Molecular Genetic Characterization of an Anthrabenzoxocinones Gene Cluster in Streptomyces Sp. FJS31-2 for the Biosynthesis of BE-24566B and Zunyimycin Ale
Abstract
:1. Introduction
2. Results
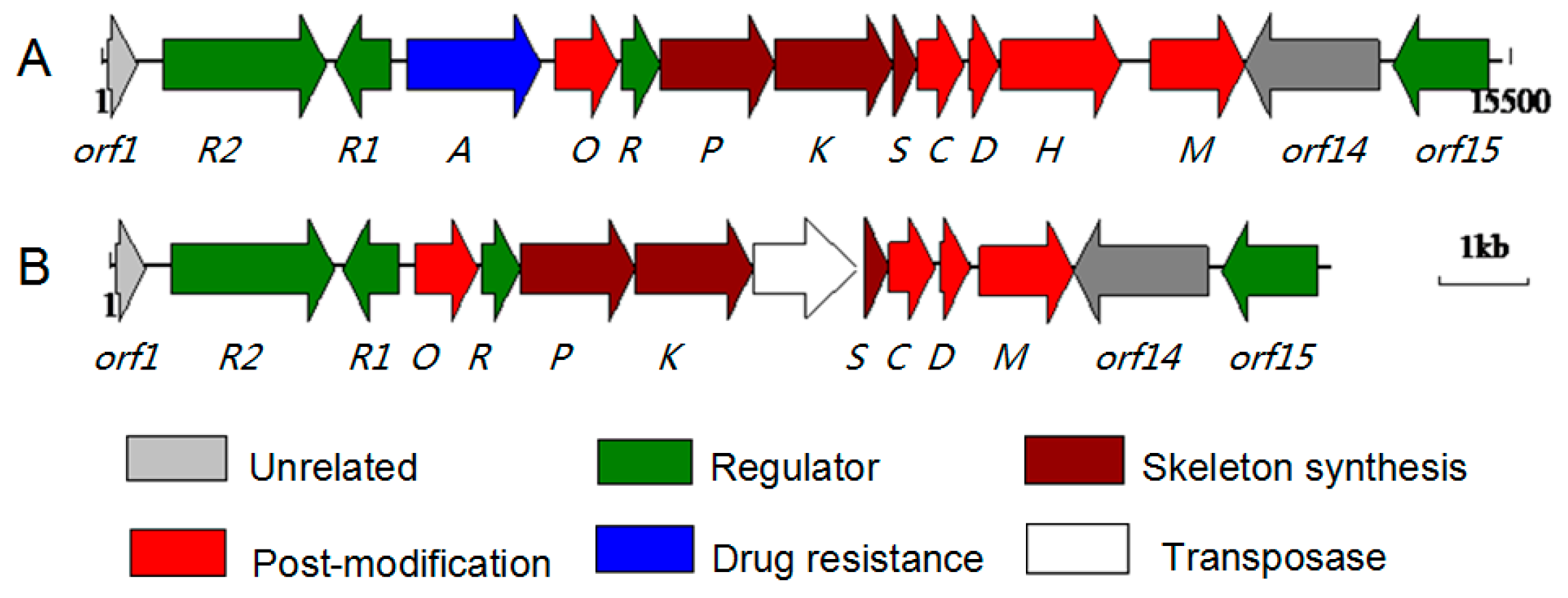
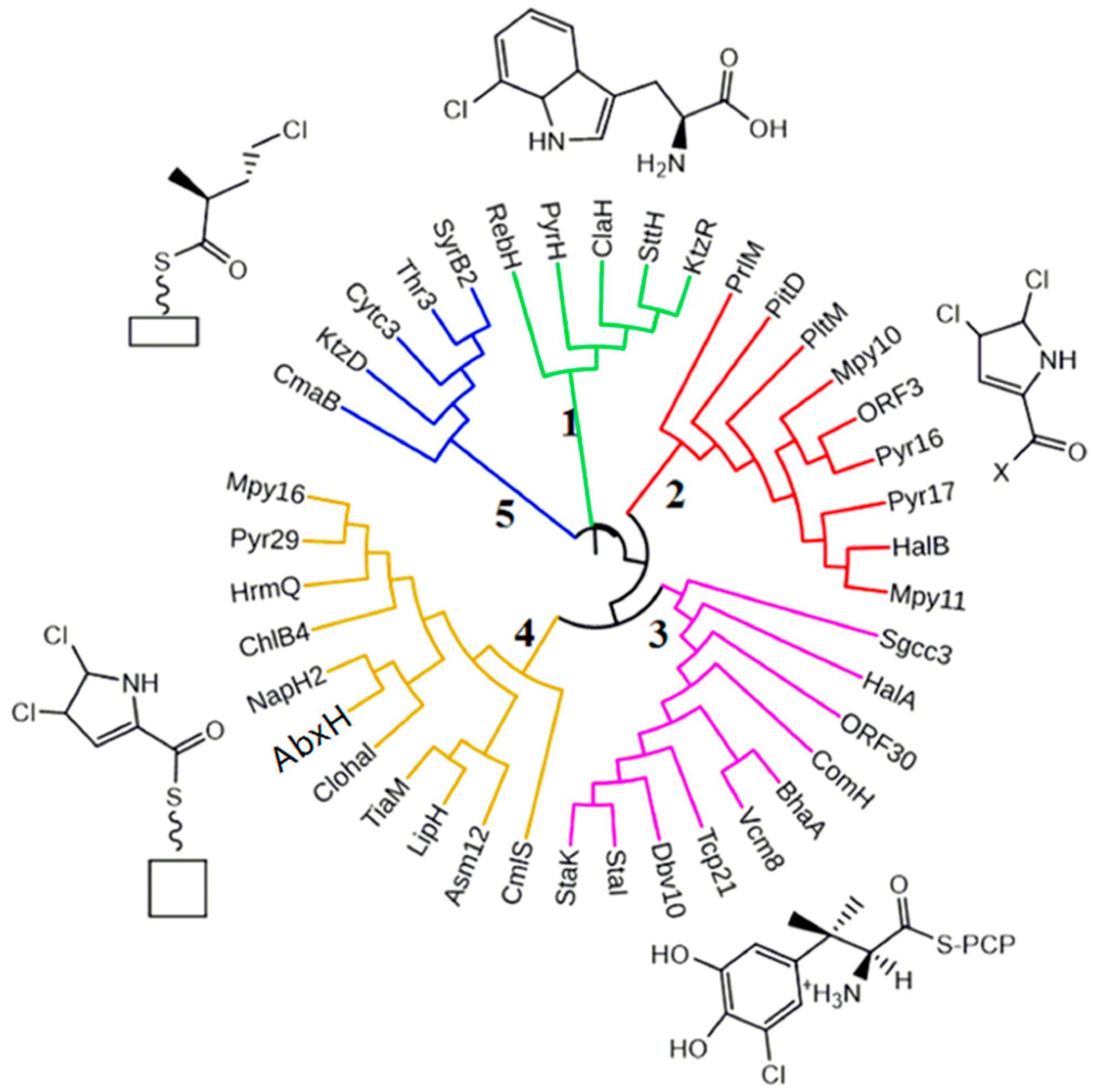
2.1. Comparative Analysis of ABX Gene Clusters
2.2. Characterization of the ABX Gene Cluster
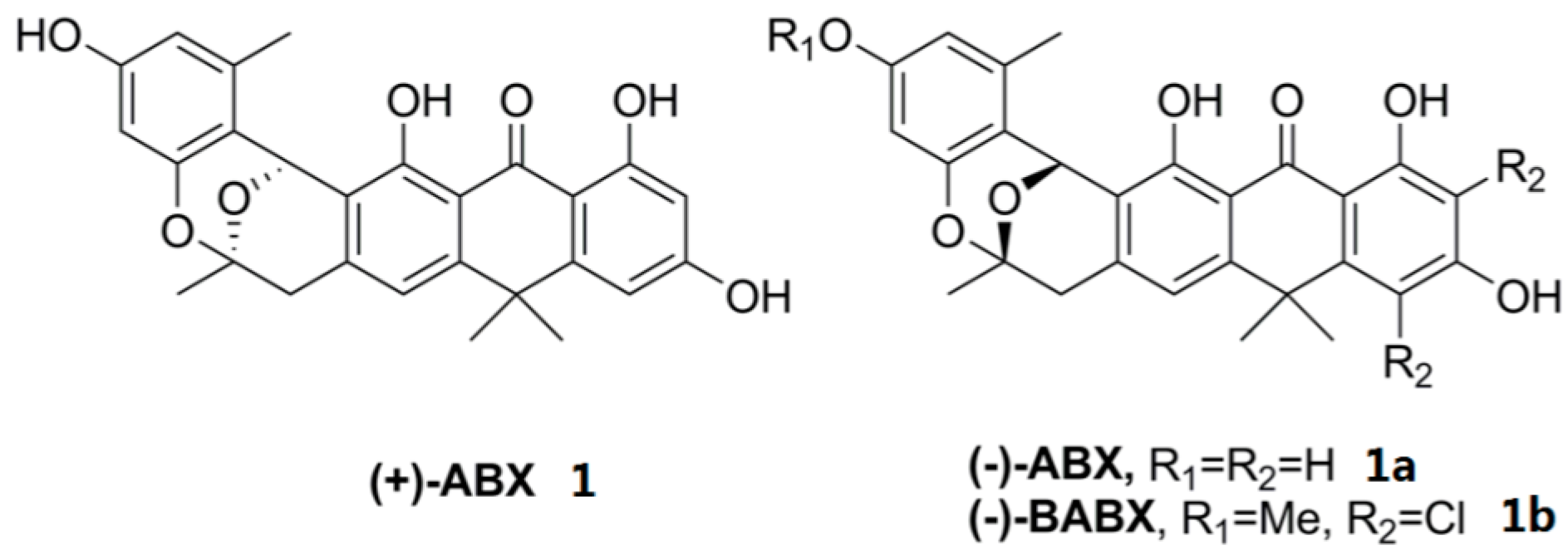
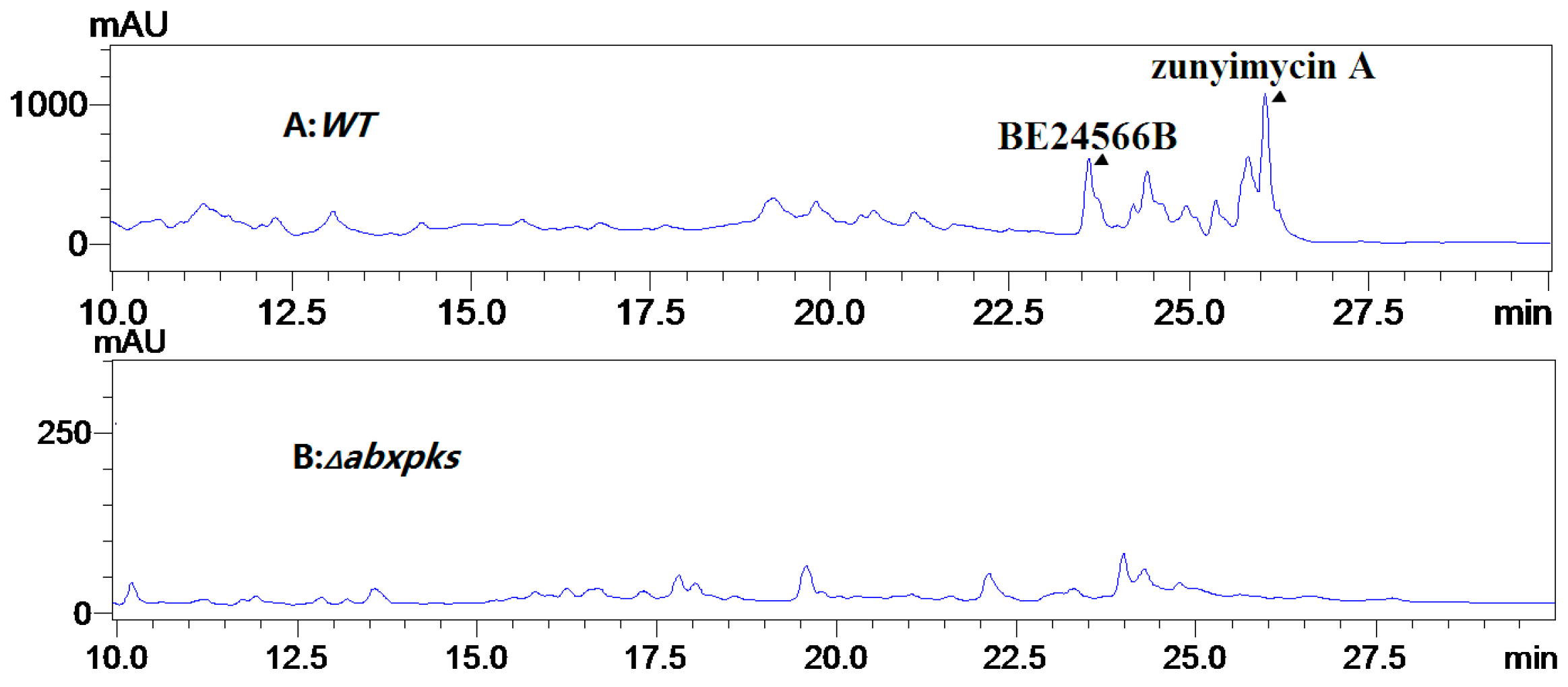
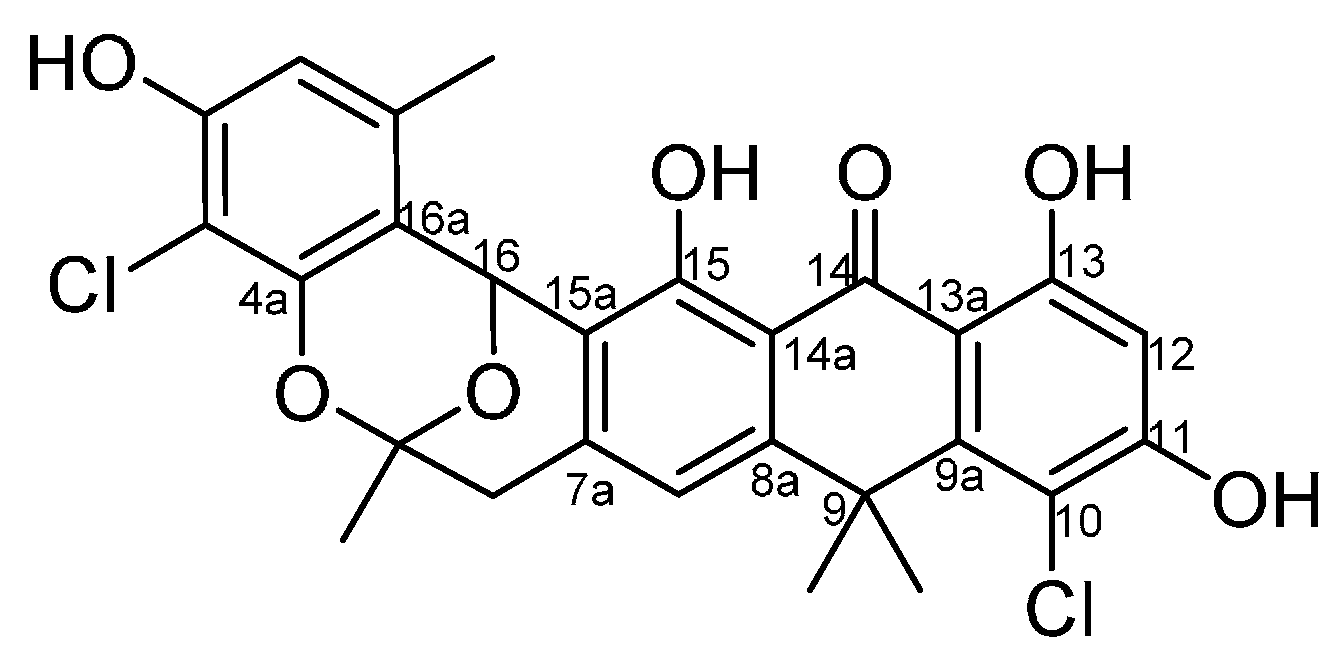
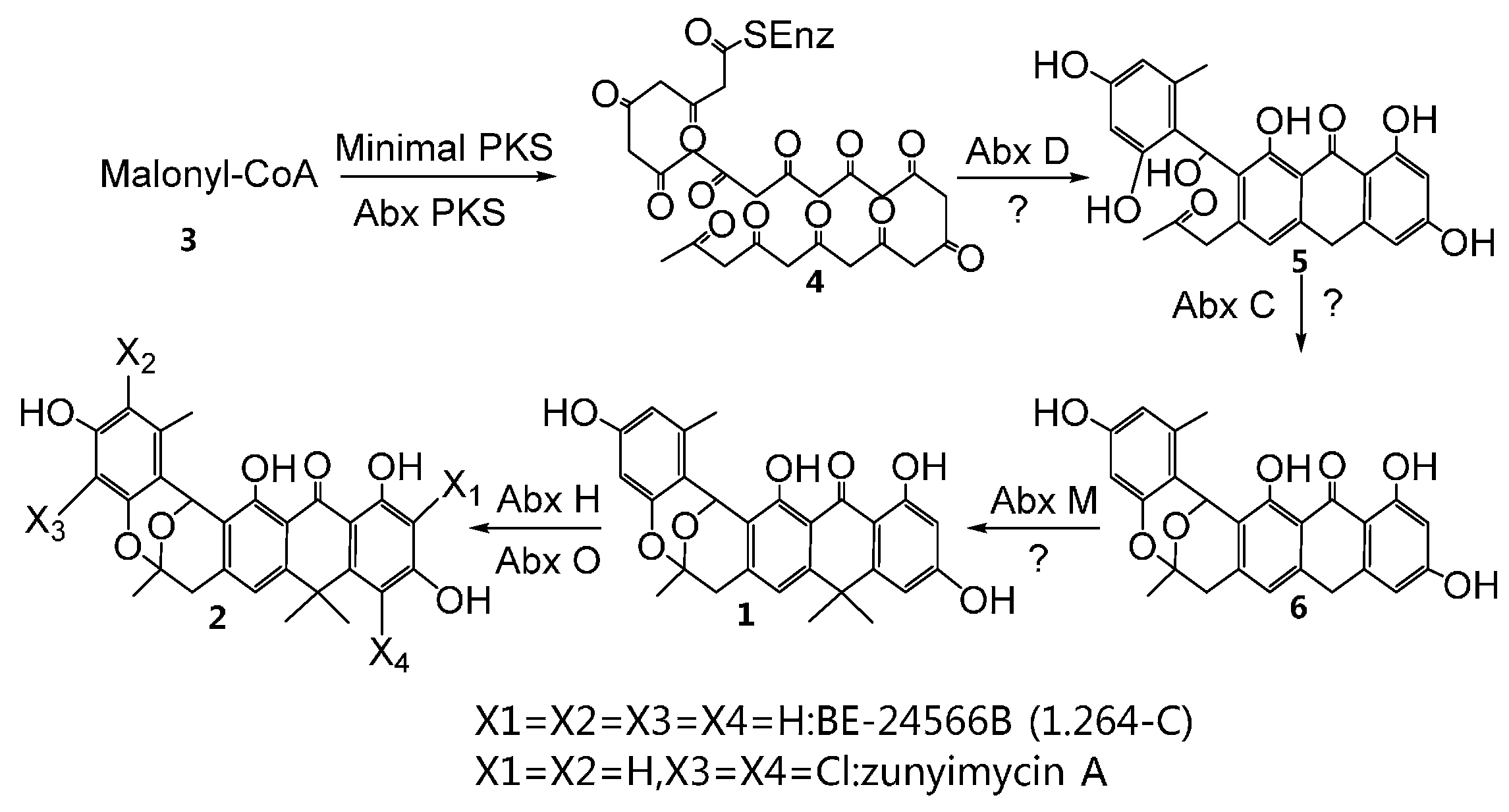
2.3. Chemical Identification of BE-24566B and Zunyimycin A
3. Discussion
4. Materials and Methods
4.1. Strains, Plasmids, and Reagents
4.2. DNA Isolation, Manipulation, and Sequencing
4.3. The Genomic Library and Subclone Library Construction and Screening
4.4. Analysis of the Anthrabenzoxocinone Biosynthetic Gene Cluster
4.5. Inactivation of the Anthrabenzoxocinone Biosynthetic Gene Cluster
4.6. Fermentation, Isolation, and Chemical Identification of BE-24566B and Zunyimycin A
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kallifidas, D.; Kang, H.S.; Brady, S.F. Tetarimycin A, an MRSA-active antibiotic identified through induced expression of environmental DNA gene clusters. J. Am. Chem. Soc. 2012, 134, 19552–19555. [Google Scholar] [CrossRef] [PubMed]
