Terretonin N: A New Meroterpenoid from Nocardiopsis sp.
Abstract
:1. Introduction
2. Results and Discussion
2.1. Isolation and Maintenance of the Producing Strain
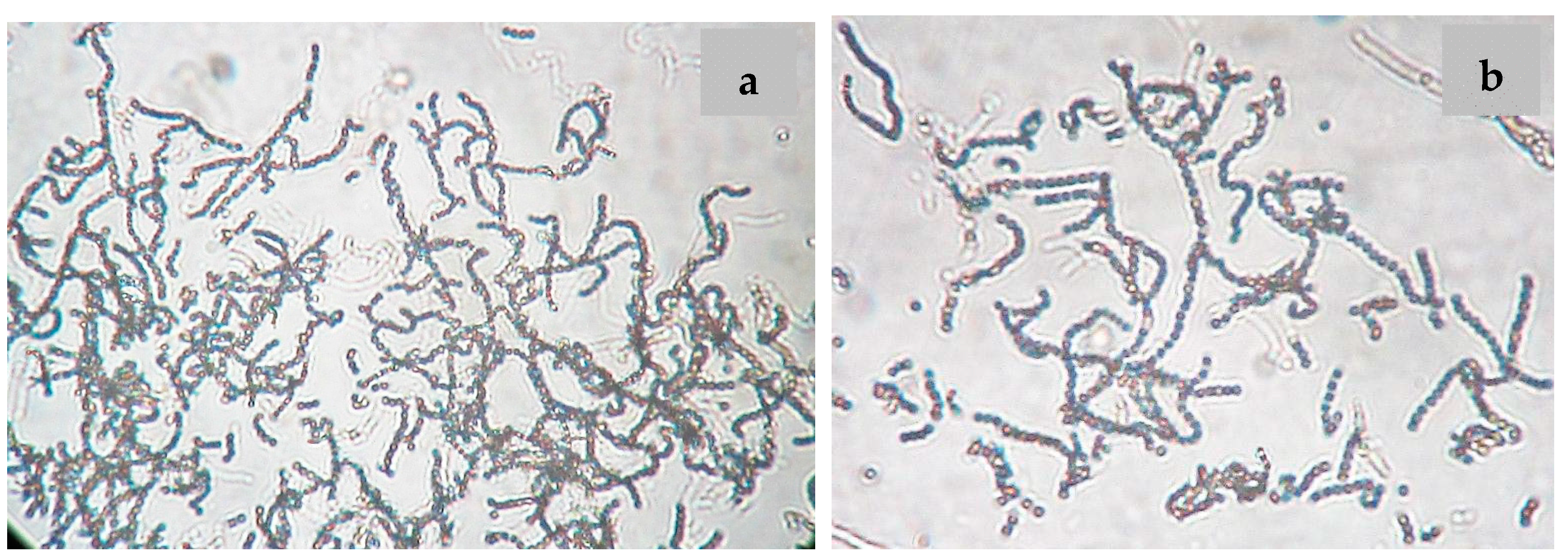
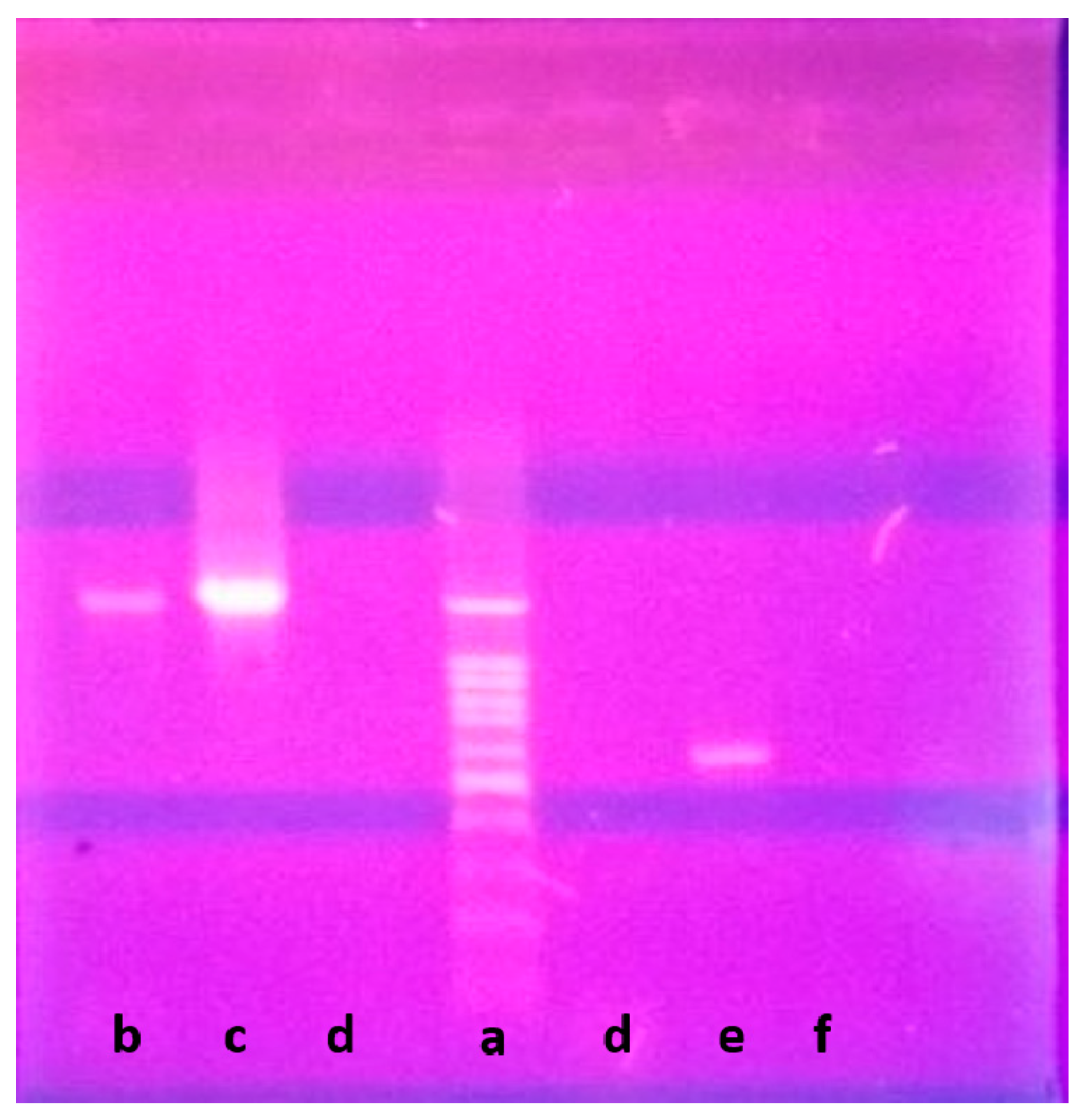
2.2. Phenotypic and Genotypic Characteristics
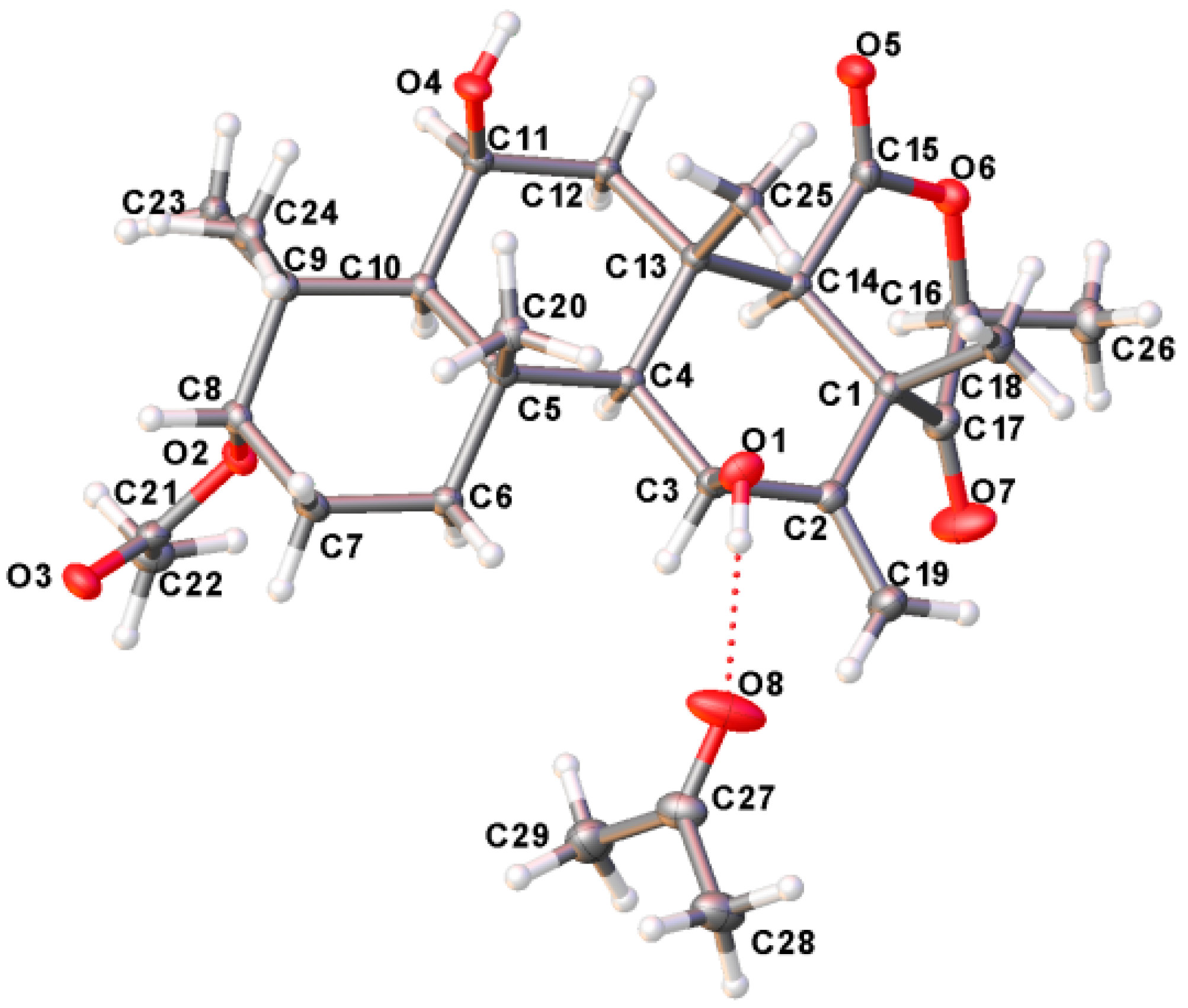
2.3. Structure Elucidation
2.4. Biological Activity
3. Materials and Methods
3.1. General Experimental Details
3.2. Isolation of Nocardiopsis sp. LGO5
3.3. DNA Isolation and 16S rRNA Gene Sequencing
3.4. Phenotypic Characterization of the Strain
3.5. Large-Scale Fermentation and Isolation
3.6. Antimicrobial Activity Assay
3.7. Cytotoxicity Assays
3.8. Crystal Data
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Laatsch, H.; AntiBase, A. Data Base for Rapid Structural Determination of Microbial Natural Products, and Annual Updates; Wiley-VCH: Weinheim, Germany. Available online: http://wwwuser.gwdg.de/~ucoc/laatsch/AntiBase.htm (accessed on 28 January 2016).
