Abstract
The present study utilizes polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) analysis using partial plastid rbcL and mitochondrial trnC–trnP gene sequences to distinguish the six representative Pyropia species produced via mariculture in Korea. The rbcL, trnC, and trnP sequences of 15 Pyropia species from the NCBI database were aligned to determine specific restriction enzyme sites of the six Pyropia species. To confirm the presence of restriction sites of eight enzymes, PCR amplicons were digested as follows: a 556 bp fragment within the rbcL region of chloroplast DNA was confirmed in P. yezoensis using BglI, whereas Tth111I, AvaII, BsrI, and BsaAI enzymes produced fragments of 664, 271, 600, and 510 bp, respectively, from the rps11–trnG region of mitochondrial DNA in P. seriata, P. dentata, P. suborbiculata, and P. haitanensis. In the case of P. pseudolinearis, HindIII, SacII, and SphI enzymes each had two cleavage sites, at positions 174 and 825, 788 and 211, and 397 and 602 bp, respectively. All six species were successfully distinguished using these eight restriction enzymes. Therefore, we propose that PCR-RFLP analysis is an efficient tool for the potential use of distinguishing between the six Pyropia species cultivated via mariculture in Korea.
1. Introduction
Red algae (Bangiales, Rhodophyta) of the genus Pyropia, including more than 75 species, are economically important mariculture crops and are available in China, Japan, and South Korea [1]. This genus is widely used as a dried sheet product, “zicai” in China, “nori” in Japan, and “gim” in South Korea, and is commonly consumed as a food source (laver) in Asian countries [2]. Pyropia is becoming increasingly popular worldwide. The economic value of Pyropia products exported in 2016 from South Korea was 353.3 million in US dollars [3,4]. Twelve species within the genus Pyropia have been recorded in South Korea [5]. Among them, four species, P. yezoensis, P. seriata, P. dentata, and P. suborbiculata, are widely distributed and extensively cultured in the South Jeolla Province, whereas P. pseudolinearis is distributed in Ulreung Island and its export accounts for about 80% of the total production of Pyropia in South Korea [3]. P. haitanensis is widely distributed and extensively cultured in the Fujian, Zhejiang, and Guangdong Provinces of China, and its export accounts for about 75% of the total production of Pyropia in China [6]. Recently, P. haitanensis has been utilized in Korean mariculture industries to develop improved cultivars that are optimized for the warm temperature zones typically used in Pyropia breeding.
The polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) method is based on the digestion of PCR amplicons with appropriate restriction enzymes to produce distinct polymorphic fragments used as markers for species identification [7,8]. This method is beginning to be used for identifying different Pyropia species or breeding cultivars that could presumably be applied for mariculture in warming ocean waters. Japanese researchers have revealed clear differences between native Pyropia species based on PCR-RFLP analysis of plastid, internal transcribed spacer (ITS), and mitochondrial DNA (mtDNA) [9,10,11]. Chinese and Korean researchers have mainly used this method to identify and discriminate among seaweed diseases and pest species [12,13]. In South Korea, Pyropia species are identified using morphological and anatomical features. However, morphological identification of Pyropia taxa is limited because of the temporal variation in morphological characteristics and surface contamination from symbiotic seaweed and bacteria. Therefore, a better method is required for efficient and rapid identification to distinguish among these Pyropia species in natural environments.
In this study, we utilized PCR-RFLP analysis to successfully develop species-specific markers for five Pyropia species collected around the south-west coastal regions of South Korea, and P. haitanenesis collected from sites in China.
