Sulfur Atom in its Bound State Is a Unique Element Involved in Physiological Functions in Mammals
Abstract
:1. Introduction
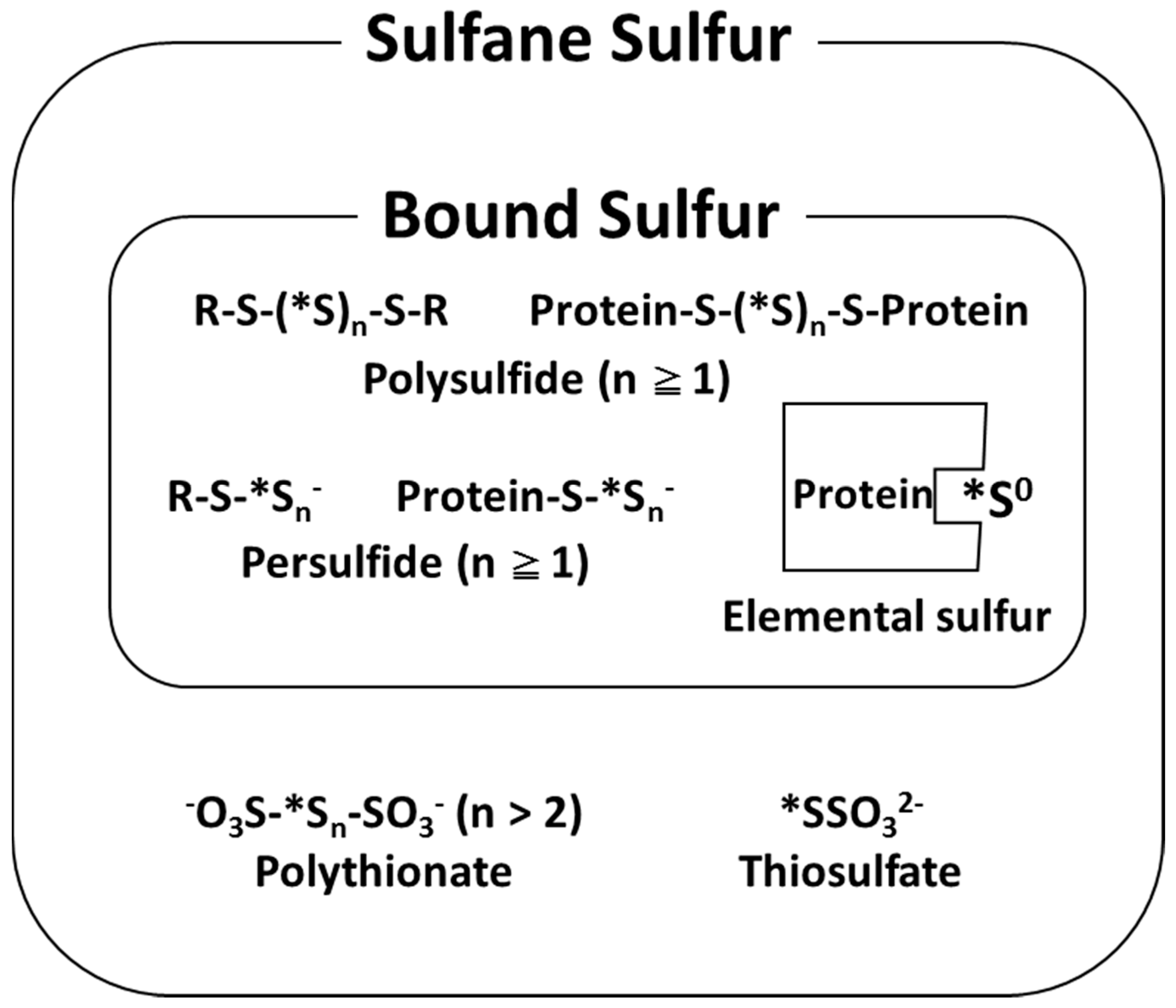
2. Analysis and Characterization of Bound Sulfur
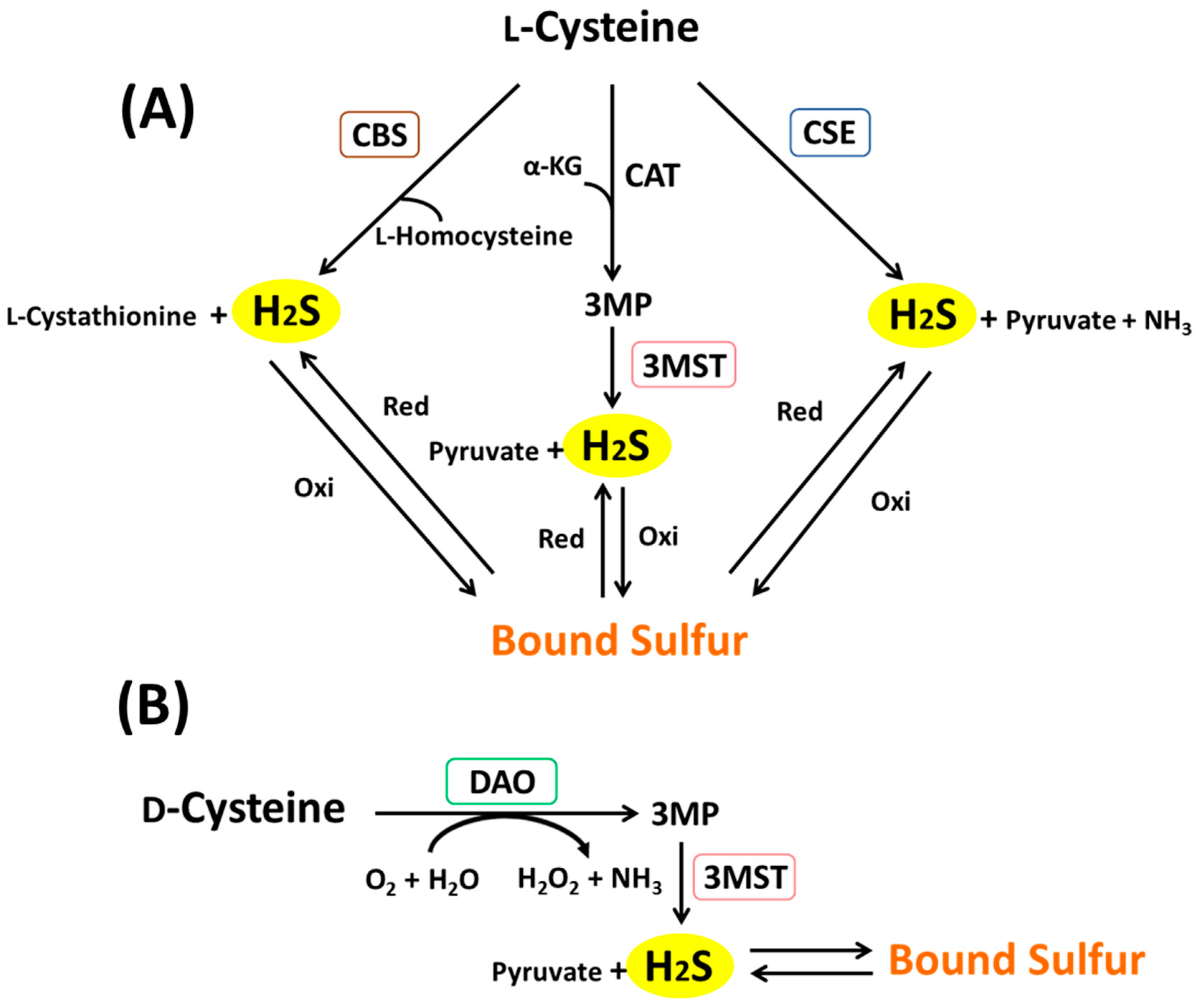
3. Metabolic Pathways of H2S and Bound Sulfur
4. The Significance of Differentiating between Hydrogen Sulfide and Bound Sulfur
4.1. The Issues with Previous Studies
4.2. An Old and New Physiological Metabolite Called Bound Sulfur
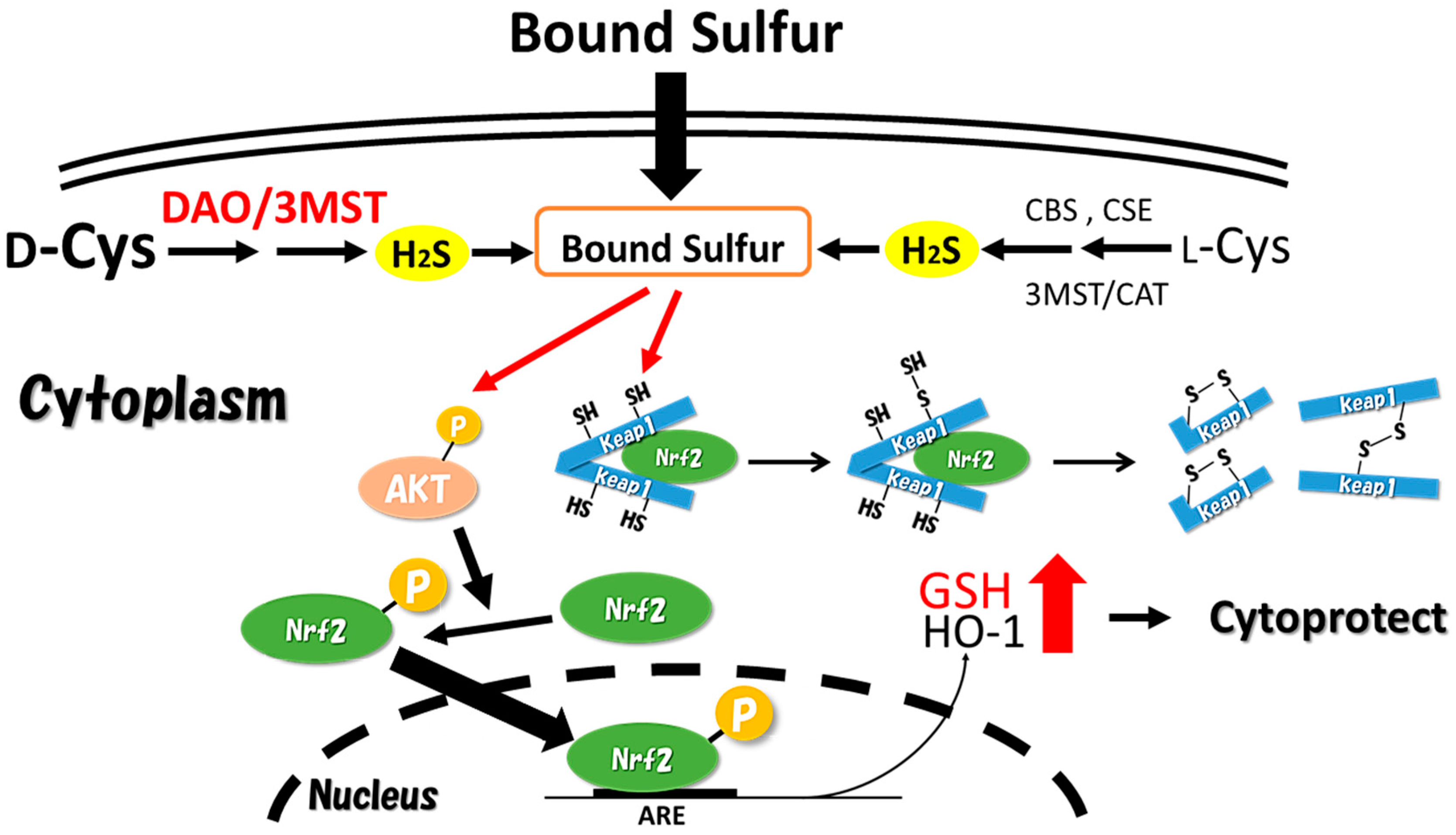
4.3. Cytoprotective Effect against Oxidative Stress
4.4. Physiological Action against Glycation Stress
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ogasawara, Y.; Ishii, K.; Togawa, T.; Tanabe, S. Determination of bound sulfur in serum by gas dialysis/high-performance liquid chromatography. Anal. Biochem. 1993, 215, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Stipanuk, M.H. Metabolism of sulfur-containing amino acids. Annu. Rev. Nutr. 1986, 6, 179–209. [Google Scholar] [CrossRef] [PubMed]
- Abe, K.; Kimura, H. The possible role of hydrogen sulfide as an endogenous neuromodulator. J. Neurosci. 1996, 16, 1066–1071. [Google Scholar] [PubMed]
- Nagy, P. Mechanistic chemical perspective of hydrogen sulfide signaling. Methods Enzymol. 2015, 554, 3–29. [Google Scholar] [PubMed]
- Kimura, H. Hydrogen polysulfide (H2Sn) signaling along with hydrogen sulfide (H2S) and nitric oxide (NO). J. Neural Transm. 2016, 123, 1235–1245. [Google Scholar] [CrossRef] [PubMed]
- Greiner, R.; Dick, T.P. Real-time assays for monitoring the influence of sulfide and sulfane sulfur species on protein thiol redox states. Methods Enzymol. 2015, 555, 57–77. [Google Scholar] [PubMed]
- Wood, J.L. Sulfane sulfur. Methods Enzymol. 1987, 143, 25–29. [Google Scholar] [PubMed]
- Ogasawara, Y.; Ishii, K.; Togawa, T.; Tanabe, S. Determination of trace amounts of sulphide in human red blood cells by high-performance liquid chromatography with fluorimetric detection after derivatization with p-phenylenediamine and iron (III). Analyst 1991, 116, 1359–1363. [Google Scholar] [CrossRef] [PubMed]