- Butler, M.S. The role of natural product chemistry in drug discovery. J. Nat. Prod. 2004, 67, 2141–2153. [Google Scholar] [CrossRef] [PubMed]
- Eliwa, E.M.; Abdel-Razek, A.S.; Frese, M.; Halawa, A.H.; El-Agrody, A.M.; Bedair, A.H.; Sewald, N.; Shaaban, M. New bioactive compounds from the marine-derived actinomycete Nocardiopsis lucentensis sp. ASMR2. Zeitschrift für Naturforschung B 2017, 72, 351–360. [Google Scholar] [CrossRef]
- Singh, S.B.; Zink, D.L.; Guan, Z.; Collado, J.; Pelaez, F.; Felock, P.J.; Hazuda, D.J. Isolation, structure and HIV-1 integrase inhibitory activity of xanthoviridicatin E and F, two novel fungal metabolites produced by penicillium chrysogenum. Helv. Chim. Acta 2003, 86, 3380–3385. [Google Scholar] [CrossRef]
- Zhang, H.W.; Song, Y.C.; Tan, R.X. Biology and chemistry of endophytes. Nat. Prod. Rep. 2006, 23, 753–771. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.C.; Li, H.; Ye, Y.H.; Shan, C.Y.; Yang, Y.M.; Tan, R.X. Endophytic naphthopyrone metabolites are co-inhibitors of xanthine oxidase, SW1116 cell and some microbial growths. FEMS Microbiol. Lett. 2004, 241, 67–72. [Google Scholar] [CrossRef] [PubMed]
- Hamed, A.; Abdel-Razek, A.S.; Frese, M.; Wibberg, D.; El-Haddad, A.F.; Ibrahim, T.M.A.; Kalinowski, J.; Sewald, N.; Shaaban, M. New Oxaphenalene Derivative from marine-derived Streptomyces griseorubens sp. ASMR4. Zeitschrift für Naturforschung B 2017, 72, 53–62. [Google Scholar] [CrossRef]
- Debnath, M.; Paul, A.K.; Bisen, P.S. Natural Bioactive Compounds and Biotechnological potential of Marine Bacteria. Curr. Pharm. Biotechnol. 2007, 8, 253–260. [Google Scholar] [CrossRef] [PubMed]
- Blunt, J.W.; Copp, B.R.; Hu, W.P.; Munro, M.H.G.; Northcote, P.T.; Prinsep, M.R. Marine Natural Products. Nat. Prod. Rep. 2007, 24, 31–86. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.-H.; Miao, F.-P.; Qiao, M.-F.; Cichewicz, R.H.; Ji, N.-Y. Terretonin, ophiobolin, and drimane terpenes with absolute configurations from an algicolous Aspergillus ustus. RSC Adv. 2013, 3, 588–595. [Google Scholar] [CrossRef]
- Yamada, Y.; Kuzuyama, T.; Komatsu, M.; Shin-ya, K.; Omura, S.; Cane, D.E.; Ikeda, H. Terpene synthases are widely distributed in bacteria. Proc. Natl. Acad. Sci. USA 2015, 112, 857–862. [Google Scholar] [CrossRef] [PubMed]
- Shirai, M.; Okuda, M.; Motohashi, K.; Imoto, M.; Furihata, K.; Matsuo, Y.; Katsuta, A.; Shizuri, Y.; Seto, H. Terpenoids produced by actinomycetes: Isolation, structural elucidation and biosynthesis of new diterpenes, gifhornenolones A and B from Verrucosispora gifhornensis YM28-088. J. Antibiot. 2010, 63, 245–250. [Google Scholar] [CrossRef] [PubMed]
- Schneider, K.; Graf, E.; Irran, E.; Nicholson, G.; Stainsby, F.M.; Goodfellow, M.; Borden, S.A.; Keller, S.; Süssmuth, R.D.; Fiedler, H.-P. Bendigoles A–C, New Steroids from Gordonia australis Acta 2299. J. Antibiot. 2008, 61, 356–364. [Google Scholar] [CrossRef] [PubMed]
- Springer, J.P.; Dorner, J.W.; Cole, R.J.; Cox, R.H. Terretonin, a toxic compound from aspergillus terreus. J. Org. Chem. 1979, 44, 4852–4854. [Google Scholar] [CrossRef]