2. Results and Discussion
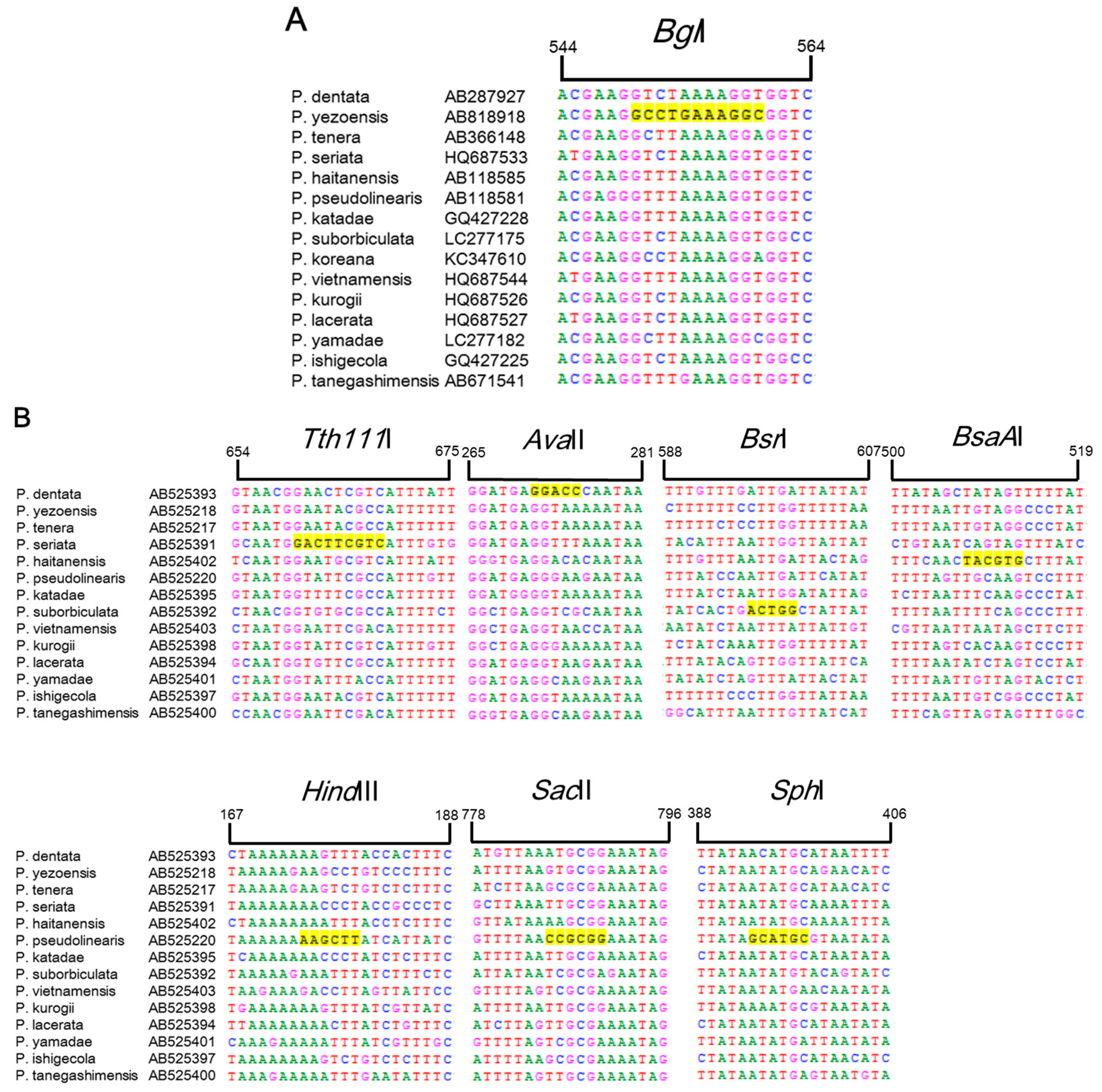
Currently, identification and discrimination between Pyropia species is primarily conducted based on classical taxonomic methods using microscopic analysis, which is limited by a lack of sufficient morphological features, time constraints, and inconsistencies between different taxonomists in South Korea [5]. Many phylogenetic analyses in other families have been based on the trnC–trnP region of mtDNA and the plastid rbcL intergenic spacer, as these two regions have high rates of variation between species [14,15]. Based on the partial sequences of rbcL and trnC–trnP for 15 Pyropia species available in the GenBank public database (National Center for Biotechnology Information (NCBI), Rockville, MD, USA), we found that six restriction enzymes were predicted to show species-specific RFLP patterns and could be used to identify the six Pyropia species using rbcL in P. yezoensis and rps11–sdh3 in P. seriata, P. pseudolinearis, P. dentate, P. suborbiculata, and P. haitanensis. As shown in Figure 1, the predicted fragment sizes for the six Pyropia species were as follows: BglI produced a 556 bp fragment from rbcL of chloroplast DNA in P. yezoensis, whereas Tth111I produced a 664 bp fragment from P. seriata, AvaII produced a 271 bp fragment from P. dentata, BsrI produced a 600 bp fragment from P. suborbiculata, and BsaAI produced a 510 bp fragment from P. haitanensis in the rps11–trnG region of mtDNA. In the case of P. pseudolinearis, HindIII, SacI, and SphI produced one cleavage site in rps11–trnG at positions 174, 788 and 397 bp, respectively.