- Arasu, M.V.; Duraipandiyan, V.; Ignacimuthu, S. Antibacterial and antifungal activities of polyketide metabolite from marine Streptomyces sp. AP-123 and its cytotoxic effect. Chemosphere 2013, 90, 479–487. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Christenson, S.D.; Standage, S.; Shen, B. Biosynthesis of the enediyne antitumor antibiotic C-1027. Science 2002, 297, 1170–1173. [Google Scholar] [CrossRef] [PubMed]
- Rui, Z.; Petrícková, K.; Skanta, F.; Pospísil, S.; Yang, Y.; Chen, C.Y.; Tsai, S.F.; Floss, H.G.; Petrícek, M.; Yu, T.W. Biochemical and genetic insights into asukamycin biosynthesis. J. Biol. Chem. 2010, 285, 24915–24924. [Google Scholar] [CrossRef] [PubMed]
- Zaleta-Rivera, K.; Charkoudian, L.K.; Ridley, C.P.; Khosla, C. Cloning, sequencing, heterologous expression, and mechanistic analysis of A-74528 biosynthesis. J. Am. Chem. Soc. 2010, 132, 9122–9128. [Google Scholar] [CrossRef] [PubMed]
- Shen, B. Polyketide biosynthesis beyond the type I, II and III polyketide synthase paradigms. Curr. Opin Chem. Biol. 2003, 7, 285–295. [Google Scholar] [CrossRef]
- Hopwood, D.A.; Khosla, C. Genes for polyketide secondary metabolic pathways in microorganisms and plants. Ciba. Found Symp. 1992, 171, 88–106. [Google Scholar] [PubMed]
