source for plants. In Proceedings of the 6th VI International Symposium on Light in Horticulture, Tsukuba, Japan, 15–19 November 2009; pp. 37–44. [Google Scholar]
- Goto, E.; Matsumoto, H.; Ishigami, Y.; Hikosaka, S.; Fujiwara, K.; Yano, A. Measurements of the Photosynthetic Rates in Vegetables under Various Qualities of Light from Light-Emitting Diodes. In Proceedings of the International Symposium on New Technologies for Environment Control, Energy-Saving and Crop Production in Greenhouse and Plant, Jeju, Korea, 6–11 October 2013; pp. 261–268. [Google Scholar]
- Bian, Z.H.; Yang, Q.C.; Liu, W.K. Effects of light quality on the accumulation of phytochemicals in vegetables produced in controlled environments: A review. J. Sci. Food Agric. 2015, 95, 869–877. [Google Scholar] [CrossRef] [PubMed]
- Ilić, Z.S.; Fallik, E. Light quality manipulation improves vegetable quality at harvest and postharvest: A review. Environ. Exp. Bot. 2017, 139, 79–90. [Google Scholar] [CrossRef]
- Nadalini, S.; Zucchi, P.; Andreotti, C. Effects of blue and red led lights on soilless cultivated strawberry growth performances and fruit quality. Eur. J. Hortic. Sci. 2017, 82, 12–20. [Google Scholar] [CrossRef]
- Yoshida, H.; Mizuta, D.; Fukuda, N.; Hikosaka, S.; Goto, E. Effects of varying light quality from single-peak blue and red light-emitting diodes during nursery period on flowering, photosynthesis, growth, and fruit yield of everbearing strawberry. Plant. Biotechnol. 2016, 33, 267–276. [Google Scholar] [CrossRef]
- Kondo, S.; Tomiyama, H.; Rodyoung, A.; Okawa, K.; Ohara, H.; Sugaya, S.; Terahara, N.; Hirai, N. Abscisic acid metabolism and anthocyanin synthesis in grape skin are affected by light emitting diode (led) irradiation at night. J. Plant. Physiol. 2014, 171, 823–829. [Google Scholar] [CrossRef] [PubMed]
- Zoratti, L.; Sarala, M.; Carvalho, E.; Karppinen, K.; Martens, S.; Giongo, L.; Häggman, H.; Jaakola, L. Monochromatic light increases anthocyanin content during fruit development in bilberry. BMC Plant. Biol. 2014, 14, 377. [Google Scholar] [CrossRef] [PubMed]
- Aaby, K.; Mazur, S.; Nes, A.; Skrede, G. Phenolic compounds in strawberry (Fragaria × ananassa duch.) fruits: Composition in 27 cultivars and changes during ripening. Food Chem. 2012, 132, 86–97. [Google Scholar] [CrossRef] [PubMed]
- Tulipani, S.; Mezzetti, B.; Capocasa, F.; Bompadre, S.; Beekwilder, J.; De Vos, C.H.R.; Capanoglu, E.; Bovy, A.; Battino, M. Antioxidants, phenolic compounds, and nutritional quality of different strawberry genotypes. J. Agric. Food Chem. 2008, 56, 696–704. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Kubota, C. Effects of supplemental light quality on growth and phytochemicals of baby leaf lettuce. Environ. Exp. Bot. 2009, 67, 59–64. [Google Scholar] [CrossRef]
- Olle, M.; Viršile, A. The effects of light-emitting diode lighting on greenhouse plant growth and quality. Agric. Food Sci. 2013, 22, 223–234. [Google Scholar]
- Nishimura, T.; Ohyama, K.; Goto, E.; Inagaki, N. Concentrations of perillaldehyde, limonene, and anthocyanin of perilla plants as affected by light quality under controlled environments. Sci. Hortic. 2009, 122, 134–137. [Google Scholar] [CrossRef]
- Mizuno, T.; Amaki, W.; Watanabe, H. Effects of Monochromatic Light Irradiation by Led on the Growth and Anthocyanin Contents in Leaves of Cabbage Seedlings. In Proceedings of the 6th VI International Symposium on Light in Horticulture, Tsukuba, Japan, 15–19 November 2009; pp. 179–184. [Google Scholar]
- Miao, L.; Zhang, Y.; Yang, X.; Xiao, J.; Zhang, H.; Zhang, Z.; Wang, Y.; Jiang, G. Colored light-quality selective plastic films affect anthocyanin content, enzyme activities, and the expression of flavonoid genes in strawberry (Fragaria × ananassa) fruit. Food Chem. 2016, 207, 93–100. [Google Scholar] [CrossRef] [PubMed]
