Thymoquinone Inhibits the Migration and Invasive Characteristics of Cervical Cancer Cells SiHa and CaSki In Vitro by Targeting Epithelial to Mesenchymal Transition Associated Transcription Factors Twist1 and Zeb1
Abstract
:1. Introduction
2. Results
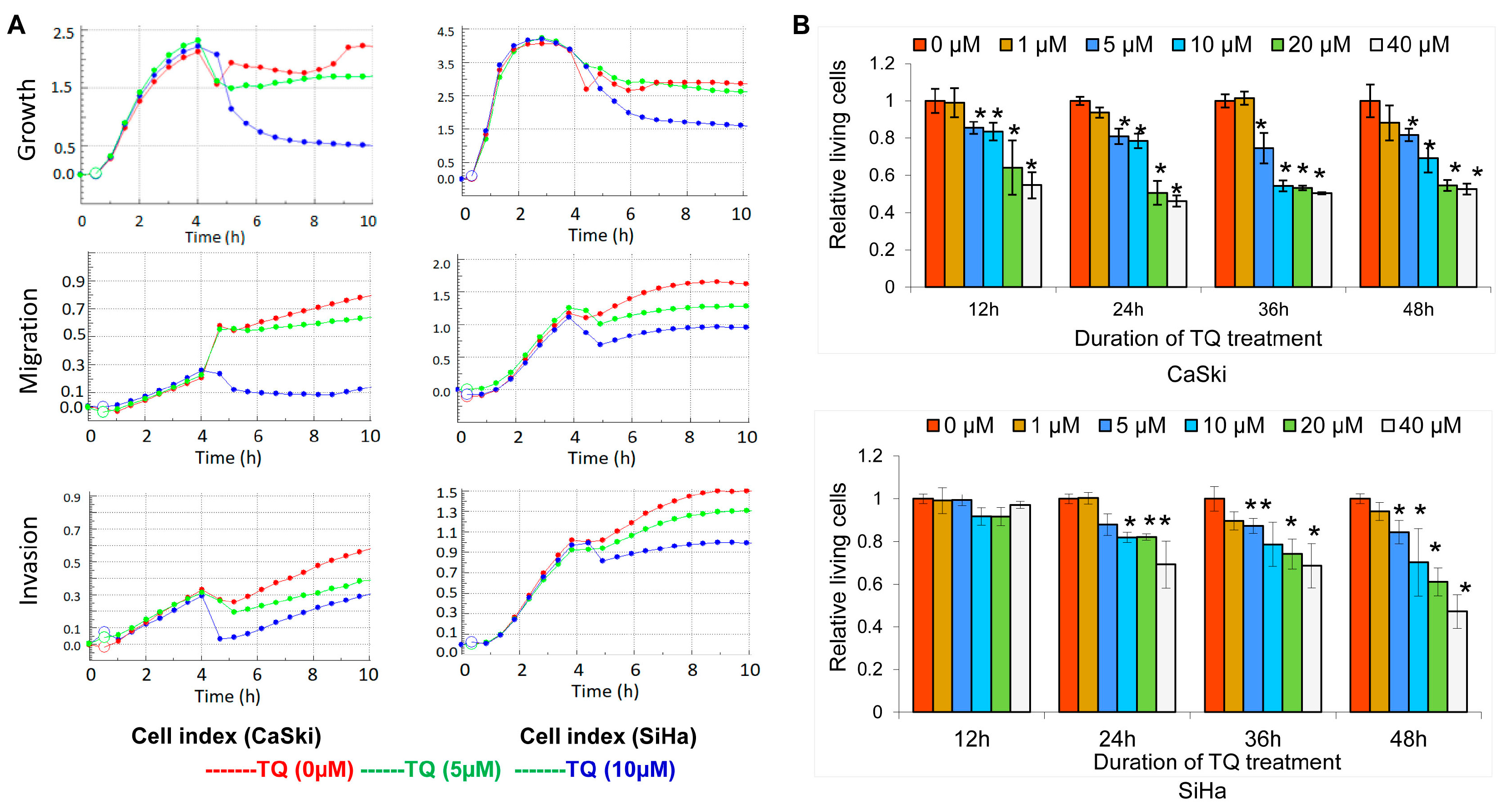
2.1. Thymoquinone Inhibits Cervical Cancer Cell Growth, Migration, and Invasion
2.2. Thymoquinone Induces Apoptosis in Cervical Cancer Cell Lines
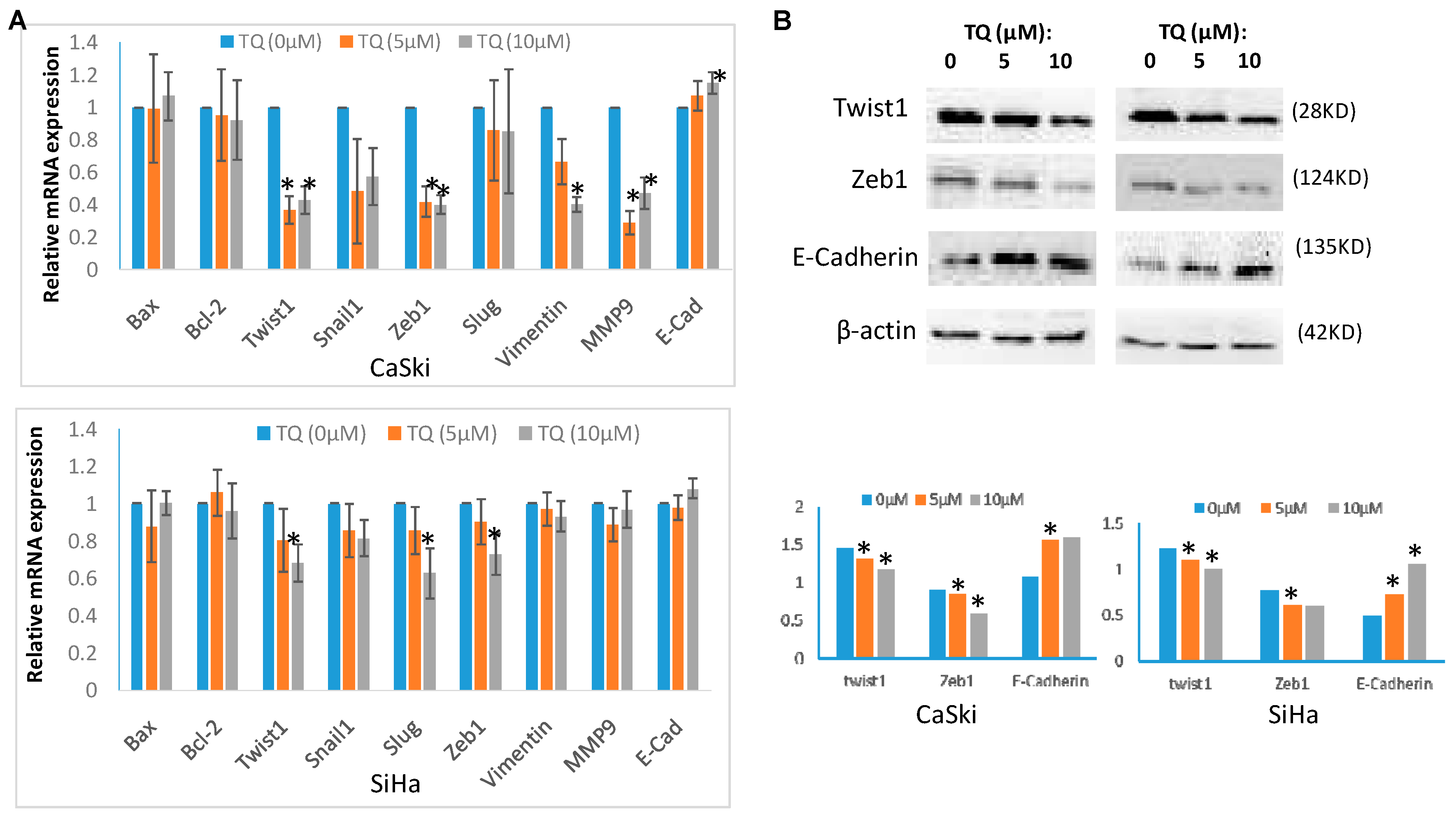
2.3. Thymoquinone Regulates EMT Associated Genes/Proteins in Cervical Cancer Cells CaSki and SiHa
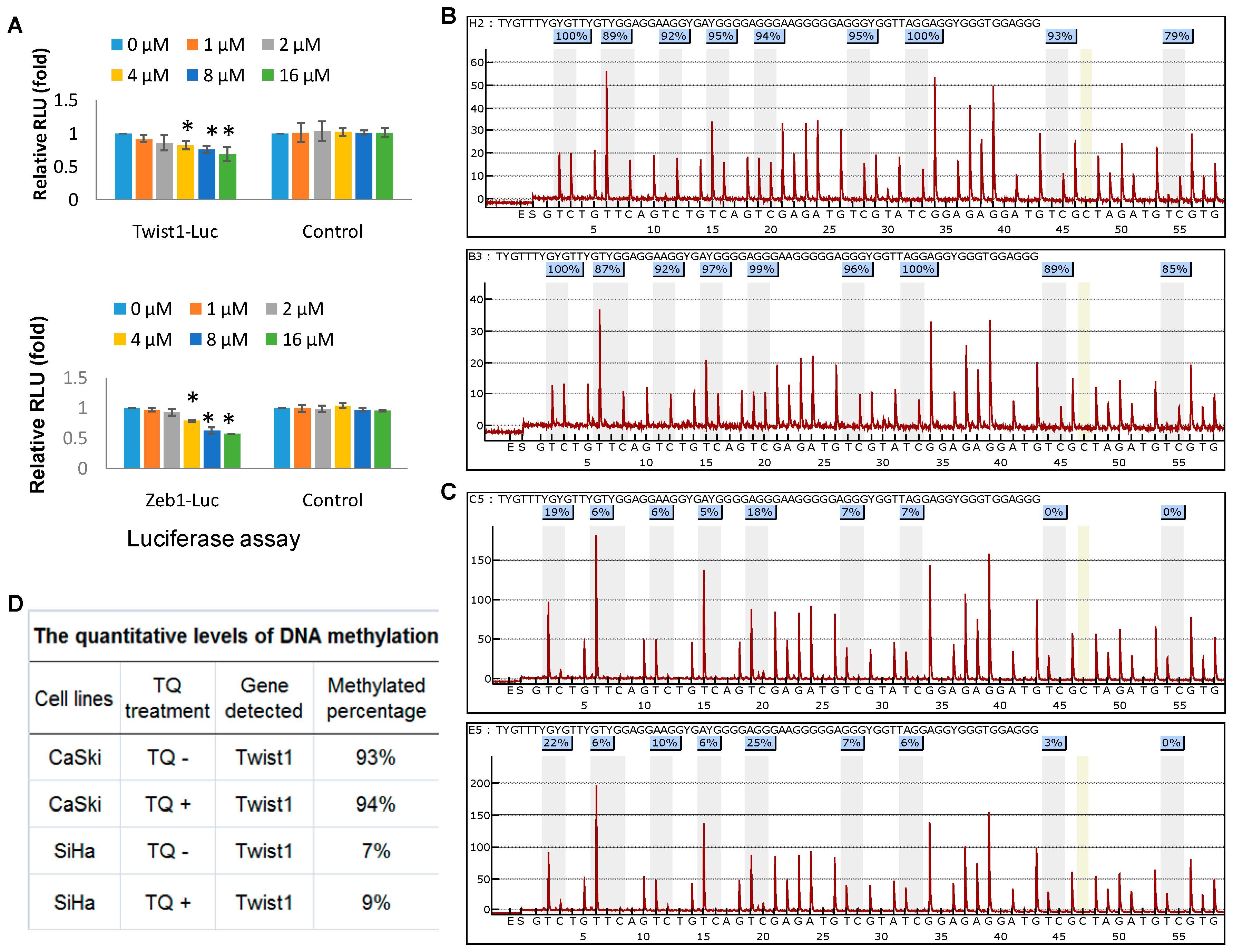
2.4. Thymoquinone Directly Targets Twist1 and Zeb1 Gene
2.5. Effects of Thymoquinone on Twist1 Promoter Methylation in Cancer Cells
3. Discussion
4. Materials and Methods
4.1. Cell Culture and Thymoquinone Treatment
4.2. Cell Viability Assay
4.3. Cell Growth, Migration and Invasion Assays
4.4. RNA Extraction, RT-PCR and qPCR Analysis
4.5. Protein Extraction and Western Blot Analysis
4.6. Luciferase Reporter Assay
4.7. Measurement of Cellular Apoptosis
4.8. Twist1 Gene Methylation Assay
4.9. Statistical Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| ACS | American Cancer Society |
| DMSO | dimethyl sulfoxide |
| FBS | fetal bovine serum |
| EMT | epithelial to mesenchymal transition |
| NCCN | National Comprehensive Cancer Network |
| RLU | Relative Light Units |
| TFs | Transcription factors |
| TQ | Thymoquinone |
References
- Rong, C.; Feng, Y.; Ye, Z. Notch is a critical regulator in cervical cancer by regulating Numb splicing. Oncol. Lett. 2017, 13, 2465–2470. [Google Scholar] [CrossRef] [PubMed]
- Lan, K.; Zhao, Y.; Fan, Y.; Ma, B.; Yang, S.; Liu, Q.; Linghu, H.; Wang, H. Sulfiredoxin May Promote Cervical Cancer Metastasis via Wnt/β-Catenin Signaling Pathway. Int. J. Mol. Sci. 2017, 18, 917. [Google Scholar] [CrossRef]
