Structure Identification and Anti-Cancer Pharmacological Prediction of Triterpenes from Ganoderma lucidum
Abstract
:1. Introduction
2. Results and Discussion
2.1. Identification of the Compounds
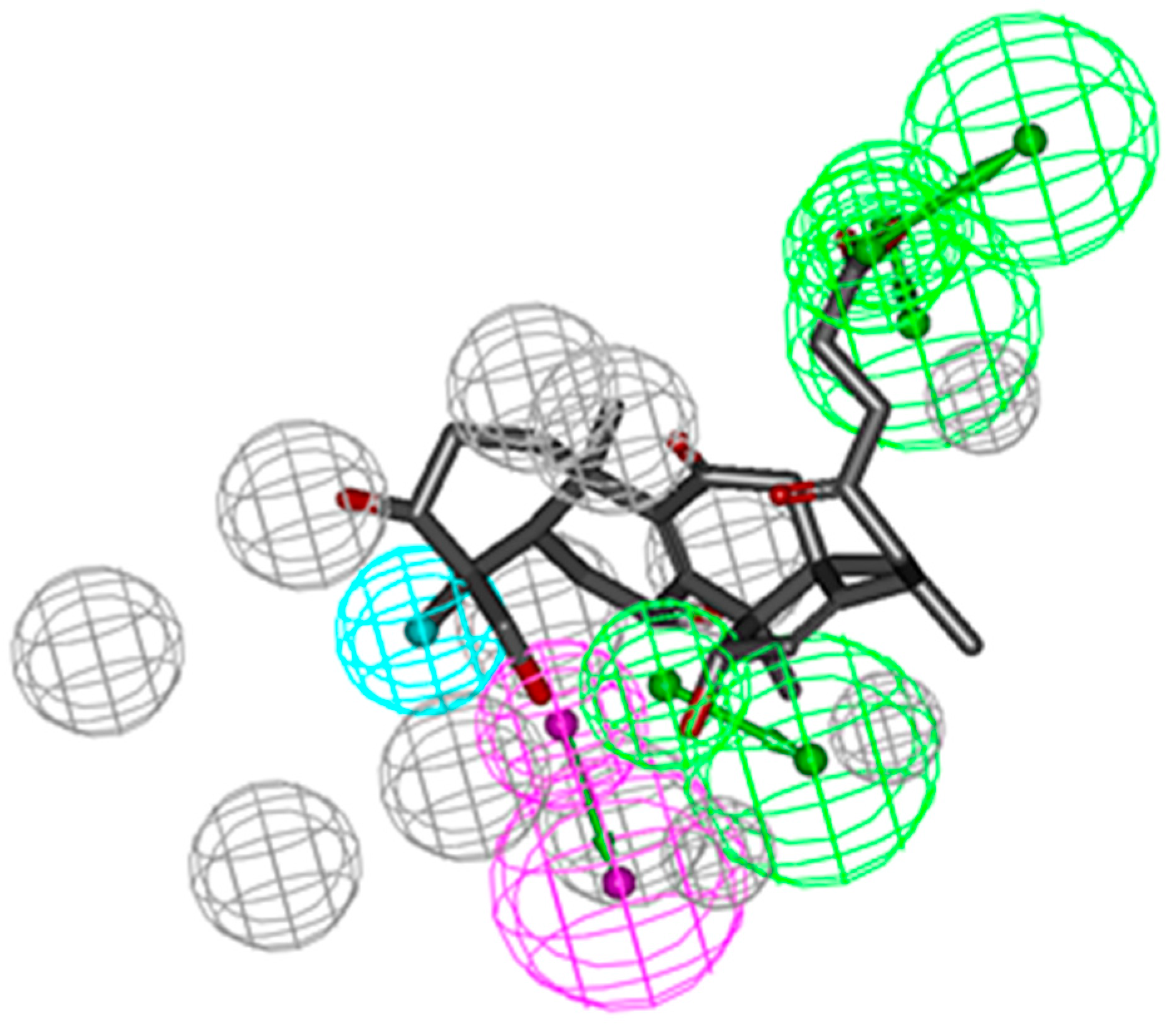
2.2. Pharmacophoric Profiling of GTs
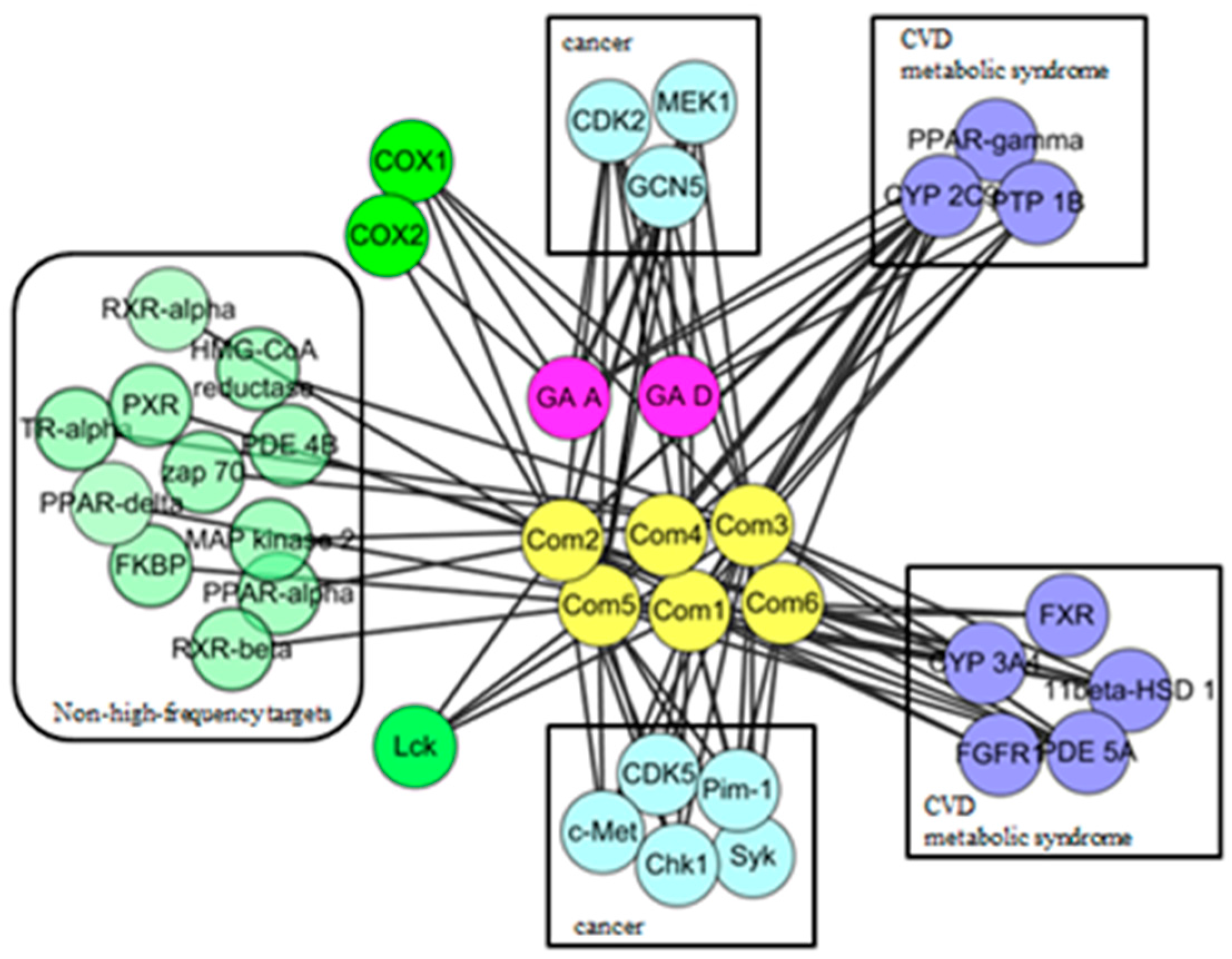
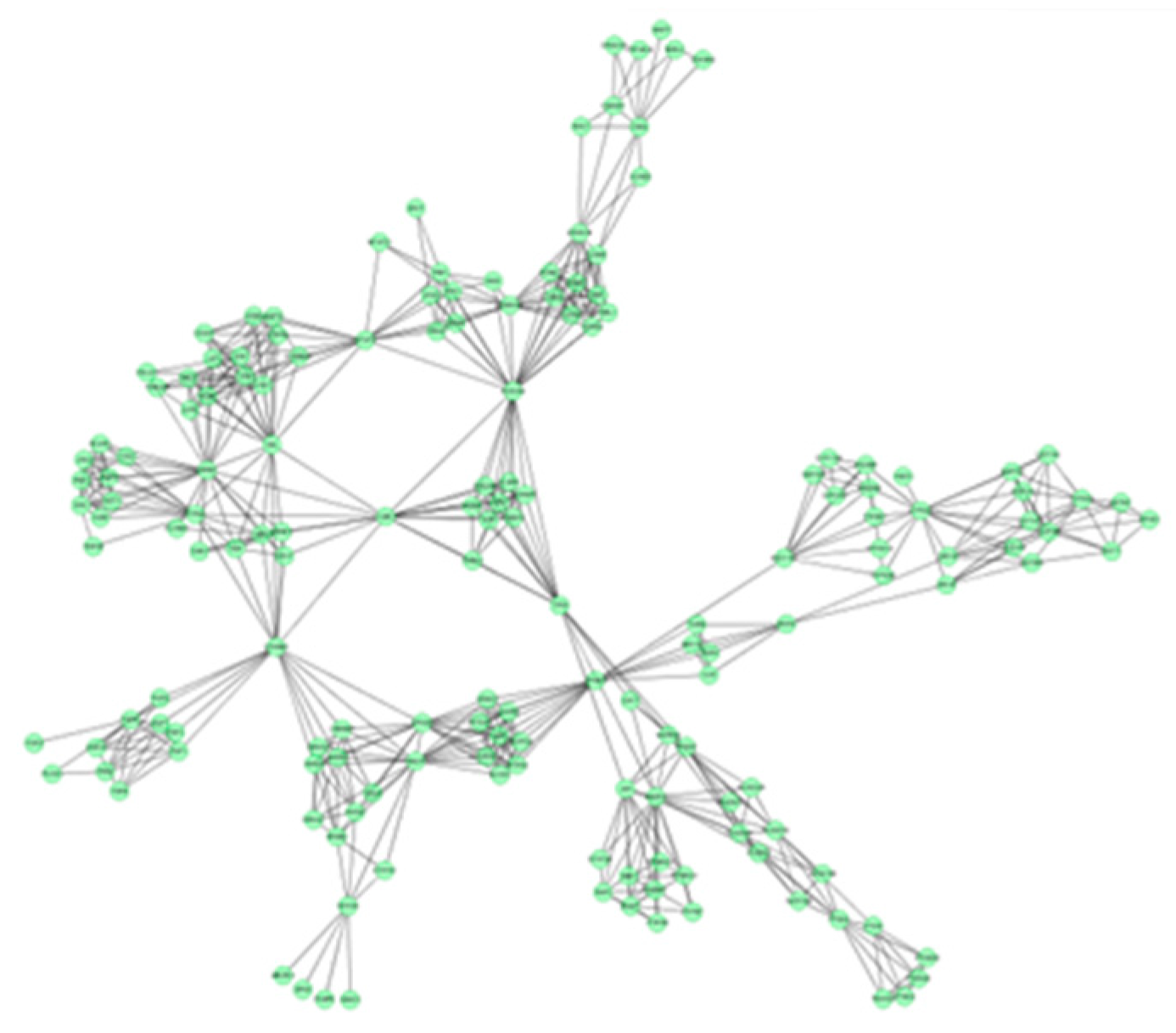
2.3. Protein Interaction Network Analysis of GTs
3. Materials and Methods
3.1. Plant Material
3.2. Extraction
3.3. Isolation
3.4. Reverse Target Identification
3.5. Network Construction and Analysis
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Boh, B. Ganoderma lucidum: A potential for biotechnological production of anti-cancer and immunomodulatory drugs. Recent Pat. Anticancer Drug. Discov. 2013, 8, 255–287. [Google Scholar] [CrossRef] [PubMed]
- Xia, Q.; Zhang, H.Z.; Sun, X.F.; Zhao, H.J.; Wu, L.F.; Zhu, D.; Yang, G.H.; Shao, Y.Y.; Zhang, X.X.; Mao, X.; Zhang, L.Z.; She, G.M. A comprehensive review of the structure elucidation and biological activity of triterpenoids from Ganoderma spp. Molecules 2014, 19, 17478–17535. [Google Scholar] [CrossRef] [PubMed]
- Feng, L.; Yuan, L.; Du, M.; Chen, Y.; Zhang, M.H.; Gu, J.F.; He, J.J.; Wang, Y.; Cao, W. Anti-Lung cancer activity through enhancement of immunomodulation and induction of cell apoptosis of total triterpenes extracted from Ganoderma luncidum (Leyss. ex Fr.) Karst. Molecules 2013, 18, 9966–9981. [Google Scholar] [CrossRef] [PubMed]
- Radwan, F.F.; Hossain, A.; God, J.M.; Leaphart, N.; Elvington, M.; Nagarkatti, M.; Tomlinson, S.; Haque, A. Reduction of myeloid-derived suppressor cells and lymphoma growth by a natural triterpenoid. J. Cell. Biochem. 2015, 116, 102–114. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.M.; Yang, X.L.; Wang, B.W.; Zhu, H.S.; Xu, J.L. Antitumor activity of ethanol-soluble and acidic components from Ganodermalucidum. Nat. Prod. Res. 2004, 16, 146–148. [Google Scholar]
- Smina, T.P.; Mathew, J.; Janardhanan, K.K.; Devasagayam, T.P. Antioxidant activity and toxicity profile of total triterpenes isolated from Ganoderma lucidum (Fr.) P. Karst occurring in South India. Environ. Toxicol. Pharmacol. 2011, 32, 438–446. [Google Scholar] [CrossRef] [PubMed]