- Gao, X.; Wang, P.; Tang, Y. Engineered polyketide biosynthesis and biocatalysis in Escherichia coli. Appl. Microbiol. Biotechnol. 2010, 88, 1233–1242. [Google Scholar] [CrossRef] [PubMed]
- Hertweck, C.; Luzhetskyy, A.; Rebets, Y.; Bechthold, A. Type II polyketide synthases: Gaining a deeper insight into enzymatic teamwork. Nat. Prod. Rep. 2007, 24, 62–90. [Google Scholar] [CrossRef] [PubMed]
- Van Lanen, S.G.; Shen, B. Advances in polyketide synthase structure and function. Curr. Opin Drug Discov. Devel. 2008, 11, 186–195. [Google Scholar] [PubMed]
- Caldara-Festin, G.; Jackson, D.R.; Barajas, J.F.; Valentic, T.R.; Patel, A.B.; Aguilar, S.; Nguyen, M.; Vo, M.; Khanna, A.; Sasaki, E.; et al. Structural and functional analysis of two di-domain aromatase/cyclases from type II polyketide synthases. Proc. Natl. Acad. Sci. USA 2015, 112, E6844–E6851. [Google Scholar] [CrossRef] [PubMed]
- Hillenmeyer, M.E.; Vandova, G.A.; Berlew, E.E.; Charkoudian, L.K. Evolution of chemical diversity by coordinated gene swaps in type II polyketide gene clusters. Proc. Natl. Acad. Sci. USA 2015, 112, 13952–13957. [Google Scholar] [CrossRef] [PubMed]
- Jackson, D.R.; Tu, S.S.; Nguyen, M.; Barajas, J.F.; Schaub, A.J.; Krug, D.; Pistorius, D.; Luo, R.; Müller, R.; Tsai, S.C. Structural Insights into Anthranilate Priming during Type II Polyketide Biosynthesis. ACS Chem. Biol. 2016, 11, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Bailey, C.B.; Pasman, M.E.; Keatinge-Clay, A.T. Substrate structure-activity relationships guide rational engineering of modular polyketide synthase ketoreductases. Chem. Commun. 2015, 52, 792–795. [Google Scholar] [CrossRef] [PubMed]
- Mugnai, M.L.; Shi, Y.; Keatinge-Clay, A.T.; Elber, R. Molecular dynamics studies of modular polyketide synthase ketoreductase stereospecificity. Biochemistry 2015, 54, 2346–2359. [Google Scholar] [CrossRef] [PubMed]
- Sciara, G.; Kendrew, S.G.; Miele, A.E.; Marsh, N.G.; Federici, L.; Malatesta, F.; Schimperna, G.; Savino, C.; Vallone, B. The structure of ActVA-Orf6, a novel type of monooxygenase involved in actinorhodin biosynthesis. EMBO J. 2003, 22, 205–215. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, K. Effective use of heterologous hosts for characterization of biosynthetic enzymes allows production of natural products and promotes new natural product discovery. Chem. Pharm. Bull. 2014, 62, 1153–1165. [Google Scholar] [CrossRef] [PubMed]
- Thompson, T.B.; Katayama, K.; Watanabe, K.; Hutchinson, C.R.; Rayment, I.J. Structural and Functional Analysis of Tetracenomycin F2 Cyclase from Streptomyces glaucescens a type II polyketide cyclase. J. Biol. Chem. 2004, 279, 37956–37963. [Google Scholar] [CrossRef] [PubMed]
