Abstract
The complete chloroplast (cp) genome of Lonicera japonica, a common ornamental and medicinal plant in North America and East Asia, was sequenced and analyzed. The length of the L. japonica cp genome is 155,078 bp, contains a pair of inverted repeat regions (IRa and IRb), of 23,774 bp each, as well as large (LSC, 88,858 bp) and small (SSC, 18,672 bp) single-copy regions. A total of 129 genes were identified in the cp genome, 16 of which were duplicated within the IR regions. Relative to other plant cp genomes, the L. japonica cp genome had a unique rearrangement between trnI-CAU and trnN-GUU. In L. japonica cpDNA, rps19, rpl2, and rpl23 move to the LSC region, from the IR region. The ycf1 pesudogene in the IR region is lost, and only one copy locates in the SSC region. Comparative cp DNA sequence analyses of L. japonica with other cp genomes reveal that the gene order, and the gene and intron contents, are slightly different. The introns in ycf2 and rps18 genes are found for the first time. Four genes (clpP, petB, petD, and rpl16) lost introns. However, its genome structure, GC content, and codon usage were similar to those of typical angiosperm cp genomes. All preferred synonymous codons were found to use codons ending with A/T. The AT-rich sequences were less abundant in the coding regions than in the non-coding ones. A phylogenetic analysis based on 71 protein-coding genes supported the idea that L. japonica is a sister of the Araliaceae species. This study identified unique characteristics of the L. japonica cp genome that contribute to our understanding of the cpDNA evolution. It offers valuable information for the phylogenetic and specific barcoding of this medicinal plant.
1. Introduction
L. japonica is a sprawling and twining liana of the genus Lonicera in Caprifoliaceae and Dipsacales. It is native to eastern Asia and was cultivated as medicinal plant with great economic value. The Lonicera genus has almost 100 species in China, and half of them have medicinal effects, including L. japonica, L. macranthoides [1], L. similis [2], L. fulvotomentosa [3], and L. hypoglauca [4]. To date, more than 140 compounds have been isolated and identified from L. japonica [5]. The dried flowers, buds, and leaves of L. japonica are widely used with other Chinese medicines in the treatment of epidemic febrile and infectious diseases, such as SARS and avian influenza [6].
Centuries ago, L. japonica was introduced to North America, South America, and Oceania as an ornamental plant [7]. Now, it is well-known in America as a horticultural plant with wind breaker and sand-fixation properties. L. japonica does not wither, even in winter, where mean temperatures are at least −1 °C, and is very effective for ecological protection in China.
However, L. japonica is the only species within the genus used as traditional Chinese Medicine, and species identification of L. japonica, from other Lonicera species, is quite difficult. Molecular barcodes based on the cp genome have shown great potential for species discrimination, especially between closely related taxa [8]. The complete chloroplast genome sequence might enhance our ability to explore reliable barcoding for accurate plant identification, at both the species and population levels [9].
In higher plants, photosynthesis occurs in the cp, to provide the essential energy needed for plant growth and survival. New leaves of L. japonica have higher photosynthetic rates than other Lonicera species, whether they are under the forest canopy or in the open [7]. The annual carbon gain for Japanese honeysuckle was much greater in different light environments [10]. However, the molecular mechanism of photosynthetic adaptability of L. japonica is still beyond our outstanding. The lack of the cp genome of L. japonica has become a bottleneck for investigating whether there are links between L. japonica’s high level adaptability and photosynthetic adaptability, as well as chloroplast function. With rapid advances in sequencing technologies, Herbgenomics provides an effective tool to uncover the genetic information of herbs and to clarify their molecular mechanisms in related biological responses [11,12].
Although the transcriptome sequences of L. japonica have been previously reported [13], this study is the first to report its cp genome sequence. Comparative analyses among cp genomes of Apiales species revealed changes in the genome sizes, as well as the loss of genes and introns. Our data will help to identify the genetic and evolutionary mechanisms required for an in-depth study of L. japonica, and will be beneficial for DNA barcoding studies in Lonicera.
2. Results and Discussion
2.1. Characteristics of L. japonica cpDNA
The library was constructed from the cpDNA of L. japonica leaves with the 454 GS FLX Titanium platform, using the manufacturer’s manual. A total of 22,185 reads were obtained, with an average length of 412 bp, yielding approximately 58× coverage of the cp genome. The complete cp genome of L. japonica is 155,078 bp in length (Accession No. KJ170923). Its genome exhibits a typical quadripartite structure that consists of a pair of IR regions (23,774 bp), separated by the LSC (88,858 bp) and SSC (18,672 bp) regions (Table 1, Figure 1).

