Proteomic Profiling of Iron Overload-Induced Human Hepatic Cells Reveals Activation of TLR2-Mediated Inflammatory Response
Abstract
:1. Introduction
2. Results
2.1. Overview of Proteomic Analysis (2D-LC-MS)
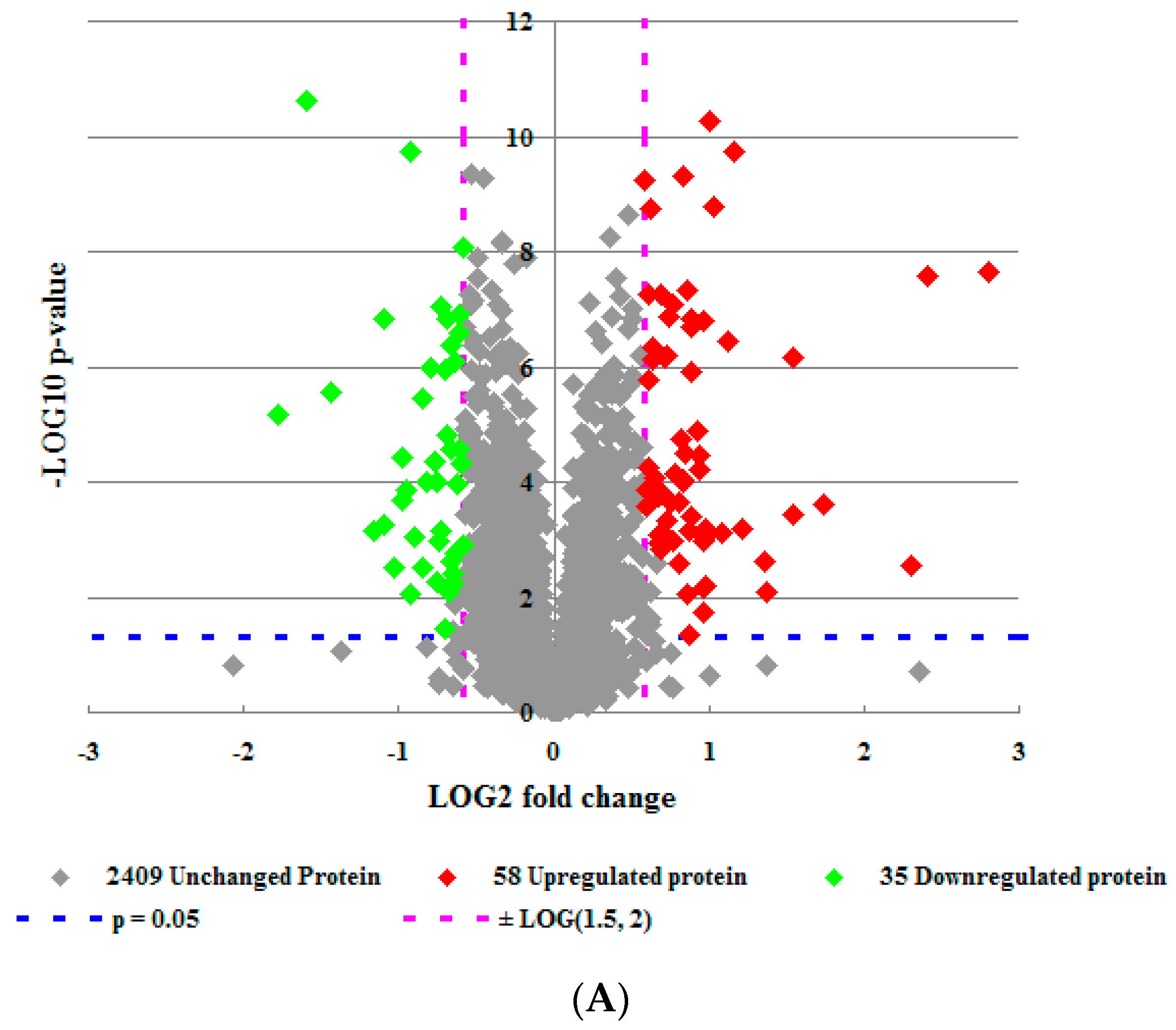
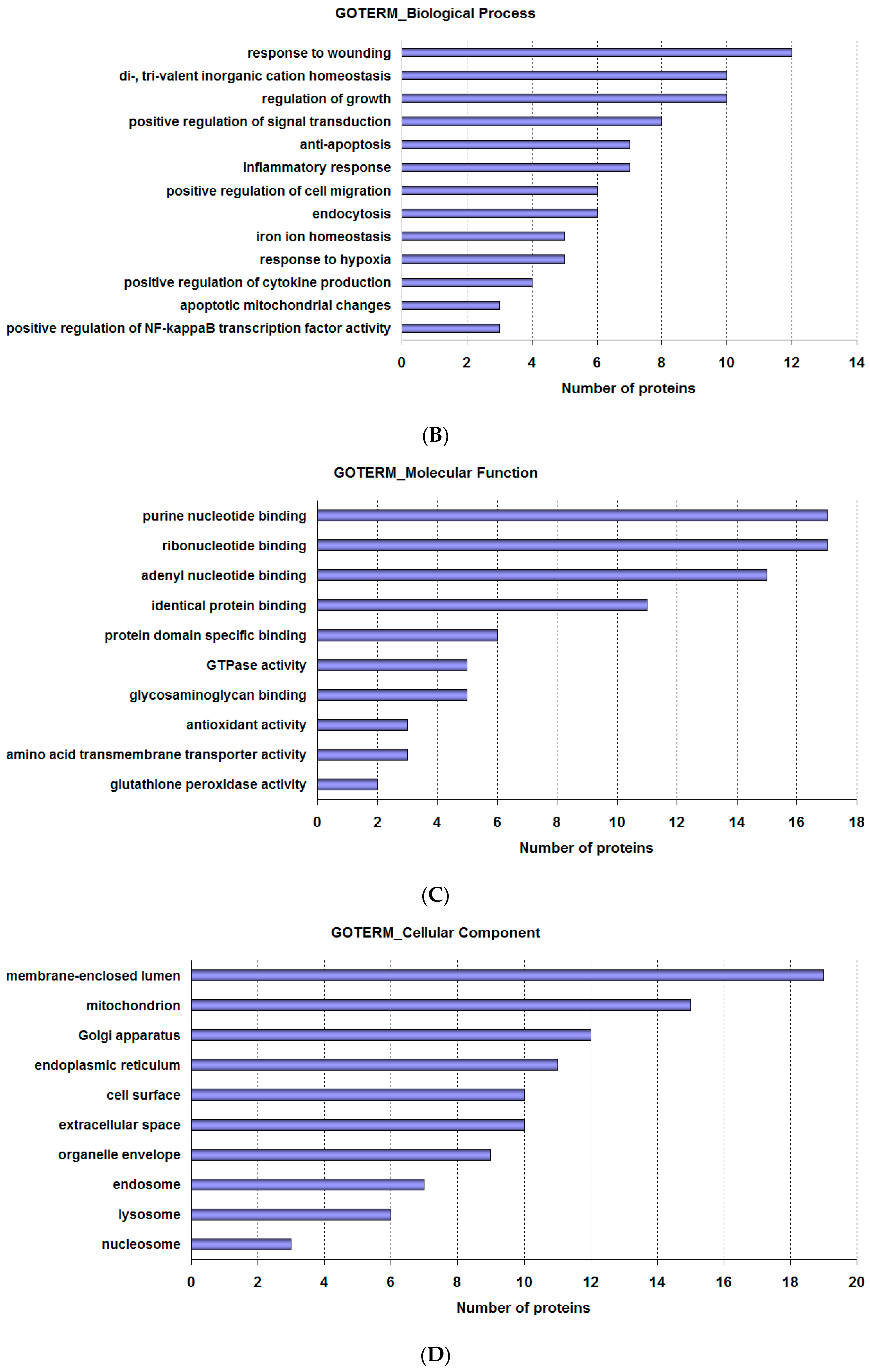
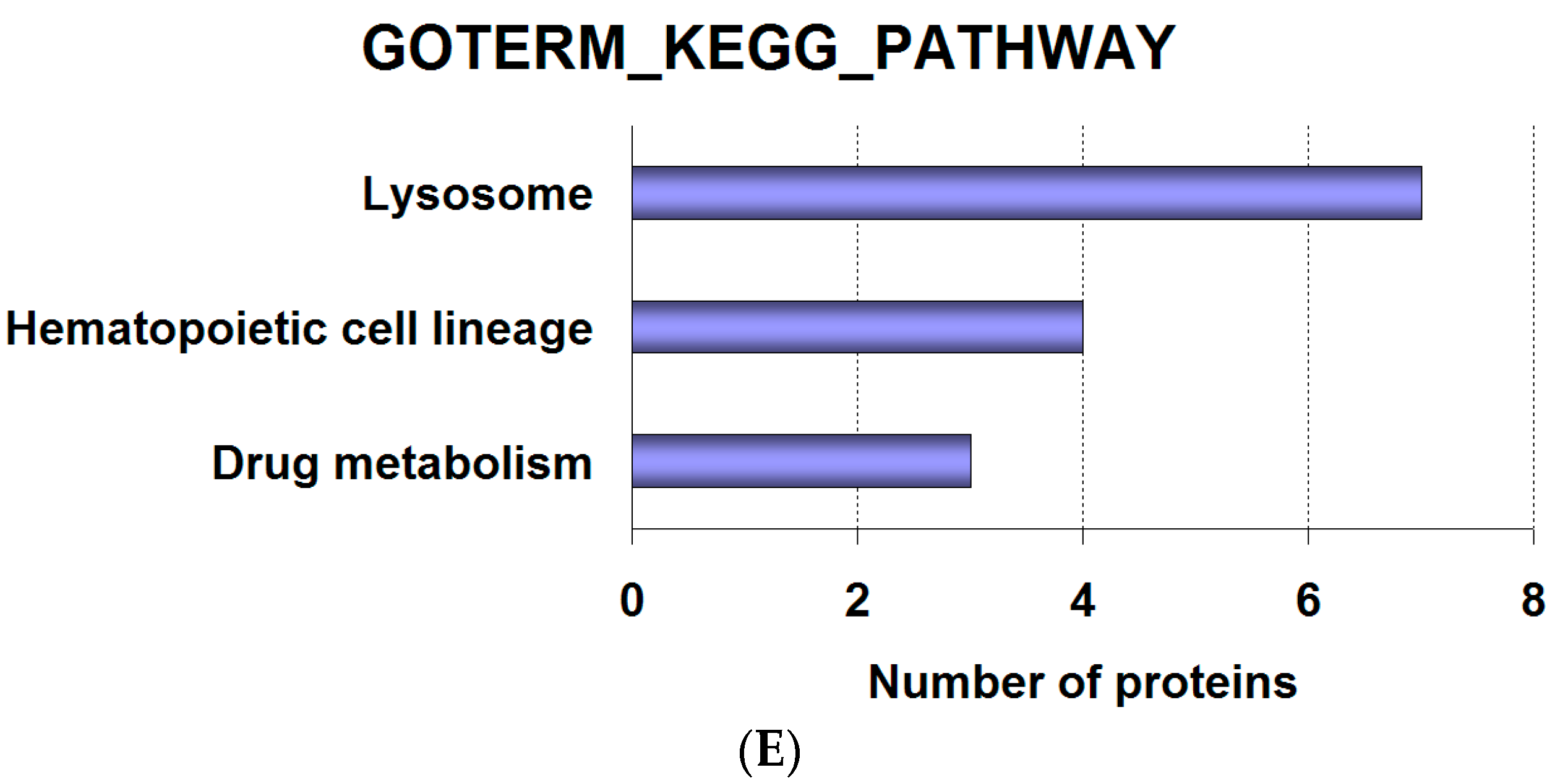
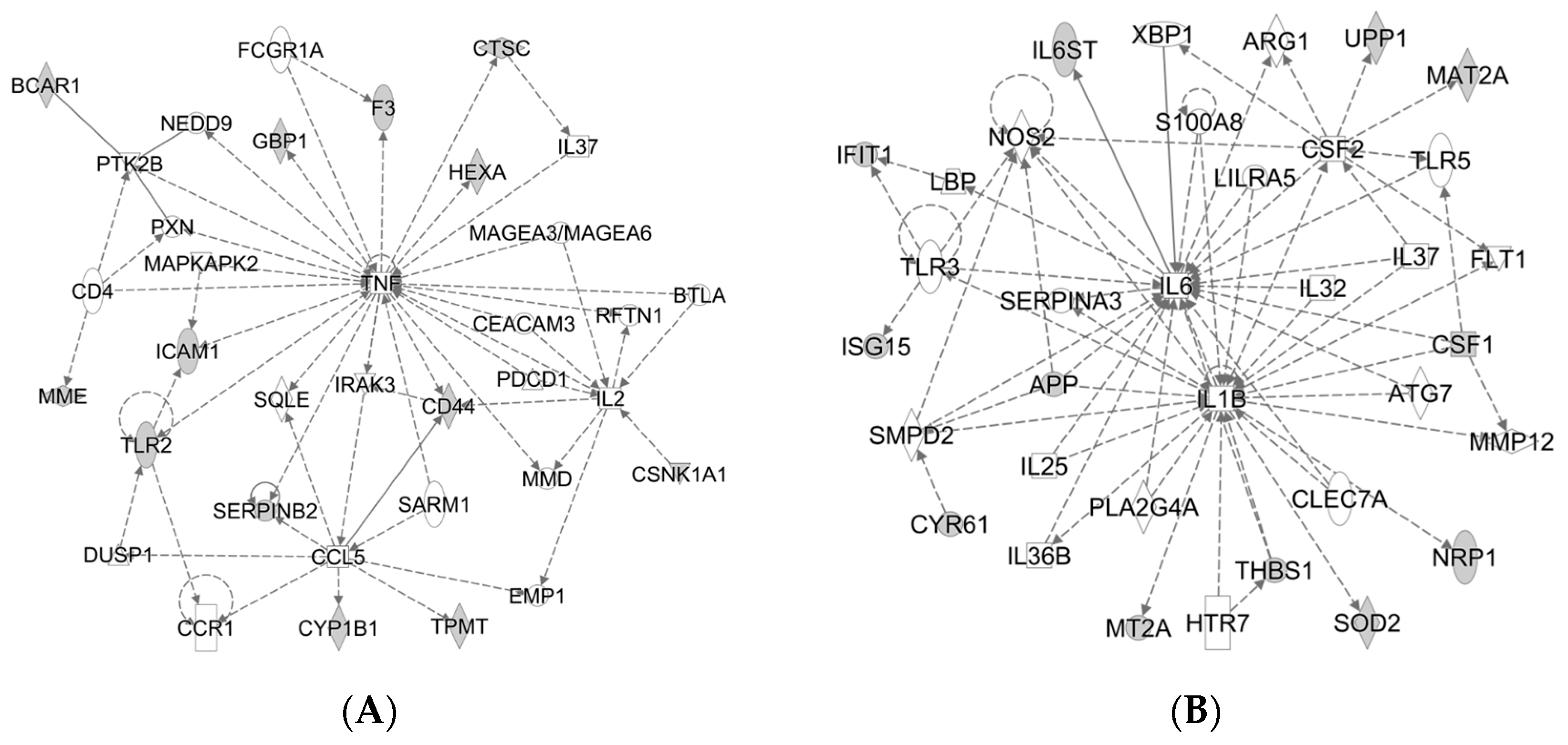
2.2. Analysis of Proteomics Data
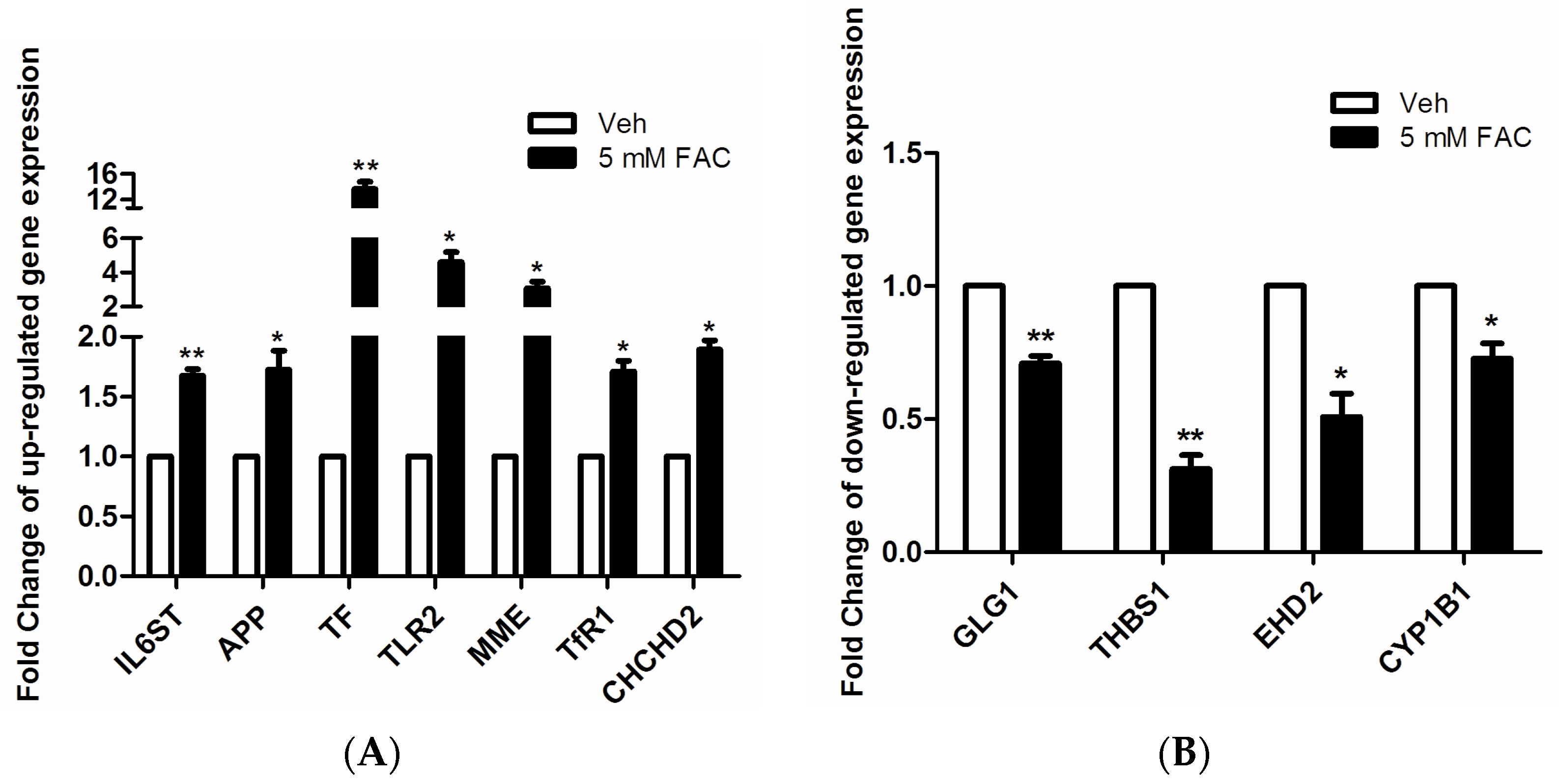
2.3. Validation of LC-MS/MS Results by RT-PCR
2.4. FAC-Induced Iron Overload Affected Iron Homeostasis Related Proteins
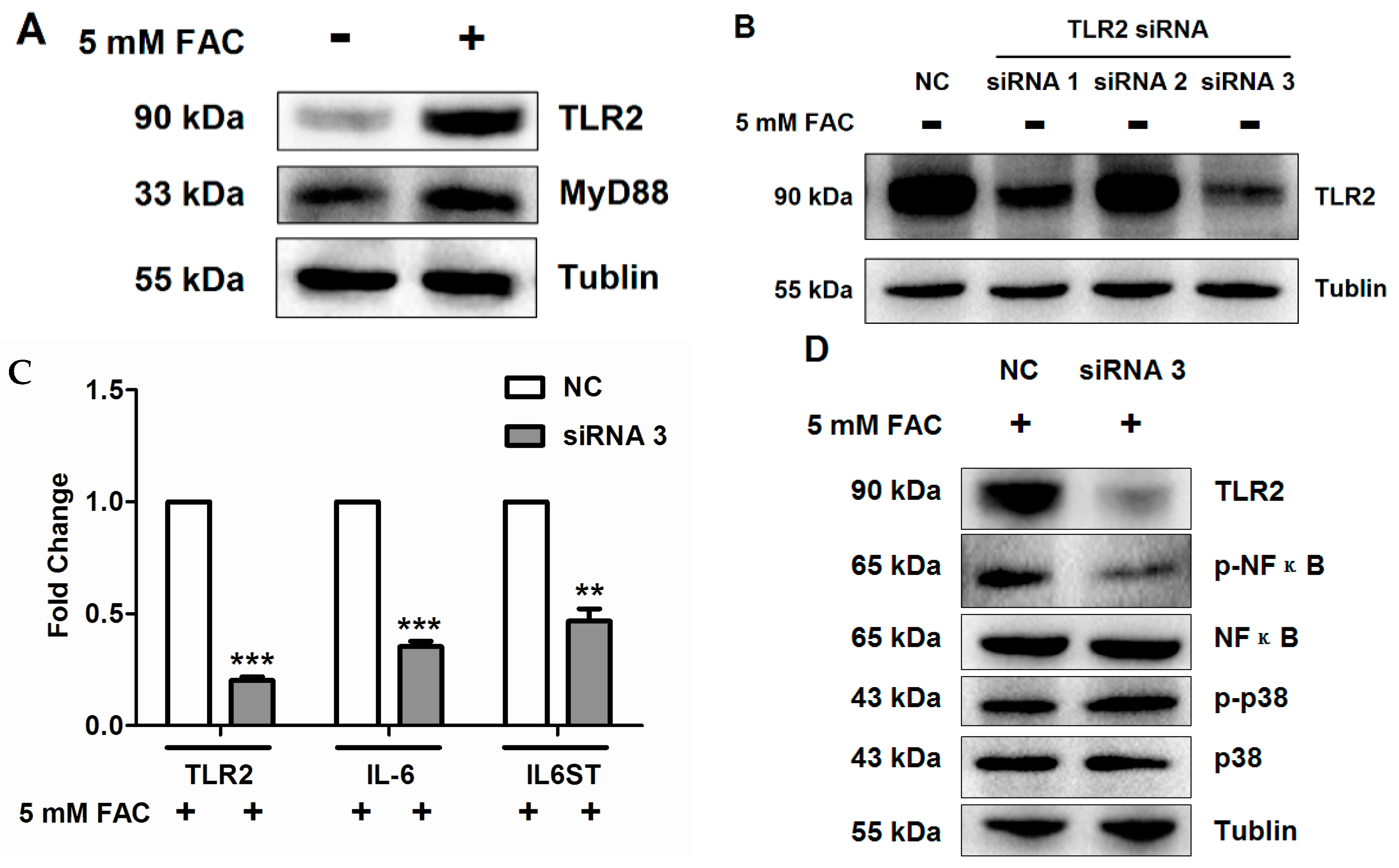
2.5. FAC Overload Triggered TLR2-Mediated Inflammatory Response
3. Discussion
4. Experimental Section
4.1. Cell Culture
4.2. In Vitro Iron Overload Model
4.3. Proteomics Analysis
4.3.1. Cell Lysis and Protein Digestion
4.3.2. Nanoflow LC-MS/MS
4.3.3. Data Analysis
4.4. Realtime-PCR
4.5. Western Blot Analysis
4.6. RNA Interference (RNAi) of Gene TLR2
4.7. Statistical Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Armand, P.; Kim, H.T.; Cutler, C.S.; Ho, V.T.; Koreth, J.; Alyea, E.P.; Soiffer, R.J.; Antin, J.H. Prognostic impact of elevated pretransplantation serum ferritin in patients undergoing myeloablative stem cell transplantation. Blood 2007, 109, 4586–4588. [Google Scholar] [CrossRef] [PubMed]
- Locasciulli, A.; Oneto, R.; Bacigalupo, A.; Socié, G.; Korthof, E.; Bekassy, A.; Schrezenmeier, H.; Passweg, J.; Führer, M. Outcome of patients with acquired aplastic anemia given first line bone marrow transplantation or immunosuppressive treatment in the last decade: A report from the European Group for Blood and Marrow Transplantation. Haematologica 2007, 9, 11–18. [Google Scholar] [CrossRef]
- Platzbecker, U.; Bornhäuser, M.; Germing, U.; Stumpf, J.; Scott, B.L.; Kröger, N.; Schwerdtfeger, R.; Böhm, A.; Kobbe, G.; Theuser, C. Red blood cell transfusion dependence and outcome after allogeneic peripheral blood stem cell transplantation in patients with de novo myelodysplastic syndrome (MDS). Biol. Blood Marrow Transplant. 2008, 14, 1217–1225. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Laissue, J.A.; Zimmermann, A. Hepatocyte apoptosis in hepatic iron overload diseases. Histol. Histopathol. 1997, 12, 367–374. [Google Scholar] [PubMed]