- Li, P.Z.; Zhang, K.C. Isolation, purification, and antimicrobial activity of ganoderic acids M1 from the fermented mycelia of Ganoderma lucidum. Nat. Prod. Res. 1999, 11, 67–70. [Google Scholar]
- Liu, J.; Shimizu, K.; Konishi, F.; Noda, K.; Kumamoto, S.; Kurashiki, K.; Kondo, R. Antiandrogenic activities of the triterpenoids fraction of Ganoderma lucidum. Food Chem. 2007, 100, 1691–1696. [Google Scholar] [CrossRef]
- Mizushina, Y.; Takahashi, N.; Hanashima, L.; Koshino, H.; Esumi, Y.; Uzawa, J.; Sugawara, F.; Sakaguchi, K. Lucidenic acid O and lactone, new terpene inhibitors of eukaryotic DNA polymerases from a basidiomycete, Ganoderma lucidum. Bioorg. Med. Chem. 1999, 7, 2047–2052. [Google Scholar] [CrossRef]
- Yan, Z.; Xia, B.; Qiu, M.H.; Li, S.D.; Xu, H.X. Fast analysis of triterpenoids in Ganoderma lucidum spores by ultra-performance liquid chromatography coupled with triple quadrupole mass spectrometry. Biomed. Chromatogr. 2013, 27, 1560–1567. [Google Scholar] [CrossRef] [PubMed]
- Wei, Z.H.; Duan, Y.Y.; Qian, Y.Q.; Guo, X.F.; Li, Y.J.; Jin, S.H.; Zhou, Z.X.; Shan, S.Y.; Wang, C.R.; Chen, X.J.; et al. Screening of Ganoderma strains with high polysaccharides and ganoderic acid contents and optimization of the fermentation medium by statistical methods. Bioprocess. Biosyst. Eng. 2014, 37, 1789–1797. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Xie, Z.P.; Huang, Z.S.; Li, H.; Wei, A.Y.; Di, J.M.; Xiao, H.J.; Zhang, Z.G.; Cai, L.H.; Tao, X.; et al. Total triterpenoids from Ganoderma Lucidum suppresses prostate cancer cell growth by inducing growth arrest and apoptosis*. J. Huazhong Univ. Sci. Technol. Med. Sci. 2015, 35, 736–741. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.Z.; Yin, R.H.; Chen, H.P.; Feng, T.; Li, Z.H.; Dong, Z.J.; Cui, B.K.; Liu, J.K. Two new triterpenoids from fruiting bodies of fungus Ganoderma lucidum. J. Asian Nat. Prod. Res. 2015, 17, 750–755. [Google Scholar] [CrossRef] [PubMed]
- Suarez-Arroyo, I.J.; Rosario-Acevedo, R.; Aguilar-Perez, A.; Clemente, P.L.; Cubano, L.A.; Serrano, J.; Schneider, R.J.; Martı´nez-Montemayor, M.M. Anti-tumor effects of Ganoderma lucidum (Reishi) in inflammatory breast cancer in in vivo and in vitro models. PLoS ONE 2013, 8, 1–11. [Google Scholar] [CrossRef]