- Togawa, T.; Ogawa, M.; Nawata, M.; Ogasawara, Y.; Kawanabe, K.; Tanabe, S. High performance liquid chromatographic determination of bound sulfide and sulfite and thiosulfate at their low levels in human serum by pre-column fluorescence derivatization with monobromobimane. Chem. Pharm. Bull. 1992, 40, 3000–3004. [Google Scholar] [CrossRef] [PubMed]
- Westley, A.M.; Westley, J. Biological sulfane sulfur. Anal. Biochem. 1991, 67, 63–67. [Google Scholar] [CrossRef]
- Hannestad, U.; Margheri, S.; Sörbo, B. A sensitive gas chromatographic method for determination of protein-associated sulfur. Anal. Biochem. 1989, 178, 394–398. [Google Scholar] [CrossRef]
- Shen, X.; Kolluru, G.K.; Yuan, S.; Kevil, C. Measurement of H2S in vivo and in vitro by the monobromobimane method. Methods Enzymol. 2015, 554, 31–45. [Google Scholar] [PubMed]
- Wintner, E.A.; Deckwerth, T.L.; Langston, W.; Bengtsson, A.; Leviten, D.; Hill, P.; Insko, M.A.; Dumpit, R.; VandenEkart, E.; Toombs, C.F.; et al. A monobromobimane-based assay to measure the pharmacokinetic profile of reactive sulphide species in blood. Br. J. Pharmacol. 2010, 160, 941–957. [Google Scholar] [CrossRef] [PubMed]
- Kimura, Y.; Toyofuku, Y.; Koike, S.; Shibuya, N.; Nagahara, N.; Lefer, D.; Ogasawara, Y.; Kimura, H. Identification of H2S3 and H2S produced by 3-mercaptopyruvate sulfurtransferase in the brain. Sci. Rep. 2015, 5, 14774. [Google Scholar] [CrossRef] [PubMed]
- Park, C.M.; Weerasinghe, L.; Day, J.J.; Fukuto, J.M.; Xian, M. Persulfides: Current knowledge and challenges in chemistry and chemical biology. Mol. Biosyst. 2015, 11, 1775–1785. [Google Scholar] [CrossRef] [PubMed]
- Mustafa, A.K.; Gadalla, M.M.; Sen, N.; Kim, S.; Mu, W.; Sadia, K.; Barrow, R.K.; Yang, G.; Wang, R.; Snyder, S.H. H2S signals through protein S-sulfhydration. Sci. Signal. 2009, 2, ra72. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Zhao, K.; Ju, Y.; Mani, S.; Cao, Q.; Puukila, S.; Khaper, N.; Wu, L.; Wang, R. Hydrogen sulfide protects against cellular senescence via S-sulfhydration of Keap1 and activation of Nrf2. Antioxid. Redox Signal. 2013, 18, 1906–1919. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Macinkovic, I.; Devarie-baez, N.O.; Pan, J.; Park, C.; Carroll, K.S.; Filipovic, M.R.; Xian, M. Detection of protein S-sulfhydration by a tag-switch technique. Angew. Chem. Int. Ed. Engl. 2014, 53, 575–581. [Google Scholar] [CrossRef] [PubMed]