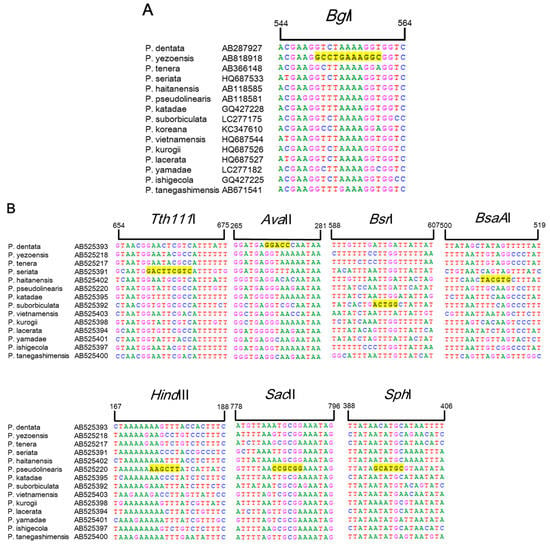
Figure 1.
Sequence alignment of rbcL (A) and rps11–trnG (B) regions from 15 Pyropia species obtained from GenBank (National Center for Biotechnology Information (NCBI), Rockville, MD, USA). The bases highlighted in yellow correspond to the Tth111I, AvaII, BsrI, BsaAI, HindIII, SacII, and SphI enzyme restriction sites.
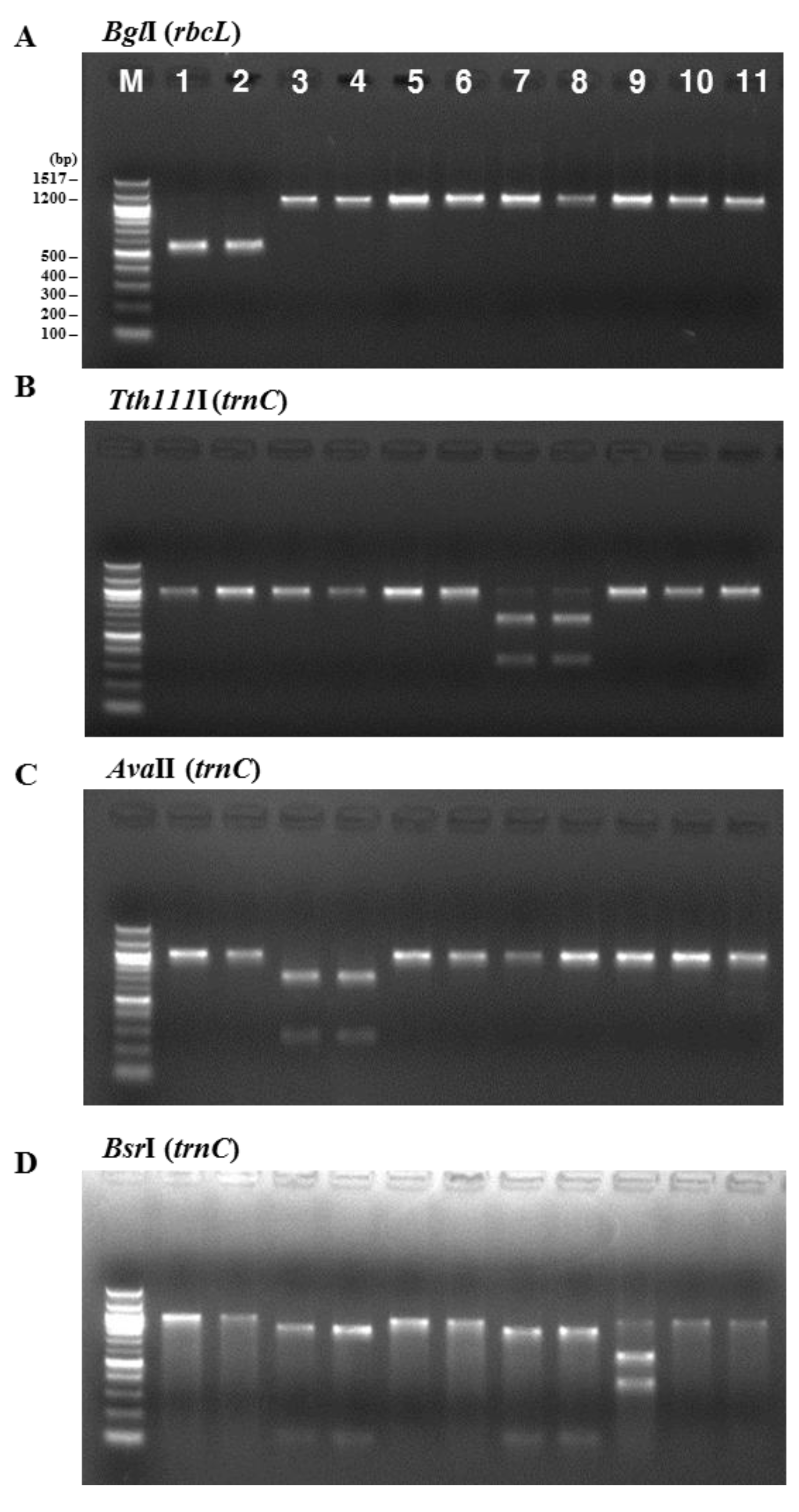
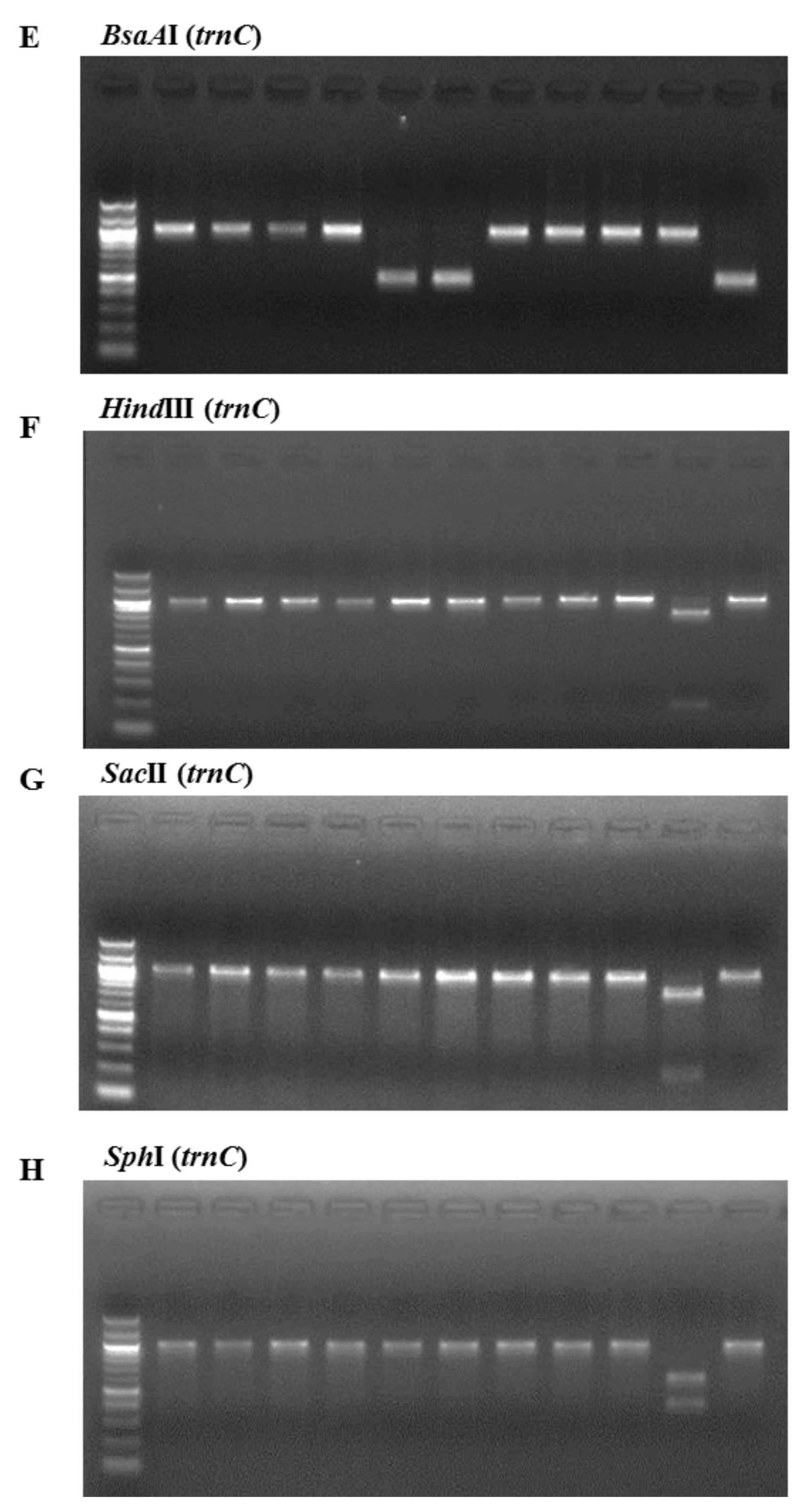
The consistency of the presence of each enzyme restriction site was confirmed by analyzing PCR-RFLPs of 11 Pyropia samples collected at coastal areas in Korea and dried laver production in China. The samples were analyzed with the