The Complete Chloroplast Genome Sequences of Aconitum pseudolaeve and Aconitum longecassidatum, and Development of Molecular Markers for Distinguishing Species in the Aconitum Subgenus Lycoctonum
Abstract
:1. Introduction
2. Results and Discussion
2.1. CP Genome Organization of A. pseudolaeve and A. longecassidatum
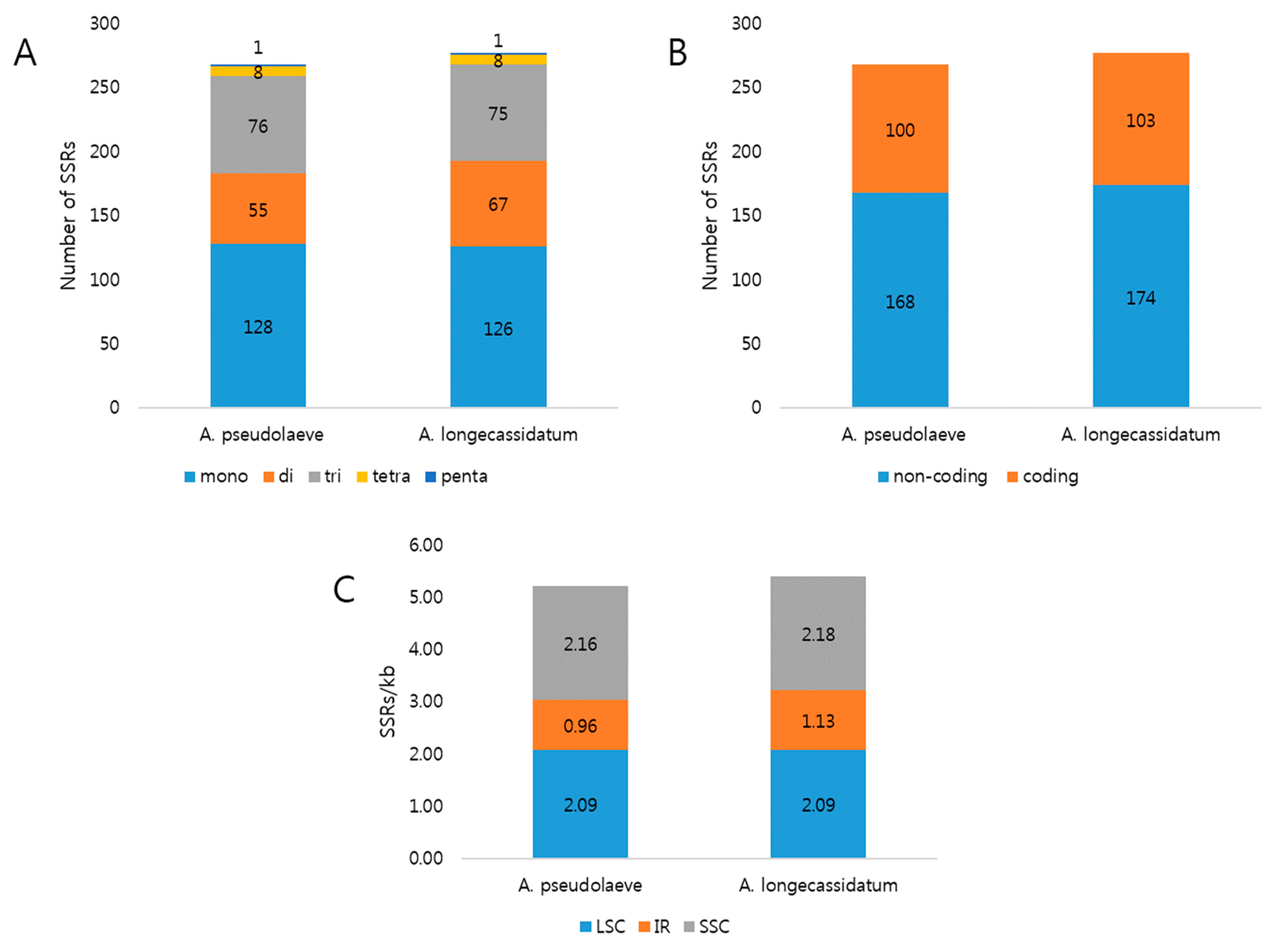
2.2. Repeat Analysis in Two Aconitum Chloroplast Genomes
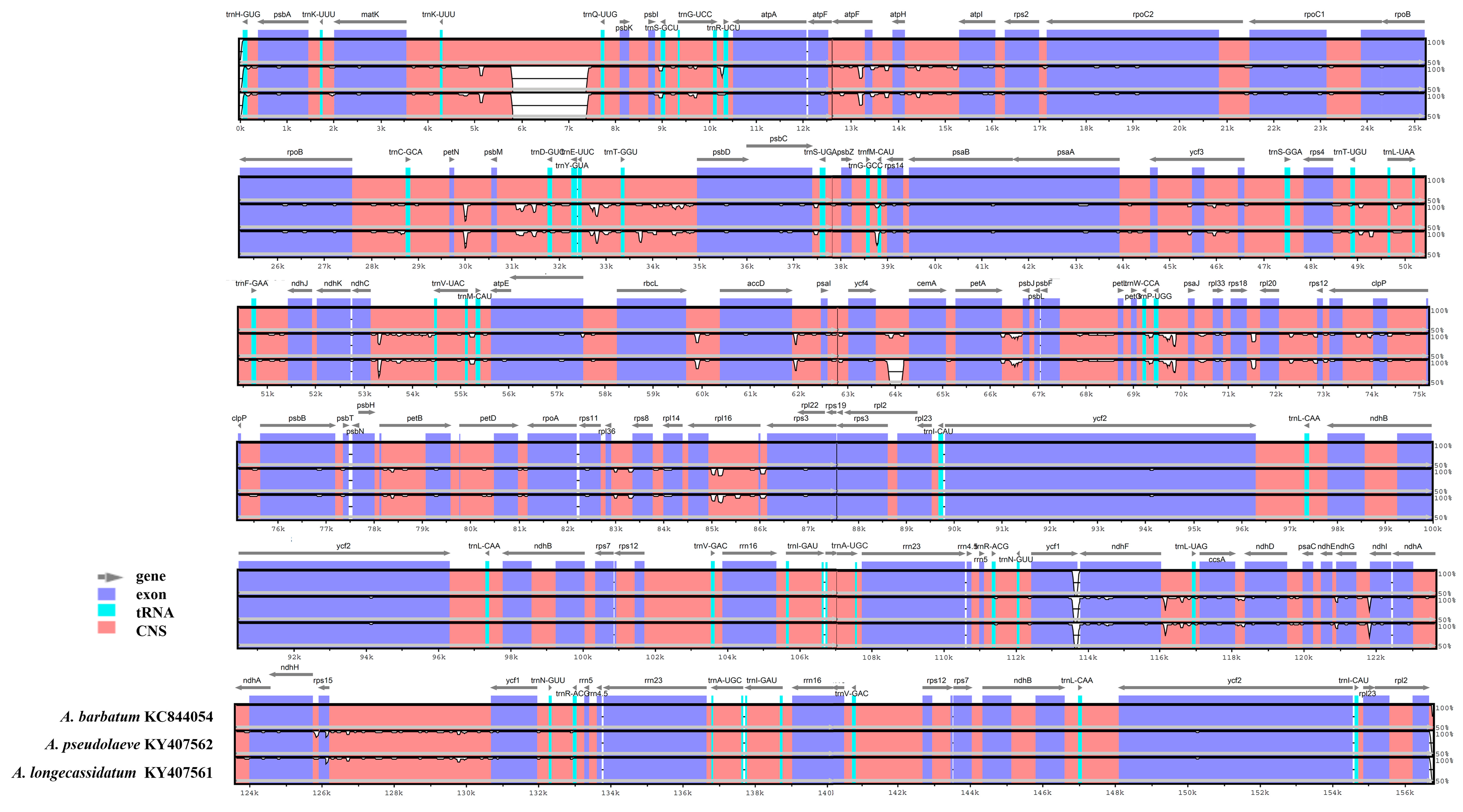
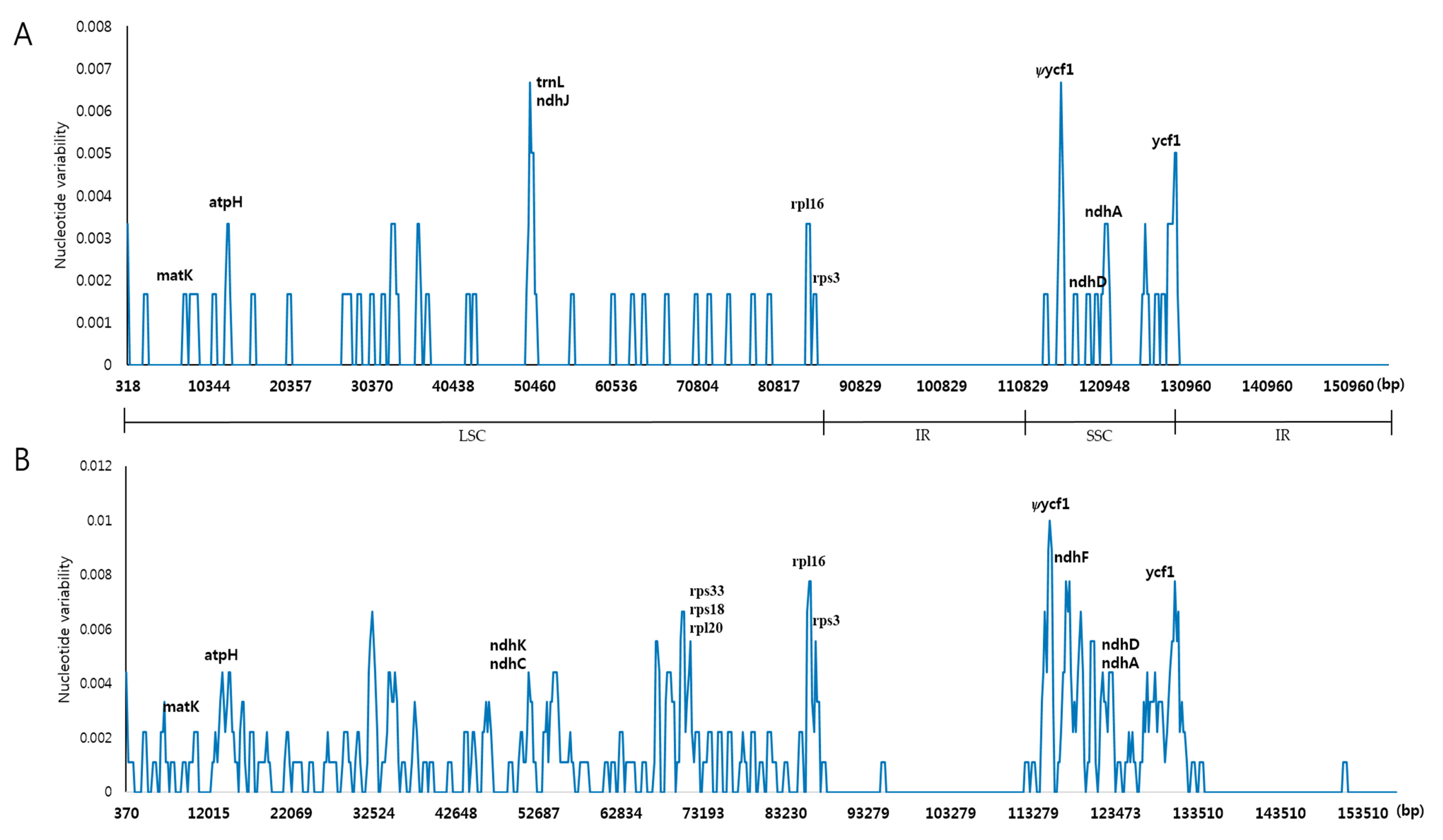
2.3. Comparison of the Chloroplast Genomes of A. pseudolaeve Nakai, A. longecassidatum Nakai and Aconitum barbatum Patrin ex Pers.