- Dóka, É.; Pader, I.; Bíró, A.; Johansson, K.; Cheng, Q.; Ballagó, K.; Prigge, J.R.; Pastor-flores, D.; Dick, T.P.; Schmidt, E.E.; et al. A novel persulfide detection method reveals protein persulfide- and polysulfide-reducing functions of thioredoxin and glutathione systems. Sci. Adv. 2016, 2, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Cuevasanta, E.; Lange, M.; Bonanata, J.; Coitiño, E.L.; Ferrer-Sueta, G.; Filipovic, M.R.; Alvarez, B. Reaction of hydrogen sulfide with disulfide and sulfenic acid to form the strongly nucleophilic persulfide. J. Biol. Chem. 2015, 290, 26866–26880. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Padovani, D.; Leslie, R.A.; Chiku, T.; Banerjee, R. Relative contributions of cystathionine β-synthase and γ-cystathionase to H2S biogenesis via alternative. J. Biol. Chem. 2009, 284, 22457–22466. [Google Scholar] [CrossRef] [PubMed]
- Shibuya, N.; Tanaka, M.; Yoshida, M.; Ogasawara, Y.; Togawa, T.; Ishii, K.; Kimura, H. 3-Mercaptopyruvate sulfurtransferase produces hydrogen sulfide and bound sulfane sulfur in the brain. Antioxid. Redox Signal. 2009, 11, 703–714. [Google Scholar] [CrossRef] [PubMed]
- Shibuya, N.; Koike, S.; Tanaka, M.; Ishigami-Yuasa, M.; Kimura, Y.; Ogasawara, Y.; Fukui, K.; Nagahara, N.; Kimura, H. A novel pathway for the production of hydrogen sulfide from d-cysteine in mammalian cells. Nat. Commun. 2013, 4, 1366. [Google Scholar] [CrossRef] [PubMed]
- Khoronenkova, S. V.; Tishkov, V.I. d-amino acid oxidase: Physiological role and applications. Biochemistry 2008, 73, 1511–1518. [Google Scholar] [CrossRef] [PubMed]
- Filipovic, M.R. Persulfidation (S-sulfhydration) and H2S. Handb. Exp. Pharmacol. 2015, 230, 29–59. [Google Scholar] [PubMed]