- Wu, G.S.; Guo, J.J.; Bao, J.L.; Li, X.W.; Chen, X.P.; Lu, L.L.; Wang, Y.T. Anti-cancer properties of triterpenoids isolated from Ganoderma lucidum—A review. Expert Opin. Investig. Drugs 2013, 22, 981–992. [Google Scholar] [CrossRef] [PubMed]
- Weng, C.J.; Chau, C.F.; Yen, G.C.; Liao, J.W.; Chen, D.H.; Chen, K.D. Inhibitory effects of Ganoderma lucidum on tumorigenesis and metastasis of human hepatoma cells in cells and animal models. J. Agric. Food Chem. 2009, 57, 5049–5057. [Google Scholar] [CrossRef] [PubMed]
- Yue, Q.X.; Song, X.Y.; Ma, C.; Feng, L.X.; Guan, S.H.; Wu, W.Y.; Yang, M.; Jiang, B.H.; Liu, X.; Cui, Y.J.; et al. Effects of triterpenes from Ganoderma lucidum on protein expression profile of HeLa cells. Phytomedicine 2010, 17, 606–613. [Google Scholar] [CrossRef] [PubMed]
- Thyagarajan, A.; Jedinak, A.; Nquyen, H.; Terry, C.; Baldridge, L.A.; Jiang, J.; Sliva, D. Triterpenes from Ganoderma lucidum induce autophagy in colon cancer through the inhibition of p38 mitogen-activated kinase (p38 MAPK). Nutr. Cancer 2010, 62, 630–640. [Google Scholar] [CrossRef] [PubMed]
- Tanrikulu, Y.; Krüger, B.; Proschak, E. The holistic integration of virtual screening in drug discovery. Drug Discov. Today 2013, 18, 358–364. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Li, H.L.; Zhang, Q.J.; Bao, X.G.; Yu, K.Q.; Luo, X.M.; Zhu, W.L.; Jiang, H.L. Pharmacophore-based virtual screening versus docking-based virtual screening: A benchmark comparison against eight targets. Acta Pharmacol. Sin 2009, 30, 1694–1708. [Google Scholar] [CrossRef] [PubMed]
- Grienke, U.; Kaserer, T.; Pfluger, F.; Mair, C.E.; Langer, T.; Schuster, D.; Rollinger, J.M. Accessing biological actions of Ganoderma secondary metabolites by in silico profiling. Phytochemistry 2014, 114, 114–124. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Jiang, P.; Zhang, W. Molecular networks for the study of TCM pharmacology. Brief. Bioinform. 2010, 11, 417–430. [Google Scholar] [CrossRef] [PubMed]