- Li, S.W.; Liu, C.M.; Guo, J.; Marcondes, A.M.; Deeg, J.; Li, X.; Guan, F. Iron overload induced by ferric ammonium citrate triggers reactive oxygen species-mediated apoptosis via both extrinsic and intrinsic pathways in human hepatic cells. Hum. Exp. Toxicol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Laport, G.G.; Sandmaier, B.M.; Storer, B.E.; Scott, B.L.; Stuart, M.J.; Lange, T.; Maris, M.B.; Agura, E.D.; Chauncey, T.R.; Wong, R.M. Reduced-intensity conditioning followed by allogeneic hematopoietic cell transplantation for adult patients with myelodysplastic syndrome and myeloproliferative disorders. Biol. Blood Marrow Tranplant. 2008, 14, 246–255. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Xu, F.; Karoopongse, E.; Marcondes, A.M.; Lee, K.; Kowdley, K.V.; Miao, C.H.; Trobridge, G.D.; Campbell, J.S.; Deeg, H.J. Allogeneic transplantation, Fas signaling, and dysregulation of hepcidin. Biol Blood Marrow Transplant. 2013, 19, 1210–1219. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sahlstedt, L.; Ebeling, F.; Von Bonsdorff, L.; Parkkinen, J.; Ruutu, T. Non-transferrin-bound iron during allogeneic stem cell transplantation. Brit. J. Haematol. 2001, 113, 836–838. [Google Scholar] [CrossRef]
- Galaris, D.; Pantopoulos, K. Oxidative stress and iron homeostasis: mechanistic and health aspects. Crit. Rev. Clin. Lab. Sci. 2008, 45, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Yakes, F.M.; Van Houten, B. Mitochondrial DNA damage is more extensive and persists longer than nuclear DNA damage in human cells following oxidative stress. Proc. Natl. Acad. Sci. USA 1997, 94, 514–519. [Google Scholar] [CrossRef] [PubMed]
- Andrews, N.C. Forging a field: The golden age of iron biology. Blood 2008, 112, 219–230. [Google Scholar] [CrossRef] [PubMed]
- Molina, M.F.; Qu, H.-Q.; Rentfro, A.R.; Nair, S.; Lu, Y.; Hanis, C.L.; McCormick, J.B.; Fisher-Hoch, S.P. Decreased expression of ATP6V1H in type 2 diabetes: A pilot report on the diabetes risk study in Mexican Americans. Biochem. Biophys. Res. Commun. 2011, 412, 728–731. [Google Scholar] [CrossRef] [PubMed]
- George, M.; Ying, G.G.; Rainey, M.A.; Solomon, A.; Parikh, P.T.; Gao, Q.; Band, V.; Band, H. Shared as well as distinct roles of EHD proteins revealed by biochemical and functional comparisons in mammalian cells and C. elegans. BMC Cell Biol. 2007, 8, 3. [Google Scholar] [CrossRef] [PubMed]