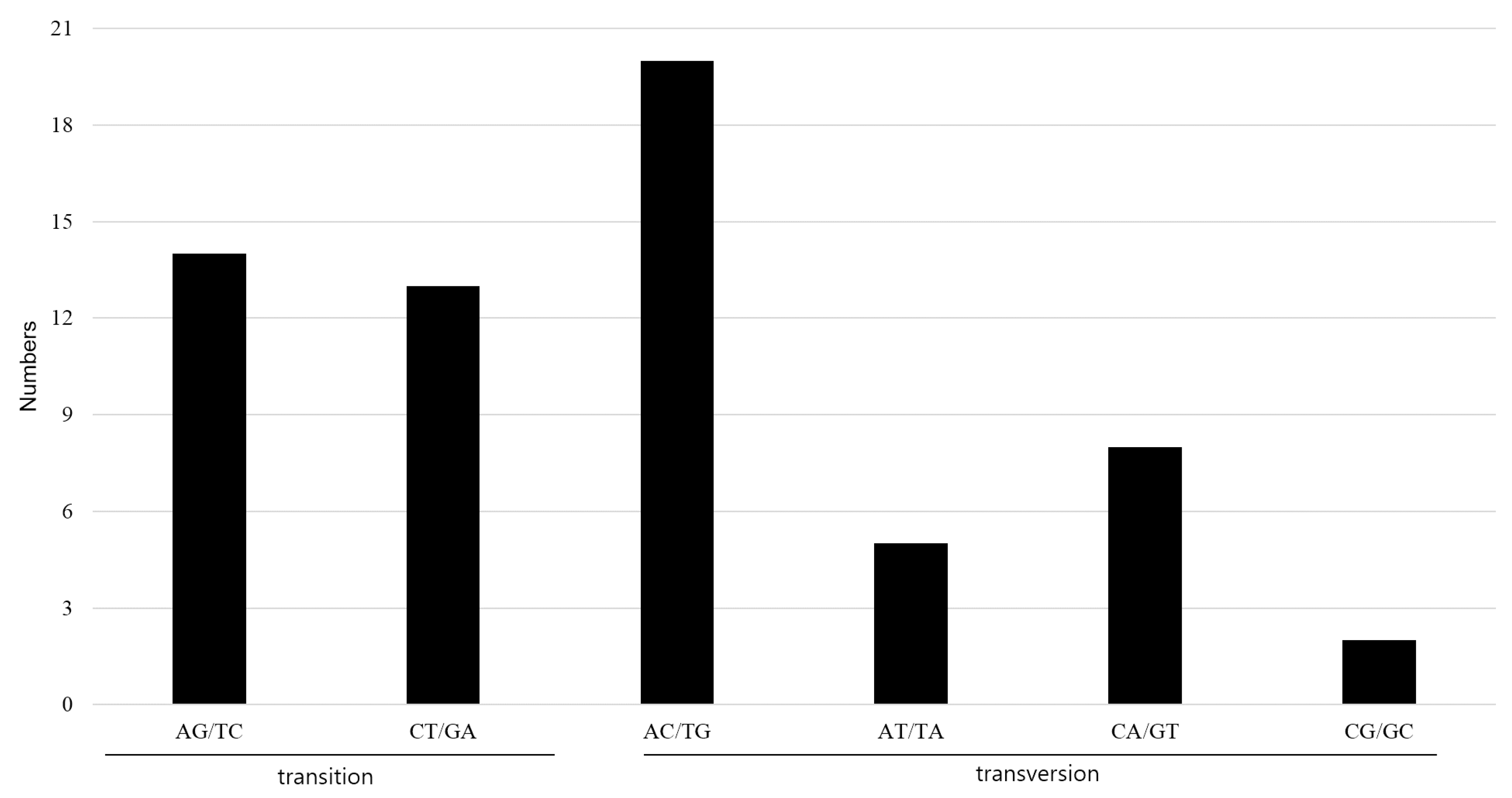
2.4. Indel and SNP Mutation between A. pseudolaeve and A. longecassidatum
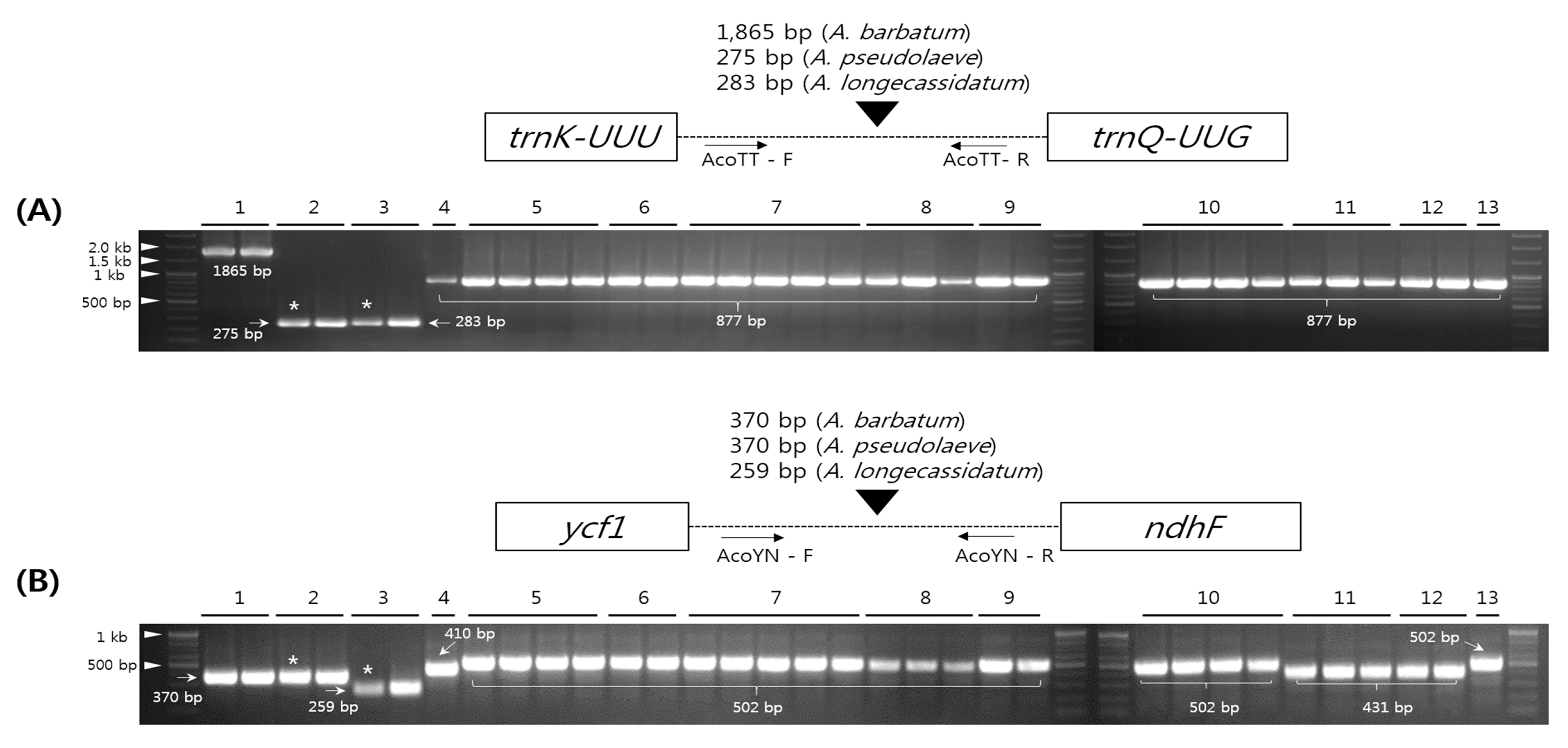
2.5. Development and Validation of the Indel Marker for Authentication of Three Species in the Aconitum Subgenus Lycoctonum
3. Materials and Methods
3.1. Plant Materials and Genome Sequencing
3.2. Assembly and Annotation of Two Aconitum Species
3.3. SSR, Tandem, and Palindromic Repeat Analysis in Two Aconitum CP Genomes
3.4. Comparative Analysis of CP Genomes of A. pseudolaeve and A. longecassidatum
3.5. Development and Validation of Indel Markers (AcoTT and AcoYN) Among Aconitum Species
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| CP | Chloroplast |
| LSC | Large single copy |
| SSC | Small single copy |
| IR | Inverted repeat |
| tRNA | Transfer RNA |
| rRNA | Ribosomal RNA |
| KIOM | Korea Institute of Oriental Medicine |
| SSRs | Simple Sequence Repeats |
| SNPs | Single-Nucleotide Polymorphisms |
| Indel | Insertion and Deletion |
References
- Wicke, S.; Schneeweiss, G.M.; dePamphilis, C.W.; Muller, K.F.; Quandt, D. The evolution of the plastid chromosome in land plants: Gene content, gene order, gene function. Plant Mol. Biol. 2011, 76, 273–297. [Google Scholar] [CrossRef] [PubMed]
- Choi, K.S.; Park, S. The complete chloroplast genome sequence of Aster spathulifolius (Asteraceae); genomic features and relationship with Asteraceae. Gene 2015, 572, 214–221. [Google Scholar] [CrossRef] [PubMed]
- Daniell, H.; Lin, C.S.; Yu, M.; Chang, W.J. Chloroplast genomes: Diversity, evolution, and applications in genetic engineering. Genome Biol. 2016, 17, 134. [Google Scholar] [CrossRef] [PubMed]
- Benson, D.A.; Karsch-Mizrachi, I.; Lipman, D.J.; Ostell, J.; Rapp, B.A.; Wheeler, D.L. Genbank. Nucleic Acids Res. 2000, 28, 15–18. [Google Scholar] [CrossRef] [PubMed]
- Provan, J.; Powell, W.; Hollingsworth, P.M. Chloroplast microsatellites: New tools for studies in plant ecology and evolution. Trends Ecol. Evol. 2001, 16, 142–147. [Google Scholar] [CrossRef]
- Huang, Y.; Li, X.; Yang, Z.; Yang, C.; Yang, J.; Ji, Y. Analysis of complete chloroplast genome sequences improves phylogenetic resolution in paris (Melanthiaceae). Front. Plant Sci. 2016, 7, 1797. [Google Scholar] [CrossRef] [PubMed]
- Park, I.; Kim, W.J.; Yang, S.; Yeo, S.M.; Li, H.; Moon, B.C. The complete chloroplast genome sequence of Aconitum coreanum and Aconitum carmichaelii and comparative analysis with other Aconitum species. PLoS ONE 2017, 12, e0184257. [Google Scholar] [CrossRef] [PubMed]
- Park, I.; Kim, W.J.; Yeo, S.M.; Choi, G.; Kang, Y.M.; Piao, R.; Moon, B.C. The complete chloroplast genome sequences of Fritillaria ussuriensis maxim. and Fritillaria cirrhosa D. Don, and comparative analysis with other Fritillaria species. Molecules 2017, 22. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Dong, W.; Liu, B.; Xu, C.; Yao, X.; Gao, J.; Corlett, R.T. Comparative analysis of complete chloroplast genome sequences of two tropical trees Machilus yunnanensis and Machilus balansae in the family Lauraceae. Front. Plant Sci. 2015, 6, 662. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Shi, C.; Liu, Y.; Mao, S.Y.; Gao, L.Z. Thirteen Camellia chloroplast genome sequences determined by high-throughput sequencing: Genome structure and phylogenetic relationships. BMC Evol. Biol. 2014, 14, 151. [Google Scholar] [CrossRef] [PubMed]