Table 1.
Base composition in the L. japonica chloroplast genome.
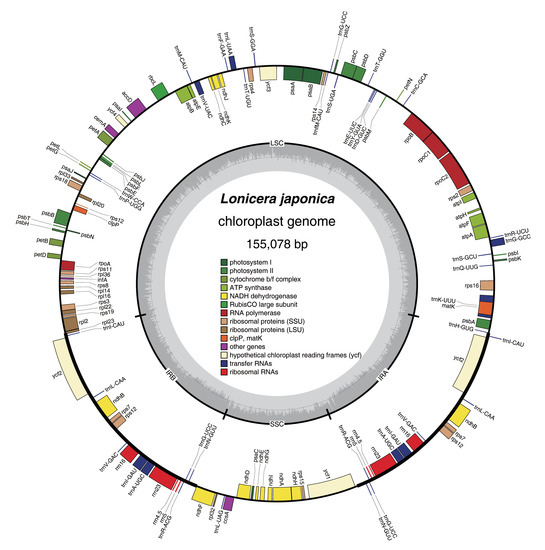
Figure 1.
Map of the L. japonica chloroplast genome. Genes drawn inside the circle are transcribed in a clockwise direction, whereas those outside the circle are transcribed in a counterclockwise direction. Genes belonging to the same functional groups have the same colors.
A total of 79 protein-coding genes, 30 tRNA genes, and 4 rRNA genes, were annotated (Table S1). These genes have been retained in several angiosperms [14,15,16,17,18]. Among these genes, eight tRNA genes, four rRNA genes, and four protein-coding genes, were duplicated in the IR regions. The LSC region contains 63 protein-coding and 22 tRNA genes, whereas the SSC region contains one tRNA gene and 12 protein-coding genes.
The majority (52%) of the L. japonica cp genome is composed of non-coding regions, including introns, intergenic spacers, and pseudogenes. The overall GC and AT content of the L. japonica cp genome was 38.6% and 61.4%, respectively. The AT content of the LSC, SSC, and IR regions was 62.9%, 66.6% and 56.5%, respectively (Table 1). Within the protein-coding regions (CDS), the AT content of the first, second, and third codon positions, is 53.7%, 61.6% and 68.8%, respectively (Table 1). The bias toward a higher AT representation at the third codon position is generally found in plant cp genomes; this bias is used to distinguish cpDNA from nuclear DNA and mitochondrial DNA [14,15,16,19]. Based on the sequences of the protein-coding and tRNA genes, the frequency of codon usage was deduced for the L. japonica cp genome and summarized in Table 2. The high AT content at the third codon position reflects a codon usage bias for A or T. The codon usage frequencies of stop codons are similarly biased to A or T at the second and third codon positions. A total of 2692 codons (10.6%) encoded for leucine, whereas 273 (1.1%) encoded for cysteine, which are the most and least prevalent amino acids, respectively. A general excess of A- and U-ending codons was noted. Except for trL-CAA, all of the types of preferred synonymous codons (RSCU > 1) ended with A or U (Table 2).

Table 2.
Codon–anticodon recognition patterns and codon usage of the L. japonica chloroplast genome.
2.2. Intron Gain and Loss
Advances in phylogenetic research have demonstrated that cp genome evolution includes nucleotide substitutions and structural changes [20,21]. A few examples of these changes, including gene or intron losses, have been found in cp genomes [22,23,24,25,26,27]. Previous study has demonstrated that introns had important roles in alternative splicing, and it has been demonstrated that the introns can significantly stabilize the transcripts in some eukaryotic lineages [28]. Additionally, orthologous genes are also believed to have lost or gained introns throughout evolution. To provide more information for further study on cp genome evolution in Caprifoliaceae and the potential functional change from variations of intron gain and loss, we analyzed the cp genome of L. japonica.
In total, there are 16 intron-containing genes, 14 of which contain one intron, and two of which (rps18 and ycf3) contain two introns (Table 3). The ribosomal protein S18 is essential for plastid translation in plant development [29]. Ycf3 is required for the stable accumulation of the photosystem I complex. In green alga Chlamydomonas reinhardtii, ycf3 and rps18 genes belong to the rps9-ycf4-ycf3-rps18 polycistronic transcriptional unit. In land plants, ycf3 and rps18 are found in different clusters [30]. The presence of two introns in the rps18 gene in the L. japonica cp genome, is rare. Similarly, the intron in ycf3 was not previously mentioned in other cp genomes. The intron gain in several L. japonica cp genes is first reported, and the intron gain in rps18 and ycf3 of L. japonica may be useful for further studies on the mechanism of photosynthesis evolution.