- Khazaei, S.; Ramachandran, V.; Abdul Hamid, R.; MohdEsa, N.; Etemad, A.; Moradipoor, S.; Ismail, P. Flower extract of Allium atroviolaceum triggered apoptosis, activated caspase-3 and down-regulated antiapoptotic Bcl-2 gene in HeLa cancer cell line. Biomed. Pharmacother. 2017, 89, 1216–1226. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Li, Y.; Liu, N.; Gao, Y.; Li, L. MiR-23b controls ALDH1A1 expression in cervical cancer stem cells. BMC Cancer 2017, 17, 292. [Google Scholar] [CrossRef] [PubMed]
- Smith, R.A.; Andrews, K.S.; Brooks, D.; Fedewa, S.A.; Manassaram-Baptiste, D.; Saslow, D.; Brawley, O.W.; Wender, R.C. Cancer screening in the United States, 2017: A review of current American Cancer Society guidelines and current issues in cancer screening. CA Cancer J. Clin. 2017, 67, 100–121. [Google Scholar] [CrossRef] [PubMed]
- Gu, J.; Hao, C.; Yan, X.; Xuan, S. Applied analysis of ultrasound-guided ilioinguinal and iliohypogastric nerve blocks in the radical surgery of aged cervical cancer. Oncol. Lett. 2017, 13, 1637–1640. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Zhang, H.; Yu, Y.; Huang, H.; Li, G.; Xu, C. Synergistic effects of a novel lipid-soluble extract from Pinelliapedatisecta Schott and cisplatin on human cervical carcinoma cell lines through the regulation of DNA damage response signaling pathway. Oncol. Lett. 2017, 13, 2121–2128. [Google Scholar] [PubMed]
- Khan, M.A.; Tania, M.; Wei, C.; Mei, Z.; Fu, S.; Cheng, J.; Xu, J.; Fu, J. Thymoquinone inhibits cancer metastasis by downregulating TWIST1 expression to reduce epithelial to mesenchymal transition. Oncotarget 2015, 6, 19580–19591. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.A.; Tania, M.; Fu, S.; Fu, J. Thymoquinone, as an anticancer molecule: From basic research to clinical investigation. Oncotarget 2017, 8, 51907–51919. [Google Scholar]
- Ahmad, A.; Husain, A.; Mujeeb, M.; Khan, S.A.; Najmi, A.K.; Siddique, N.A.; Damanhouri, Z.A.; Anwar, F.; Kishore, K. A review on therapeutic potential of Nigella sativa: A miracle herb. Asian Pac. J. Trop. Biomed. 2013, 3, 337–352. [Google Scholar] [CrossRef]