- Bielsa, B.; Jiwan, D.; i Marti, A.F.; Dhingra, A.; Rubio-Cabetas, M. Detection of SNP and validation of a SFP indel (deletion) in inverted repeat region of the prunus species chloroplast genome. Sci. Hortic. 2014, 168, 108–112. [Google Scholar] [CrossRef]
- Cai, Z.; Guisinger, M.; Kim, H.-G.; Ruck, E.; Blazier, J.C.; McMurtry, V.; Kuehl, J.V.; Boore, J.; Jansen, R.K. Extensive reorganization of the plastid genome of Trifolium subterraneum (Fabaceae) is associated with numerous repeated sequences and novel DNA insertions. J. Mol. Evol. 2008, 67, 696–704. [Google Scholar] [CrossRef] [PubMed]
- Yin, P.; Kang, J.; He, F.; Qu, L.-J.; Gu, H. The origin of populations of Arabidopsis thaliana in china, based on the chloroplast DNA sequences. BMC Plant Biol. 2010, 10, 22. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.; Lee, S.C.; Lee, J.; Lee, H.O.; Joh, H.J.; Kim, N.H.; Park, H.S.; Yang, T.J. Comprehensive survey of genetic diversity in chloroplast genomes and 45 s nrDNAs within Panax ginseng species. PLoS ONE 2015, 10, e0117159. [Google Scholar] [CrossRef] [PubMed]
- Cho, K.S.; Yun, B.K.; Yoon, Y.H.; Hong, S.Y.; Mekapogu, M.; Kim, K.H.; Yang, T.J. Complete chloroplast genome sequence of tartary buckwheat (Fagopyrum tataricum) and comparative analysis with common buckwheat (F. esculentum). PLoS ONE 2015, 10, e0125332. [Google Scholar] [CrossRef] [PubMed]
- Cho, K.S.; Cheon, K.S.; Hong, S.Y.; Cho, J.H.; Im, J.S.; Mekapogu, M.; Yu, Y.S.; Park, T.H. Complete chloroplast genome sequences of Solanum commersonii and its application to chloroplast genotype in somatic hybrids with Solanum tuberosum. Plant Cell Rep. 2016, 35, 2113–2123. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Fu, D.; Li, L.Q.; Bartholomew, B.; Brach, A.R.; Dutton, B.E.; Gilbert, M.G.; Kadota, Y.; Robinson, O.R.; Tamura, M.; et al. Ranunculaceae. In Flora of China; Wu, Z.Y., Raven, P.H., Eds.; Science Press: Beijing, China; Missouri Botanical Garden Press: St. Louis, MO, USA, 2001; Volume 6, pp. 149–222. [Google Scholar]
- National Institute of Biological Resources (NIBR). Endemic Species of Korea. 2013. Available online: https://species.nibr.go.kr/index.do (accessed on 15 May 2017).
- Floral of Korea Editorial Committee (FKEC). The Genera of Vascular Plants of Korea; Academy Publishing Co.: Seoul, Korea, 2007; pp. 176–182. [Google Scholar]
- Korea Institute of Oriental Medicine (KIOM). Defining Dictionary for Medicinal Herbs. 2016. Available online: http://boncho.kiom.re.kr/codex/ (accessed on 15 May 2017).
- Lee, S.-Y.; Kwon, O.-J.; Park, J.-H.; Lee, J.-Y. Anti-aging and anti-diabetes effects of Aconitum pesudo-laeve var. Erectum extracts. J. Life Sci. 2013, 23, 616–621. [Google Scholar]
- Lee, J.-Y.; Lee, S.Y.; Jeoun, H.; Youn, J.Y. Cosmetic Composition Comprising the Extract of Aconitum pseudolaeve var. erectum Nakai as Active Ingredient. Patent No. KR20130040413A, 24 April 2013. [Google Scholar]