Table 3.
Genes with introns in the L. japonica chloroplast genome, including the exon and intron length.
Compared to other cp genes, the introns in the clpP, petB, petD, and rpl16 genes, were lost in the L. japonica cp genome. The rpl16 intron is a highly stable component of angiosperm cp genomes; this intron is absent from very few taxa, namely the Geraniaceae, Goodeniaceae, and Plumbaginaceae families [31]. Similarly, previous studies have shown that introns are also absent in the clpP gene of the Jasminum nudiflorum cp genome [22,31]. The intron loss of petB, petD, and rpl16 was first found in the lineages of Asterids. Introns are important in the regulation of gene expression. They can enhance the gene expression level, on the special position, in the specific time [16]. Some introns are known to enhance, or are required for, normal levels of mRNA transcription, processing, and transport. Several unicellular eukaryotes appear to be under selection pressure to lose introns. However, no studies between intron loss and gene expression, using transcriptome data from L. japonica, have been published.
The intron density in eukaryote genomes varies by more than three orders of magnitude. Therefore, extensive intron gain and/or intron loss must have occurred during evolution. A common partial explanation for the range of intron densities, is the stochastic accumulation of introns in large eukaryote genomes during their evolution from an intron-poor ancestor. We still need more experimental information to reveal whether the variation of the introns in the L. japonica cp genome is related to the adaptability to stress.
2.3. Comparison with Other cp Genomes in the Order Apiales
Both L. japonica and Kolkwitzia amabilis [32] belong to the order Dipsacales (Figure S1). Apiales and Dipsacales are both Asterids. Some cp genomes in the Apiales clade have been reported, such as those of the Eleutherococcus senticosus [16], Daucus carota [17], and Panax ginseng [18] chloroplast. These representative cp genome sequences of Apiales were selected for comparison with those of L. japonica and K. amabilis. The overall sequence identification of the cp genomes was plotted using mVISTA, with the annotation of L. japonica as a reference (Figure 2). The length of the LSC and IR regions was the main difference between genomes (Table S2). The comparison showed that the two IR regions were less divergent than the LSC and SSC regions. Coding and non-coding regions were present, and the most divergent regions among the four cp genomes were localized to the intergenic spacers. These highly divergent regions were included in the alignment.
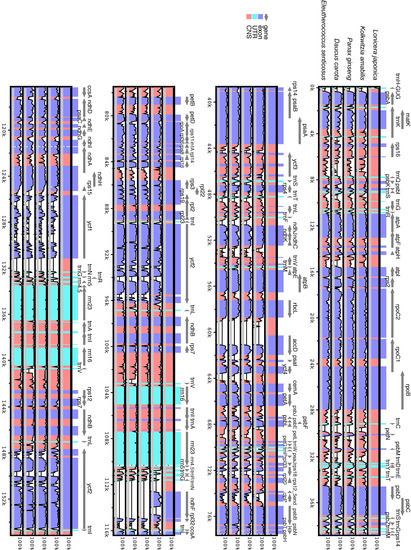
Figure 2.
Comparison of four chloroplast genomes using mVISTA program. Grey arrows and thick black lines above the alignment indicate the orientation of genes and the position of the IR regions, respectively. Purple bars represent exons, blue ones represent UTRs, and pink ones represent non-coding sequences (CNS). A cut-off of 70% identity was used for the plots. The Y-scale axis represents the percent identity within 50%–100%. Genome regions are color-coded as either protein-coding exons, rRNAs, tRNAs, or conserved noncoding sequences (CNS).
