Cloning, 3D Modeling and Expression Analysis of Three Vacuolar Invertase Genes from Cassava (Manihot Esculenta Crantz)
Abstract
:1. Introduction
2. Results and Discussion
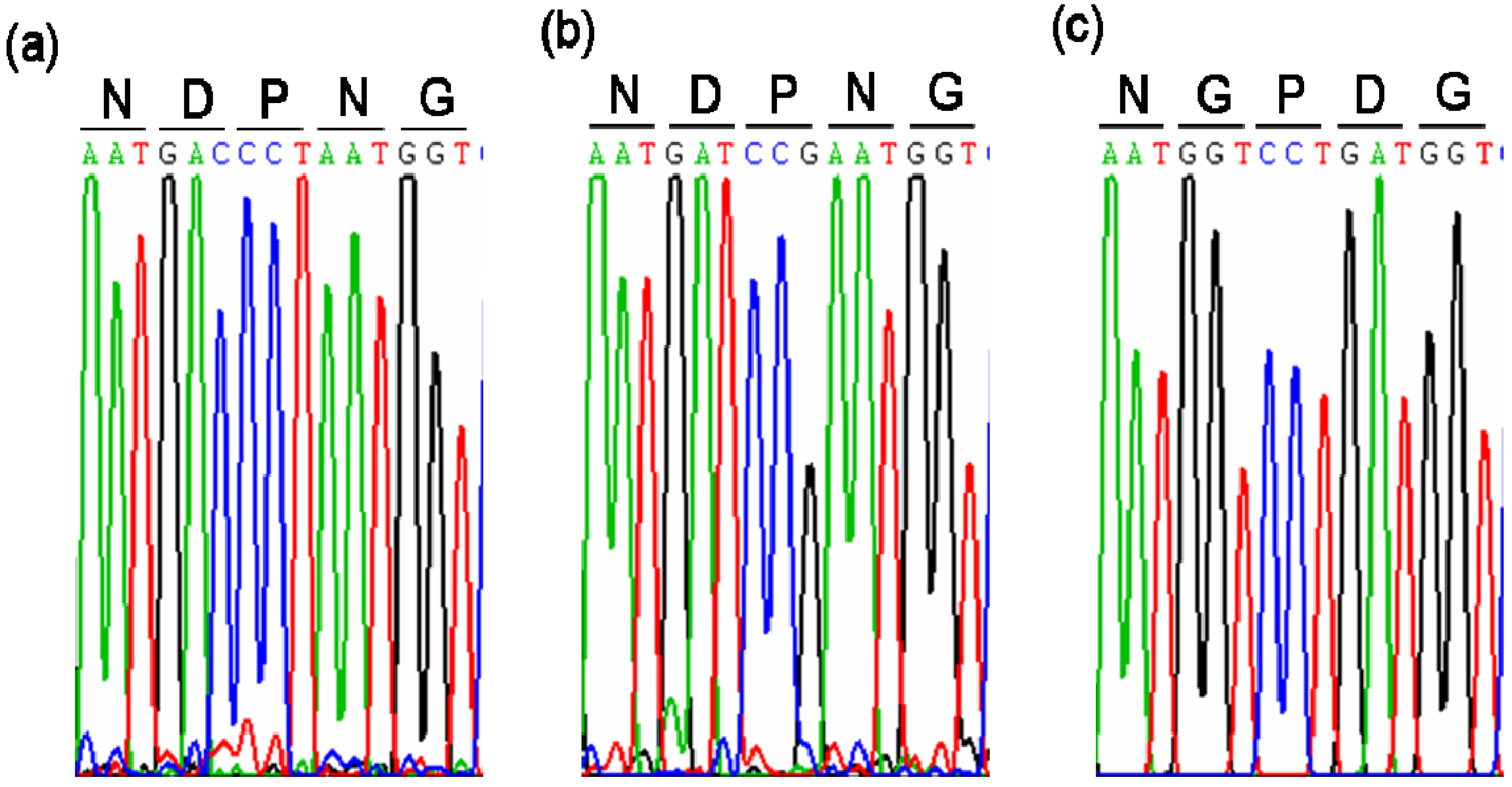
2.1. Cloning and Sequence Analysis of MeVINVs

