2-Keto-D-Gluconate-Yielding Membrane-Bound D-Glucose Dehydrogenase from Arthrobacter globiformis C224: Purification and Characterization
Abstract
:1. Introduction
2. Results and Discussion
2.1. Extraction and Purification of GlcDH from A. globiformis C224
| Step | Protein (mg) | Enzyme Activity (Units) | Specific Activity (Units/mg) | Recovery (%) | Purification (Fold) |
|---|---|---|---|---|---|
| Membrane fraction | 9758.1 ± 121.3 | 4730.6 ± 96.5 | 0.5 | 100.0 | 1.0 |
| Triton X-114 phase separation | 6004.5 ± 96.1 | 4396.4 ± 79.8 | 0.7 | 92.9 | 1.5 |
| Acetone precipitation | 1103.7 ± 30.6 | 1524.9 ± 34.2 | 1.4 | 32.2 | 2.9 |
| PEG precipitation | 408.8 ± 17.4 | 846.3 ± 12.3 | 2.1 | 17.9 | 4.3 |
| Ethanol precipitation | 33.6 ± 1.2 | 268.2 ± 6.4 | 7.9 | 5. 7 | 16.6 |
| Hydroxylapatite fraction | 1.8 ± 0.1 | 159.4 ± 2.1 | 88.1 | 3.4 | 183.5 |

2.2. Type Identification of GlcDH from A. globiformis C224
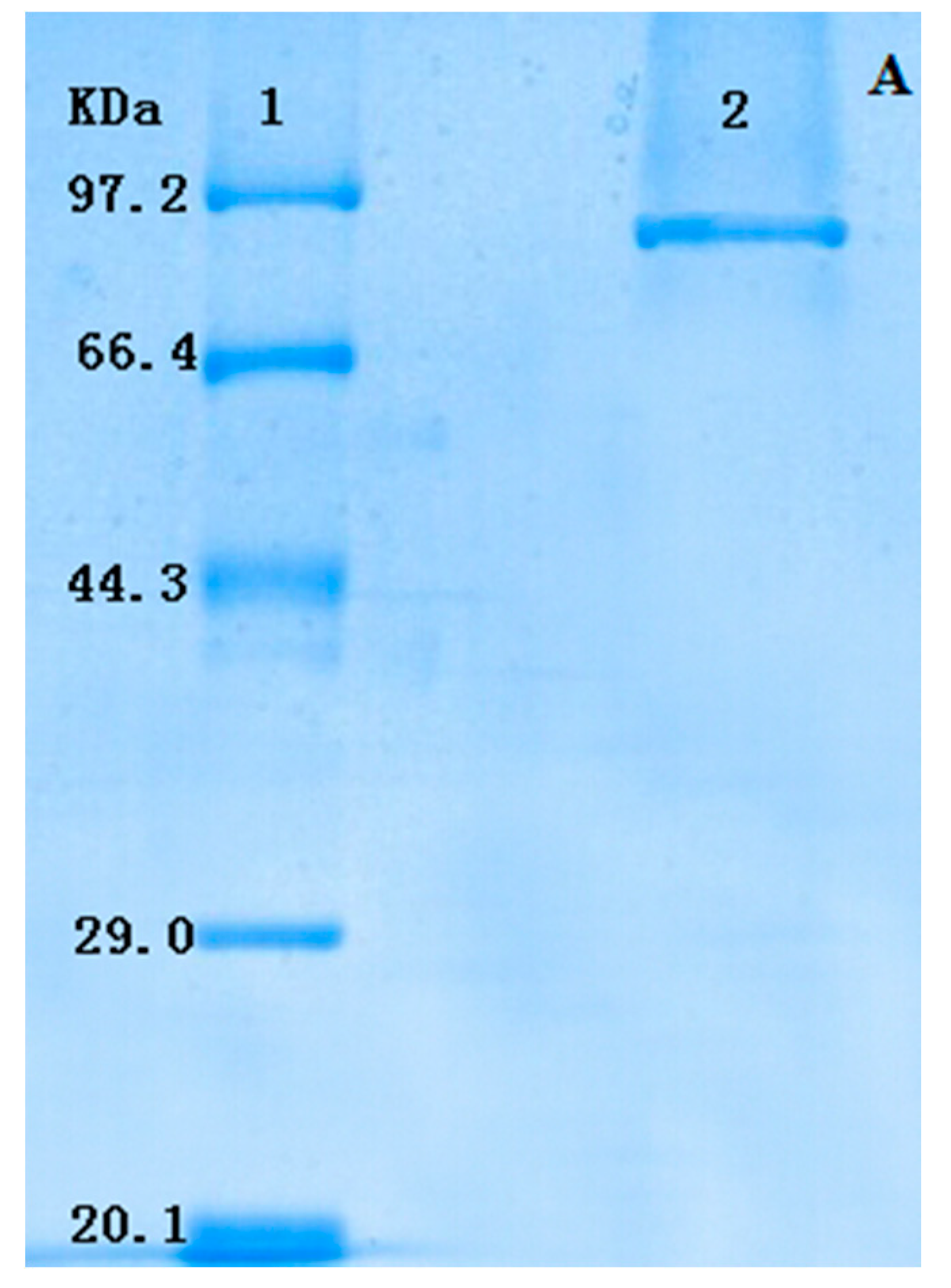

| Protein | Similar Sequence in the Database of the NCBI Blast | Strains | Mass (Da) | Score |
|---|---|---|---|---|
| Gi|104780479 glucose dehydrogenase | R.LLALDPDTGAEIWR.Y | Pseudomonas entomophila L48 | 87,067 | 142 |
| R.GIGPFTAGGYYSTSPAAITR.S | ||||
| Gi|339489095 glucose dehydrogenase | R.TEHGDRYSPLR.Q | Pseudomonas putida S16 | 86,935 | 117 |
| R.LLALDPDTGAEIWR.F | ||||
| R.GVSYYDENRYVSR.D | ||||
| Gi|167035354 PQQ-dependent glucose dehydrogenase | R.TEHGDRYSPLR.Q | Pseudomonas putida GB-1 | 86,939 | 114 |
| R.LLALDPDTGAEIWR.F | ||||
| R.GVSYYDENRYVSR.D | ||||