- Kadomura-Ishikawa, Y.; Miyawaki, K.; Noji, S.; Takahashi, A. Phototropin 2 is involved in blue light-induced anthocyanin accumulation in Fragaria × ananassa fruits. J. Plant. Res. 2013, 126, 847–857. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Cao, S.; Shi, L.; Chen, W.; Su, X.; Yang, Z. Blue light irradiation affects anthocyanin content and enzyme activities involved in postharvest strawberry fruit. J. Agric. Food Chem. 2014, 62, 4778–4783. [Google Scholar] [CrossRef] [PubMed]
- Möglich, A.; Yang, X.; Ayers, R.A.; Moffat, K. Structure and function of plant photoreceptors. Annu. Rev. Plant. Biol. 2010, 61, 21–47. [Google Scholar] [CrossRef] [PubMed]
- Viczián, A.; Klose, C.; Ádám, É.; Nagy, F. New insights of red light-induced development. Plant. Cell. Environ. 2017, 40, 2457–2468. [Google Scholar] [CrossRef] [PubMed]
- Cashmore, A.R.; Jarillo, J.A.; Wu, Y.J.; Liu, D. Cryptochromes: Blue light receptors for plants and animals. Science 1999, 284, 760–765. [Google Scholar] [CrossRef] [PubMed]
- Kang, B.; Grancher, N.; Koyffmann, V.; Lardemer, D.; Burney, S.; Ahmad, M. Multiple interactions between cryptochrome and phototropin blue-light signalling pathways in Arabidopsis thaliana. Planta 2008, 227, 1091–1099. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M.; Cashmore, A.R. The blue-light receptor cryptochrome 1 shows functional dependence on phytochrome a or phytochrome b in Arabidopsis thaliana. Plant. J. 1997, 11, 421–427. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M.; Jarillo, J.A.; Smirnova, O.; Cashmore, A.R. The cry1 blue light photoreceptor of Arabidopsis interacts with phytochrome a in vitro. Mol. Cell. 1998, 1, 939–948. [Google Scholar] [CrossRef]
- Mas, P.; Devlin, P.F.; Panda, S.; Kay, S.A. Functional interaction of phytochrome b and cryptochrome 2. Nature 2000, 408, 207–211. [Google Scholar] [CrossRef] [PubMed]
- Jenkins, G.I. The uv-b photoreceptor UVR8: From structure to physiology. Plant. Cell 2014, 26, 21–37. [Google Scholar] [CrossRef] [PubMed]
- Morales, L.O.; Brosché, M.; Vainonen, J.; Jenkins, G.I.; Wargent, J.J.; Sipari, N.; Strid, Å.; Lindfors, A.V.; Tegelberg, R.; Aphalo, P.J. Multiple roles for uv resistance locus8 in regulating gene expression and metabolite accumulation in Arabidopsis under solar ultraviolet radiation. Plant. Physiol. 2013, 161, 744–759. [Google Scholar] [CrossRef] [PubMed]
- Yamagishi, M. A novel R2R3-MYB transcription factor regulates light-mediated floral and vegetative anthocyanin pigmentation patterns in lilium regale. Mol. Breed. 2016, 36, 3. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, L.; Zhang, J.; Yu, B.; Wang, J.; Wang, D. The myb transcription factor stmyba1 from potato requires light to activate anthocyanin biosynthesis in transgenic tobacco. J. Plant. Biol. 2017, 60, 93–101. [Google Scholar] [CrossRef]
- Azuma, A.; Yakushiji, H.; Koshita, Y.; Kobayashi, S. Flavonoid biosynthesis-related genes in grape skin are differentially regulated by temperature and light conditions. Planta 2012, 236, 1067–1080. [Google Scholar] [CrossRef] [PubMed]
- Feng, F.; Li, M.; Ma, F.; Cheng, L. Phenylpropanoid metabolites and expression of key genes involved in anthocyanin biosynthesis in the shaded peel of apple fruit in response to sun exposure. Plant. Physiol. Biochem. 2013, 69, 54–61. [Google Scholar] [CrossRef] [PubMed]
- Feng, S.; Wang, Y.; Yang, S.; Xu, Y.; Chen, X. Anthocyanin biosynthesis in pears is regulated by a R2R3-MYB transcription factor pymyb10. Planta 2010, 232, 245–255. [Google Scholar] [CrossRef] [PubMed]
- Kadomura-Ishikawa, Y.; Miyawaki, K.; Takahashi, A.; Masuda, T.; Noji, S. Light and abscisic acid independently regulated famyb10 in Fragaria × ananassa fruit. Planta 2015, 241, 953–965. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, N.H.; Jeong, C.Y.; Kang, G.H.; Yoo, S.D.; Hong, S.W.; Lee, H. Mybd employed by HY5 increases anthocyanin accumulation via repression of MYBl2 in Arabidopsis. Plant. J. 2015, 84, 1192–1205. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Du, H.; Zhai, R.; Song, L.; Ma, F.; Xu, L. Transcriptome analysis reveals candidate genes related to color fading of ‘red bartlett’(Pyrus communis L.). Front. Plant. Sci. 2017, 8, 455. [Google Scholar] [CrossRef] [PubMed]