above restriction enzymes. Figure 2 illustrates the band sizes obtained by digestion using BglI for rbcL and Tth111I, AvaII, BsaAI, BsrI, HindIII, SacI, and SphI for rps11–trnG. BglI produced a 556 bp fragment in the P. yezoensis samples and was used for their identification (Figure 2A). The following enzymes were used to identify P. seriata, P. dentata, and P. haitanensis: Tth111I produced two fragments (664 bp and 333 bp) in P. seriata (Figure 2B), AvaII produced two fragments (271 bp and 732 bp) in P. dentata (Figure 2C), and BsaAI produced one fragment (510 bp) in P. haitanensis (Figure 2E). BsrI was used to identify P. suborbiculata samples and produced two fragments (600 bp and 401 bp) in this species, whereas BsrI produced two fragments (119 bp and 844 bp) in both P. seriata and P. dentata (Figure 2D). Among four Pyropia samples that were origianlly identified as P. dentata using morphological and anatomical features, two samples were confirmed as P. haitanensis using our PCR-RFLP method (Figure 2E). We confirmed the ability of this PCR-RFLP assay to differentiate between P. dentata and P. haitanensis. Furthermore, three enzymes of HindIII, SacII, and SphI were used to identify P. pseudolinearis samples and produced two fragments of the 174 and 825 bp, 788 and 211 bp, and 397 and 602 bp, respectively (Figure 2F–H, Table 1).