- Salih, B.; Sipahi, T.; OybakDönmez, E. Ancient nige-lla seeds from BoyalıHöyük in north-central Turkey. J. Ethnopharmacol. 2009, 124, 416–420. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.A.; Chen, H.C.; Tania, M.; Zhang, D.Z. Anticancer activities of Nigella sativa (black cumin). Afr. J. Tradit. Complement. Altern. Med. 2011, 8, 226–232. [Google Scholar] [CrossRef] [PubMed]
- Imani, S.; Wei, C.; Cheng, J.; Khan, M.A.; Fu, S.; Yang, L.; Tania, M.; Zhang, X.; Xiao, X.; Zhang, X.; et al. MicroRNA-34a targets epithelial to mesenchymal transition-inducing transcription factors (EMT-TFs) and inhibits breast cancer cell migration and invasion. Oncotarget 2017, 8, 21362–21379. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Qin, L.; He, T.; Qin, J.; Hong, J.; Wong, J.; Liao, L.; Xu, J. The TWIST/Mi2/NuRD protein complex and its essential role in cancer metastasis. Cell Res. 2011, 21, 275–289. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Yang, M.; Gan, L.; He, T.; Xiao, X.; Stewart, M.D.; Liu, X.; Yang, L.; Zhang, T.; Zhao, Y.; et al. DLX4 upregulates TWIST and enhances tumor migration, invasion and metas-tasis. Int. J. Biol. Sci. 2012, 8, 1178–1187. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.A.; Chen, H.C.; Zhang, D.; Fu, J. Twist: A molecular target in cancer therapeutics. Tumour Biol. 2013, 34, 2497–2506. [Google Scholar] [CrossRef] [PubMed]
- Tania, M.; Khan, M.A.; Fu, J. Epithelial to mesenchymal transition inducing transcription factors and metastatic cancer. Tumour Biol. 2014, 35, 7335–7342. [Google Scholar] [CrossRef] [PubMed]
- Cheng, K.Y.; Hao, M. Mammalian Target of Rapamycin (mTOR) Regulates Transforming Growth Factor-β1 (TGF-β1)-Induced Epithelial-Mesenchymal Transition via Decreased Pyruvate Kinase M2 (PKM2) Expression in Cervical Cancer Cells. Med. Sci. Monit. 2017, 23, 2017–2028. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, A.; Loughner, C.L.; Swamynathan, S.; Swamynathan, S.K. KLF4 Plays an Essential Role in Corneal Epithelial Homeostasis by Promoting Epithelial Cell Fate and Suppressing Epithelial-Mesenchymal Transition. Investig. Ophthalmol. Vis. Sci. 2017, 58, 2785–2795. [Google Scholar] [CrossRef] [PubMed]