- Lee, J.-Y.; Choi, Y.-H.; Lee, S.-M.; Lee, S.-Y.; Jun, H.-J. Physiological activity of Aconitum pseudolaeve var. erectum Nakai and its effect on perfume on an electroencephalogram (EEG). J. Life Sci. 2012, 22, 1214–1223. [Google Scholar] [CrossRef]
- Qian, J.; Song, J.; Gao, H.; Zhu, Y.; Xu, J.; Pang, X.; Yao, H.; Sun, C.; Li, X.; Li, C.; et al. The complete chloroplast genome sequence of the medicinal plant Salvia miltiorrhiza. PLoS ONE 2013, 8, e57607. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Zhang, X.; Liu, G.; Yin, Y.; Chen, K.; Yun, Q.; Zhao, D.; Al-Mssallem, I.S.; Yu, J. The complete chloroplast genome sequence of date palm (Phoenix dactylifera l.). PLoS ONE 2010, 5, e12762. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhan, D.F.; Jia, X.; Mei, W.L.; Dai, H.F.; Chen, X.T.; Peng, S.Q. Complete chloroplast genome sequence of Aquilaria sinensis (Lour.) Gilg and evolution analysis within the Malvales order. Front. Plant Sci. 2016, 7, 280. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Li, Q.; Li, Y.; Qian, J.; Han, J. Chloroplast genome of Aconitum barbatum var. puberulum (Ranunculaceae) derived from CCS reads using the pacbio RS platform. Front. Plant Sci. 2015, 6, 42. [Google Scholar] [CrossRef] [PubMed]
- Zuo, L.H.; Shang, A.Q.; Zhang, S.; Yu, X.Y.; Ren, Y.C.; Yang, M.S.; Wang, J.M. The first complete chloroplast genome sequences of Ulmus species by de novo sequencing: Genome comparative and taxonomic position analysis. PLoS ONE 2017, 12, e0171264. [Google Scholar] [CrossRef] [PubMed]
- Lim, C.E.; Kim, G.B.; Baek, S.; Han, S.M.; Yu, H.J.; Mun, J.H. The complete chloroplast genome of Aconitum chiisanense Nakai (Ranunculaceae). Mitochondrial DNA 2015, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Echt, C.S.; DeVerno, L.; Anzidei, M.; Vendramin, G. Chloroplast microsatellites reveal population genetic diversity in red pine, Pinus resinosa Ait. Mol. Ecol. 1998, 7, 307–316. [Google Scholar] [CrossRef]
- Nishikawa, T.; Vaughan, D.A.; Kadowaki, K.-I. Phylogenetic analysis of Oryza species, based on simple sequence repeats and their flanking nucleotide sequences from the mitochondrial and chloroplast genomes. Theor. Appl. Genet. 2005, 110, 696–705. [Google Scholar] [CrossRef] [PubMed]
- Nie, X.; Lv, S.; Zhang, Y.; Du, X.; Wang, L.; Biradar, S.S.; Tan, X.; Wan, F.; Weining, S. Complete chloroplast genome sequence of a major invasive species, crofton weed (Ageratina adenophora). PLoS ONE 2012, 7, e36869. [Google Scholar] [CrossRef] [PubMed]
- Cho, K.-S.; Hong, S.-Y.; Yun, B.-K.; Won, H.-S.; Yoon, Y.-H.; Kwon, K.-B.; Mekapogu, M. Application of indel markers based on the chloroplast genome sequences for authentication and traceability of tartary and common buckwheat. Czech J. Food Sci. 2017, 35, 122–130. [Google Scholar]
- Tang, J.; Xia, H.A.; Cao, M.; Zhang, X.; Zeng, W.; Hu, S.; Tong, W.; Wang, J.; Wang, J.; Yu, J. A comparison of rice chloroplast genomes. Plant Physiol. 2004, 135, 412–420. [Google Scholar] [CrossRef] [PubMed]