2.4. IR Contraction in the L. japonica cp Genome
The contraction and expansion at the borders of the IR regions are common evolutionary events and represent the main reasons for the size variation of cp genomes [33]. The ends of the IRa and IRb regions, as well as the gene length, differ among plant lineages. Detailed comparisons of IR-SSC and IR-LSC boundaries among the five Asterid cp genomes, are presented in Figure 3. Similar to the N. tabacum [34] and Penthorum chinense [35], the rps19 gene existed in the LSC region. However, some unique structural differences exist: the contraction of the inverted repeat region, to exclude the rpl2, rpl23, and ycf1 genes, typically exists in the IR region in other angiosperm cp genomes; however, they were not excluded from the LSC region in L. japonica. Moreover, rps19, rpl2, rpl23, and ycf1 pseudogenes were found in L. japonica. In Spirogyra maxima, trnI-rpl23-rpl2-rps19- is a large operon of angiosperm chloroplast genomes. The rpl23 gene cluster of Spirogyra contains a distinct eubacterial promoter sequence, upstream of rpl23, which is the first gene of the green algal rpl23 gene cluster. This sequence is completely absent in angiosperms, but is present in non-flowering plants. The results imply that, in the rpl23 gene cluster, early charophytes had at least two promoters, one which was upstream of trnI, and another which was upstream of rpl23, which partially or completely lost its function in land plants [36]. The IRb/SSC border is generally located between the ycf1 pseudogene and the ndhF gene. However, the ycf1 pseudogene was absent in L. japonica. Ycf1 pseudogenes have been proved useful for analyzing cp genome variation in higher plants and algae, even though their function is not thoroughly known. Ycf1 and ycf2 are essential for plant survival [37]. A combined analysis of the chloroplast genome and transcriptome of Deschampsia antarctica Desv, indicated that the rps19 gene was one of the most abundant transcripts in the chloroplast’s genome [38]. The portion of the ndhF gene located in the IRb region was 8 bp long. The rps19 and ycf1 genes were not found in the IR region, and the two pseudogenes were absent in the cp genome. These features are reported for Asterid plants for the first time.
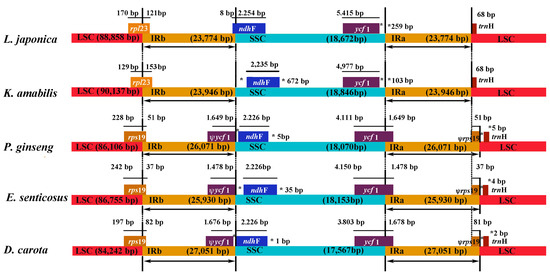
Figure 3.
Comparison of the borders of the LSC, SSC, and IR regions among five chloroplast genomes. Ψ: pseudogenes, *: the distance from the edge.
The cp genome size displayed among the examined Apiales species was compared. The length of the IR (23,774 bp) in L. japonica was 3278 bp smaller than that of D. carota, 2157 bp smaller than that of E. senticosus, and 2298 bp smaller than that of P. ginseng. These differences could be attributed to the loss of the rpl2 and ycf15 genes, as well as the ycf1 and rps19 pseudogenes in L. japonica IR regions. However, the length of the whole genome was not significantly different among the five Asterid cp genomes. The genome of L. japonica (155,078 bp) was 834 bp smaller than that of D. carota, 1691 bp smaller than that of E. senticosus, and 1241 bp smaller than that of P. ginseng. Non-functional DNA was likewise rapidly deleted, resulting in the failure of pseudogenes to accumulate, despite the high rates of pseudogenes.
2.5. Phylogenetic Analysis
The gene content of cpDNA is highly conserved among most land plants. The cp genome sequence is a useful resource for studying the taxonomic status of the genus Lonicera in the angiosperm clade, and for analyzing evolutionary relationships within the family [22]. To obtain a reasonable phylogenetic status of Lonicera, we performed multiple sequence alignments of protein coding genes, from a variety of plant plastomes. A total of 15 complete cp genomes represented six families, within five orders. Phylogenetic analysis was performed on a 71-gene data matrix, using MP and ML methods. MP analysis resulted in a single tree with a length of 17,973, a consistency index (CI) of 0.8080, and a retention index (RI) of 0.8285 (Figure 4). Bootstrap analysis showed that 12 out of the 13 nodes had bootstrap values >95%.
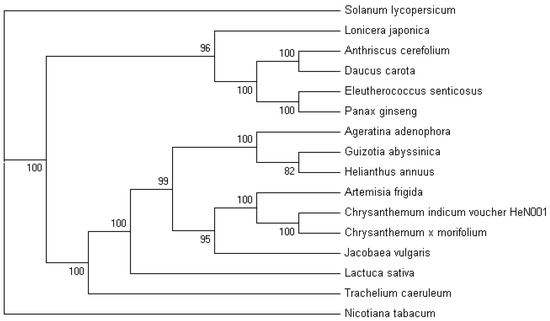
Figure 4.
MP phylogenetic tree of the Apiales clade based on 71 protein-coding genes. The MP tree has a length of 17,973, with a consistency index of 0.8080 and a retention index of 0.8285. Numbers above each node are the bootstrap support values. Solanum lycopersicum and Nicotiana tabacum were set as the outgroups.