- Passeri, V.; Koes, R.; Quattrocchio, F.M. New challenges for the design of high value plant products: Stabilization of anthocyanins in plant vacuoles. Front. Plant. Sci. 2016, 7, 153. [Google Scholar] [CrossRef] [PubMed]
- Prior, R.L.; Fan, E.; Ji, H.; Howell, A.; Nio, C.; Payne, M.J.; Reed, J. Multi-laboratory validation of a standard method for quantifying proanthocyanidins in cranberry powders. J. Sci. Food Agric. 2010, 90, 1473–1478. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Yu, H.W.; Wang, X.R.; Xie, X.L.; Yue, X.Y.; Tang, H.R. An alternative cetyltrimethylammonium bromide-based protocol for rna isolation from blackberry (rubus L.). Genet. Mol. Res. 2012, 11, 1773–1782. [Google Scholar] [CrossRef] [PubMed]
- Hirakawa, H.; Shirasawa, K.; Kosugi, S.; Tashiro, K.; Nakayama, S.; Yamada, M.; Kohara, M.; Watanabe, A.; Kishida, Y.; Fujishiro, T. Dissection of the octoploid strawberry genome by deep sequencing of the genomes of fragaria species. DNA Res. 2014, 21, 169–181. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Sevilla, J.F.; Vallarino, J.G.; Osorio, S.; Bombarely, A.; Posé, D.; Merchante, C.; Botella, M.A.; Amaya, I.; Valpuesta, V. Gene expression atlas of fruit ripening and transcriptome assembly from rna-seq data in octoploid strawberry (Fragaria × ananassa). Sci. Rep. 2017, 7, 13737. [Google Scholar] [CrossRef] [PubMed]
- Shulaev, V.; Sargent, D.J.; Crowhurst, R.N.; Mockler, T.C.; Folkerts, O.; Delcher, A.L.; Jaiswal, P.; Mockaitis, K.; Liston, A.; Mane, S.P.; et al. The genome of woodland strawberry (Fragaria vesca). Nat. Genet. 2011, 43, 109–116. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Williams, B.A.; Pertea, G.; Mortazavi, A.; Kwan, G.; Van Baren, M.J.; Salzberg, S.L.; Wold, B.J.; Pachter, L. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 2010, 28, 511–515. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B (Methodol.) 1995, 57, 289–300. [Google Scholar]
Sample Availability: Samples of the compounds are available from the authors upon request. |
| Sample Name | Raw Reads | Clean Reads | Clean Bases | Error Rate (%) | Q20 (%) | Q30 (%) | GC Content (%) | Total Mapped (%) |
|---|---|---|---|---|---|---|---|---|
| W251 | 51,106,150 | 49,645,282 | 7.45 G | 0.02 | 96.75 | 92.19 | 46.8 | 48.01 |
| W252 | 49,154,200 | 47,925,342 | 7.19 G | 0.01 | 96.85 | 92.33 | 47.32 | 48.49 |
| B251 | 56,427,402 | 54,927,816 | 8.24 G | 0.02 | 96.7 | 92.09 | 46.89 | 47.57 |
| B252 | 48,846,890 | 47,634,610 | 7.15 G | 0.02 | 96.57 | 91.86 | 46.61 | 47.80 |
| R251 | 48,997,928 | 47,876,282 | 7.18 G | 0.02 | 96.57 | 91.84 | 46.92 | 47.96 |
| R252 | 47,859,022 | 46,555,194 | 6.98 G | 0.01 | 96.84 | 92.38 | 47.18 | 49.01 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; Jiang, L.; Li, Y.; Chen, Q.; Ye, Y.; Zhang, Y.; Luo, Y.; Sun, B.; Wang, X.; Tang, H. Effect of Red and Blue Light on Anthocyanin Accumulation and Differential Gene Expression in Strawberry (Fragaria × ananassa). Molecules 2018, 23, 820. https://doi.org/10.3390/molecules23040820
Zhang Y, Jiang L, Li Y, Chen Q, Ye Y, Zhang Y, Luo Y, Sun B, Wang X, Tang H. Effect of Red and Blue Light on Anthocyanin Accumulation and Differential Gene Expression in Strawberry (Fragaria × ananassa). Molecules. 2018; 23(4):820. https://doi.org/10.3390/molecules23040820
Chicago/Turabian StyleZhang, Yunting, Leiyu Jiang, Yali Li, Qing Chen, Yuntian Ye, Yong Zhang, Ya Luo, Bo Sun, Xiaorong Wang, and Haoru Tang. 2018. "Effect of Red and Blue Light on Anthocyanin Accumulation and Differential Gene Expression in Strawberry (Fragaria × ananassa)" Molecules 23, no. 4: 820. https://doi.org/10.3390/molecules23040820
APA StyleZhang, Y., Jiang, L., Li, Y., Chen, Q., Ye, Y., Zhang, Y., Luo, Y., Sun, B., Wang, X., & Tang, H. (2018). Effect of Red and Blue Light on Anthocyanin Accumulation and Differential Gene Expression in Strawberry (Fragaria × ananassa). Molecules, 23(4), 820. https://doi.org/10.3390/molecules23040820