- Suo, Z.; Zhang, C.; Zheng, Y.; He, L.; Jin, X.; Hou, B.; Li, J. Revealing genetic diversity of tree peonies at micro-evolution level with hyper-variable chloroplast markers and floral traits. Plant Cell Rep. 2012, 31, 2199–2213. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.; Xu, C.; Cheng, T.; Lin, K.; Zhou, S. Sequencing angiosperm plastid genomes made easy: A complete set of universal primers and a case study on the phylogeny of saxifragales. Genome Biol. Evol. 2013, 5, 989–997. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.; Xu, C.; Li, D.; Jin, X.; Li, R.; Lu, Q.; Suo, Z. Comparative analysis of the complete chloroplast genome sequences in Psammophytic haloxylon species (Amaranthaceae). PeerJ 2016, 4, e2699. [Google Scholar] [CrossRef] [PubMed]
- Redwan, R.M.; Saidin, A.; Kumar, S.V. Complete chloroplast genome sequence of MD-2 pineapple and its comparative analysis among nine other plants from the subclass Commelinidae. BMC Plant Biol. 2015, 15, 196. [Google Scholar] [CrossRef] [PubMed]
- Suo, Z.; Jia, Z.; Lu, Q.; Pan, B.; Jin, X.; Xu, G.; Peng, X.; Sun, H.; Tao, Y. Distinguishing Haloxylon persicum and H. Ammodendron (Haloxylon Bunge, Amaranthaceae) using DNA marker. AASRI Procedia 2012, 1, 305–310. [Google Scholar] [CrossRef]
- Delcher, A.L.; Salzberg, S.L.; Phillippy, A.M. Using mummer to identify similar regions in large sequence sets. Curr. Protoc. Bioinform. 2003. [CrossRef]
- Wyman, S.K.; Jansen, R.K.; Boore, J.L. Automatic annotation of organellar genomes with DOGMA. Bioinformatics 2004, 20, 3252–3255. [Google Scholar] [CrossRef] [PubMed]
- Lohse, M.; Drechsel, O.; Bock, R. OrganellarGenomeDRAW (OGDRAW): A tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Curr. Genet. 2007, 52, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. Mega6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Benson, G. Tandem repeats finder: A program to analyze DNA sequences. Nucleic Acids Res. 1999, 27, 573–580. [Google Scholar] [CrossRef] [PubMed]
- Thiel, T. Misa—Microsatellite Identification Tool. 2003. Available online: http://pgrc.ipk-gatersleben.de/misa/ (accessed on 10 June 2017).
- Warburton, P.E.; Giordano, J.; Cheung, F.; Gelfand, Y.; Benson, G. Inverted repeat structure of the human genome: The X-chromosome contains a preponderance of large, highly homologous inverted repeats that contain testes genes. Genome Res. 2004, 14, 1861–1869. [Google Scholar] [CrossRef] [PubMed]
- Frazer, K.A.; Pachter, L.; Poliakov, A.; Rubin, E.M.; Dubchak, I. Vista: Computational tools for comparative genomics. Nucleic Acids Res. 2004, 32, W273–W279. [Google Scholar] [CrossRef] [PubMed]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. Mafft: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the A. pseudolaeve, A. longecassidatum, A. voluvile var. pubescens, A. jaluense var. triphyllum, A. kusnezoffii, A. jaluense, A. austrokoreense and A. chiisanense are available from the authors and the herbarium of KIOM. |
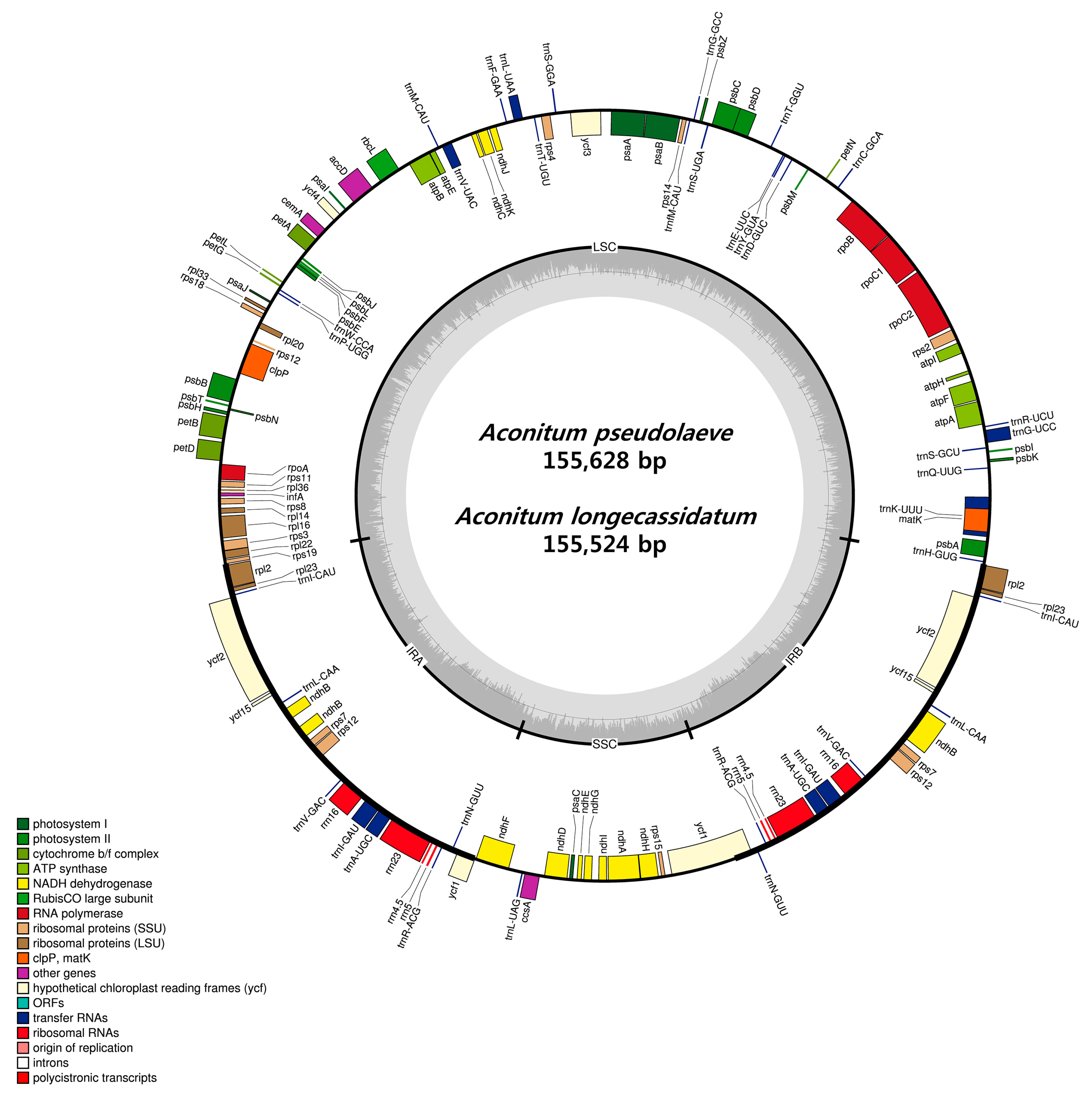
| Species | Aconitum pseudolaeve | Aconitum longecassidatum |
|---|---|---|
| Total CP genome size (bp) | 155,628 | 155,524 |
| LSC region (bp) | 86,683 | 86,466 |