- Romero-Durán, F.J.; Alonso, N.; Yañez, M.; Caamaño, O.; García-Mera, X.; González-Díaz, H. Brain-inspired cheminformatics of drug-target brain interactome, synthesis, and assay of TVP1022 derivatives. Neuropharmacology 2016, 103, 270–278. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Niu, X.; Lu, C.; Jiang, M.; Xiao, G.G.; Lu, A. The effect of curcumin on human bronchial epithelial cells exposed to fine particulate matter: A predictive analysis. Molecules 2012, 17, 12406–12426. [Google Scholar] [CrossRef] [PubMed]
- Zheng, S.; Ren, Z.; Zhang, Y.; Qiao, Y. Anti-inflammatory mechanism research of tanshinone II A by module-based network analysis. Biomed. Mater. Eng. 2014, 24, 3815–3824. [Google Scholar] [PubMed]
- Gill, B.S.; Kumar, S. Differential algorithms-assisted molecular modeling-based identification of mechanistic binding of ganoderic acids. Med. Chem. Res. 2015, 24, 3483–3493. [Google Scholar] [CrossRef]
- Yue, Q.X.; Cao, Z.W.; Guan, S.H.; Liu, X.H.; Tao, L.; Wu, W.Y.; Li, Y.X.; Yang, P.Y.; Liu, X.; Guo, D.A. Proteomics characterization of the cytotoxicity mechanism of ganoderic acid D and computer-automated estimation of the possible drug target network. Mol. Cell. Proteom. 2008, 7, 949–961. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.L.; Ma, Q.Y.; Huang, S.Z.; Guo, Z.K.; Ma, H.X.; Guo, J.C.; Dai, H.F.; Zhao, Y.X. Three new lanostanoid triterpenes from the fruiting bodies of Ganoderma tropicum. J. Asian Prod. Res. 2013, 15, 357–362. [Google Scholar] [CrossRef]
- Zhang, S.S.; Wang, Y.G.; Ma, Q.Y.; Huang, S.Z.; Hu, L.L.; Dai, H.F.; Yu, Z.F.; Zhao, Y.X. Three new lanostanoids from the Mushroom Ganoderma tropicum. Molecules 2015, 20, 3281–3289. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.L.; Ma, Q.J.; Huang, S.Z.; Guo, Z.K.; Ma, H.X.; Guo, J.C.; Dai, H.F.; Zhao, Y.X. A new nortriterpenoid from the fruiting bodies of Ganoderma tropicum. Phytochem. Lett. 2014, 7, 11–13. [Google Scholar] [CrossRef]
- Lu, Z.Q.; Chen, G.T.; Zhang, J.Q.; Huang, H.L.; Guan, S.H.; Guo, D.A. Four new lanostane triterpenoids from Euphorbia humifusa. Helv. Chim. Acta 2007, 90, 2245–2250. [Google Scholar] [CrossRef]
- Takaishi, Y.; Ohashi, T.; Tomimatsu, T. Ergosta-7,22-dien-3β-ol glycoside from Tylopilus Neofelleus. Phytochemistry 1989, 28, 945–947. [Google Scholar] [CrossRef]
- Wu, X.L.; Lin, S.; Zhu, C.G.; Zhao, F.; Yu, Y.; Yue, Z.G.; Liu, B.; Yang, Y.C.; Dai, J.G.; Shi, J.G. Studies on constituents of cultures of fungus Phellinusigniarius. China J. Chin. Mater. Med. 2011, 36, 875–880. [Google Scholar]