| Gi|26988177 glucose dehydrogenase | R.LLALDPDTGAEIWR.Y | Pseudomonas putida KT2440 | 86,926 | 100 |
| R.GVSYYDENRYVSR.D |
2.3. Enzymatic Properties of Purified Membrane-Bound GlcDH from A. globiformis C224
2.3.1. Substrate Specificity
| Substrate | Concentration of Substrate (mM) | Membrane Enzyme (U/mL) | Purified EnzymeRelative Activity (%) |
|---|---|---|---|
| d-Glucose | 33.0 | 21.9 ± 0.9 | 100.0 ± 1.6 |
| d-Gluconate | 33.0 | 82.2 ± 3.8 | 0 |
| Maltose | 33.0 | 3.1 ± 0.1 | 17.0 ± 0.8 |
| d-Sorbose | 33.0 | 0 | 0 |
| d-Galactose | 33.0 | 3.8 ± 0.1 | 20.0 ± 0.9 |
| d-Mannose | 33.0 | 0 | 0 |
| d-Fructose | 33.0 | 0 | 0 |
| d-Arabinose | 33.0 | 2.1 ± 0.1 | 8.0 ± 0.5 |
| Malic acid | 33.0 | 0 | 0 |
| Sucrose | 33.0 | 0 | 0 |
| Citric acid | 33.0 | 0 | 0 |
| d-Xylose | 33.0 | 5.3 ± 0.2 | 22.0 ± 0.9 |
2.3.2. Kinetic Studies of GlcDH from A. globiformis C224
Single Substrate Kinetics
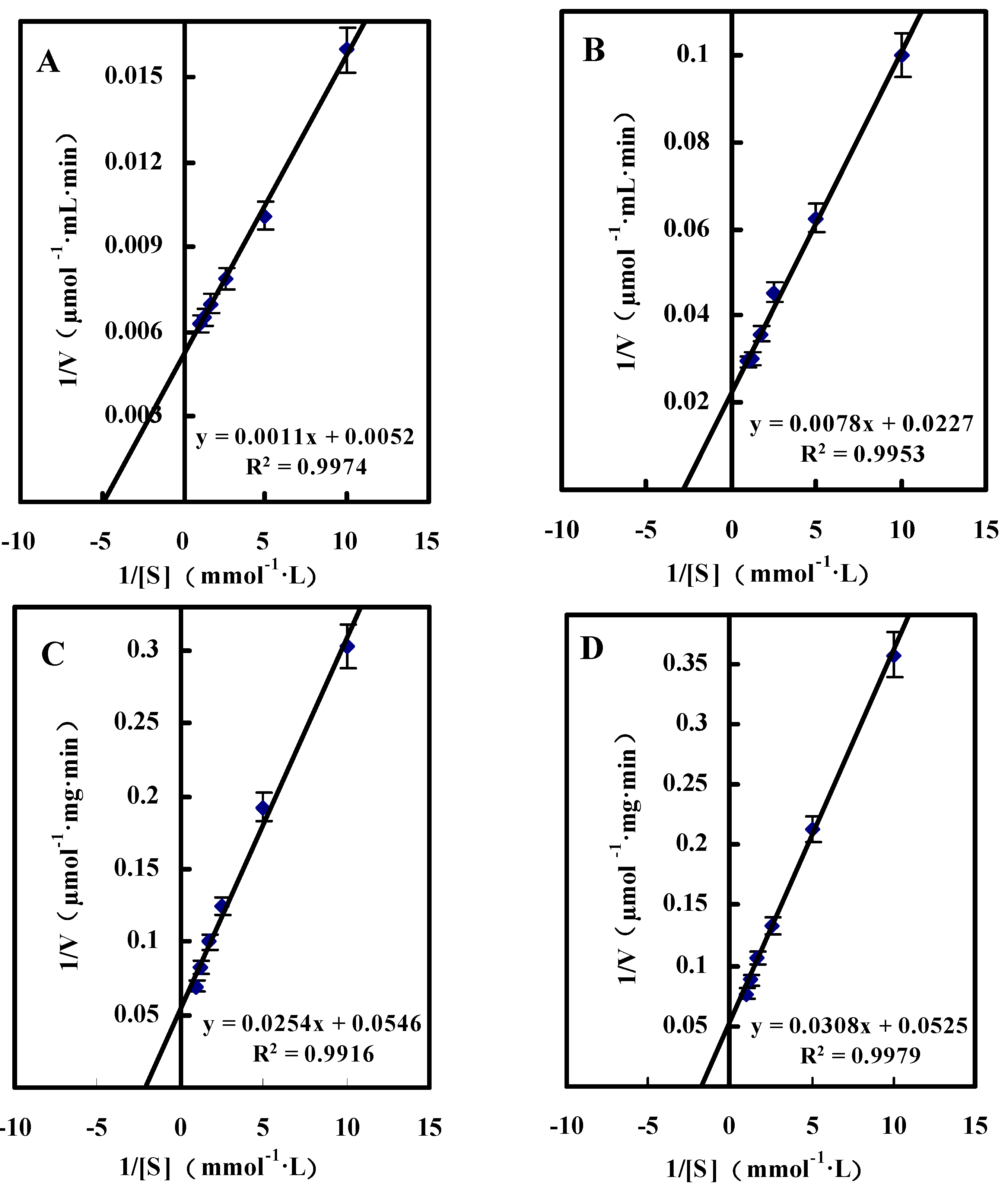
| Substrate | Km (mM) | Vmax (μmol/mg·min) |
|---|---|---|
| d-Glucose | 0.21 ± 0.01 | 192.31 ± 9.45 |
| d-Xylose | 0.34 ± 0.02 | 44.05 ± 1.94 |
| d-Galactose | 0.46 ± 0.02 | 18.32 ± 0.76 |
| Maltose | 0.59 ± 0.03 | 19.05 ± 0.85 |
Dual Substrates Kinetics
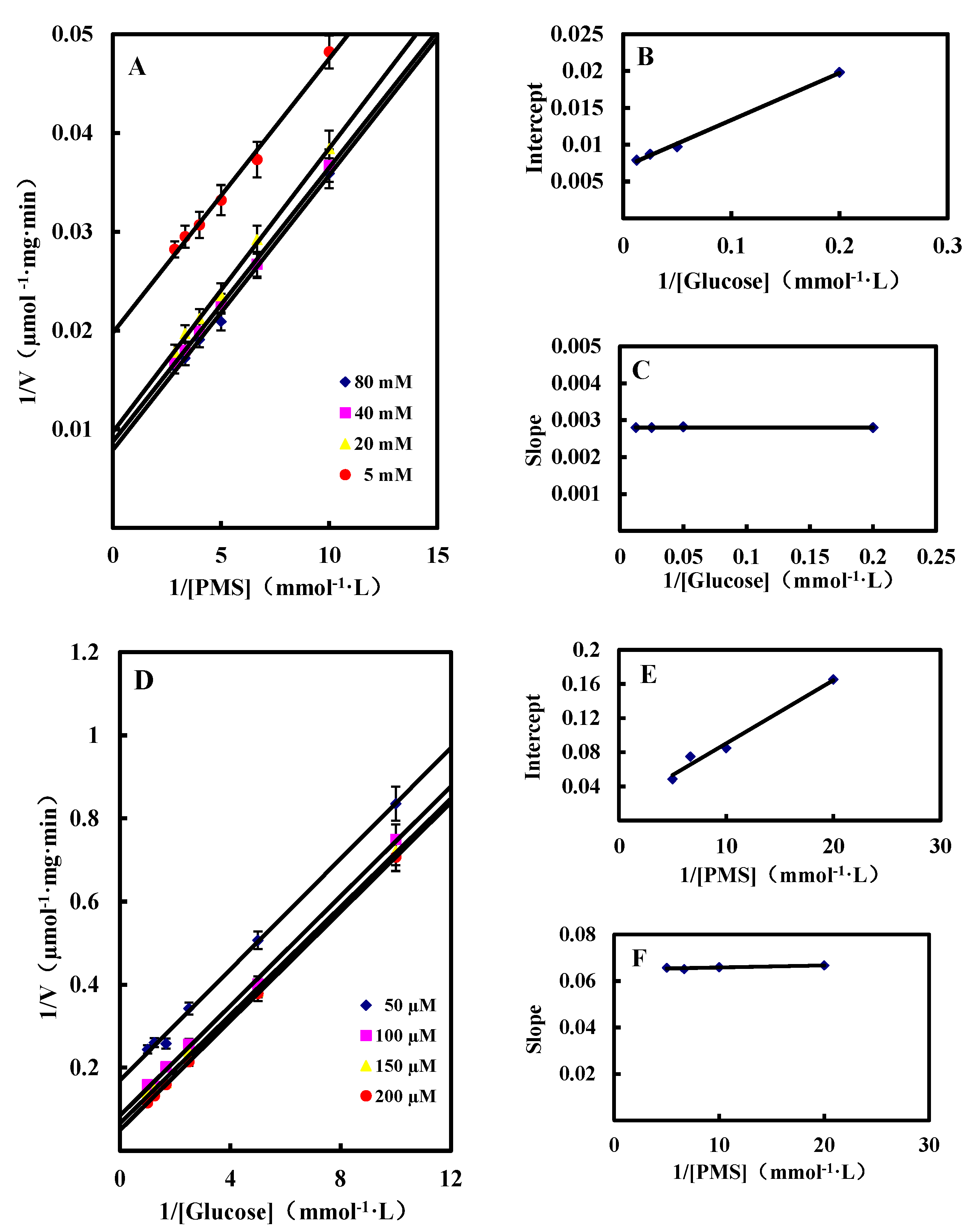
Inhibition by Gluconic Acid
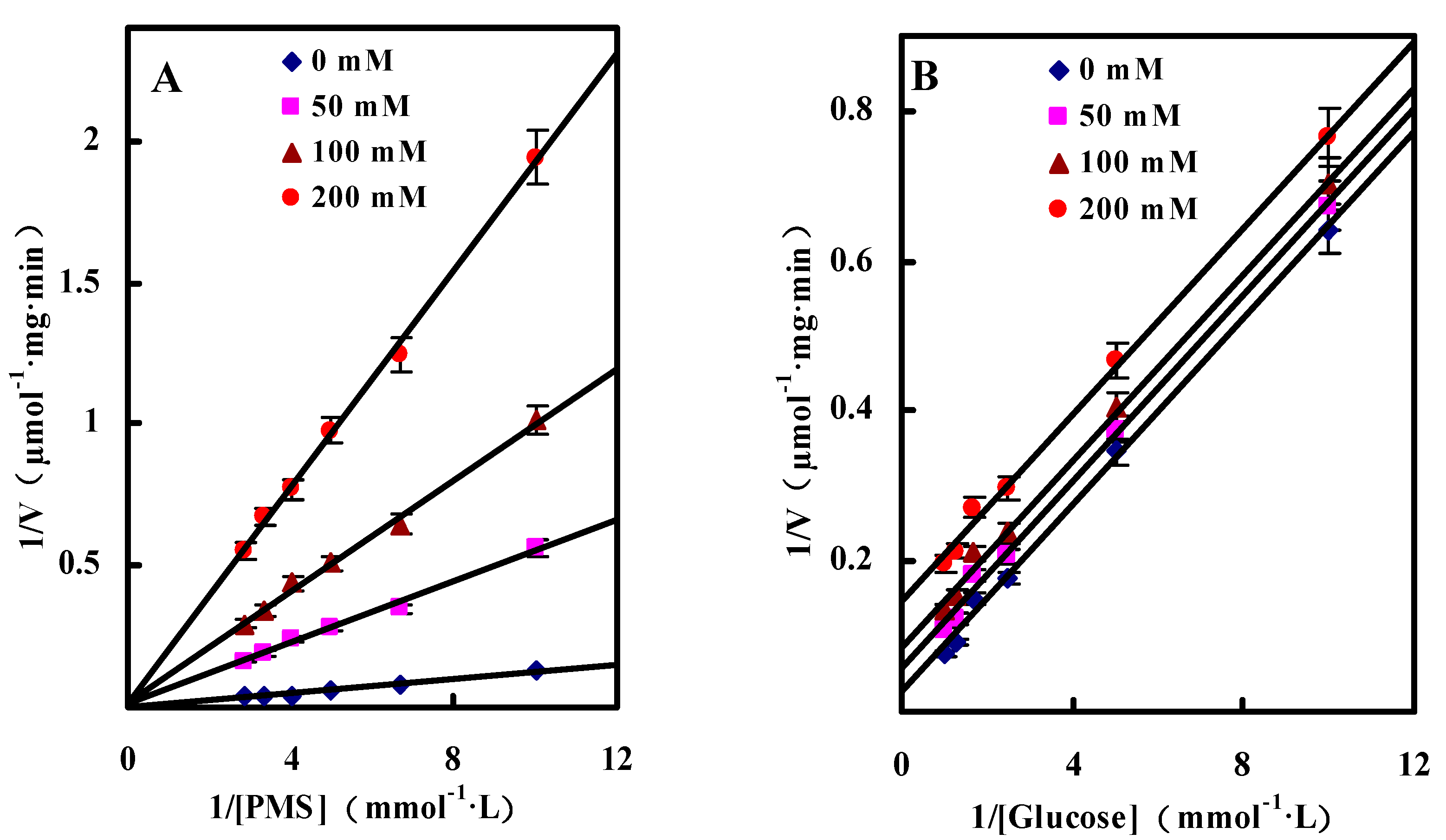
2.3.3. Effect of pH on GlcDH Activity and Stability
| Properties | Escherichia coli [11] | Gluconobacter Suboxydans [12] | Bacillus Thuringiensis [13] | Pseudomonas sp. [14] | Acinetobacter Calcoaceticus [15] | Arthrobacter Globiformis(This Study) |
|---|---|---|---|---|---|---|