- Li, G.-Y.; Li, B.-G.; Yang, T.; Yin, J.-H.; Qi, H.-Y.; Liu, G.-Y.; Zhang, G.-L. Sesterterpenoids, Terretonins A–D, and an Alkaloid, Asterrelenin, from Aspergillus terreus. J. Nat. Prod. 2005, 68, 1243–1246. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.B.; Drada-Ebel, R.; Ebel, E.R.; Schulz, W.B.; Draeger, S.; Mueller, W.E.G.; Wary, V.; Lin, W.H.; Proksch, P. Ophiobolin sesterterpenoids and pyrrolidine alkaloids from the sponge-derived fungus Aspergillus ustus. Helv. Chim. Acta 2011, 94, 623–631. [Google Scholar] [CrossRef]
- Liu, H.B.; Edrada-Ebel, R.; Ebel, R.; Schulz, W.B.; Draeger, S.; Mueller, W.E.G.; Wary, V.; Lin, W.H.; Proksch, P. Drimane Sesquiterpenoids from the Fungus Aspergillus ustus Isolated from the Marine Sponge Suberites domuncula. J. Nat. Prod. 2009, 72, 1585–1588. [Google Scholar] [CrossRef] [PubMed]
- Kikuchi, H.; Tsukitani, Y.; Kobayshi, M.; Kitagawa, I. Marine Natural Products. XI. An Antiinflammatory Scalarane-type Bishomosesterterpene, Foliaspongin, from the Okinawan Marine Sponge Phyllospongia foliascens (PALLAS). Chem. Pharm. Bull. 1983, 31, 552–556. [Google Scholar] [CrossRef]
- Charan, R.D.; Mckee, T.C.; Boyd, M.R. Thorectandrols C, D, and E, New Sesterterpenes from the Marine Sponge Thorectandra sp. J. Nat. Prod. 2002, 65, 492–495. [Google Scholar] [CrossRef] [PubMed]
- Terem, B.; Scheuer, P.J. Scalaradical derivatives from the nurdibranch chromodoris youngbleuthi and the sponge Spongia Oceania. Tetrahedron 1986, 42, 4409–4412. [Google Scholar] [CrossRef]
- Kazlauskas, R.; Murphy, P.T.; Wells, R.J. Five new C26 tetracyclic terpenes from a sponge (Lendenfeldia sp.). Aust. J. Chem. 1982, 35, 51–59. [Google Scholar] [CrossRef]
- Bowden, B.F.; Coll, J.C.; Li, H.; Cambie, R.C.; Kernan, M.R.; Bergquist, P.R. New Cytotoxic Scalarane Sesterterpenes from the Dictyoceratid Sponge Strepsichordaia lendenfeldi. J. Nat. Prod. 1992, 55, 1234–1240. [Google Scholar] [CrossRef] [PubMed]
- Komai, S.-I.; Hosoe, T.; Itabashi, T.; Nozawa, K.; Okada, K.; de Campos Takaki, G.M.; Yaguchi, T.; Takizawa, K.; Fukushima, K.; Kawai, K.-I. Two New Meroterpenoids, Penisimplicin A and B, Isolated from Penicillium simplicissimum. Chem. Pharm. Bull. 2005, 53, 1114–1117. [Google Scholar] [CrossRef] [PubMed]
- Shaaban, M. Bioactive Secondary Metabolites from Marine and Terrestrial Bacteria: Isoquinolinequinones, Bacterial Compounds with a Novel Pharmacophor. Ph.D. Thesis, Georg-August University, Göttingen, Germany, 2004. [Google Scholar]
- Agzamova, M.A.; Isaev, I.M.; Rakhmatov, K.A. Steroids and Glycosides from Astragalus turczaninowii. Chem. Nat. Compd. 2017, 53, 398–399. [Google Scholar] [CrossRef]
- Ma, Q.; Ding, W.; Chen, Z.; Ma, Z. Bisamides and rhamnosides from mangrove actinomycete Streptomyces sp. SZ-A15. Nat. Prod. Res. 2017. [Google Scholar] [CrossRef] [PubMed]
- Nagia, M.M.S.; El-Metwally, M.M.; Shaaban, M.; El-Zalabani, S.M.; Hanna, A.G. Four butyrolactones and diverse bioactive secondary metabolites from terrestrial Aspergillus flavipes MM2: Isolation and structure determination. Org. Med. Chem. Lett. 2012, 2, 9. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Shaaban, M.; Shaaban, K.A.; Abd-Alla, H.I.; Hanna, A.G.; Laatsch, H. Dendrophen, a Novel Glycyrrhetyl Amino Acid from Dendronephthya hemprichi. Zeitschrift für Naturforschung B 2011, 66, 425–432. [Google Scholar] [CrossRef]
- Aroma of Basil, Scent of Pine. Bacteria Could Be Rich Source for Making Terpenes. 22 December 2014 Media Contact: Kevin Stacey 401-863-3766. Available online: https://news.brown.edu/articles/2014/12/terpenes (accessed on 25 January 2018).