- Meyron-Holtz, E.G.; Ghosh, M.C.; Rouault, T.A. Mammalian tissue oxygen levels modulate iron-regulatory protein activities in vivo. Science 2004, 306, 2087–2090. [Google Scholar] [CrossRef] [PubMed]
- Fleming, R.E.; Ponka, P. Iron overload in human disease. N. Engl. J. Med. 2012, 366, 348–359. [Google Scholar] [CrossRef] [PubMed]
- Wong, B.X.; Tsatsanis, A.; Lim, L.Q.; Adlard, P.A.; Bush, A.I.; Duce, J.A. β-Amyloid precursor protein does not possess ferroxidase activity but does stabilize the cell surface ferrous iron exporter ferroportin. PLoS ONE 2014, 9, e114174. [Google Scholar] [CrossRef] [PubMed]
- Schuermann, A.; Helker, C.S.M.; Herzog, W. Metallothionein 2 regulates endothelial cell migration through transcriptional regulation of vegfc expression. Angiogenesis 2015, 18, 463–475. [Google Scholar] [CrossRef] [PubMed]
- Puukila, S.; Bryan, S.; Laakso, A.; Abdel-Malak, J.; Gurney, C.; Agostino, A.; Belló-Klein, A.; Prasad, K.; Khaper, N. Secoisolariciresinol diglucoside abrogates oxidative stress-induced damage in cardiac iron overload condition. PLoS ONE 2015, 10, e0122852. [Google Scholar] [CrossRef] [PubMed]
- Agah, A.; Kyriakides, T.R.; Lawler, J.; Bornstein, P. The lack of thrombospondin-1 (TSP1) dictates the course of wound healing in double-TSP1/TSP2-null mice. Am. J. Pathol. 2002, 161, 831–839. [Google Scholar] [CrossRef]
- Karbowski, M.; Jeong, S.-Y.; Youle, R.J. Endophilin B1 is required for the maintenance of mitochondrial morphology. J. Cell. Biol. 2004, 166, 1027–1039. [Google Scholar] [CrossRef] [PubMed]
- Lugrin, J.; Rosenblatt-Velin, N.; Parapanov, R.; Liaudet, L. The role of oxidative stress during inflammatory processes. Biol. Chem. 2014, 395, 203–230. [Google Scholar] [CrossRef] [PubMed]
- Rajamani, U.; Jialal, I. Hyperglycemia induces Toll-like receptor-2 and -4 expression and activity in human microvascular retinal endothelial cells: implications for diabetic retinopathy. J. Diabetes Res. 2014, 2014, 790902. [Google Scholar] [CrossRef] [PubMed]
- Oliveira-Nascimento, L.; Massari, P.; Wetzler, L.M. The role of TLR2 in infection and immunity. Front. Immunol. 2012, 3. [Google Scholar] [CrossRef] [PubMed]
- Guida, C.; Altamura, S.; Klein, F.A.; Galy, B.; Boutros, M.; Ulmer, A.J.; Hentze, M.W.; Muckenthaler, M.U. A novel inflammatory pathway mediating rapid hepcidin-independent hypoferremia. Blood 2015, 12, 2265–2275. [Google Scholar] [CrossRef] [PubMed]