- Szynglarewicz, B.; Kasprzak, P.; Donizy, P.; Biecek, P.; Halon, A.; Matkowski, R. Epithelial-mesenchymal transition inducer Snail1 and invasive potential of intraductal breast cancer. J. Surg. Oncol. 2017, 116, 696–705. [Google Scholar] [CrossRef] [PubMed]
- Prieto-García, E.; Díaz-García, C.V.; García-Ruiz, I.; Agulló-Ortuño, M.T. Epithelial-to-mesenchymal transition in tumor progression. Med. Oncol. 2017, 34, 122. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, G.; Rath, B. Mesenchymal-Epithelial Transition and Circulating Tumor Cells in Small Cell Lung Cancer. Adv. Exp. Med. Biol. 2017, 994, 229–245. [Google Scholar] [PubMed]
- Hu, Y.; Xie, H.; Liu, Y.; Liu, W.; Liu, M.; Tang, H. miR-484 suppresses proliferation and epithelial-mesenchymal transition by targeting ZEB1 and SMAD2 in cervical cancer cells. Cancer Cell Int. 2017, 17, 36. [Google Scholar] [CrossRef] [PubMed]
- Rojas-Puentes, L.; Cardona, A.F.; Carranza, H.; Vargas, C.; Jaramillo, L.F.; Zea, D.; Cetina, L.; Wills, B.; Ruiz-Garcia, E.; Arrieta, O. Epithelial-mesenchymal transition, proliferation, and angiogenesis in locally advanced cervical cancer treated with chemoradiotherapy. Cancer Med. 2016, 5, 1989–1999. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; Sun, F.; Dong, P.; Watari, H.; Yue, J.; Yu, M.F.; Lan, C.Y.; Wang, Y.; Ma, Z.B. iASPP induces EMT and cisplatin resistance in human cervical cancer through miR-20a-FBXL5/BTG3 signaling. J. Exp. Clin. Cancer Res. 2017, 36, 48. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Zhang, J.; Hu, G.; Liu, L.; Liang, W. Hypoxia and TGF-β1 induced PLOD2 expression improve the migration and invasion of cervical cancer cells by promoting epithelial-to-mesenchymal transition (EMT) and focal adhesion formation. Cancer Cell Int. 2017, 17, 54. [Google Scholar] [CrossRef] [PubMed]