| Molecular mass of subunits (Da) | 88,000 | 87,000 | 25,000 26,000 | 90,000 | 47,500 48,000 | 87,000 |
| Optimum pH with ferricyanide | 3.5 | 3.0 | ND | 4.5 | ND | 5.0 |
| With DCIP-PMS | 6.0 | 6.0 | 8.0 | 6.0 | 6.0 | ND |
| Optimum temperature (°C) | ND | ND | 55 | ND | ND | 45 |
| Substrate range | Glucose; Mannose; Galactose; Fructose; Rhamnose; Xylose | Glucose; Maltose | Glucose; 2-d-deoxy-d-glucose | Glucose; Mannose; Galactose; Xylose; Maltose; Rhamnose | Glucose; Xylose; Arabinose; Lactose; Galactose; Melibiose; Cellobiose; Maltose | Glucose; Xylose; Galactose; Maltose |
| Km (mM) | ND | ND | 14 (Glucose, pH 8.0); 12.2 (2-d-deoxy-d-glucose, pH 8.0) | 0.69 (Glucose, pH 4.5); 1.6 (Glucose, pH 6.0) | 22 (Glucose, pH 6.0) | 0.21 (Glucose, pH 5.0); 0.34 (Xylose, pH 5.0); 0.46 (Galactose, pH 5.0); 0.59 (Maltose, pH 5.0) |

2.3.4. Effect of Temperature on GlcDH Activity and Stability
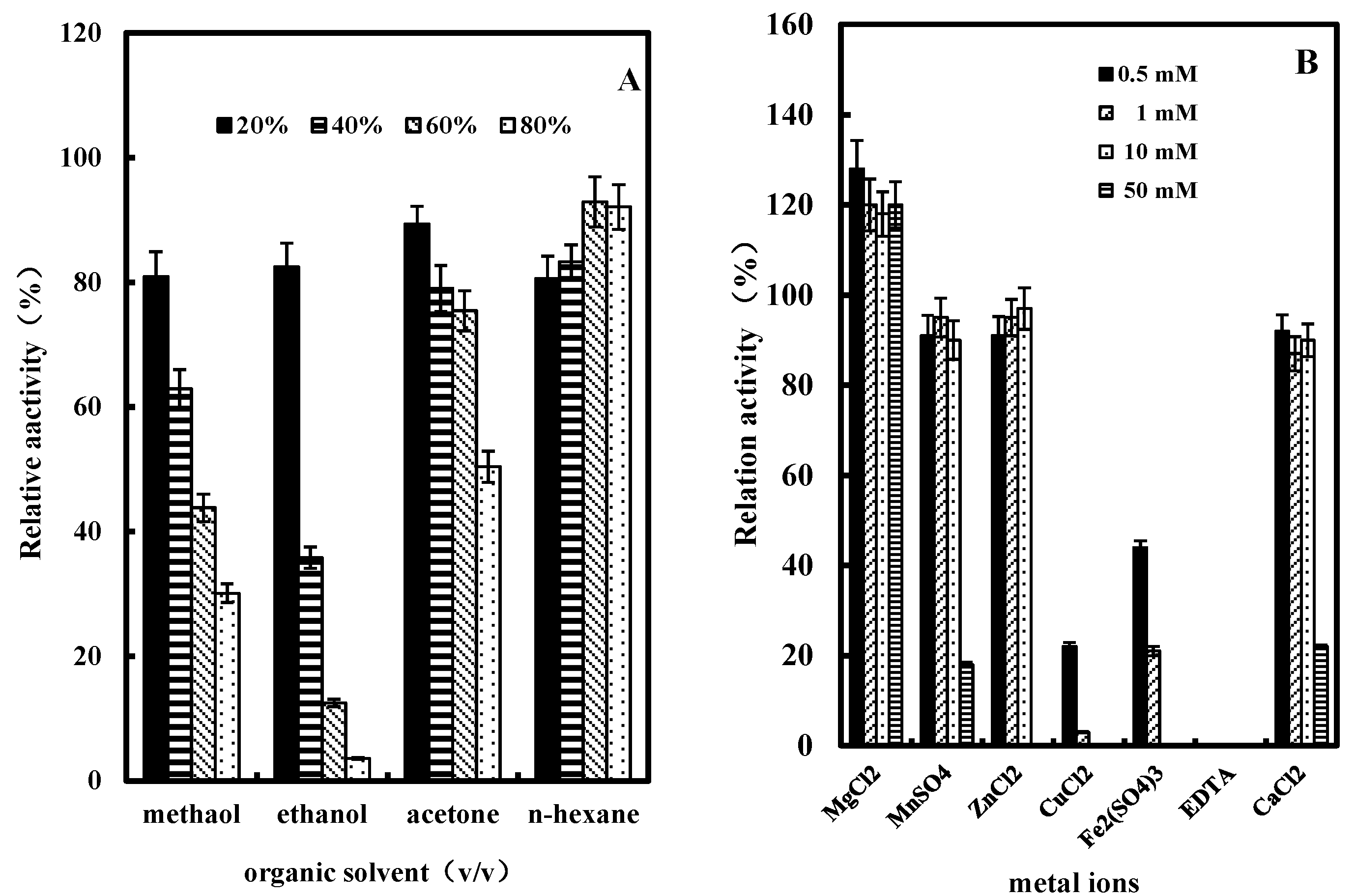
2.3.5. Effect of Organic Solvents on GlcDH Activity
2.3.6. Effect of Metal Ions or EDTA on GlcDH Activity
3. Experimental Section
3.1. Chemicals
3.2. Microorganism and Medium
3.3. Preparation of Cell Culture
3.4. Extraction and Purification of GlcDH from A. globiformis C224
3.5. Polyacrylamide Gel Electrophoresis
3.6. MALDI-TOF-MS Analysis
3.7. Assay of GlcDH Activity and Protein Determination
3.8. Enzymatic Properties
3.8.1. Substrate Specificity
3.8.2. Effect of pH, Temperature, Organic Solvents or Metal Ions on GlcDH Activity
3.9. Statistical Analysis