- Jiang, J.F.; Lu, J.Y.; Lu, D.; Liang, Z.J.; Li, L.C.; Ouyang, S.S.; Kong, X.Q.; Jiang, H.L.; Shen, B.R.; Luo, C. Investigation of the acetylation mechanism by GCN5 histone acetyltransferase. PLoS ONE 2012, 7, 539–539. [Google Scholar] [CrossRef] [PubMed]
- Martinez, E.; Kundu, T.K.; Fu, J.; Roeder, R.G. A human SPT3-TAFII31-GCN5-Lacetylase complex distinct from transcription factor IID. J. Biol. Chem. 1998, 273, 23781–23785. [Google Scholar] [CrossRef] [PubMed]
- Brand, M.; Yamamoto, K.; Staub, A.; Tora, L. Identification of TATA-binding protein-free TAFII-containing complex subunits suggests a role in nucleosome acetylation and signal transduction. J. Biol. Chem. 1999, 274, 18285–18289. [Google Scholar] [CrossRef] [PubMed]
- Kazantseva, J.; Palm, K. Diversity in TAF proteomics: Consequences for cellular differentiation and migration. Int. J. Mol. Sci. 2014, 15, 16680–16697. [Google Scholar] [CrossRef] [PubMed]
- Chohan, T.A.; Qian, H.; Pan, Y.; Chen, J.Z. Cyclin-dependent kinase-2 as a target for cancer therapy: Progress in the development of CDK2 inhibitors as anti-cancer agents. Curr. Med. Chem. 2014, 22, 237–263. [Google Scholar] [CrossRef]
- Moore, J.D.; Yang, J.; Truant, R.; Kornbluth, S. Nuclear import of Cdk/cyclin complexes: Identification of distinct mechanisms for import of Cdk2/cyclinE and Cdc2/cyclinB1. J. Cell. Biol. 1999, 144, 213–224. [Google Scholar] [CrossRef] [PubMed]
- Etemadmoghadam, D.; Au-Yeung, G.; Wall, M.; Mitchell, C.; Kansara, M.; Loehrer, E.; Batzios, C.; George, J.; Ftouni, S.; Weir, B.A.; et al. Resistance to CDK2 inhibitors is associated with selection of polyploid cells in CCNE1-amplified ovarian cancer. Clin. Cancer. Res. 2013, 19, 5960–5971. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.Y.; Liu, D.; Ye, M.; Han, J.; Deng, S.; Ma, X.C.; Zhao, Y.Y.; Zhang, B.J.; Shen, X.; Che, Q.M. Structural characterization of minor metabolites and pharmacokinetics of ganoderic acid C2 in rat plasma by HPLC coupled with electrospray ionization tandem mass spectrometry. J. Pharm. Biomed. Anal. 2013, 65, 64–73. [Google Scholar] [CrossRef] [PubMed]