- Abdelmaksoud-Dammak, R.; Chamtouri, N.; Triki, M.; Saadallah-Kallel, A.; Ayadi, W.; Charfi, S.; Khabir, A.; Ayadi, L.; Sallemi-Boudawara, T.; Mokdad-Gargouri, R. Overexpression of miR-10b in colorectal cancer patients: Correlation with TWIST-1 and E-cadherin expression. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Li, T.; Lu, Y.; Shen, G.; Guo, H.; Wu, J.; Lei, C.; Du, F.; Zhou, F.; Zhao, X.; et al. miR-2392 suppresses metastasis and epithelial-mesenchymal transition by targeting MAML3 and WHSC1 in gastric cancer. FASEB J. 2017, 31, 3774–3786. [Google Scholar] [CrossRef] [PubMed]
- Yeh, T.C.; Huang, T.T.; Yeh, T.S.; Chen, Y.R.; Hsu, K.W.; Yin, P.H.; Lee, H.C.; Tseng, L.M. miR-151-3p Targets TWIST1 to Repress Migration of Human Breast Cancer Cells. PLoS ONE 2016, 11, e0168171. [Google Scholar] [CrossRef] [PubMed]
- Li, J.P.; Liu, Y.; Yin, Y.H. ARHGAP1 overexpression inhibits proliferation, migration and invasion of C-33A and SiHa cell lines. Onco Targets Ther. 2017, 10, 691–701. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Dong, X.; Hu, B.; Wang, X.J.; Wang, Q.; Wang, W.L. The effects of Micro-429 on inhibition of cervical cancer cells through targeting ZEB1 and CRKL. Biomed. Pharmacother. 2016, 80, 311–321. [Google Scholar] [CrossRef] [PubMed]
- Enderle-Ammour, K.; Bader, M.; Ahrens, T.D.; Franke, K.; Timme, S.; Csanadi, A.; Hoeppner, J.; Kulemann, B.; Maurer, J.; Reiss, P.; et al. Form follows function: Morphological and immunohistological insights into epithelial-mesenchymal transition characteristics of tumor buds. Tumour Biol. 2017, 39. [Google Scholar] [CrossRef] [PubMed]
- Taha, M.M.; Sheikh, B.Y.; Salim, L.Z.; Mohan, S.; Khan, A.; Kamalidehghan, B.; Ahmadipour, F.; Abdelwahab, S.I. Thymoquinone induces apoptosis and increase ROS in ovarian cancer cell line. Cell Mol. Biol. (Noisy-le-grand) 2016, 62, 97–101. [Google Scholar]
- Dastjerdi, M.N.; Mehdiabady, E.M.; Iranpour, F.G.; Bahramian, H. Effect of Thymoquinone on P53 Gene Expression and Consequence Apoptosis in Breast Cancer Cell Line. Int. J. Prev. Med. 2016, 7, 66. [Google Scholar] [CrossRef] [PubMed]
- Guida, M.S.; Abd El-Aal, A.; Kafafy, Y.; Salama, S.F.; Badr, B.M.; Badr, G. Thymoquinone Rescues T Lymphocytes from Gamma Irradiation-Induced Apoptosis and Exhaustion by Modulating Pro-Inflammatory Cytokine Levels and PD-1, Bax, and Bcl-2 Signaling. Cell. Physiol. Biochem. 2016, 38, 786–800. [Google Scholar] [CrossRef] [PubMed]