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Zhou, Q.; Wei, Z.; Sun, W.J.; Yu, S.L.; Li, Z.B. Research progress on production technology of erythorbic acid. Food Sci. 2008, 29, 647–651. (In Chinese) [Google Scholar]
- Wei, Z.; Yu, S.L.; Sun, W.J.; Zhou, Q.; Li, Z.B. Research progress on fermentation production of 2-keto-d-gluconic acid. Food Sci. 2008, 29, 636–639. (In Chinese) [Google Scholar]
- Sun, W.J.; Yun, Q.Q.; Zhou, Y.Z.; Cui, F.J. Continuous 2-keto-gluconic acid (2KGA) production from corn starch hydrolysate by Pseudomonas fluorescens AR4. Biochem. Eng. J. 2013, 77, 97–102. [Google Scholar] [CrossRef]
- Elfari, M.; Ha, S.W.; Bremus, C.; Merfort, M. A Gluconobacter oxydans mutant converting glucose almost quantitatively to 5-keto-d-gluconic acid. Appl. Microbiol. Biotechol. 2005, 66, 668–674. [Google Scholar] [CrossRef]
- Toyama, H.; Furuya, N.; Saichana, I. Membrane-bound, 2-Keto-D-Gluconate-Yielding d-gluconate dehydrogenase from “Gluconobacter dioxyacetonicus” IFO 3271: Molecular properties and gene disruption. Appl. Environ. Microbiol. 2007, 73, 6551–6556. [Google Scholar] [CrossRef] [PubMed]
- Matsushita, K.; Fujii, Y.; Ano, Y. 5-Keto-d-Gluconate production is catalyzed by a quinoprotein glycerol dehydrogenase, major polyol dehydrogenase, in Gluconobacter species. Appl. Environ. Microbiol. 2003, 69, 1959–1966. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.J.; Zhou, Y.Z.; Zhou, Q.; Cui, F.J. Semi-continuous production of 2-keto-gluconic acid by Pseudomonas fluorescens AR4 from rice starch hydrolysate. Bioresour. Technol. 2012, 110, 546–551. [Google Scholar] [CrossRef] [PubMed]