| IR region (bp) | 51,854 | 52,108 |
| SSC region (bp) | 17,091 | 16,950 |
| GC content (%) | 38.0 | 38.1 |
| LSC (%) | 36.1 | 36.1 |
| IR (%) | 43.1 | 43.0 |
| SSC (%) | 32.6 | 32.7 |
| Number of genes | 112 | 112 |
| Protein-coding genes | 78 | 78 |
| rRNAs | 4 | 4 |
| tRNAs | 30 | 30 |
| Gene Products of Aconitum Species | |
|---|---|
| Photosystem I | psaA, B, C, I, J |
| Photosystem II | psbA, B, C, D, E, F, H, I, J, K, L, M, N, T, Z |
| Cytochrome b6_f | petA, B (1), D (1), G, L, N |
| ATP synthase | atpA, B, E, F (1), H, I |
| Rubisco | rbcL |
| NADH oxidoreductase | ndhA (1), B (1),(3), C, D, E, F, G, H, I, J, K |
| Large subunit ribosomal proteins | rpl2 (1),(3), 14, 16 (1), 20, 22, 23 (3), 32, 33, 36 |
| Small subunit ribosomal proteins | rps2, 3, 4, 7 (3), 8, 11, 12 (2)–(4), 14, 15, 18, 19 |
| RNA polymerase | rpoA, B, C1 (1), C2 |
| Unknown function protein-coding gene | ycf1 (3), 2 (3), 3 (2), 4 |
| Other genes | accD, ccsA, cemA, clpP (2), infA, matK |
| Ribosomal RNAs | rrn16 (3), rrn23 (3), rrn4.5 (3), rrn5 (3) |
| Transfer RNAs | trnA-UGC (1),(3), trnC-GCA, trnD-GUC, trnE-UUC, trnF-GAA, trnG-UCC (1), trnG-GCC, trnH-GUG, trnI-CAU (3), trnI-GAU (1),(3) trnK-UUU (1), trnL-UAA (1), trnL-UAG, trnL-CAA (3), trnM-CAU, trnfM-CAU, trnN-GUU (3), trnP-UGG, trnQ-UUG, trnR-ACG (3), trnR-UCU, trnS-GCU, trnS-GGA, trnS-UGA, trnT-GGU, trnT-UGU, trnV-UAC (1), trnV-GAC (3), trnW-CCA, trnY-GUA |
| Gene | Region | Exon I | Intron I | Exon II | Intron II | Exon III |
|---|---|---|---|---|---|---|
| trnk-UUU | LSC | 37 | 2526 (2525) | 35 | ||
| trnG-UCC | LSC | 23 | 744 (747) | 48 | ||
| atpF | LSC | 145 | 733 (735) | 410 | ||
| rpoC1 | LSC | 432 | 757 | 1635 (1611) | ||
| ycf3 | LSC | 124 | 730 (729) | 230 | 761 (762) | 153 |
| trnL-UAA | LSC | 35 | 501 (495) | 50 | ||
| trnV-UAC | LSC | 39 | 597 | 37 | ||
| rps12 * | LSC | 114 | - | 232 | - | 26 |
| clpP | LSC | 71 | 833 (830) | 292 | 651 (661) | 246 |
| petB | LSC | 6 | 801 | 642 | ||
| petD | LSC | 8 | 704 | 496 | ||
| rpl16 | LSC | 9 | 1045 (1054) | 399 | ||
| rpl2 | IR | 391 | 667 | 434 | ||
| ndhB | IR | 777 | 702 | 756 | ||
| trnI-GAU | IR | 42 | 937 | 35 | ||
| trnA-UGC | IR | 38 | 802 | 35 | ||
| ndhA | SSC | 553 | 1006 (1004) | 539 |
| No. | Location | Region | Motif | Repeat Number | |
|---|---|---|---|---|---|
| A. pseudolaeve | A. longecassidatum | ||||
| 1 | trnH-psbA | IGS | A | 9 | 8 |
| 2 | trnK-matK | IGS | T | 13 | 12 |
| 3 | trnK-trnQ | IGS | A | 12 | 10 |
| 4 | trnK-trnQ | IGS | T | 9 | 10 |
| 5 | trnG | intron | T | 15 | 18 |
| 6 | atpF | intron | T | 16 | 14 |
| 7 | psbM-trnD | IGS | T | 16 | 11 |
| 8 | psbM-trnD | IGS | AT | 18 | 12 |
| 9 | trnS-psbZ | IGS | A | 11 | 10 |
| 10 | trnG-trnfM | IGS | A | 18 | 19 |
| 11 | psaA-ycf3 | IGS | T | 9 | 10 |
| 12 | ycf3 | intron | T | 11 | 12 |
| 13 | ycf3 | intron | A | 9 | 8 |
| 14 | rps4-trnT | IGS | A | 12 | 11 |
| 15 | trnF-ndhJ | IGS | T | 8 | 10 |
| 16 | ndhC-trnV | IGS | A | 9 | 11 |
| 17 | ndhC-trnV | IGS | A | 14 | 9 |
| 18 | accD-psaI | IGS | A | 9 | 8 |
| 19 | accD-psaI | IGS | A | 13 | 12 |
| 20 | psaI-ycf4 | IGS | T | 10 | 11 |