- Iskender, B.; Izgi, K.; Hizar, E.; Jauch, J.; Arslanhan, A.; Yuksek, E.H.; Canatan, H. Inhibition of epithelial-mesenchymal transition in bladder cancer cells via modulation of mTOR signalling. Tumour Biol. 2016, 37, 8281–8291. [Google Scholar] [CrossRef] [PubMed]
- Kou, B.; Liu, W.; Zhao, W.; Duan, P.; Yang, Y.; Yi, Q.; Guo, F.; Li, J.; Zhou, J.; Kou, Q. Thymoquinone inhibits epithelial-mesenchymal transition in prostate cancer cells by negatively regulating the TGF-β/Smad2/3 signaling pathway. Oncol. Rep. 2017, 38, 3592–3598. [Google Scholar] [CrossRef] [PubMed]
- Peng, L.; Yuan, X.; Jiang, B.; Tang, Z.; Li, G.C. LncRNAs: Key players and novel insights into cervical cancer. Tumour Biol. 2016, 37, 2779–2788. [Google Scholar] [CrossRef] [PubMed]
- Ji, F.; Wuerkenbieke, D.; He, Y.; Ding, Y. Long Noncoding RNA HOTAIR: An Oncogene in Human Cervical Cancer Interacting with MicroRNA-17-5p. Oncol. Res. 2017. [Google Scholar] [CrossRef] [PubMed]
- Battistelli, C.; Cicchini, C.; Santangelo, L.; Tramontano, A.; Grassi, L.; Gonzalez, F.J.; de Nonno, V.; Grassi, G.; Amicone, L.; Tripodi, M. The Snail repressor recruits EZH2 to specific genomic sites through the enrollment of the lncRNA HOTAIR in epithelial-to-mesenchymal transition. Oncogene 2017, 36, 942–955. [Google Scholar] [CrossRef] [PubMed]
- Fu, S.; Cheng, J.; Wei, C.; Yang, L.; Xiao, X.; Zhang, D.; Stewart, M.D.; Fu, J. Development of diagnostic SCAR markers for genomic DNA amplifications in breast carcinoma by DNA cloning of high-GC RAMP-PCR fragments. Oncotarget 2017, 8, 43866–43877. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Not available. |

| Genes | 5′-3′ Sequences |
|---|---|
| Q18S-48L: | GCAATTATTCCCCATGAACG |
| Q18S-48R: | GGGACTTAATCAACGCAAGC |
| Twist1-6L: | GGCATCACTATGGACTTTCTCTATT |
| Twist1-6R: | GGCCAGTTTGATCCCAGTATT |
| Snail1-11L: | GCTGCAGGACTCTAATCCAGA |
| Snail1-11R: | ATCTCCGGAGGTGGGATG |
| Slug-26L: | TGCACCCTCGGATACCTG |