- Teng, W.H.; Sun, W.J.; Yu, B.; Cui, F.J. Continuous conversion of rice starch hydrolysate to 2-keto-d-gluconic acid by Arthrobacter globiformis C224. Biotechnol. Bioprocess Eng. 2013, 18, 709–714. [Google Scholar] [CrossRef]
- Iwamoto, R.; Tanimura, R.; Ikehara, K. Purification and characterization of 2-keto-d-galactonate reductase from Psedomonas fluorescens. J. Mol. Catal. B: Enzym. 2007, 47, 43–50. [Google Scholar] [CrossRef]
- Dewanti, A.R.; Duine, J.A. Reconstitution of Membrane-Integrated Quinoprotein Glucose Dehydrogenase Apoenzyme with PQQ and the Holoenzyme’s Mechanism of Action. Biochemistry 1998, 37, 6810–6818. [Google Scholar] [CrossRef] [PubMed]
- Ameyama, M.; Nonobe, M.; Shinagawa, E. Purication and Characterization of the Quinoprotein d-Glucose Dehydrogenase Apoenzyme from Escherichia coli. Agric. Biol. Chem. 1986, 50, 49–57. [Google Scholar] [CrossRef]
- Ameyama, M.; Shinagawa, E.; Matsushita, K. d-Glucose Dehydrogenase of Gluconobacter suboxydans: Solubilization, Purification and Characterization. Agric. Biol. Chem. 1981, 45, 851–861. [Google Scholar] [CrossRef]
- Boontim, N.; Yoshimune, K.; Lumyong, S. Purification and characterization of d-glucose dehydrogenase from Bacillus thuringiensis M15. Ann. Microbiol. 2004, 54, 481–492. [Google Scholar]
- Matsushita, K.; Ohno, Y.; Shinagawa, E. Membrane-bound d-Glucose Dehydrogenase from Pseudomonas sp.: Solubilization, Purification and Characterization. Agric. Biol. Chem. 1980, 44, 1505–1512. [Google Scholar] [CrossRef]