| 21 | ycf4-cemA | IGS | T | 10 | 9 |
| 22 | petA-psbJ | IGS | T | 10 | 9 |
| 23 | psbE-petL | IGS | T | 9 | 8 |
| 24 | psbE-petL | IGS | A | 9 | 8 |
| 25 | psbE-petL | IGS | T | 11 | 10 |
| 26 | rps12-clpP | IGS | ATT | 12 | 9 |
| 27 | clpP | intron | A | 13 | 11 |
| 28 | clpP | intron | A | 14 | 11 |
| 29 | rpl16 | intron | A | 12 | 15 |
| 30 | rpl16-rps3 | IGS | T | 11 | 14 |
| 31 | ndhF-trnL | IGS | A | 12 | 11 |
| 32 | ndhF-trnL | IGS | T | 12 | 11 |
| 33 | ccsA-ndhD | IGS | T | 13 | 10 |
| 34 | ndhD-psaC | IGS | A | 8 | 9 |
| 35 | ndhG | Exon | T | 11 | 10 |
| 36 | ndhA | intron | A | 11 | 9 |
| Species | Position | Repeat Unit Length (bp) | Repeat Units Sequences | Region |
|---|---|---|---|---|
| A. pseudolaeve | IGS (trnE-trnT) | 31 | TCTATTTCTTATTTCTATATATTCTAATGAT | LSC |
| IGS (petA-psbJ) | 33 | GTAAGAATAAGAACTCAATGGACCTTGCCCCTC | LSC | |
| IGS (psbT-psbN) | 28 | TTGAAGTAAAGTAATGAGCCTCCCATAT | LSC | |
| IGS (petD-rpoA) | 24 | ATGTATCTAGGGACTAGTCCCTTC | LSC | |
| Exon (ycf2) | 24 | AGATCCATTAGATAATGAACTATT | IR | |
| Exon (ycf15) | 21 | TGGTTGTTCGCCGTTCAAGAA | IR | |
| A. longecassidatum | IGS (trnE-trnT) | 31 | TCTATTTCTTATTTCTATATATTCTAATGAT | LSC |
| IGS (petA-psbJ) | 33 | GTAAGAATAAGAACTCAATGGACCTTGCCCCTC | LSC | |
| IGS (psbT-psbN) | 28 | TTGAAGTAAAGTAATGAGCCTCCCATAT | LSC | |
| IGS (petD-rpoA) | 24 | ATGTATCTAGGGACTAGTCCCTTC | LSC | |
| Exon (ycf2) | 24 | AGATCCATTAGATAATGAACTATT | IR | |
| Exon (ycf15) | 21 | TGGTTGTTCGCCGTTCAAGAA | IR |
| Primer Name | Primer Sequence (5′ > 3′) | Position |
|---|---|---|
| AcoTT-F | TGC TTA CGA AGT TGT TCC GGC T | trnK-trnQ |
| AcoTT-R | CAC AAA CCA AAT CCG AGT ACC GA | |
| AcoYN-F | GAT GGA ATC GTC CAT CGC GT | ycf1-ndhF |
| AcoYN-R | TGT AAG TGG AGG ACG GAT CTC T |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Park, I.; Yang, S.; Choi, G.; Kim, W.J.; Moon, B.C. The Complete Chloroplast Genome Sequences of Aconitum pseudolaeve and Aconitum longecassidatum, and Development of Molecular Markers for Distinguishing Species in the Aconitum Subgenus Lycoctonum. Molecules 2017, 22, 2012. https://doi.org/10.3390/molecules22112012
Park I, Yang S, Choi G, Kim WJ, Moon BC. The Complete Chloroplast Genome Sequences of Aconitum pseudolaeve and Aconitum longecassidatum, and Development of Molecular Markers for Distinguishing Species in the Aconitum Subgenus Lycoctonum. Molecules. 2017; 22(11):2012. https://doi.org/10.3390/molecules22112012
Chicago/Turabian StylePark, Inkyu, Sungyu Yang, Goya Choi, Wook Jin Kim, and Byeong Cheol Moon. 2017. "The Complete Chloroplast Genome Sequences of Aconitum pseudolaeve and Aconitum longecassidatum, and Development of Molecular Markers for Distinguishing Species in the Aconitum Subgenus Lycoctonum" Molecules 22, no. 11: 2012. https://doi.org/10.3390/molecules22112012
APA StylePark, I., Yang, S., Choi, G., Kim, W. J., & Moon, B. C. (2017). The Complete Chloroplast Genome Sequences of Aconitum pseudolaeve and Aconitum longecassidatum, and Development of Molecular Markers for Distinguishing Species in the Aconitum Subgenus Lycoctonum. Molecules, 22(11), 2012. https://doi.org/10.3390/molecules22112012