- Yamada, Y.; Arima, S.; Nagamitsu, T.; Johmoto, K.; Uekusa, H.; Eguchi, T.; Shin-ya, K.; Cane, D.E.; Ikeda, H. Novel terpenes generated by heterologous expression of bacterial terpene synthase genes in an engineered Streptomyces host. J. Antibiot. 2015, 68, 385–394. [Google Scholar] [CrossRef] [PubMed]
- Cane, D.E.; Ikeda, H. Exploration and mining of the bacterial terpenome. Acc. Chem. Res. 2012, 45, 463–472. [Google Scholar] [CrossRef] [PubMed]
- Zhu, D.; Seo, M.J.; Ikeda, H.; Cane, D.E. Genome mining in streptomyces. Discovery of an unprecedented P450-catalyzed oxidative rearrangement that is the final step in the biosynthesis of pentalenolactone. J. Am. Chem. Soc. 2011, 133, 2128–2131. [Google Scholar] [CrossRef] [PubMed]
- Doud, E.H.; Perlstein, D.L.; Wolpert, M.; Cane, D.E.; Walker, S. Two distinct mechanisms for TIM barrel prenyltransferases in bacteria. J. Am. Chem. Soc. 2011, 133, 1270–1273. [Google Scholar] [CrossRef] [PubMed]
- Takamatsu, S.; Lin, X.; Nara, A.; Komatsu, M.; Cane, D.E.; Ikeda, H. Characterization of a silent sesquiterpenoid biosynthetic pathway in Streptomyces avermitilis controlling epi-isozizaene albaflavenone biosynthesis and isolation of a new oxidized epi-isozizaene metabolite. Microb. Biotechnol. 2011, 4, 184–191. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Chou, W.K.; Hopson, R.; Cane, D.E. Genome mining in Streptomyces clavuligerus: Expression and biochemical characterization of two new cryptic sesquiterpene synthases. Chem. Biol. 2011, 18, 32–37. [Google Scholar] [CrossRef] [PubMed]
- Aaron, J.A.; Lin, X.; Cane, D.E.; Christianson, D.W. Structure of epi-isozizaene synthase from Streptomyces coelicolor A3(2), a platform for new terpenoid cyclization templates. Biochemistry 2010, 49, 1787–1797. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, Y.; Awakawa, T.; Itoh, T.; Wakimoto, T.; Kushiro, T.; Fujii, I.; Ebizuka, Y.; Abe, I. Terretonin Biosynthesis Requires Methylation as Essential Step for Cyclization. ChemBioChem 2012, 13, 1738–1741. [Google Scholar] [CrossRef] [PubMed]