| Gene Name | Accession Number | Genomic Location | ORF Length (bp) | Length (aa) | pI | MW(kDa) |
|---|---|---|---|---|---|---|
| MeVINV1 | JX291158 | scaffold07035: 858738–862530 | 1962 | 653 | 5.75 | 73.3 |
| MeVINV2 | JQ792174 | scaffold11581: 118337–121973 | 1983 | 660 | 5.26 | 73.5 |
| MeVINV3 | JQ792173 | scaffold01127: 398782–401926 | 1920 | 639 | 4.60 | 71.5 |

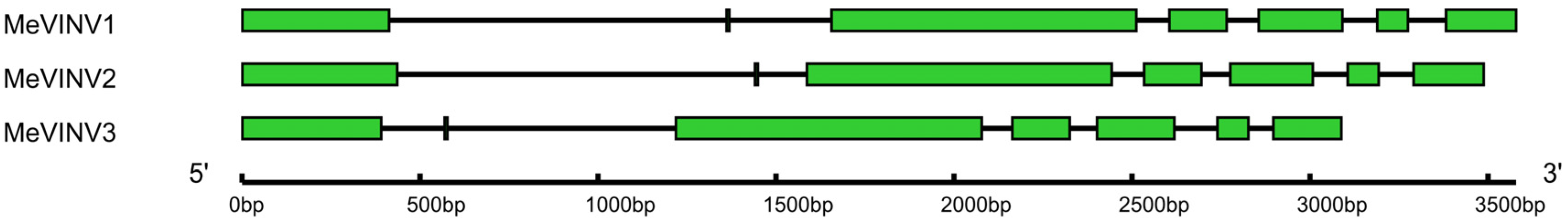
2.2. Structure Analysis of the MeVINV Family Genes