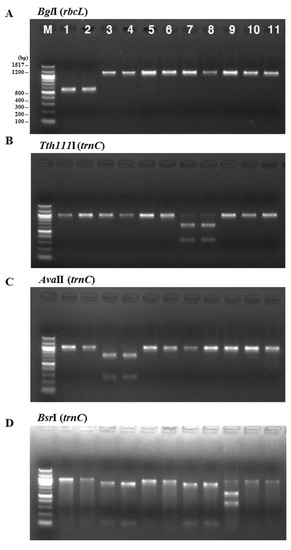
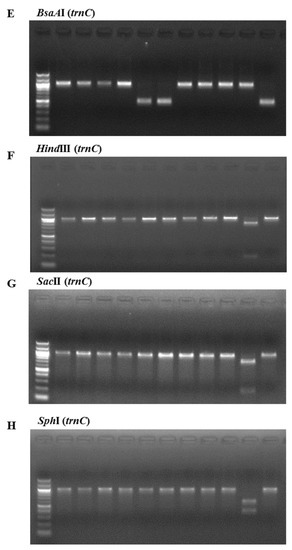
Figure 2.
PCR-RFLP profiles of (A) the partial region of rbcL from all tested species digested with BglI; and the partial region of mitochondrial DNA including trnC–trnP digested with (B) Tth111I; (C) AvaII; (D) BsrI; (E) BsaAI; (F) HindIII; (G) SacII; and (H) SphI in P. yezoensis, P. seriata, P. dentata, P. suborbiculata, P. haitanensis, and P. pseudolinearis. Numbers indicate Pyropia samples as described in the Table 2.

Table 1.
The fragment size (bps) for restriction digests of partial rbcL and rps11–trnG region in six Pyropia species.
The different restriction fragment patterns obtained for each species were analyzed to develop a discrimination method using PCR-RFLP analysis without the need for sequencing. Futhermore, our PCR-RFLP method showed that a higher sensitiviy and specificity to six Pyropia species found in Korean mariculture industires can be determined within a short time (approximately 2.5 h) compared to traditional morphological techiniques.
Most studies on PCR-RFLP analysis of Pyropia species have been conducted in Japan. Touhata et al. [16] demonstrated that the dried Pyropia products produed in China, Japan, and South Korea could be identified using PCR-RFLP analysis of the ITS-1 region. Abe et al. [9] distinguished 18 Pyropia species from Japan using restriction enzymes on mtDNA related to the atp6 gene and partial mtDNA including the trnC, rps11, sdh3, trnG, trnN, trnP, and rns genes. However, these methods are hampered by the need for interpretation of complex RFLP patterns as well as the inter-laboratory differences and are relatively difficult to apply in the field.
The six species used in this study were successfully distinguished using a combination of eight restriction enzymes (BglI, Tth111I, AvaII, BsrI, BsaAI, HindIII, SacII, and SphI ) and two plastid and mitochodrial primer sets. These primer sets were designed based on the partial plastid and mtDNA sequences of 15 Pyropia species. As our study used only one or two samples per species, further studies should use a larger sample size as well as refined Pyropia markers based on a comparison with East Asian Pyropia species.
3. Materials and Methods
3.1. Sequence Analysis
To rapidly identify the six Pyropia species, a PCR-RFLP method was designed base on the rbcL and trnC–trnP gene sequences from all the publicly available Pyropia species in the NCBI GenBank database (Table 2). These sequences were aligned by multiple sequence alignment (MSA) (Clustal Omega tool, EMBL-EBI, www.ebi.ac.uk) [17] and MEGA version 7.0 [18]. Scanning for the various restriction enzyme sites in the reference sequences was carried out using Restriction Mapper, version 3 (http://restrictionmapper.org). Restriction enzymes were selected based on the potential of showing variation between the reference sequences.

Table 2.
DNA sequences of 15 Pyropia species used for RFLP analysis.