| Slug-26R: | ACATTTGGATCACAGAGGCATA |
| Zeb1-31L: | TGACTATCAAAAGGAAGTCAATGG |
| Zeb1-31R: | GTGCAGGAGGGACCTCTTTA |
| E-Cadherin-84L: | TGGAGGAATTCTTGCTTTGC |
| E-Cadherin-84R: | CGCTCTCCTCCGAAGAAAC |
| Vimentin-56L: | TGGTCTAACGGTTTCCCCTA |
| Vimentin-56R: | GACCTCGGAGCGAGAGTG |
| MMP9-6L: | GAACCAATCTCACCGACAGG |
| MMP9-6R: | GCCACCCGAGTGTAACCATA |
| BCL2-2L: | GTGGTTGGCTTACACATGGA |
| BCL2-2R: | CACCAGGGCCAAACTGAG |
| BAX-55L: | CAAGACCAGGGTGGTTGG |
| BAX-55R: | CACTCCCGCCACAAAGAT |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, J.; Khan, M.A.; Wei, C.; Cheng, J.; Chen, H.; Yang, L.; Ijaz, I.; Fu, J. Thymoquinone Inhibits the Migration and Invasive Characteristics of Cervical Cancer Cells SiHa and CaSki In Vitro by Targeting Epithelial to Mesenchymal Transition Associated Transcription Factors Twist1 and Zeb1. Molecules 2017, 22, 2105. https://doi.org/10.3390/molecules22122105
Li J, Khan MA, Wei C, Cheng J, Chen H, Yang L, Ijaz I, Fu J. Thymoquinone Inhibits the Migration and Invasive Characteristics of Cervical Cancer Cells SiHa and CaSki In Vitro by Targeting Epithelial to Mesenchymal Transition Associated Transcription Factors Twist1 and Zeb1. Molecules. 2017; 22(12):2105. https://doi.org/10.3390/molecules22122105
Chicago/Turabian StyleLi, Jun, Md. Asaduzzaman Khan, Chunli Wei, Jingliang Cheng, Hanchun Chen, Lisha Yang, Iqra Ijaz, and Junjiang Fu. 2017. "Thymoquinone Inhibits the Migration and Invasive Characteristics of Cervical Cancer Cells SiHa and CaSki In Vitro by Targeting Epithelial to Mesenchymal Transition Associated Transcription Factors Twist1 and Zeb1" Molecules 22, no. 12: 2105. https://doi.org/10.3390/molecules22122105
APA StyleLi, J., Khan, M. A., Wei, C., Cheng, J., Chen, H., Yang, L., Ijaz, I., & Fu, J. (2017). Thymoquinone Inhibits the Migration and Invasive Characteristics of Cervical Cancer Cells SiHa and CaSki In Vitro by Targeting Epithelial to Mesenchymal Transition Associated Transcription Factors Twist1 and Zeb1. Molecules, 22(12), 2105. https://doi.org/10.3390/molecules22122105