- Schuster, D. 3D pharmacophores as tools for activity profiling. Drug. Discov. Today Technol. 2010, 7, e205–e211. [Google Scholar] [CrossRef] [PubMed]
- Sakkiah, S.; Thangapandian, S.; John, S.; Kwon, Y.J.; Lee, K.W. 3D-QSAR pharmacophore based virtual screening and molecular docking for identification of potential HSP90 inhibitors. Eur. J. Med. Chem. 2010, 45, 2132–2140. [Google Scholar] [CrossRef] [PubMed]
- PharmaDB. Available online: http:// www.inteligand.com/pharmdb/ (accessed on 10 July 2015).
- Steindl, T.M.; Schuster, D.; Laggner, C.; Langer, T. Parallel screening: anovel concept in pharmacophoremodeling and virtual screening. J. Chem. Inf. Model. 2006, 46, 2146–2157. [Google Scholar] [CrossRef] [PubMed]
- Schuster, D.; Waltenberger, B.; Kirchmair, J.; Distinto, S.; Markt, P.; Stuppner, H.; Rollinger, J.M.; Wolber, G. Predicting cyclooxygenase inhibition by three-dimensional pharmacophoricprofiling. Part I: Model generation, validation and applicability in ethnopharmacology. Mol. Inform. 2010, 29, 75–86. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- String 9.1. Available online: http://string-db.org (accessed on 19 August 2015).
- Assenov, Y.; Ramirez, F.; Schelhorn, S.E.; Lengauer, T.; Albrecht, M. Computing topological parameters of biological networks. Bioinformatics 2008, 24, 282–284. [Google Scholar] [CrossRef] [PubMed]
- Bader, G.D.; Hogue, C.W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform. 2003, 4, 1–27. [Google Scholar] [CrossRef]
- Maere, S.; Heymans, K.; Kuiper, M. BiNGO: A Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 2005, 21, 3448–3449. [Google Scholar] [CrossRef] [PubMed]
- Sample Availability: Samples of the compounds (1–24) are available from the authors.

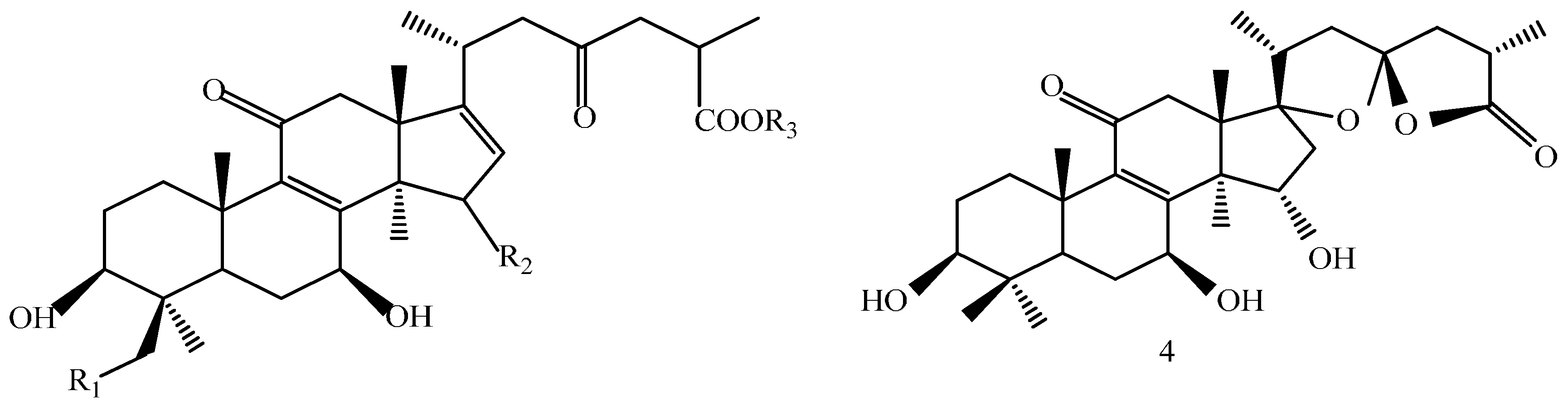
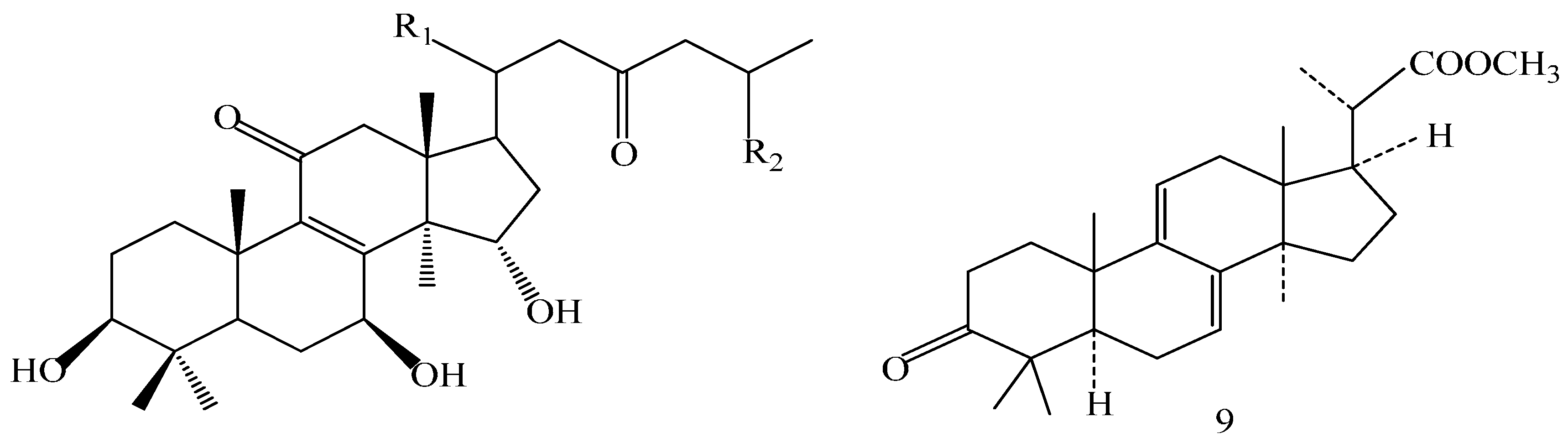
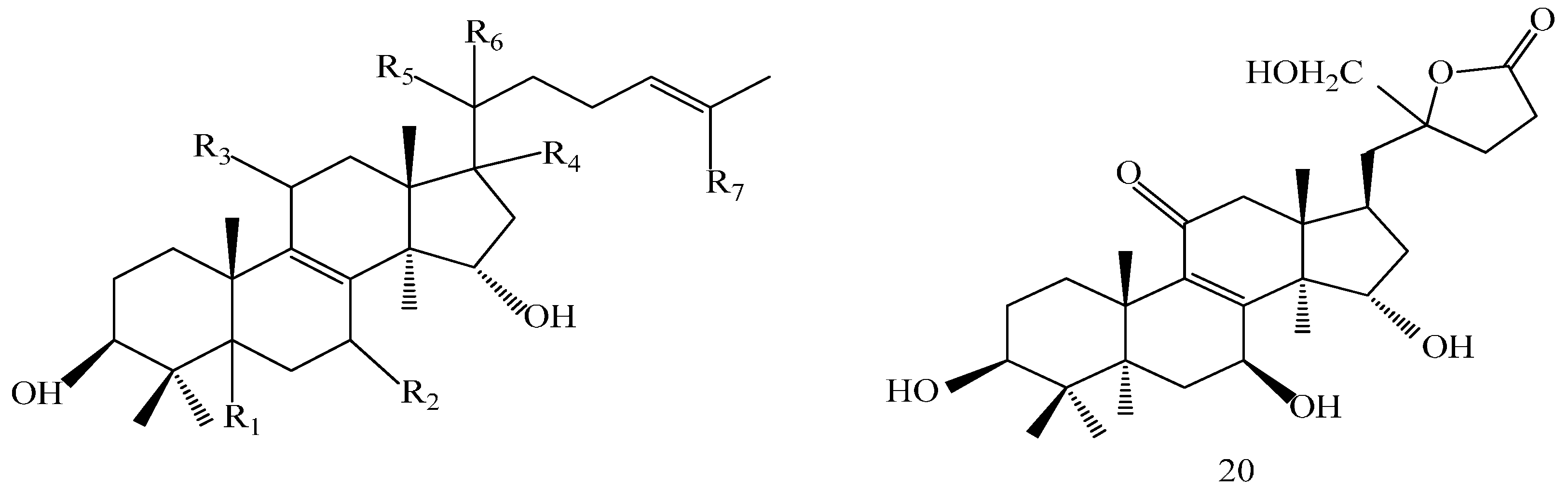
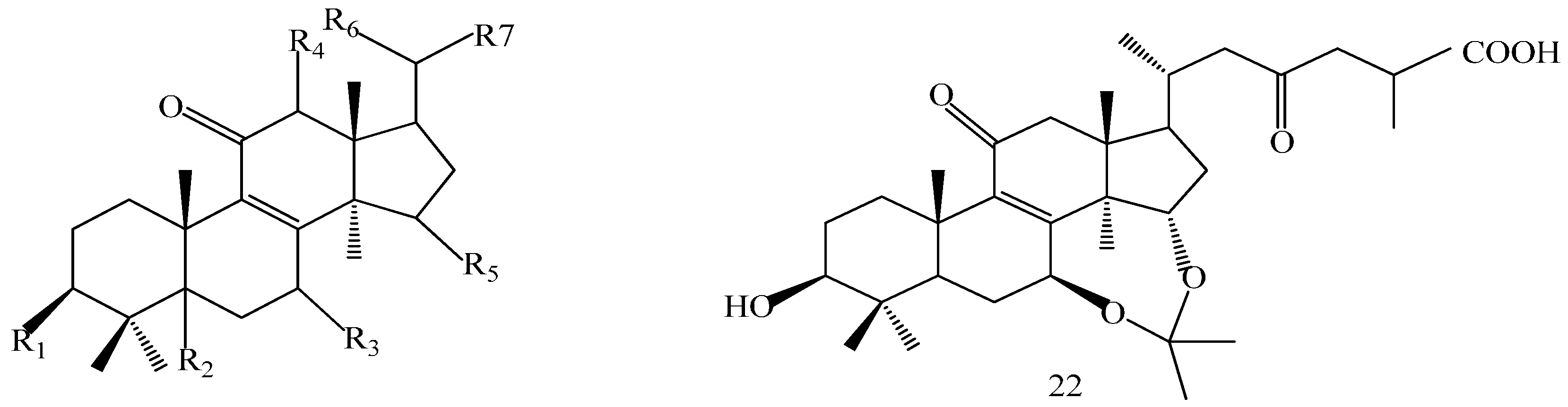
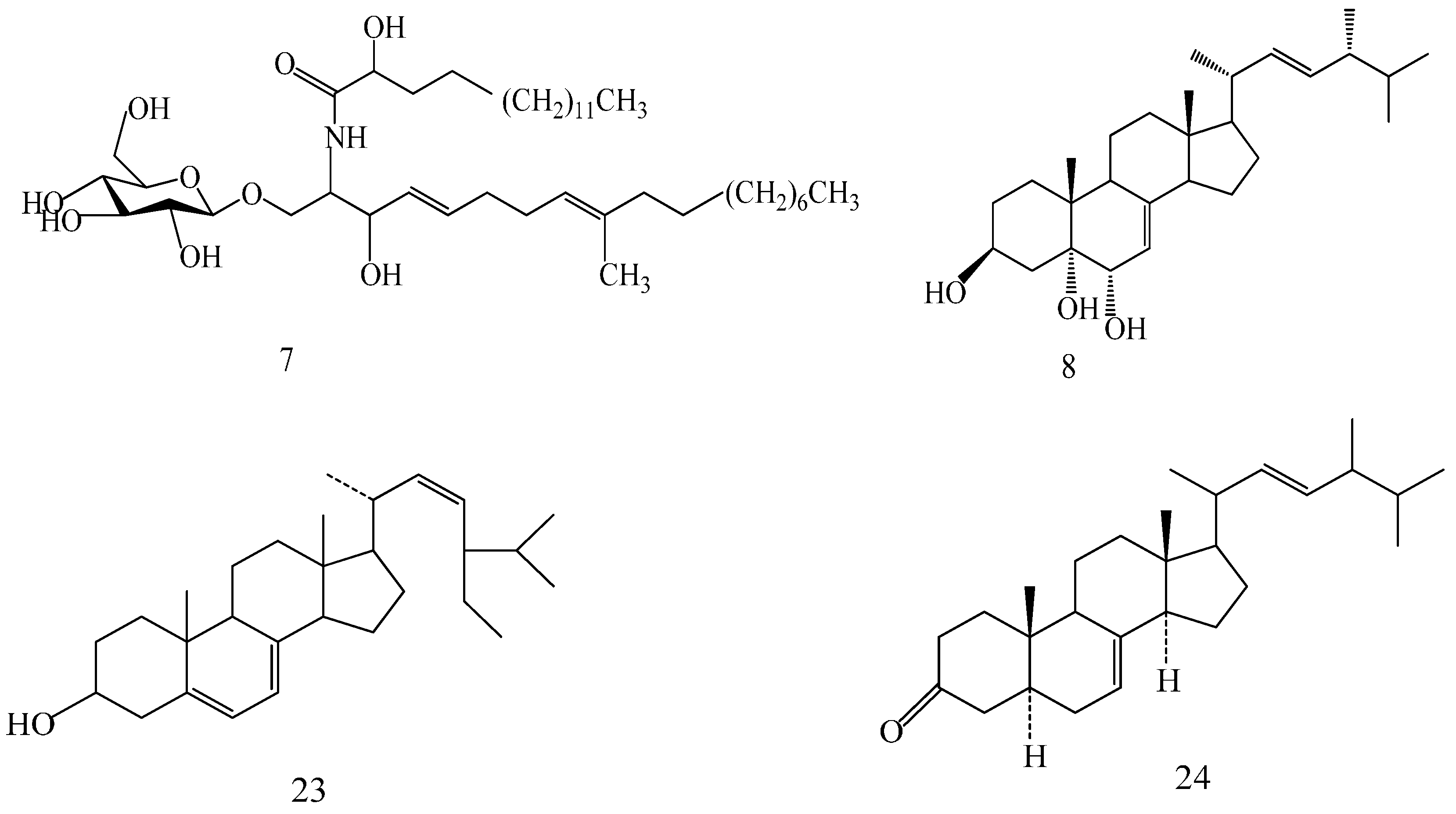
| Compound | R1 | R2 | R3 |
|---|---|---|---|
| 1 | H | β-OH | CH3 |
| 2 | H | β-OH | H |
| 3 | -OH | α-OH | H |
| Comp. | R1 | R2 |
|---|---|---|
| 5 | α-CH3 | OH |
| 16 | α-CH3 | COOH |
| 17 | β-CH3 | COOCH3 |
| 19 | α-CH3 | COOCH3 |
| 21 | β-CH3 | COOH |
| Comp. | R1 | R2 | R3 | R4 | R5 | R6 | R7 |
|---|---|---|---|---|---|---|---|