- Kelstrup, C.D.; Jersie-Christensen, R.R.; Batth, T.S.; Arrey, T.N.; Kuehn, A.; Kellmann, M.; Olsen, J.V. Rapid and deep proteomes by faster sequencing on a benchtop quadrupole ultra-high-field Orbitrap mass spectrometer. J. Proteome Res. 2014, 13, 6187–6195. [Google Scholar] [CrossRef] [PubMed]
- Kelstrup, C.D.; Young, C.; Lavallee, R.; Nielsen, M.L.; Olsen, J.V. Optimized fast and sensitive acquisition methods for shotgun proteomics on a quadrupole orbitrap mass spectrometer. J. Proteome Res. 2012, 11, 3487–3497. [Google Scholar] [CrossRef] [PubMed]
- Scheltema, R.A.; Hauschild, J.P.; Lange, O.; Hornburg, D.; Denisov, E.; Damoc, E.; Kuehn, A.; Makarov, A.; Mann, M. The Q Exactive HF, a Benchtop mass spectrometer with a pre-filter, high-performance quadrupole and an ultra-high-field Orbitrap analyzer. Mol. Cell. Proteom. 2014, 13, 3698–3708. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 2008, 26, 1367–1372. [Google Scholar] [CrossRef] [PubMed]
- Cox, J.; Neuhauser, N.; Michalski, A.; Scheltema, R.A.; Olsen, J.V.; Mann, M. Andromeda: A peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 2011, 10, 1794–1805. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Sample Availability: Not Appliable.

| Ratio 1 | Protein IDs | Protein Description | Gene Name | MW [kDa] 2 |
|---|---|---|---|---|
| −1.781 | Q92896 | Golgi apparatus protein 1 | GLG1 | 135 |
| −1.602 | Q9NZN4 | EH domain-containing protein 2 | EHD2 | 61 |
| −1.437 | P09972 | Fructose-bisphosphate aldolase C | ALDOC | 39 |
| −1.091 | P42126 | Isoform 2 of Enoyl-CoA delta isomerase 1, mitochondrial | ECI1 | 31 |
| −1.09 | P07996 | Thrombospondin-1 | THBS1 | 129 |
| −0.983 | P09603 | Macrophage colony-stimulating factor 1 | CSF1 | 60 |
| −0.977 | Q92522 | Histone H1x | H1FX | 22 |
| −0.956 | O95197 | Isoform 3 of Reticulon-3 | RTN3 | 26 |
| −0.927 | P21399 | Cytoplasmic aconitate hydratase | ACO1 | 98 |
| −0.895 | Q15418 | Isoform 3 of Ribosomal protein S6 kinase α-1 | RPS6KA1 | 73 |
| −0.845 | P07203 | Glutathione peroxidase 1 | GPX1 | 22 |
| −0.814 | Q8TED1 | Probable glutathione peroxidase 8 | GPX8 | 24 |
| −0.793 | Q16678 | Cytochrome P450 1B1 | CYP1B1 | 61 |
| −0.768 | Q6EMK4 | Vasorin | VASN | 72 |