- Dickschat, J.S. Bacterial terpene cyclases. Nat. Prod. Rep. 2016, 33, 87–110. [Google Scholar] [CrossRef] [PubMed]
- Sakoulas, G.; Nam, S.-J.; Loesgen, S.; Fenuical, W.; Gensen, P.R.; Nizet, V.; Hensler, M. Novel bacterial metabolite merhochlorin A demonstrates in vitro activities against multi-drug resistant methicillin-resistant Staphylococcus aureus. PLoS ONE 2012, 7, e29439. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.-J.; Knox, B.P.; Chiang, Y.-M.; Lo, H.-C.; Sanchez, J.-F.; Lee, K.-H.; Oakley, B.-R.; Bruno, K.S.; Wang, C.C.C. Molecular Genetic Characterization of a Cluster in A. terreus for Biosynthesis of the Meroterpenoid Terretonin. Org. Lett. 2012, 14, 5684–5687. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Bharti, A.; Gusain, O.; Bisht, G.S. An improved method for isolation of genomic DNA from filamentous actinomycetes. J. Eng. Technol. Manag. 2010, 2, 10–13. [Google Scholar]
- Tresner, H.D.; Hayes, J.A.; Backus, E.J. Differential tolerance of streptomycetes to sodium chloride as a taxonomic aid. Appl. Microbiol. 1968, 16, 1134–1136. [Google Scholar] [PubMed]
- Bauer, A.W.; Kirby, W.M.; Sherris, J.C.; Truck, M. Antibiotic susceptibility testing by a standardized single disk method. Am. J. Clin. Pathol. 1966, 45, 493–496. [Google Scholar] [PubMed]
Sample Availability: Sample of compound 1 is available from the authors. |
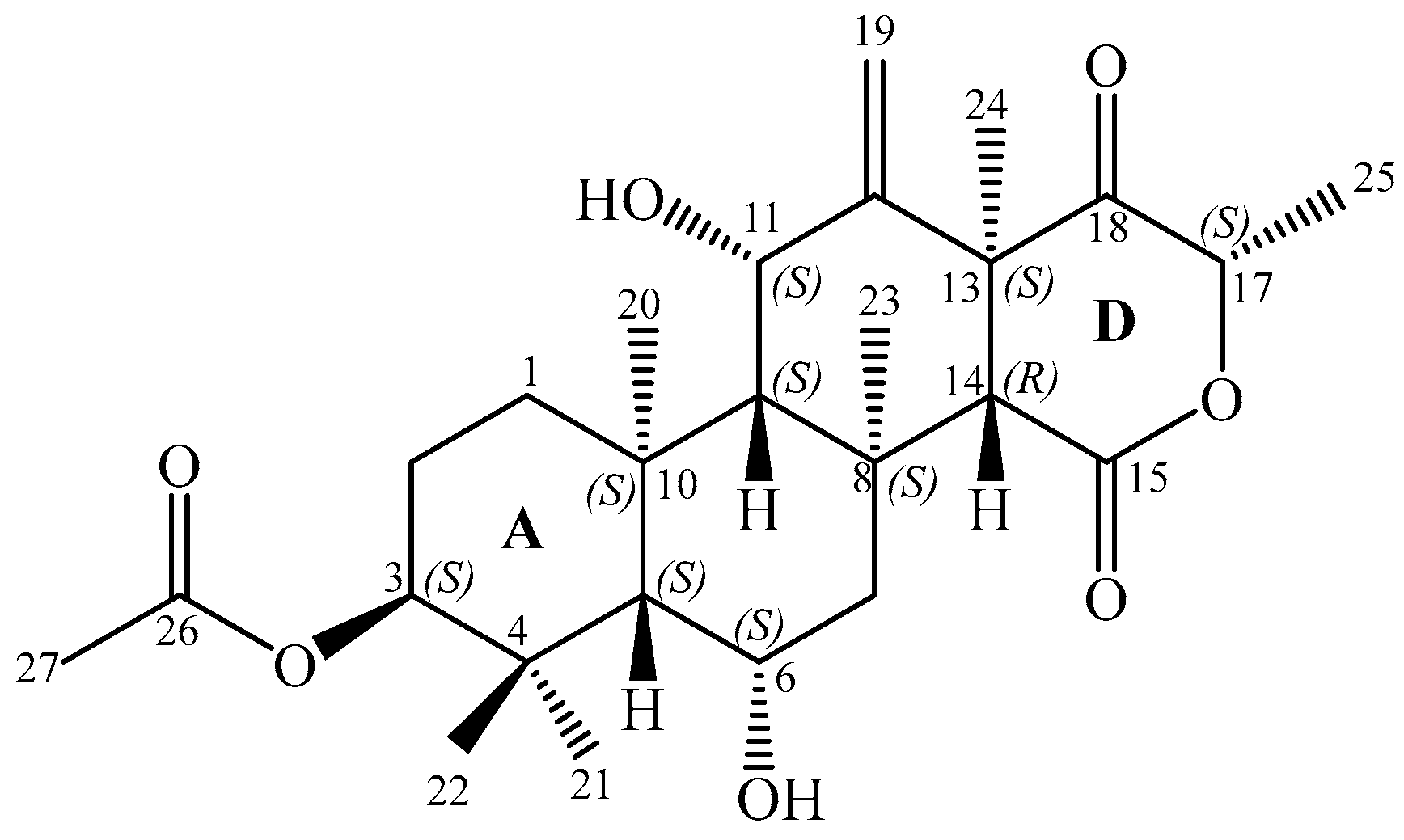


 ) and key HMBC (
) and key HMBC (  ) correlations of terretonin N (1).
) correlations of terretonin N (1).
 ) correlations of terretonin N (1).