- Fritzsche, K.; Ishida, K.; Hertweck, C. Orchestration of discoid polyketide cyclization in the resistomycin pathway. J. Am. Chem. Soc. 2008, 130, 8307–8316. [Google Scholar] [CrossRef] [PubMed]
- Lopez, P.; Hornung, A.; Welzel, K.; Unsin, C.; Wohlleben, W.; Weber, T.; Pelzer, S. Isolation of the lysolipin gene cluster of Streptomyces tendae Tü 4042. Gene 2010, 461, 5–14. [Google Scholar] [CrossRef] [PubMed]
- Akey, D.L.; Gehret, J.J.; Khare, D.; Smith, J.L. Insights from the sea: Structural biology of marine polyketide synthases. Nat. Prod. Rep. 2012, 29, 1038–1049. [Google Scholar] [CrossRef] [PubMed]
- Zhu, T.; Cheng, X.; Liu, Y.; Deng, Z.; You, D. Deciphering and engineering of the final step halogenase for improved chlortetracycline biosynthesis in industrial Streptomyces aureofaciens. Metab. Eng. 2013, 19, 69–78. [Google Scholar] [CrossRef] [PubMed]
- Siitonen, V.; Claesson, M.; Patrikainen, P.; Aromaa, M.; Mäntsälä, P.; Schneider, G.; Metsä-Ketelä, M. Identification of late-stage glycosylation steps in the biosynthetic pathway of the anthracycline nogalamycin. ChemBioChem 2012, 13, 120–128. [Google Scholar] [CrossRef] [PubMed]
- Ma, M.; Kwong, T.; Lim, S.K.; Ju, J.; Lohman, J.R.; Shen, B. Post-polyketide synthase steps in iso-migrastatin biosynthesis, featuring tailoring enzymes with broad substrate specificity. J. Am. Chem. Soc. 2013, 135, 2489–2492. [Google Scholar] [CrossRef] [PubMed]