- Mishanina, T.V.; Libiad, M.; Banerjee, R. Biogenesis of reactive sulfur species for signaling by hydrogen sulfide oxidation pathways. Nat. Chem. Biol. 2015, 11, 457–464. [Google Scholar] [CrossRef] [PubMed]
- Wedmann, R.; Onderka, C.; Wei, S.; Szijártó, I.A.; Miljkovic, J.L.; Mitrovic, A.; Lange, M.; Savitsky, S.; Yadav, P.K.; Torregrossa, R.; et al. Improved tag-switch method reveals that thioredoxin acts as depersulfidase and controls the intracellular levels of protein persulfidation. Chem. Sci. 2016, 7, 3414–3426. [Google Scholar] [CrossRef] [PubMed]
- Goodwin, L.R.; Francom, D.; Dieken, F.P.; Taylor, J.D.; Warenycia, M.W.; Reiffenstein, R.J.; Dowling, G. Determination of sulfide in brain tissue by gas dialysis/ion chromatography: Postmortem studies and two case reports. J. Anal. Toxicol. 1989, 13, 105–109. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Ndisang, J.F.; Wang, R. Modulation of endogenous production of H2S in rat tissues. Can. J. Physiol. Pharmacol. 2003, 81, 848–853. [Google Scholar] [CrossRef] [PubMed]
- Savage, J.C.; Gould, D.H. Determination of sulfide in brain tissue and rumen fluid by ion-interaction reversed-phase high-performance liquid chromatography. J. Chromatogr. B Biomed. Sci. Appl. 1990, 526, 540–545. [Google Scholar] [CrossRef]
- Furne, J.; Saeed, A.; Levitt, M.D. Whole tissue hydrogen sulfide concentrations are orders of magnitude lower than presently accepted values. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2008, 295, 1479–1485. [Google Scholar] [CrossRef] [PubMed]
- Yuan, S.; Patel, R.P.; Kevil, C.G. Working with nitric oxide and hydrogen sulfide in biological systems. Am. J. Physiol. Lung Cell. Mol. Physiol. 2015, 308, L403–L415. [Google Scholar] [CrossRef] [PubMed]
- Hughes, M.N.; Centelles, M.N.; Moore, K.P. Making and working with hydrogen sulfide: The chemistry and generation of hydrogen sulfide in vitro and its measurement in vivo: A review. Free Radic. Biol. Med. 2009, 47, 1346–1353. [Google Scholar] [CrossRef] [PubMed]
- Greiner, R.; Pálinkás, Z.; Bäsell, K.; Becher, D.; Antelmann, H.; Nagy, P.; Dick, T.P. Polysulfides link H2S to protein thiol oxidation. Antioxid. Redox Signal. 2013, 19, 1749–1765. [Google Scholar] [CrossRef] [PubMed]