3. Materials and Methods
3.1. DNA Sequencing, Genome Assembly, and Validation
Fresh L. japonica leaves were collected from cultivated fields in Zhengcheng, Shandong Province, China. The samples used in this study were obtained from a local company (Jintai Yaoye Co., Ltd., Linyi, China). The total chloroplast DNA (cpDNA) was extracted from approximately 100 g of leaves via a sucrose gradient centrifugation method that was improved by Li et al. [39]. The cpDNA concentration for each sample was estimated by measuring A260 with an ND-2000 spectrometer (Nanodrop Technologies, Wilmington, DE, USA), whereas visual approximation was performed using gel electrophoresis. Pure cpDNA was used to construct shotgun libraries with the 454 GS FLX Titanium platform, according to the manufacturer’s instructions. This results in approximately 58× coverage of the cp genome. The obtained Sff-file was pre-processed, including the trimming of low-quality (Q < 20) and short (L < 50 bp) reads. The trimmed and cleaned reads were used for sequence assembly with the GS FLX De Novo Assembler Software (Newbler V2.6). To verify the assembly, four junction regions between the IR regions and LSC/SSC were confirmed by PCR amplifications and Sanger sequencing, with the primers listed in Table S3. The final cp genome sequence of L. japonica was then submitted to GenBank (Accession Number: KJ170923).
3.2. Gene Annotation and Sequence Analyses
Gene annotation was performed using BLAST and DOGMA [40]. The tRNA genes were identified using DOGMA and tRNAscanSE [41]. The circular cp genome map was drawn using the OGDRAW program [42]. To analyze the characteristics of the variations in synonymous codon usage, by neglecting the influence of amino acid composition, the relative synonymous codon usage values (RSCU), codon usage, and AT content, were determined using MEGA5.2 [43].
3.3. Genome Comparison
MUMmer [44] was used to perform pairwise cp genomic alignment. The mVISTA [45] program in the Shuffle-LAGAN mode [46], was used to compare the cp genome of L. japonica with the cp genomes of K. amabilis, P. ginseng, D. carota, and E. senticosus (KT966716, AY582139, DQ898156 and NC_016430), with the annotation of L. japonica as the reference. REPuter [47] was used to visualize the forward and inverted repeats.
3.4. Phylogenetic Analysis
A total of 15 complete cp genome sequences were downloaded from the NCBI Organelle Genome Resources database. For the phylogenetic analysis, a set of 71 protein-coding genes that were common in the 16 analyzed genomes, was used. Maximum parsimony (MP) analysis was performed with PAUP*4.0b10 [48], using a heuristic search combined with the random addition of 1000 replicates and tree bisection-reconnection (TBR) branch swapping, in the Multrees option. Bootstrap analysis was also performed with 1000 replicates and TBR branch swapping. Solanum lycopersicum and Nicotiana tabacum were set as outgroups.
4. Conclusions
High-throughput pyrosequencing technology was used to describe the completely sequenced L. japonica cp genome, which is a very important medicinal plant in East Asia. Compared to the cp genomes of three Apiales species, the cp genome of L. japonica has a relatively small size. Several genes were absent in the IR region, including the rps19, rpl2, and ycf1 pseudogenes. This absence may be attributed to the obvious contraction of the IR region in L. japonica. Phylogenetic relationships among 15 angiosperms strongly supported the known classification of L. japonica. The data presented in this study can facilitate the biological identification of this important medicinal plant. Our data reveal that L. japonica cpDNA possesses several unique features that contribute to our current understanding of cpDNA evolution in seed plants. Additionally, other Dipsacales plastomes need to be sequenced to determine whether the atypical characteristics of L. japonica cpDNA are shared by all Dipsacales species, or if these characteristics are the unusual genomic features of a very unique plant.
Supplementary Materials
Supplementary materials can be accessed at: http://www.mdpi.com/1420-3049/22/2/249/s1.
Acknowledgments
The authors would like to thank Jingyuan Song and Hongmei Luo for their valuable comments and suggestions. This work was supported by grants from the National Natural Science Foundation of China (No. 81102761).