3.2. Sample Collection and DNA Isolation
Table 3 lists the six Pyropia species collected from China and South Korea used in this study. All samples were identified based on morphological characteristics. Cultures of the conchocelis stage of each specimen were maintained at the Ocean & Fisheries Science Institute, Haenam Branch, Jeollanamdo, Korea.

Table 3.
Pyropia samples used in this study.
Prior to DNA extraction, conchocelis-stage samples of all strains (~300 mg wet weight) were washed twice with distilled water and frozen using liquid nitrogen, and then mechanically homogenized using a pestle tissue grinder. Genomic DNA was extracted from ground conchocelis-stage samples using the Qiagen DNeasy Plant Mini Kit (Qiagen, Valencia, CA, USA) according to the manufacturer’s instructions.
3.3. PCR Amplification and Restriction Digestion
PCR was performed using two 18S rDNA internal transcribed regions within rbcL; forward primer 5′-GCGAACGTTACGAATCTGGAGT-3′ and reverse primer 5′-ATGCTACTGGTACAACTTTACGTA-3′ (product size: 1107 bp), trnC–P; forward primer 5′-GTCAGTTCGAATCTGGCCCTAGTTT-3′ and reverse primer 5′-ATTCCCTTGCACCCAAAGCAAGTAC-3′ (approximately product size: 1005 bp). The PCR reactions were conducted in a 50 μL mixture containing 25 μL premix (Ex Taq Version 2.0, TaKaRa, Otsu, Japan), 50 ng genomic DNA templates, and 1 μL (10 pM) forward and reverse primers. The mixtures were denatured at 95 °C for 5 min and amplified with 40 cycles of 95 °C for 30 s, 55 °C for 20 s, and 72 °C for 30 s, with a final extension at 72 °C for 5 min. The PCR amplified products were electrophoresed on 1.5% agarose gels for 30 to 40 min.
Two μL of each PCR amplified rbcL and trnC–P product(concentration ranging from 0.6 to 1 μg/μL) was digested in 2 μL of 10× buffer, 1 unit of restriction enzyme, and 15.8 μL of distilled water in a final volume of 20 μL, followed by incubation at 37 °C for 1 h. The digested fragments were separated by electrophoresis on 1.5% agarose gels stained with ethidium bromide, and fragment patterns were visualized under UV light.
Acknowledgments
This research was supported by the Support Program for Creative Industry Institutes (Commercial Bio-technology Sophistication Platform Construction Program, R0003950) funded by the Ministry of Trade, Industry & Energy (MOTIE, Korea).
Author Contributions
Y.K., S.-J.C., and C.C. conceived and designed the experiments; Y.K. performed the experiments; S.-J.C. collected the Pyropia samples; Y.K. analyzed the data and wrote the manuscript. All authors read and approved the manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Vergés, A.; Sánchez, N.; Peteiro, C.; Polo, L.; Brodie, J. Pyropia suborbiculata (Bangiales, Rhodophyta): First records from the northeastern Atlantic and Mediterranean of this North Pacific species. Phycologia 2013, 52, 121–129. [Google Scholar] [CrossRef]
- Hallmann, A. Algae biotechnology–green cell-factories on the rise. Curr. Biotechnol. 2015, 4, 389–415. [Google Scholar] [CrossRef]
- K-Fish Information Portal. Statistical Database for Seasoned Laver Export. Available online: http://www.kfishinfo.net (accessed on 25 November 2017).