- Kojiri, K.; Nakajima, S.; Fuse, A.; Suzuki, H.; Suda, H. BE-24566B, a new antibiotic produced by streptomyces violaceusniger. J. Antibiot. 1995, 48, 1506–1508. [Google Scholar] [CrossRef] [PubMed]
- Herath, K.B.; Jayasuriya, H.; Guan, Z.; Schulman, M.; Ruby, C.; Sharma, N.; MacNaul, K.; Menke, J.G.; Kodali, S.; Galgoci, A.; et al. Anthrabenzoxocinones from Streptomyces sp. as liver X receptor ligands and antibacterial agents. J. Nat. Prod. 2005, 68, 1437–1440. [Google Scholar] [CrossRef] [PubMed]
- Kodali, S.; Galgoci, A.; Young, K.; Painter, R.; Silver, L.L.; Herath, K.B.; Singh, S.B.; Cully, D.; Barrett, J.F.; Schmatz, D.; Wang, J. Determination of selectivity and efficacy of fatty acid synthesis inhibitors. J. Biol. Chem. 2005, 280, 1669–1677. [Google Scholar] [CrossRef] [PubMed]
- Lam, Y.K.T.; Hensens, O.; Helms, G.; Williams, D.J.; Nallin, M.; Smith, J.; Gartner, S.; Rodriguez, L.H.; Stevens-Miles, S. L-755,805, a new polyketide endothelin binding inhibitor from an actinomycete. Tetrahedron Lett. 1995, 36, 2013–2016. [Google Scholar] [CrossRef]