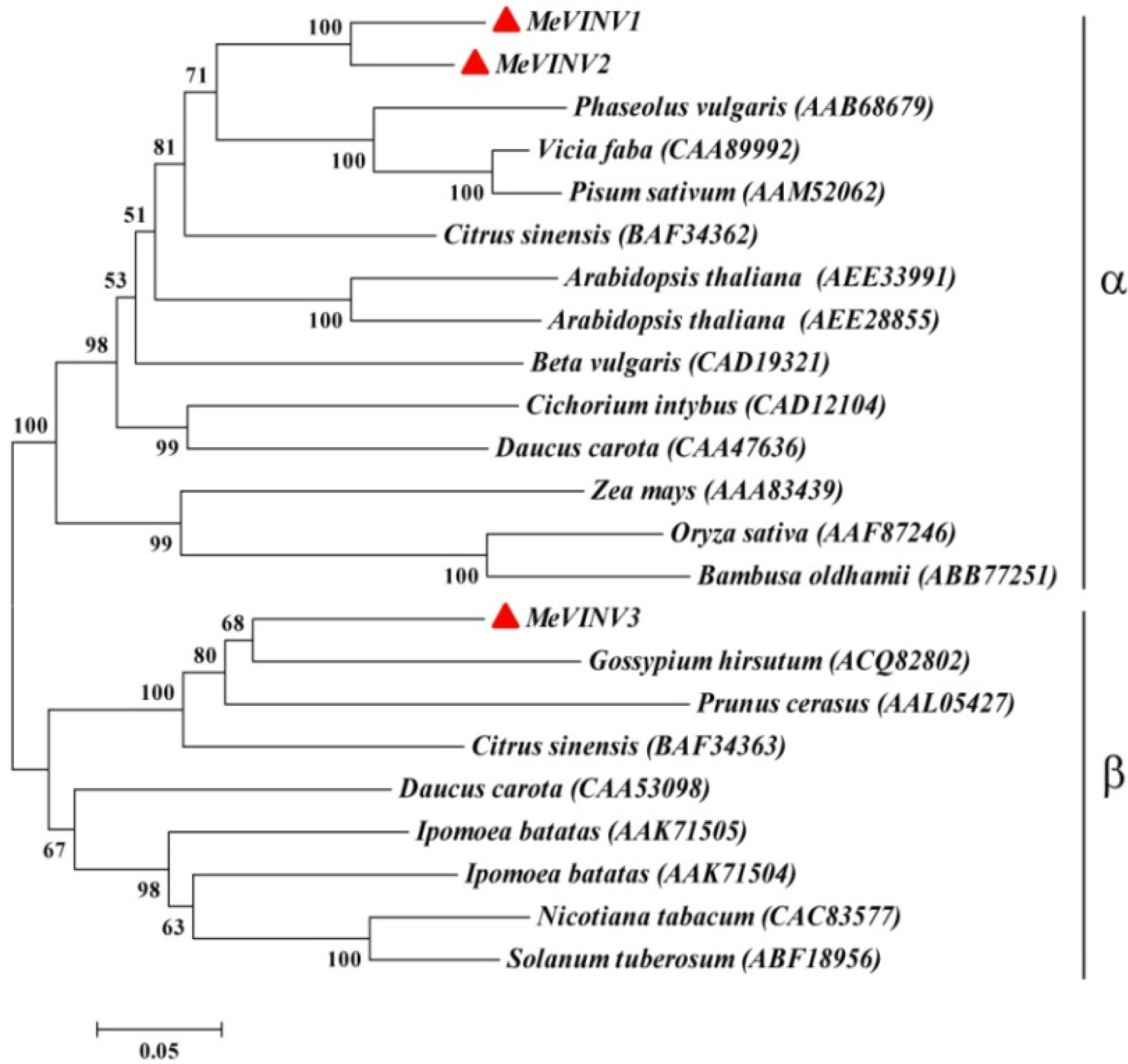
2.3. Phylogenetic Analysis of MeVINVs
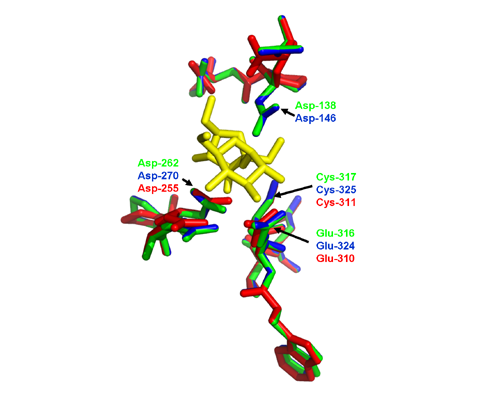
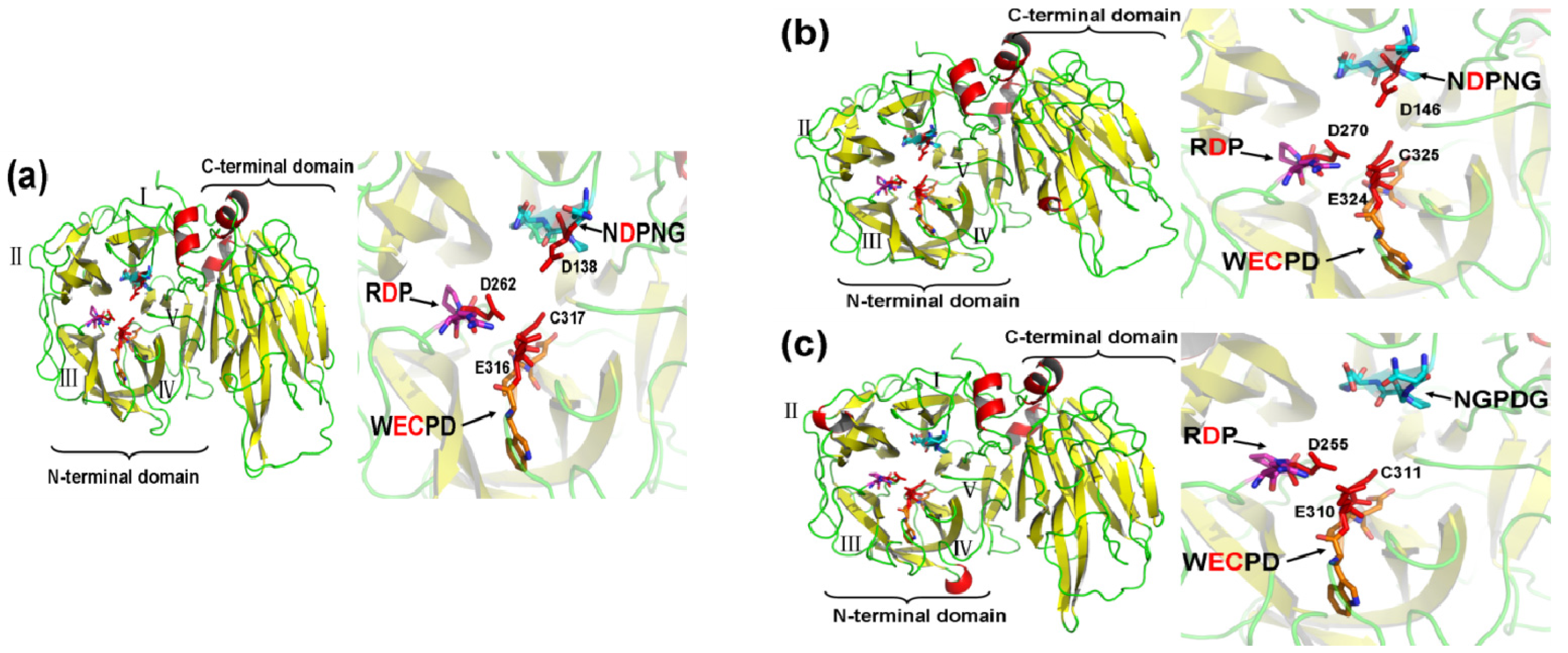
2.4. Three-Dimensional (3D) Structure of MeVINVs

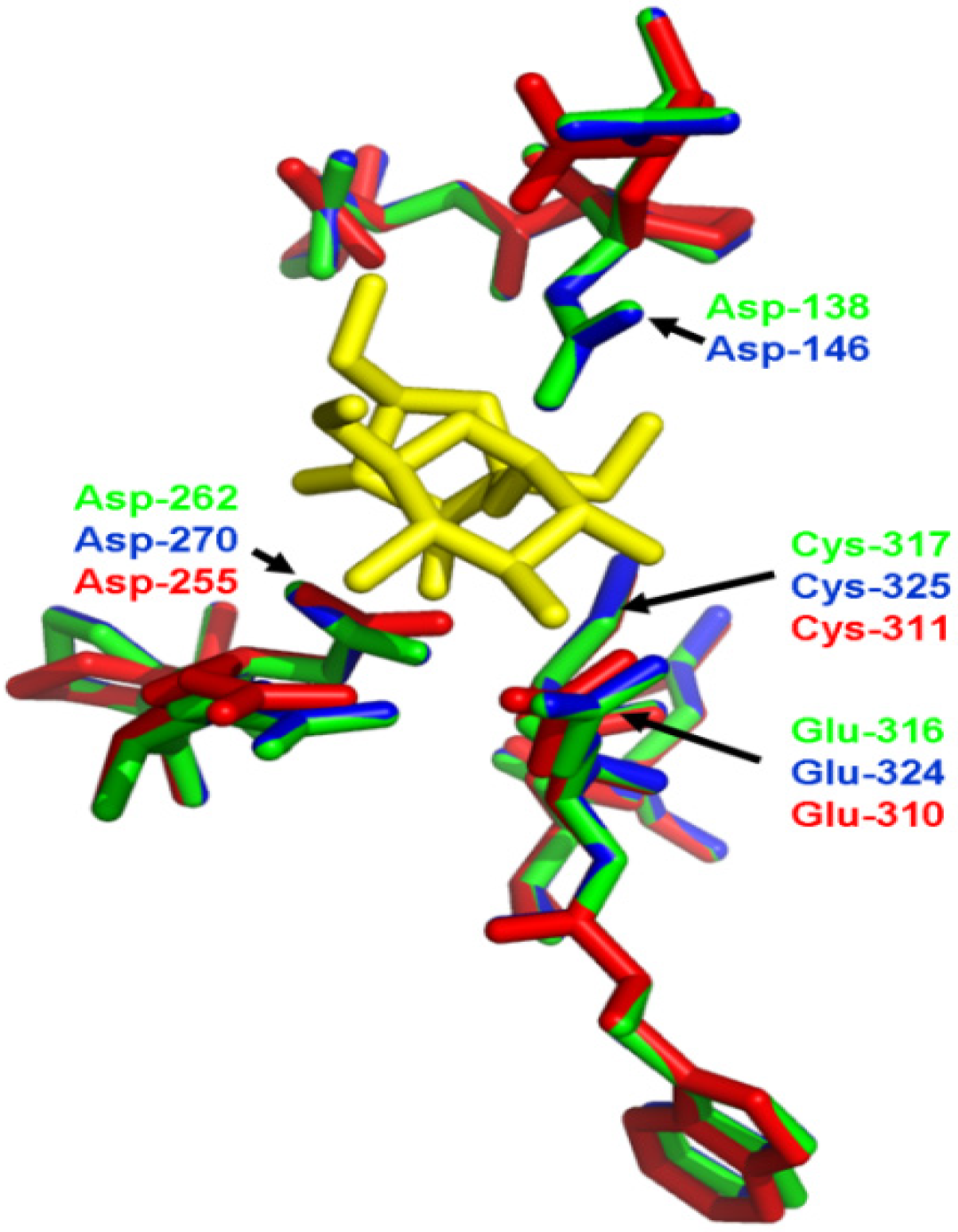