| −0.749 | P51580 | Thiopurine S-methyltransferase | TPMT | 28 |
| −0.745 | Q969V3 | Isoform 2 of Nicalin | NCLN | 63 |
| −0.733 | O00622 | Protein CYR61 | CYR61 | 42 |
| −0.729 | P06865 | β-Hexosaminidase | HEXA | 58 |
| −0.7 | P04179 | Superoxide dismutase [Mn], mitochondrial | SOD2 | 25 |
| −0.693 | P17342 | Isoform 2 of atrial natriuretic peptide receptor 3 | NPR3 | 60 |
| −0.685 | P07602 | Proactivator polypeptide | PSAP | 58 |
| −0.661 | O43865 | Putative adenosylhomocysteinase 2 | AHCYL1 | 59 |
| −0.657 | P52630 | Signal transducer and activator of transcription 2 | STAT2 | 97 |
| −0.657 | P51553 | Isocitrate dehydrogenase [NAD] subunit gamma, mitochondrial | IDH3G | 43 |
| −0.648 | P61970 | Nuclear transport factor 2 | NUTF2 | 14 |
| −0.647 | Q9ULZ3 | Apoptosis-associated speck-like protein containing a CARD (PYD and CARD domain containing) | PYCARD | 22 |
| −0.639 | P30626 | Sorcin | SRI | 18 |
| −0.637 | P0DMM9 | Sulfotransferase 1A3 | SULT1A3 | 34 |
| −0.617 | Q9H845 | Acyl-CoA dehydrogenase family member 9, mitochondrial | ACAD9 | 69 |
| −0.615 | P53634 | Dipeptidyl peptidase 1 | CTSC | 52 |
| −0.613 | Q6P587 | Acylpyruvase FAHD1, mitochondrial | FAHD1 | 25 |
| −0.609 | P16403 | Histone H1.2 | HIST1H1C | 21 |
| −0.602 | P62805 | Histone H4 | HIST1H4A | 11 |
| −0.59 | P04844 | Isoform 2 of Dolichyl-diphosphooligosaccharide—protein glycosyltransferase subunit 2 | RPN2 | 68 |
| −0.588 | Q13228 | Selenium-binding protein 1 | SELENBP1 | 52 |
| 0.586 | P05362 | Intercellular adhesion molecule 1 | ICAM1 | 58 |
| 0.597 | Q9Y6H1 | Coiled-coil-helix-coiled-coil-helix domain-containing protein 2, mitochondrial | CHCHD2 | 16 |
| 0.601 | Q9Y371 | Endophilin-B1 | SH3GLB1 | 41 |
| 0.61 | P05161 | Ubiquitin-like protein ISG15 | ISG15 | 18 |
| 0.614 | P31689 | DnaJ homolog subfamily A member 1 | DNAJA1 | 45 |
| 0.614 | P11387 | DNA topoisomerase 1 | TOP1 | 91 |
| 0.621 | P31153 | S-adenosylmethionine synthase isoform type-2 | MAT2A | 44 |
| 0.636 | Q14498 | Isoform 3 of RNA-binding protein 39 | RBM39 | 57 |
| 0.636 | O75534 | Isoform Short of Cold shock domain-containing protein E1 | CSDE1 | 86 |
| 0.643 | P02786 | Transferrin receptor protein 1 | TfR1 | 85 |
| 0.653 | O75976 | Carboxypeptidase D | CPD | 153 |
| 0.656 | Q9BQ52 | Zinc phosphodiesterase ELAC protein 2 | ELAC2 | 92 |