- Dokter, P.; Frank, J.; Duine, A. Purification and characterization of quinoprotein glucose dehydrogenase from Acinetobacter calcoaceticus L.M.D. 79.41. Biochem. J. 1986, 239, 163–167. [Google Scholar] [PubMed]
- Seddon, A.M.; Curnow, P.; Booth, P.J. Membrane proteins, lipids and detergents: Not just a soap opera. Biochim. Biophys. Acta 2004, 1666, 105–117. [Google Scholar] [CrossRef] [PubMed]
- Mathias, R.A.; Chen, Y.S.; Kapp, E.A. Triton X-114 phase separation in the isolation and purification of mouse liver microsomal membrane proteins. Methods 2011, 54, 396–406. [Google Scholar] [CrossRef] [PubMed]
- Misquitta, Y.; Caffrey, M. Detergents destabilize the cubic phase of monoolein: Implications for membrane protein crystallization. Biophys. J. 2003, 85, 3084–3096. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.F.; Liu, R.; Yin, Y.Q. Triton X-114 based cloud point extraction: A thermoreversible approach for separation/concentration and dispersion of nanomaterials in the aqueous phase. Chem. Commun. 2009, 25, 1514–1516. [Google Scholar] [CrossRef]
- Qin, W.; Huang, Y.; Ding, Y.W. Surfactant distribution in the clouding of TX114. Chin. J. Chem. Eng. 2008, 16, 722–725. [Google Scholar] [CrossRef]
- Cavallaro, A.S.; Mahony, D.; Commins, M.; Mahony, T.J. Endotoxin-free purification for the isolation of Bovine Viral Diarrhoea Virus E2 protein from insoluble inclusion body aggregates. Microb. Cell Fact. 2011, 10, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Målen, H.; Pathak, S. Definition of novel cell envelope associated proteins in Triton X-114 extracts of Mycobacterium tuberculosis H37Rv. BMC Microbiol. 2010, 10, 132. [Google Scholar] [CrossRef] [PubMed]