- Winter, J.M.; Moffitt, M.C.; Zazopoulos, E.; McAlpine, J.B.; Dorrestein, P.C.; Moore, B.S. Molecular basis for chloronium-mediated meroterpene cyclization: Cloning, sequencing, and heterologous expression of the napyradiomycin biosynthetic gene cluster. J. Biol. Chem. 2007, 282, 16362–16368. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Zhu, H.; Ruan, J.; Qian, W.; Fang, X.; Shi, Z.; Li, Y.; Li, S.; Shan, G.; Kristiansen, K.; et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 2010, 20, 265–272. [Google Scholar] [CrossRef] [PubMed]
- Shirling, E.B.; Gottlieb, D. Methods for characterizing Streptomyces species. Int. J. Syst. Bacteriol. 1966, 16, 313–340. [Google Scholar] [CrossRef]
- Hopwood, D.A. Genetic manipulation of Streptomyces polyketide synthase genes for novel secondary metabolite production. FEMS Microbiol. Rev. 1995, 16, 233–234. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Manual; Cold Spring Harbor: New York, NY, USA, 2001. [Google Scholar]
- Blin, K.; Medema, M.H.; Kazempour, D.; Fischbach, M.A.; Breitling, R.; Takano, E.; Weber, T. antiSMASH 2.0—A versatile platform for genome mining of secondary metabolite producers. Nucleic Acids Res. 2013. [Google Scholar] [CrossRef] [PubMed]
- Darling, A.E.; Mau, B.; Perna, N.T. Progressive mauve: Multiplegenome alignment with gene gain, loss and rearrangement. PLoS ONE 2010, 5, e11147. [Google Scholar] [CrossRef] [PubMed]
- Sample Availability: Samples of the compounds zunyimycin A and BE24566B are available from the authors.