- Ock, Y.S. The research on the development steps and facing problems of Korean Japanese laver industry. J. Fish. Bus. Adm. 2011, 42, 113–130. (In Korean) [Google Scholar]
- Hwang, M.-S.; In-Gyu, L. Character analysis and numerical taxonomy of Porphyra (Bangiales, Rhodophyta) from Korea. Algae 2002, 17, 217–233. [Google Scholar] [CrossRef]
- Xie, C.; Li, B.; Xu, Y.; Ji, D.; Chen, C. Characterization of the global transcriptome for Pyropia haitanensis (Bangiales, Rhodophyta) and development of cSSR markers. BMC Genom. 2013, 14, 107. [Google Scholar] [CrossRef] [PubMed]
- Wolf, C.; Rentsch, J.; Hübner, P. PCR−RFLP analysis of mitochondrial DNA: A reliable method for species identification. J. Agric. Food Chem. 1999, 47, 1350–1355. [Google Scholar] [CrossRef] [PubMed]
- Wolf, C.; Burgener, M.; Hübner, P.; Lüthy, J. PCR-RFLP analysis of mitochondrial DNA: Differentiation of fish species. LWT-Food Sci. Technol. 2000, 33, 144–150. [Google Scholar] [CrossRef]
- Abe, M.; Kobayashi, M.; Fujiyoshi, E.; Tamaki, M.; Kikuchi, N.; Murase, N. Use of PCR-RFLP for the discrimination of Japanese Porphyra and Pyropia species (Bangiales, Rhodophyta). J. Appl. Phycol. 2013, 25, 225–232. [Google Scholar] [CrossRef]
- Kyosuke, N.; Kobiyama, A.; Aruga, Y. Confirmation of cultivated Porphyra tenera (Bangiales, Rhodophyta) by polymerase chain reaction restriction fragment length polymorphism analyses of the plastid and nuclear DNA. Phycol. Res. 2005, 53, 296–302. [Google Scholar] [CrossRef]
- Niwa, K.; Iida, S.; Kato, A.; Kawai, H.; Kikuchi, N.; Kobiyama, A.; Aruga, Y. Genetic diversity and introgression in two cultivated species (Porphyra yezoensis and Porphyra tenera) and closely related wild species of Porphyra (Bangiales, Rhodophyta). J. Phycol. 2009, 45, 493–502. [Google Scholar] [CrossRef] [PubMed]
- Xiao, J.; Li, Y.; Song, W.; Wang, Z.; Fu, M.; Li, R.; Zhang, X.; Zhu, M. Discrimination of the common macroalgae (Ulva and Blidingia) in coastal waters of Yellow Sea, northern China, based on restriction fragment-length polymorphism (RFLP) analysis. Harmful Algae 2013, 27, 130–137. [Google Scholar] [CrossRef]
- Lee, S.J.; Hwang, M.S.; Park, M.A.; Baek, J.M.; Ha, D.S.; Lee, J.E.; Lee, S.R. Molecular identification of the algal pathogen Pythium chondricola (Oomycetes) from Pyropia yezoensis (Rhodophyta) using ITS and cox1 markers. Algae 2015, 30, 217. [Google Scholar] [CrossRef]
- Shaw, J.; Lickey, E.B.; Schilling, E.E.; Small, R.L. Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: The tortoise and the hare III. Am. J. Bot. 2007, 94, 275–288. [Google Scholar] [CrossRef] [PubMed]
- Tiina, S.; George, M. Predicting plastid marker variation: Can complete plastid genomes from closely related species help? PLoS ONE 2013, 8, e82266. [Google Scholar] [CrossRef]
- Touhata, K.; Namikoshi, A.; Suzuki, T.; Iguchi, J.; Mizusawa, N.; Hara, T.; Imamura, S.; Yabu, T.; Yamashita, Y.; Yamashita, M. Origin identification of dried seaweed product “nori” by PCR–RFLP analysis of Pyropia yezoensis in the internal transcribed spacer ITS-1 region. Fish. Sci. 2013, 79, 865–875. [Google Scholar] [CrossRef]
- Li, W.; Cowley, A.; Uludag, M.; Gur, T.; McWilliam, H.; Squizzato, S.; Park, Y.M.; Buso, N.; Lopez, R. The EMBL-EBI bioinformatics web and programmatic tools framework. Nucleic Acids Res. 2015, 43, W580–W584. [Google Scholar] [CrossRef] [PubMed]
- Sudhir, K.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
Sample Availability: Genomic DNA samples are available from the authors. |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).