- Målen, H.; Berven, F.S. Membrane and membrane-associated proteins in Triton X-114 extracts of Mycobacterium bovis BCG identified using a combination of gel-based and gel-free fractionation strategies. Proteomics 2008, 8, 1859–1870. [Google Scholar] [CrossRef] [PubMed]
- Bacth, J.A.; Sadoff, H.L. Aerobic sporulating bacteria. I. glucose dehydrogenase of Bacillus cereus. J. Bacteriol. 1962, 83, 699–707. [Google Scholar] [PubMed]
- Romero, M.D.; Calvo, L.; Alba, C. A kinetic study of isoamyl acetate synthesis by immobilized lipase-catalyzed acetylation in n-hexane. J. Biotechnol. 2007, 127, 269–277. [Google Scholar] [CrossRef] [PubMed]
- Goodenough-Lashua, D.M.; Garcia, G.A. tRNA-guanine transglycosy lase from E. coli: Ping-pong kinetic mechanism is consistent with nucleophilic catalysis. Bioorg. Chem. 2003, 31, 331–344. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.J.; Zhao, F.M.; Guo, J.Q. Selection of phage-resistant mutants from 2-keto-d-gluconic acid producing strain Arthrobacter globiformis K1022. Food Fermn. Ind. 2002, 28, 36–39. [Google Scholar]
- Jiang, L.; He, L. Fountoulakis M. Comparison of protein precipitation methods for sample preparation prior to proteomic analysis. J. Chromatogr. A 2004, 1023, 317–320. [Google Scholar] [CrossRef] [PubMed]
- Berber, I. Characterization of Bacillus species by numerical analysis of their SDS-PAGE protein profiles. J. Mol. Cell Biol. 2004, 3, 33–37. [Google Scholar]