| 0.67 | Q9UI12 | ATPase, H+ transporting, lysosomal 50/57kDa, V1 subunit H, isoform CRA_c | ATP6V1H | 52 |
| 0.678 | P48729 | Casein kinase 1, alpha 1, isoform CRA_g | CSNK1A1 | 38 |
| 0.681 | P08473 | Neprilysin | MME | 86 |
| 0.685 | Q9H2H9 | Sodium-coupled neutral amino acid transporter 1 | SLC38A1 | 54 |
| 0.692 | P08195 | Isoform 2 of 4F2 cell-surface antigen heavy chain | SLC3A2 | 58 |
| 0.702 | Q9H5Q4 | Dimethyladenosine transferase 2, mitochondrial | TFB2M | 45 |
| 0.712 | P29317 | Ephrin type-A receptor 2 | EPHA2 | 108 |
| 0.714 | O14786 | Neuropilin-1 | NRP1 | 101 |
| 0.723 | P11279 | Lysosome-associated membrane glycoprotein 1 | LAMP1 | 39 |
| 0.726 | Q96PD2 | Discoidin, CUB and LCCL domain-containing protein 2 | DCBLD2 | 85 |
| 0.747 | P16070 | CD44 antigen (Fragment) | CD44 | 20 |
| 0.754 | P46734 | Dual specificity mitogen-activated protein kinase kinase 3 | MAP2K3 | 39 |
| 0.765 | P08754 | Guanine nucleotide-binding protein G(k) subunit alpha | GNAI3 | 41 |
| 0.766 | Q9H5V8 | CUB domain-containing protein 1 | CDCP1 | 93 |
| 0.777 | Q13641 | Trophoblast glycoprotein | TPBG | 46 |
| 0.809 | P02795 | Metallothionein-2 | MT2A | 6 |
| 0.811 | Q14978 | Nucleolar and coiled-body phosphoprotein 1 | NOLC1 | 74 |
| 0.825 | P52926 | High mobility group protein HMGI-C | HMGA2 | 11 |
| 0.833 | P08962 | CD63 antigen | CD63 | 16 |
| 0.837 | P32455 | Interferon-induced guanylate-binding protein 1 | GBP1 | 68 |
| 0.837 | P04183 | Thymidine kinase, cytosolic | TK1 | 25 |
| 0.848 | Q16831 | Uridine phosphorylase 1 | UPP1 | 34 |
| 0.858 | P21980 | Protein-glutamine gamma-glutamyltransferase 2 | TGM2 | 69 |
| 0.875 | P35080 | Profilin-2 | PFN2 | 10 |
| 0.882 | P13726 | Tissue factor | F3 | 33 |
| 0.885 | P56945 | Breast cancer anti-estrogen resistance protein 1 | BCAR1 | 93 |
| 0.891 | O15118 | Niemann-Pick C1 protein | NPC1 | 142 |
| 0.892 | Q14669 | E3 ubiquitin-protein ligase TRIP12 | TRIP12 | 220 |
| 0.92 | Q12996 | Cleavage stimulation factor subunit 3 | CSTF3 | 83 |
| 0.935 | Q9H4A6 | Golgi phosphoprotein 3 | GOLPH3 | 34 |
| 0.942 | Q16850 | Isoform 2 of lanosterol 14-α demethylase | CYP51A1 | 46 |
| 0.962 | P11802 | Cyclin-dependent kinase 4 | CDK4 | 34 |
| 0.97 | Q9Y3E0 | Vesicle transport protein GOT1B | GOLT1B | 14 |
| 0.975 | Q9Y3D8 | Transcription initiation factor TFIID subunit 9 | AK6 | 17 |