| S. sp. FJS31-2 | S. sp. FXJ1.264 | Identit % | Deduced Function | ||
|---|---|---|---|---|---|
| Protein | Size aa | Protein | Size aa | ||
| Orf1 | 87 | 1.264DNAGM006278 | 87 | 93 | Hypothetical protein. |
| 1.264DNAGM006279 | 319 | Cation efflux family | |||
| AbxR2 | 616 | 1.264DNAGM006280 | 616 | 91 | SARP transcriptional regulator |
| AbxR2 | 207 | 1.264DNAGM006281 | 219 | 88 | TetR transcriptional regulator |
| AbxA | 490 | MFS transporter | |||
| AbxO | 227 | 1.264DNAGM006282 | 231 | 90.5 | Monooxygenase |
| AbxR | 141 | 1.264DNAGM006283 | 141 | 93.6 | AraC transcriptional regulator |
| AbxP | 421 | 1.264DNAGM006284 | 421 | 96 | Beta-ketoacyl_synthase, t2PKS |
| AbxK | 426 | 1.264DNAGM006285 | 424 | 90.3 | Beta-ketoacyl_synthase, t2PKS |
| 1.264DNAGM006286 | 394 | Transposase | |||
| AbxS | 85 | 1.264DNAGM006287 | 88 | 86.4 | Acyl carrier protein |
| AbxC | 167 | 1.264DNAGM006288 | 168 | 86.3 | cyclase/dehydrase |
| AbxD | 113 | 1.264DNAGM006289 | 113 | 88.5 | Polyketide synthesis cyclase |
| AbxH | 436 | tryptophan halogenase | |||
| AbxM | 343 | 1.264DNAGM006290 | 349 | 91.4 | O-methyltransferase |
| Orf14 | 491 | 1.264DNAGM006291 | 490 | 91.0 | Lipase maturation factor |
| Orf15 | 348 | 1.264DNAGM006292 | 348 | 94.5 | LacI transcription regulator |
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lü, Y.; Yue, C.; Shao, M.; Qian, S.; Liu, N.; Bao, Y.; Wang, M.; Liu, M.; Li, X.; Wang, Y.; et al. Molecular Genetic Characterization of an Anthrabenzoxocinones Gene Cluster in Streptomyces Sp. FJS31-2 for the Biosynthesis of BE-24566B and Zunyimycin Ale. Molecules 2016, 21, 711. https://doi.org/10.3390/molecules21060711
Lü Y, Yue C, Shao M, Qian S, Liu N, Bao Y, Wang M, Liu M, Li X, Wang Y, et al. Molecular Genetic Characterization of an Anthrabenzoxocinones Gene Cluster in Streptomyces Sp. FJS31-2 for the Biosynthesis of BE-24566B and Zunyimycin Ale. Molecules. 2016; 21(6):711. https://doi.org/10.3390/molecules21060711
Chicago/Turabian StyleLü, Yuhong, Changwu Yue, Meiyun Shao, Shengyan Qian, Ning Liu, Yuxin Bao, Miao Wang, Minghao Liu, Xiaoqian Li, Yinyin Wang, and et al. 2016. "Molecular Genetic Characterization of an Anthrabenzoxocinones Gene Cluster in Streptomyces Sp. FJS31-2 for the Biosynthesis of BE-24566B and Zunyimycin Ale" Molecules 21, no. 6: 711. https://doi.org/10.3390/molecules21060711
APA StyleLü, Y., Yue, C., Shao, M., Qian, S., Liu, N., Bao, Y., Wang, M., Liu, M., Li, X., Wang, Y., & Huang, Y. (2016). Molecular Genetic Characterization of an Anthrabenzoxocinones Gene Cluster in Streptomyces Sp. FJS31-2 for the Biosynthesis of BE-24566B and Zunyimycin Ale. Molecules, 21(6), 711. https://doi.org/10.3390/molecules21060711