- Nagy, P.; Pálinkás, Z.; Nagy, A.; Budai, B.; Tóth, I.; Vasa, A. Chemical aspects of hydrogen sulfide measurements in physiological samples. Biochim. Biophys. Acta 2014, 1840, 876–891. [Google Scholar] [CrossRef] [PubMed]
- Westley, J. Rhodanese. Adv. Enzymol. Relat. Areas Mol. Biol. 1973, 39, 327–368. [Google Scholar] [PubMed]
- Westley, J. Rhodanese and the sulfane pool. In Enzymatic Basis of Detoxication; Jacob, W.B., Ed.; Academic Press: New York, NY, USA, 1980; Volume 2, pp. 245–262. [Google Scholar]
- Westley, J.; Adler, H.; Westley, L.; Nishida, C. The sulfurtransferases. Fundam. Appl. Toxicol. 1983, 3, 377–382. [Google Scholar] [CrossRef]
- Westley, J. Mammalian cyanide detoxification with sulphane sulphur. CIBA Found. Symp. 1988, 140, 201–212. [Google Scholar] [PubMed]
- Ogasawara, Y.; Isoda, S.; Tanabe, S. Tissue and subcellular distribution of bound and acid-labile sulfur, and the enzymic capacity for sulfide production in the rat. Biol. Pharm. Bull. 1994, 17, 1535–1542. [Google Scholar] [CrossRef] [PubMed]
- Ishigami, M.; Hiraki, K.; Umemura, K.; Ogasawara, Y.; Ishii, K.; Kimura, H. A source of hydrogen sulfide and a mechanism of its release in the brain. Antioxid. Redox Signal. 2009, 11, 205–214. [Google Scholar] [CrossRef] [PubMed]
- Enokido, Y.; Suzuki, E.; Iwasawa, K.; Namekata, K.; Okazawa, H.; Kimura, H.; Medical, T.; Medical, T. Cystathionine β-synthase, a key enzyme for homocysteine metabolism, is preferentially expressed in the radial glia/astrocyte lineage of developing mouse CNS. FASEB J. 2005, 24, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Nagahara, N.; Ito, T.; Kitamura, H.; Nishino, T. Tissue and subcellular distribution of mercaptopyruvate sulfurtransferase in the rat: Confocal laser fluorescence and immunoelectron microscopic studies combined with biochemical analysis. Histochem. Cell Biol. 1998, 110, 243–250. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Chan, S.J.; Ng, Y.K.; Wong, P.T.H. Brain 3-mercaptopyruvate sulfurtransferase (3MST): Cellular localization and downregulation after acute stroke. PLoS ONE 2013, 8, e67322. [Google Scholar] [CrossRef] [PubMed]