Author Contributions
X.L. and S.C. conceived and designed the experiments; L.H., Z.S. and X.L. performed the experiments; L.H., J.Q. and X.X. analyzed the data; L.H. wrote the paper; X.L. revised the paper.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Wang, J.; Zhao, X.Z.; Qi, Q.; Tao, L.; Zhao, Q.; Mu, R.; Gu, H.Y.; Wang, M.; Feng, X.; Guo, Q.L. Macranthoside B, a hederagenin saponin extracted from Lonicera macranthoides and its anti-tumor activities in vitro and in vivo. Food Chem. Toxicol. 2009, 47, 1716–1721. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.M.; Wang, T.Z.; Wang, Z.X. Studies on chemical constituents in dried buds of Lonicera similis Hemsl. Zhongguo Zhong Yao Za Zhi 2001, 26, 45–47. [Google Scholar] [PubMed]
- Tang, D.; Li, H.J.; Li, P.; Wen, X.D.; Qian, Z.M. Interaction of bioactive components caffeoylquinic Acid derivatives in Chinese medicines with bovine serum albumin. Chem. Pharm. Bull. 2008, 56, 360–365. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.Y.; Tseng, C.P.; Tsai, K.C.; Lin, C.F.; Wen, C.Y.; Tsay, H.S.; Sakamoto, N.; Tseng, C.H.; Cheng, J.C. Bioactivity-guided screening identifies pheophytin a as a potent anti-hepatitis C virus compound from Lonicera hypoglauca Miq. Biochem. Biophys. Res. Commun. 2009, 385, 230–235. [Google Scholar] [CrossRef] [PubMed]
- Shang, X.F.; Pan, H.; Li, M.X.; Miao, X.L.; Ding, H. Lonicera japonica Thunb.: Ethnopharmacology, phytochemistry and pharmacology of an important traditional Chinese medicine. J. Ethnopharmacol. 2011, 138, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Muluye, R.A.; Bian, Y.H.; Alemu, P.N. Anti-inflammatory and Antimicrobial effects of Heat-Clearing Chinese herbs: A Current Review. J. Tradit. Complement. Med. 2014, 4, 93–98. [Google Scholar] [CrossRef] [PubMed]
- Schierenbeck, K.A. Japanese Honeysuckle (Lonicera japonica) as an invasive species; history, ecology, and context. Crit. Rev. Plant Sci. 2004, 23, 391–400. [Google Scholar] [CrossRef]
- Yao, H.; Song, J.Y.; Liu, C.; Luo, K.; Han, J.P.; Li, Y.; Pang, X.H.; Xu, H.X.; Zhu, Y.J.; Xiao, P.G.; et al. Use of ITS2 region as the universal DNA barcode for plants and animals. PLoS ONE 2010, 5, e13102. [Google Scholar] [CrossRef] [PubMed]
- Li, X.W.; Yang, Y.; Henry, R.J.; Rossetto, M.; Wang, Y.T.; Chen, S.L. Plant DNA barcoding: From gene to genome. Biol. Rev. 2015, 90, 157–166. [Google Scholar] [CrossRef] [PubMed]
- Schierenbeck, K.A.; Marshall, J.D. Seasonal and diurnal patterns of photosynthetic gas exchange for Lonicera sempervirens and L. japonica (Caprifoliaceae). Am. J. Bot. 1993, 80, 1292–1299. [Google Scholar] [CrossRef]
- Chen, S.L.; Song, J.Y. Herbgenomics. China J. Chin. Mater. Med. 2016, 41, 3881–3889. [Google Scholar]
- Chen, S.L.; Song, J.Y.; Sun, C.; Xu, J.; Zhu, Y.J.; Verpoorte, R.; Fan, T.P. Herbal genomics: Examining the biology of traditional medicines. Science 2015, 347, S27–S29. [Google Scholar]
- He, L.; Xu, X.L.; Li, Y.; Li, C.F.; Zhu, Y.J.; Yan, H.X.; Sun, Z.Y.; Sun, C.; Song, J.Y.; Bi, Y.A.; et al. Transcriptome analysis of buds and leaves using 454 pyrosequencing to discover genes associated with the biosynthesis of active ingredients in Lonicera japonica Thunb. PLoS ONE 2013, 8, e62922. [Google Scholar] [CrossRef] [PubMed]
- Nie, X.; Lv, S.; Zhang, Y.; Du, X.; Wang, L.; Biradar, S.S.; Tan, X.; Wan, F.; Weining, S. Complete chloroplast genome sequence of a major invasive species, crofton weed (Ageratina adenophora). PLoS ONE 2012, 7, e36869. [Google Scholar] [CrossRef] [PubMed]
- Yi, D.K.; Kim, K.J. Complete chloroplast genome sequences of important oilseed crop Sesamum indicum L. PLoS ONE 2012, 7, e35872. [Google Scholar] [CrossRef] [PubMed]