) correlations of terretonin N (1).
| 1 | |
|---|---|
| Appearance | Colourless solid |
| Rf (Silica gel G/UV254, (CH2Cl2/5% MeOH)) | 0.54 |
| Staining with anisaldehyde/ sulfuric acid | Pink and later as violet |
| Molecular formula | C26H38O7 (462) |
| (+)-ESI-MS: m/z (%) | 485 ([M + Na]+, 100), 947 ([2M + Na]+, 0.58) |
| (−)-ESI-MS: m/z (%) | 461 ([M − H]−, 62), 891 ([2M − H]−, 4) |
| (−)-HR-ESI-MS: m/z | |
| Found | 461.2547 [M − H]− |
| Calcd. | 461.2545 [M − H]− |
| [α] | −114 (c = 0.1, MeOH) |
| Position | δC [ppm] | δH [ppm] (m, J [Hz]) | Position | δC [ppm] | δH [ppm] (m, J [Hz]) |
|---|---|---|---|---|---|
| 1 | 35.4 | 1.65 (m), 1.21 (m) | 14 | 59.1 | 2.58 (s) |
| 2 | 22.5 | 1.95 (m), 1.65 (m) | 15 | 169.4 | |
| 3 | 79.4 | 4.55 (t, 3.0) | 17 | 76.8 | 4.81 (m) |
| 4 | 37.6 | 18 | 206.8 | ||
| 5 | 51.5 | 1.25 (m) | 19 | 114.5 | 5.16 (s), 5.06 (s) |
| 6 | 68.1 | 4.35 (t, 3.0) | 20 | 18.8 | 1.68 (s) |
| 7 | 50.3 | 2.74 (dd, 14.5, 2.7), 1.19 (m) | 21 | 27.8 | 0.86 (s) |
| 8 | 37.1 | 22 | 23.7 | 1.28 (s) | |
| 9 | 61.3 | 0.95 (d, 2.4) | 23 | 19.0 | 2.03 (s) |
| 10 | 38.1 | 24 | 24.2 | 1.62 (s) | |
| 11 | 76.0 | 4.81 (m) | 25 | 14.6 | 1.38 (d, 6.4) |
| 12 | 147.9 | 26 | 170.6 | ||
| 13 | 50.0 | 27 | 21.3 | 1.96 (s) |
| EC a | BS b | Psa c | Ml d | Stw e | Sta f | Psae g | Ca h | Sac i | |
|---|---|---|---|---|---|---|---|---|---|
| LGO5 extract | 7 | 8 | 9 | 11 | 8 | 10 | 13 | 7 | 14 |
| Gentamycin | 19 | 18 | 20 | 16 | 14 | 20 | 20 | 18 | 15 |
| EC a | BS b | Psa c | Ml d | Stw e | |
|---|---|---|---|---|---|
| Terretonin N | 8 | – | 7 | – | 15 |
| Gentamycin | 19 | 18 | 20 | 16 | 14 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hamed, A.; Abdel-Razek, A.S.; Frese, M.; Stammler, H.G.; El-Haddad, A.F.; Ibrahim, T.M.A.; Sewald, N.; Shaaban, M. Terretonin N: A New Meroterpenoid from Nocardiopsis sp. Molecules 2018, 23, 299. https://doi.org/10.3390/molecules23020299
Hamed A, Abdel-Razek AS, Frese M, Stammler HG, El-Haddad AF, Ibrahim TMA, Sewald N, Shaaban M. Terretonin N: A New Meroterpenoid from Nocardiopsis sp. Molecules. 2018; 23(2):299. https://doi.org/10.3390/molecules23020299
Chicago/Turabian StyleHamed, Abdelaaty, Ahmed S. Abdel-Razek, Marcel Frese, Hans Georg Stammler, Atef F. El-Haddad, Tarek M. A. Ibrahim, Norbert Sewald, and Mohamed Shaaban. 2018. "Terretonin N: A New Meroterpenoid from Nocardiopsis sp." Molecules 23, no. 2: 299. https://doi.org/10.3390/molecules23020299
APA StyleHamed, A., Abdel-Razek, A. S., Frese, M., Stammler, H. G., El-Haddad, A. F., Ibrahim, T. M. A., Sewald, N., & Shaaban, M. (2018). Terretonin N: A New Meroterpenoid from Nocardiopsis sp. Molecules, 23(2), 299. https://doi.org/10.3390/molecules23020299