| 0.983 | Q9UBB6 | Isoform 2 of neurochondrin | NCDN | 77 |
| 1.003 | P17844 | Probable ATP-dependent RNA helicase DDX5 | DDX5 | 107 |
| 1.028 | P05120 | Plasminogen activator inhibitor 2 | SERPINB2 | 47 |
| 1.085 | P11234 | Ras-related protein Ral-B | RALB | 23 |
| 1.127 | Q96QD8 | Sodium-coupled neutral amino acid transporter 2 | SLC38A2 | 56 |
| 1.162 | P52292 | Importin subunit alpha-1 | KPNA2 | 58 |
| 1.539 | P40189 | Interleukin-6 receptor subunit beta | IL6ST | 104 |
| 1.54 | Q06481 | Amyloid-like protein 2 | APLP2 | 76 |
| 1.742 | P05067 | Gamma-secretase C-terminal fragment 59 | APP | 81 |
| 2.309 | P09914 | Interferon-induced protein with tetratricopeptide repeats 1 | IFIT1 | 55 |
| 2.408 | P02787 | Serotransferrin | TF | 78 |
| 2.809 | O60603 | Toll-like receptor 2 | TLR2 | 90 |
| Biological Process | Related Gene |
|---|---|
| Response to wounding | CD44; F3; DCBLD2; GPX1; MAP2K3; NRP1; SERPINB2; SOD2; THBS1; TLR2; TF; TfR1 |
| Di-, tri-valent inorganic cation homeostasis | ACO1; APP; APLP2; IL6ST; MT2A; SRI; SOD2; TF; TfR1; TGM2 |
| Regulation of growth | CD44; AK6; APP; CSF1; CYR61; DCBLD2; HMGA2; IDH3G; NRP1; BCAR1 |
| Positive regulation of signal transduction | F3; CSF1; GPX1; GOLPH3; GOLT1B; IL6ST; THBS1; TGM2 |
| Anti-apoptosis | SH3GLB1; F3; GPX1; SERPINB2; SOD2; THBS1; TGM2 |
| Inflammatory response | CD44; F3; MAP2K3; THBS1; TLR2; TF; TfR1 |
| Positive regulation of cell migration | F3; CSF1; ICAM1; IL6ST; BCAR1; THBS1 |
| Endocytosis | ATP6V1H; EHD2; NPC1; APP; THBS1; TfR1 |
| Iron ion homeostasis | ACO1; SRI; SOD2; TF; TfR1 |
| Response to hypoxia | ALDOC; SOD2; THBS1; TF; TfR1 |
| Positive regulation of cytokine production | PYCARD; IL6ST; THBS1; TLR2 |
| Apoptotic mitochondrial changes | SH3GLB1; GPX1; SOD2 |
| Positive regulation of NF-κB transcription factor activity | PYCARD; ICAM1; TLR2 |
| Gene | Primer | Sequence (5’–3’) | Product Size (bp) | Annealing Temperature (°C) |
|---|---|---|---|---|
| IL6ST | Forward | GTGAGTGGGATGGTGGAAGG | 78 | 60 |
| Reverse | CAAACTTGTGTGTTGCCCATTC | |||
| APP | Forward | GCCCTGCGGAATTGACAAG | 144 | 60 |
| Reverse | CCATCTGCATAGTCTGTGTCTG | |||
| TF | Forward | GGTGGCAGAGTTCTATGGGTC | 172 | 60 |
| Reverse | ACAGTAAAGTAAGCCTATGGGGA | |||
| TLR2 | Forward | ATCCTCCAATCAGGCTTCTCT | 118 | 60 |
| Reverse | GGACAGGTCAAGGCTTTTTACA | |||