- Yi, D.K.; Lee, H.L.; Sun, B.Y.; Chung, M.Y.; Kim, K.J. The complete chloroplast DNA sequence of Eleutherococcus senticosus (Araliaceae); comparative evolutionary analyses with other three asterids. Mol. Cells 2012, 33, 497–508. [Google Scholar] [CrossRef] [PubMed]
- Ruhlman, T.; Lee, S.B.; Jansen, R.K.; Hostetler, J.B.; Tallon, L.J.; Town, C.D.; Daniell, H. Complete plastid genome sequence of Daucus carota: Implications for biotechnology and phylogeny of angiosperms. BMC Genom. 2006, 7, 222. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.J.; Lee, H.L. Complete chloroplast genome sequences from Korean ginseng (Panax schinseng Nees) and comparative analysis of sequence evolution among 17 vascular plants. DNA Res. 2004, 11, 247–261. [Google Scholar] [CrossRef] [PubMed]
- Clegg, M.T.; Gaut, B.S.; Learn, G.H.; Morton, B.R. Rates and patterns of chloroplast DNA evolution. Proc. Natl. Acad. Sci. USA 1994, 91, 6795–6801. [Google Scholar] [CrossRef] [PubMed]
- Yue, F.; Cui, L.; de Pamphilis, C.W.; Moret, B.M.; Tang, J. Gene rearrangement analysis and ancestral order inference from chloroplast genomes with inverted repeat. BMC Genom. 2008, 9, S25. [Google Scholar] [CrossRef] [PubMed]
- Haberle, R.C.; Fourcade, H.M.; Boore, J.L.; Jansen, R.K. Extensive rearrangements in the chloroplast genome of Trachelium caeruleum are associated with repeats and tRNA genes. J. Mol. Evol. 2008, 66, 350–361. [Google Scholar] [CrossRef] [PubMed]
- Jansen, R.K.; Cai, Z.Q.; Raubeson, L.A.; Daniell, H.; dePamphilis, C.W.; Leebeans-Mack, J.; Müller, K.F.; Guisinger-Bellian, M.; Haberle, R.C.; Hansen, A.K.; et al. Analysis of 81 genes from 64 plastid genomes resolves relationships in angiosperms and identifies genome-scale evolutionary patterns. Proc. Natl. Acad. Sci. USA 2007, 104, 19369–19374. [Google Scholar] [CrossRef] [PubMed]
- Ueda, M.; Fujimoto, M.; Arimura, S.; Murata, J.; Tsutsumi, N.; Kadowaki, K. Loss of the rpl32 gene from the chloroplast genome and subsequent acquisition of a preexisting transit peptide within the nuclear gene in Populus. Gene 2007, 402, 51–56. [Google Scholar] [CrossRef] [PubMed]
- Downie, S.R.; Llanas, E.; KatzDownie, D.S. Multiple independent losses of the rpoC1 intron in angiosperm chloroplast DNA’s. Syst. Bot. 1996, 21, 135–151. [Google Scholar] [CrossRef]
- Downie, S.R.; Olmstead, R.G.; Zurawski, G.; Soltis, D.E.; Soltis, P.S. Six independent losses of the chloroplast DNA rpl2 intron in dicotyledons: Molecular and phylogenetic implications. Evolution 1991, 45, 1245–1259. [Google Scholar] [CrossRef]
- Graveley, B.R. Alternative splicing: Increasing diversity in the proteomic world. Trends Genet. 2001, 17, 100–107. [Google Scholar] [CrossRef]
- Guisinger, M.M.; Chumley, T.W.; Kuehl, J.V.; Boore, J.L.; Jansen, R.K. Implications of the plastid genome sequence of Typha (Typhaceae, Poales) for understanding genome evolution in Poaceae. J. Mol. Evol. 2010, 70, 149–166. [Google Scholar] [CrossRef] [PubMed]
- Daniell, H.; Wurdack, K.J.; Kanagaraj, A.; Lee, S.B.; Saski, C.; Jansen, R.K. The complete nucleotide sequence of the cassava (Manihot esculenta) chloroplast genome and the evolution of atpF in Malpighiales: RNA editing and multiple losses of a group II intron. Theor. Appl. Genet. 2008, 116, 723–737. [Google Scholar] [CrossRef] [PubMed]
- Rogalski, M.; Ruf, S.; Bock, R. Tobacco plastid ribosomal protein S18 is essential for cell survival. Nucleic Acids Res. 2006, 34, 4537–4545. [Google Scholar] [CrossRef] [PubMed]
- Boudreau, E.; Takahashi, Y.; Lemieux, C.; Turmel, M.; Rochaix, J.D. The chloroplast ycf3 and ycf4 open reading frames of Chlamydomonas reinhardtii are required for the accumulation of the photosystem I complex. EMBO J. 1997, 16, 6095–6104. [Google Scholar] [CrossRef] [PubMed]