- Ogasawara, Y.; Suzuki, T.; Ishii, K.; Tanabe, S. Modification of liver cytosol enzyme activities promoted in vitro by reduced sulfur species generated from cystine with γ-cystathionase. Biochim. Biophys. Acta 1997, 1334, 33–43. [Google Scholar] [CrossRef]
- Koike, S.; Ogasawara, Y.; Shibuya, N.; Kimura, H.; Ishii, K. Polysulfide exerts a protective effect against cytotoxicity caused by t-buthylhydroperoxide through Nrf2 signaling in neuroblastoma cells. FEBS Lett. 2013, 587, 3548–3555. [Google Scholar] [CrossRef] [PubMed]
- Kimura, Y.; Mikami, Y.; Osumi, K.; Tsugane, M.; Oka, J.I.; Kimura, H. Polysulfides are possible H2S-derived signaling molecules in rat brain. FASEB J. 2013, 27, 2451–2457. [Google Scholar] [CrossRef] [PubMed]
- Jarosz, A.P.; Wei, W.; Gauld, J.W.; Auld, J.; Özcan, F.; Aslan, M.; Mutus, B. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) is inactivated by S-sulfuration in vitro. Free Radic. Biol. Med. 2015, 89, 512–521. [Google Scholar] [CrossRef] [PubMed]
- Jenner, P. Oxidative stress in Parkinson’s disease. Ann. Neurol. 2003, 53, 26–38. [Google Scholar] [CrossRef] [PubMed]
- Markesbery, W.R. Oxidative stress hypothesis in Alzheimer’s disease. Free Radic. Biol. Med. 1997, 23, 134–147. [Google Scholar] [CrossRef]
- Itoh, K.; Chiba, T.; Takahashi, S.; Ishii, T.; Igarashi, K.; Katoh, Y.; Oyake, T.; Hayashi, N.; Satoh, K.; Hatayama, I.; et al. An Nrf2 / small Maf heterodimer mediates the induction of phase II detoxifying enzyme genes through antioxidant response elements. Biochem. Biophys. Res. Commun. 1997, 322, 313–322. [Google Scholar] [CrossRef]
- Calvert, J.W.; Jha, S.; Gundewar, S.; Elrod, J.W.; Ramachandran, A.; Pattillo, C.B.; Kevil, C.G.; Lefer, D.J. Hydrogen sulfide mediates cardioprotection through Nrf2 signaling. Circ. Res. 2009, 105, 365–374. [Google Scholar] [CrossRef] [PubMed]