| MME | Forward | GATCGCACTCTATGCAACCTAC | 83 | 60 |
| Reverse | TGTTTTGGATCAGTCGAGCAG | |||
| TfR1 | Forward | ACCATTGTCATATACCCGGTTCA | 219 | 60 |
| Reverse | CAATAGCCCAAGTAGCCAATCAT | |||
| CHCHD2 | Forward | ACACATTGGGTCACGCCATTA | 201 | 60 |
| Reverse | GCACCTCATTGAAACCCTCACA | |||
| GLG1 | Forward | CCAAGATGACGGCCATCATTT | 103 | 60 |
| Reverse | AGCCGAATACTGCCACATTTC | |||
| THBS1 | Forward | TGCTATCACAACGGAGTTCAGT | 108 | 60 |
| Reverse | GCAGGACACCTTTTTGCAGATG | |||
| EHD2 | Forward | TCCGCAAACTCAACCCTTTC | 78 | 60 |
| Reverse | TCTCCAGGACCTGATTAGGGA | |||
| CYP1B1 | Forward | ACGTACCGGCCACTATCACT | 142 | 60 |
| Reverse | CTCCCCACGACCTGATCCA | |||
| TfR2 | Forward | AATGTCCTGAGTCAGAGCGG | 141 | 55.9 |
| Reverse | GGCAGACTCTACTGGAGCGA | |||
| IL6 | Forward | CCAGAGCTGTGCAGATGAGT | 99 | 57.8 |
| Reverse | CATTTGTGGTTGGGTCAGG | |||
| STAT3 | Forward | ACACACGGTACCTGGAGCAG | 172 | 57.8 |
| Reverse | TACTGCTGGTCAATCTCTCCC | |||
| Ferritin | Forward | ATCAACCTGGAGCTCTACGC | 155 | 57.8 |
| Reverse | TGGTTCTGCAGCTTCATCAG | |||
| HAMP | Forward | CTGTTTTCCCACAACAGAC | 230 | 59.8 |
| Reverse | CCTTCCTTATTTATTCCTGC | |||
| β-actin | Forward | CTTCCTGGGCATGGAGTC | 232 | 60 |
| Reverse | GCCGATCCACACGGAGTA |
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, X.; Li, S.; Lu, M.; Yang, G.; Shen, Y.; Zhou, X. Proteomic Profiling of Iron Overload-Induced Human Hepatic Cells Reveals Activation of TLR2-Mediated Inflammatory Response. Molecules 2016, 21, 322. https://doi.org/10.3390/molecules21030322
Li X, Li S, Lu M, Yang G, Shen Y, Zhou X. Proteomic Profiling of Iron Overload-Induced Human Hepatic Cells Reveals Activation of TLR2-Mediated Inflammatory Response. Molecules. 2016; 21(3):322. https://doi.org/10.3390/molecules21030322
Chicago/Turabian StyleLi, Xiang, Shiwei Li, Mize Lu, Guohua Yang, Yunfeng Shen, and Xin Zhou. 2016. "Proteomic Profiling of Iron Overload-Induced Human Hepatic Cells Reveals Activation of TLR2-Mediated Inflammatory Response" Molecules 21, no. 3: 322. https://doi.org/10.3390/molecules21030322
APA StyleLi, X., Li, S., Lu, M., Yang, G., Shen, Y., & Zhou, X. (2016). Proteomic Profiling of Iron Overload-Induced Human Hepatic Cells Reveals Activation of TLR2-Mediated Inflammatory Response. Molecules, 21(3), 322. https://doi.org/10.3390/molecules21030322