- Chorev, M.; Carmel, L. The function of introns. Front. Genet. 2012, 3, 55. [Google Scholar] [CrossRef] [PubMed]
- Drescher, A.; Ruf, S.; Calsa, T.; Carrer, H.; Bock, R. The two largest chloroplast genome-encoded open reading frames of higher plants are essential genes. Plant J. 2000, 22, 97–104. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.L.; Jansen, R.K.; Chumley, T.W.; Kim, K.J. Gene relocations within chloroplast genomes of Jasminum and Menodora (Oleaceae) are due to multiple, overlapping inversions. Mol. Biol. Evol. 2007, 24, 1161–1180. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.J.; Cheng, C.L.; Chang, C.C.; Wu, C.L.; Su, T.M.; Chaw, S.M. Dynamics and evolution of the inverted repeat-large single copy junctions in the chloroplast genomes of monocots. BMC Evol. Biol. 2008, 8, 36. [Google Scholar] [CrossRef] [PubMed]
- Shinozaki, K.; Ohme, M.; Tanaka, M.; Wakasugi, T.; Hayashida, N.; Matsubayashi, T.; Zaita, N.; Chunwongse, J.; Obokata, J.; Yamaguchi-Shinozaki, K.; et al. The complete nucleotide sequence of the tobacco chloroplast genome: Its gene organization and expression. EMBO J. 1986, 5, 2043–2049. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.P.; Xu, C.; Cheng, T.; Zhou, S.L. Complete chloroplast genome of Sedum sarmentosum and chloroplast genome evolution in Saxifragales. PLoS ONE 2013, 8, e77965. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Manhart, J.R. The chloroplast rpl23 gene cluster of Spirogyra maxima (Charophyceae) shares many similarities with the Angiosperm rpl23 Operon. Algae 2002, 17, 59–68. [Google Scholar] [CrossRef]
- Lee, J.; Kang, Y.; Shin, S.C.; Park, H.; Lee, H. Combined analysis of the chloroplast genome and transcriptome of the Antarctic vascular plant Deschampsia antarctica Desv. PLoS ONE 2014, 9, e92501. [Google Scholar] [CrossRef] [PubMed]
- Li, X.W.; Hu, Z.G.; Lin, X.H.; Li, Q.; Gao, H.H.; Luo, G.A.; Chen, S.L. High-throughput pyrosequencing of the complete chloroplast genome of Magnolia officinalis and its application in species identification. Acta Pharm. Sin. 2012, 47, 124–130. [Google Scholar]
- Wyman, S.K.; Jansen, R.K.; Boore, J.L. Automatic annotation of organellar genomes with DOGMA. Bioinformatics 2004, 20, 3252–3255. [Google Scholar] [CrossRef] [PubMed]
- Schattner, P.; Brooks, A.N.; Lowe, T.M. The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res. 2005, 33, W686–W689. [Google Scholar] [CrossRef] [PubMed]
- Lohse, M.; Drechsel, O.; Bock, R. Organellar Genome DRAW (OGDRAW): A tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Curr. Genet. 2007, 52, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Kurtz, S.; Phillippy, A.; Delcher, A.L.; Smoot, M.; Shumway, M.; Antonescu, C.; Salzberg, S.L. Versatile and open software for comparing large genomes. Genome Biol. 2004, 5, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Mayor, C.; Brudno, M.; Schwartz, J.R.; Poliakov, A.; Rubin, E.M.; Frazer, K.A.; Pachter, L.S.; Dubchak, I. VISTA: Visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 2000, 16, 1046–1047. [Google Scholar] [CrossRef] [PubMed]
- Frazer, K.A.; Pachter, L.; Poliakov, A.; Rubin, E.M.; Dubchak, I. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 2004, 32, W273–W279. [Google Scholar] [CrossRef] [PubMed]
- Kurtz, S.; Choudhuri, J.V.; Ohlebusch, E.; Schleiermacher, C.; Stoye, J.; Giegerich, R. REPuter: The manifold applications of repeat analysis on a genomic scale. Nucleic Acids. Res. 2001, 29, 4633–4642. [Google Scholar] [CrossRef] [PubMed]
- Swofford, D.L. PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods); Version 4.0b10; Sinauer Associates: Sunderland, MA, USA, 2003. [Google Scholar]
- Sample Availability: Not available.
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).