- Baker, M.A.; Lane, D.J.R.; Ly, J.D. VDAC1 is a transplasma membrane NADH-ferricyanide reductase. J. Biol. Chem. 2004, 279, 4811–4819. [Google Scholar] [CrossRef] [PubMed]
- Sample Availability: Samples of the compounds 2-ketogluconic acid (2KGlcA) and Glucose dehydrogenase (GlcDH) are available from the authors.
© 2015 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xue, Q.; Wei, Z.; Sun, W.; Cui, F.; Yu, S.; Zhou, Q.; Liu, J. 2-Keto-D-Gluconate-Yielding Membrane-Bound D-Glucose Dehydrogenase from Arthrobacter globiformis C224: Purification and Characterization. Molecules 2015, 20, 846-862. https://doi.org/10.3390/molecules20010846
Xue Q, Wei Z, Sun W, Cui F, Yu S, Zhou Q, Liu J. 2-Keto-D-Gluconate-Yielding Membrane-Bound D-Glucose Dehydrogenase from Arthrobacter globiformis C224: Purification and Characterization. Molecules. 2015; 20(1):846-862. https://doi.org/10.3390/molecules20010846
Chicago/Turabian StyleXue, Qing, Zhuan Wei, Wenjing Sun, Fengjie Cui, Silian Yu, Qiang Zhou, and Jingze Liu. 2015. "2-Keto-D-Gluconate-Yielding Membrane-Bound D-Glucose Dehydrogenase from Arthrobacter globiformis C224: Purification and Characterization" Molecules 20, no. 1: 846-862. https://doi.org/10.3390/molecules20010846
APA StyleXue, Q., Wei, Z., Sun, W., Cui, F., Yu, S., Zhou, Q., & Liu, J. (2015). 2-Keto-D-Gluconate-Yielding Membrane-Bound D-Glucose Dehydrogenase from Arthrobacter globiformis C224: Purification and Characterization. Molecules, 20(1), 846-862. https://doi.org/10.3390/molecules20010846