- Peake, B.F.; Nicholson, C.K.; Lambert, J.P.; Hood, R.L.; Amin, H.; Amin, S.; Calvert, J.W. Hydrogen sulfide preconditions the db/db diabetic mouse heart against ischemia-reperfusion injury by activating Nrf2 signaling in an Erk-dependent manner. Am. J. Physiol. Heart Circ. Physiol. 2013, 304, 1215–1224. [Google Scholar] [CrossRef] [PubMed]
- Hourihan, J.M.; Kenna, J.G.; Hayes, J.D. The gasotransmitter hydrogen sulfide induces Nrf2-target genes by inactivating the Keap1 ubiquitin ligase substrate adaptor through formation of a disulfide bond between Cys-226 and Cys-613. Antioxid. Redox Signal. 2013, 19, 465–481. [Google Scholar] [CrossRef] [PubMed]
- Toohey, J.I. The conversion of H2S to sulfane sulfur. Nat. Rev. Mol. Cell Biol. 2012, 13, 803. [Google Scholar] [CrossRef] [PubMed]
- Ida, T.; Sawa, T.; Ihara, H.; Tsuchiya, Y.; Watanabe, Y.; Kumagai, Y. Reactive cysteine persulfides and S-polythiolation regulate oxidative stress and redox signaling. Proc. Natl. Acad. Sci. USA 2014, 111, 7606–7611. [Google Scholar] [CrossRef] [PubMed]
- Rabbani, N.; Thornalley, P.J. Dicarbonyl stress in cell and tissue dysfunction contributing to ageing and disease. Biochem. Biophys. Res. Commun. 2015, 458, 221–226. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, N.; Fukatsu, R.; Tsuzuki, K.; Koike, T.; Wakayama, I.; Yanagihara, R.; Garruto, R. Advanced glycation end products in Alzheimer’s disease and other neurodegenerative diseases. Am. J. Pathol. 1998, 153, 1149–1155. [Google Scholar] [CrossRef]
- Arai, M.; Yuzawa, H.; Nohara, I.; Ohnishi, T.; Obata, N.; Iwayama, Y.; Haga, S.; Toyota, T.; Ujike, H.; Arai, M.; et al. Enhanced carbonyl stress in a subpopulation of schizophrenia carbonyl stress in schizophrenia. Arch. Gen. Psychiatry 2010, 67, 589–597. [Google Scholar] [CrossRef] [PubMed]
- Arai, M.; Miyashita, M.; Kobori, A.; Toriumi, K.; Horiuchi, Y.; Itokawa, M. Carbonyl stress and schizophrenia. Psychiatry Clin. Neurosci. 2014, 68, 655–665. [Google Scholar] [CrossRef] [PubMed]
- Chang, T.; Untereiner, A.; Liu, J.; Wu, L. Interaction of methylglyoxal and hydrogen sulfide in rat vascular smooth muscle cells. Antioxid. Redox Signal. 2010, 12, 1093–1100. [Google Scholar] [CrossRef] [PubMed]