| 6 | β-H | α-OH | =O | β-H | α-CH3 | β-H | CH3 |
| 13 | α-H | =O | H | α-H | α-CH3 | β-H | COOH |
| 15 | α-H | β-OH | =O | α-H | β-CH3 | β-OH | COOH |
| 18 | β-H | β-OH | =O | β-H | α-CH3 | β-H | COOH |
| Comp. | R1 | R2 | R3 | R4 | R5 | R6 | R7 |
|---|---|---|---|---|---|---|---|
| 10 | β-OH | α-H | β-OH | β-OH | =O | β-CH3 | (CH2)2COOH |
| 11 | =O | α-H | β-OH | β-H | α-OH | α-CH3 | =O |
| 12 | β-OH | α-H | =O | β-H | =O | α-CH3 | =O |
| 14 | β-OH | β-H | β-OH | β-H | α-OH | α-CH3 | CH2OH |
| Compound | Target | Fitvalue | Hit * | Compound | Target | Fitvalue | Hit * |
|---|---|---|---|---|---|---|---|
| Com3 | CDK2 | 0.891 | 13 | Com6 | CYP3A4 | 0.792 | 5 |
| Com2 | CDK2 | 0.817 | 12 | Com5 | CYP3A4 | 0.763 | 5 |
| Com5 | CDK2 | 0.824 | 11 | GA-A | CDK2 | 0.753 | 5 |
| Com1 | CDK2 | 0.796 | 11 | Com3 | 11beta-HSD 1 | 0.83 | 4 |
| Com2 | PPAR γ | 0.787 | 10 | Com2 | COX1 | 0.824 | 4 |
| Com3 | PPAR γ | 0.789 | 7 | Com3 | PTP 1B | 0.724 | 4 |
| Com5 | PPAR γ | 0.833 | 6 | Com2 | c-Met | 0.722 | 4 |
| Com2 | CYP3A4 | 0.909 | 5 | Com2 | Lck | 0.666 | 4 |
| Com1 | CYP3A4 | 0.902 | 5 | Com3 | PPAR γ | 0.635 | 4 |
| Com3 | CYP3A4 | 0.883 | 5 | GA-D | CDK2 | 0.707 | 3 |
| ID | Target | Number of Compounds | Gene Name | Possibly Relevant Diseases |
|---|---|---|---|---|
| 1 | CYP2C9 | 8 | CYP2C9 | ADMET |
| 2 | CDK2 | 6 | CDK2 | cancer |
| 3 | GCN5 | 6 | KAT2A | cancer |
| 4 | PPAR γ | 5 | PPARG | obesity |
| 5 | COX1 | 5 | PTGS1 | pain, cardiovascular diseases |
| 6 | MEK1 | 5 | MAP2K1 | cancer |
| 7 | CYP3A4 | 5 | CYP A4 | ADMET |
| 8 | 11beta-HSD 1 | 5 | HSD11B1 | diabetes, obesity |
| 9 | PDE5A | 5 | PDE 5A | cardiovascular diseases |
| 10 | PTP1B | 4 | PTPN1 | diabetes, obesity |
| 11 | Syk | 4 | Syk | cancer |
| 12 | Lck | 4 | Lck | autoimmune suppression |
| 13 | c-Met | 4 | MET | cancer |
| 14 | Chk1 | 4 | Chk1 | cancer |
| 15 | Pim-1 | 4 | Pim-1 | cancer |
| 16 | CDK5 | 3 | CDK5 | cognitive defects, cancer |
| 17 | COX2 | 2 | Ptgs2 | pain, inflammation |
| 18 | FXR | 2 | NR1H4 | ADMET, bile acid-induced hepatotoxicity |
| 19 | FGFR1 | 2 | FGFR1 | diabetic retinopathy, cardiovascular diseases, cancer |
| Modules | p−Value | GO Terms |
|---|---|---|
| Module 1 | 1.69 × 10−21 | histone acetylation |
| Module 2 | 1.83 × 10−13 | cellular lipid metabolic process |
| Module 3 | 1.89 × 10−18 | interphase of mitotic cell cycle |
| Module 4 | 1.47 × 10−22 | activation of protein kinase A activity |
| 7.75 × 10−20 | cellular response to glucagon stimulus | |
| Module 5 | 1.88 × 10−12 | antigen receptor−mediated signaling pathway |
| 1.21 × 10−11 | positive regulation of response to stimulus | |
| Module 6 | 7.82 × 10−14 | transmembrane receptor protein tyrosine kinase signaling pathway |
| Module 7 | 1.57 × 10−17 | fibroblast growth factor receptor signaling pathway |
| Module 8 | 1.03 × 10−13 | transcription initiation from RNA polymerase II promoter |
| Module 9 | 8.98 × 10−17 | prostaglandin biosynthetic process |
| Module 10 | 1.11 × 10−15 | xenobiotic metabolic process |
| Module 11 | 1.30 × 10−07 | positive regulation of Rac protein signal transduction |
| 4.46 × 10−06 | negative regulation of apoptotic process | |
| Module 12 | 1.02 × 10−10 | glucocorticoid biosynthetic process |
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shao, Y.; Qiao, L.; Wu, L.; Sun, X.; Zhu, D.; Yang, G.; Zhang, X.; Mao, X.; Chen, W.; Liang, W.; et al. Structure Identification and Anti-Cancer Pharmacological Prediction of Triterpenes from Ganoderma lucidum. Molecules 2016, 21, 678. https://doi.org/10.3390/molecules21050678
Shao Y, Qiao L, Wu L, Sun X, Zhu D, Yang G, Zhang X, Mao X, Chen W, Liang W, et al. Structure Identification and Anti-Cancer Pharmacological Prediction of Triterpenes from Ganoderma lucidum. Molecules. 2016; 21(5):678. https://doi.org/10.3390/molecules21050678
Chicago/Turabian StyleShao, Yanyan, Liansheng Qiao, Lingfang Wu, Xuefei Sun, Dan Zhu, Guanghui Yang, Xiaoxue Zhang, Xin Mao, Wenjing Chen, Wenyi Liang, and et al. 2016. "Structure Identification and Anti-Cancer Pharmacological Prediction of Triterpenes from Ganoderma lucidum" Molecules 21, no. 5: 678. https://doi.org/10.3390/molecules21050678
APA StyleShao, Y., Qiao, L., Wu, L., Sun, X., Zhu, D., Yang, G., Zhang, X., Mao, X., Chen, W., Liang, W., Zhang, Y., & Zhang, L. (2016). Structure Identification and Anti-Cancer Pharmacological Prediction of Triterpenes from Ganoderma lucidum. Molecules, 21(5), 678. https://doi.org/10.3390/molecules21050678