- Ogasawara, Y.; Tanaka, R.; Koike, S.; Horiuchi, Y.; Miyashita, M.; Arai, M. Determination of methylglyoxal in human blood plasma using fluorescence high performance liquid chromatography after derivatization with 1,2-diamino-4,5-methylenedioxybenzene. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2016, 1029–1030, 102–105. [Google Scholar] [CrossRef] [PubMed]
- Koike, S.; Kayama, T.; Yamamoto, S.; Komine, D.; Tanaka, R.; Nishimoto, S.; Suzuki, T.; Kishida, A.; Ogasawara, Y. Polysulfides protect SH-SY5Y cells from methylglyoxal-induced toxicity by suppressing protein carbonylation: A possible physiological scavenger for carbonyl stress in the brain. Neurotoxicology 2016, 55, 13–19. [Google Scholar] [CrossRef] [PubMed]
- King, A.L.; Polhemus, D.J.; Bhushan, S.; Otsuka, H.; Kondo, K.; Nicholson, C.K. Hydrogen sulfide cytoprotective signaling is endothelial nitric oxide synthase-nitric oxide dependent. Proc. Natl. Acad. Sci. USA 2014, 111, 3182–3187. [Google Scholar] [CrossRef] [PubMed]
- Robert, K.; Nehmé, J.; Bourdon, E.; Pivert, G.; Friguet, B.; Delcayre, C.; Delabar, J.M.; Janel, N. Cystathionine beta synthase deficiency promotes oxidative stress, fibrosis, and steatosis in mice liver. Gastroenterology 2005, 128, 1405–1415. [Google Scholar] [CrossRef] [PubMed]

© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Koike, S.; Ogasawara, Y. Sulfur Atom in its Bound State Is a Unique Element Involved in Physiological Functions in Mammals. Molecules 2016, 21, 1753. https://doi.org/10.3390/molecules21121753
Koike S, Ogasawara Y. Sulfur Atom in its Bound State Is a Unique Element Involved in Physiological Functions in Mammals. Molecules. 2016; 21(12):1753. https://doi.org/10.3390/molecules21121753
Chicago/Turabian StyleKoike, Shin, and Yuki Ogasawara. 2016. "Sulfur Atom in its Bound State Is a Unique Element Involved in Physiological Functions in Mammals" Molecules 21, no. 12: 1753. https://doi.org/10.3390/molecules21121753
APA StyleKoike, S., & Ogasawara, Y. (2016). Sulfur Atom in its Bound State Is a Unique Element Involved in Physiological Functions in Mammals. Molecules, 21(12), 1753. https://doi.org/10.3390/molecules21121753