2.5. The Differential Expression Analysis of MeVINVs in Cassava Organs or Tissues

2.6. The Differential Expression of MeVINVs During Cassava Tuber Root Development
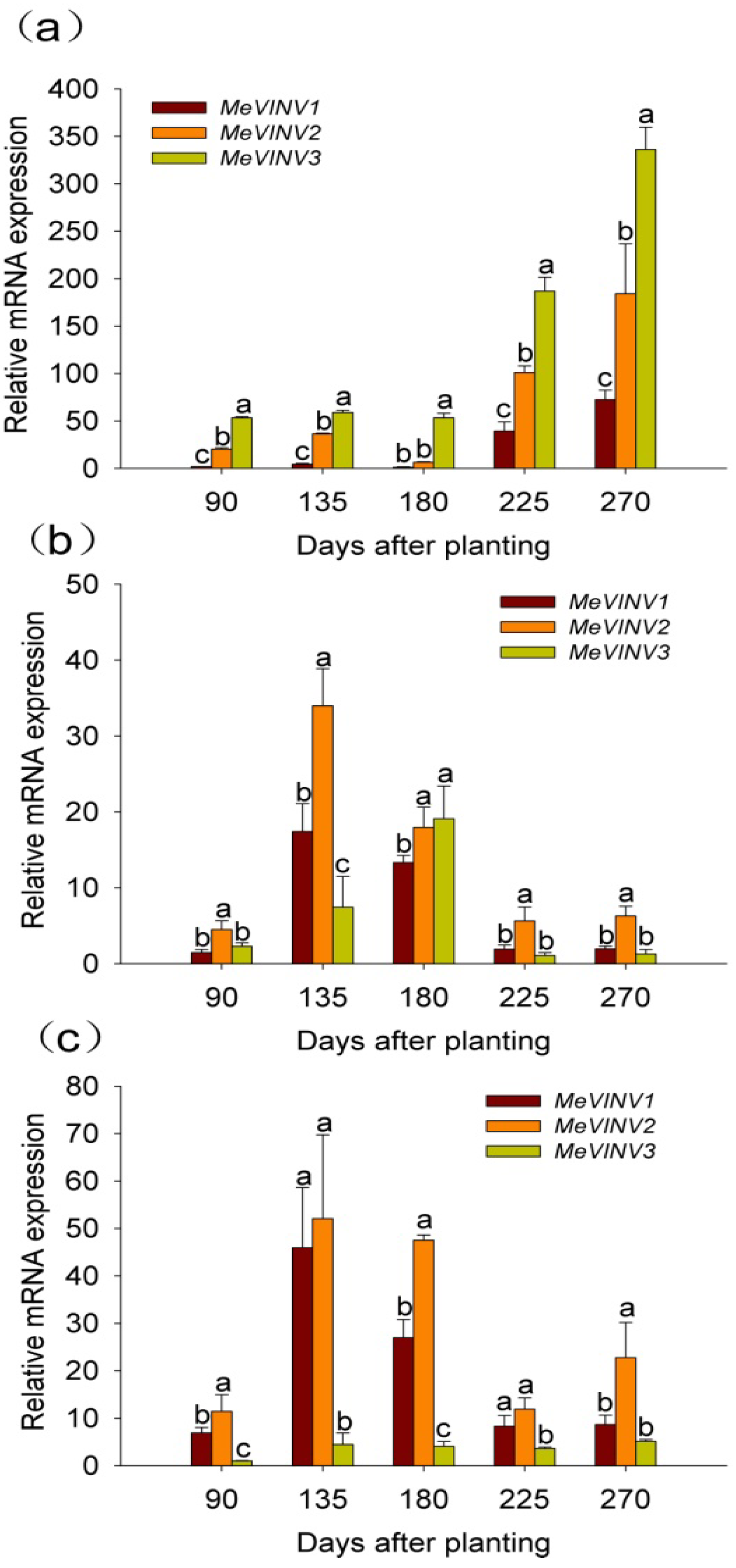
2.7. Discussion
3. Experimental
3.1. Plant Materials
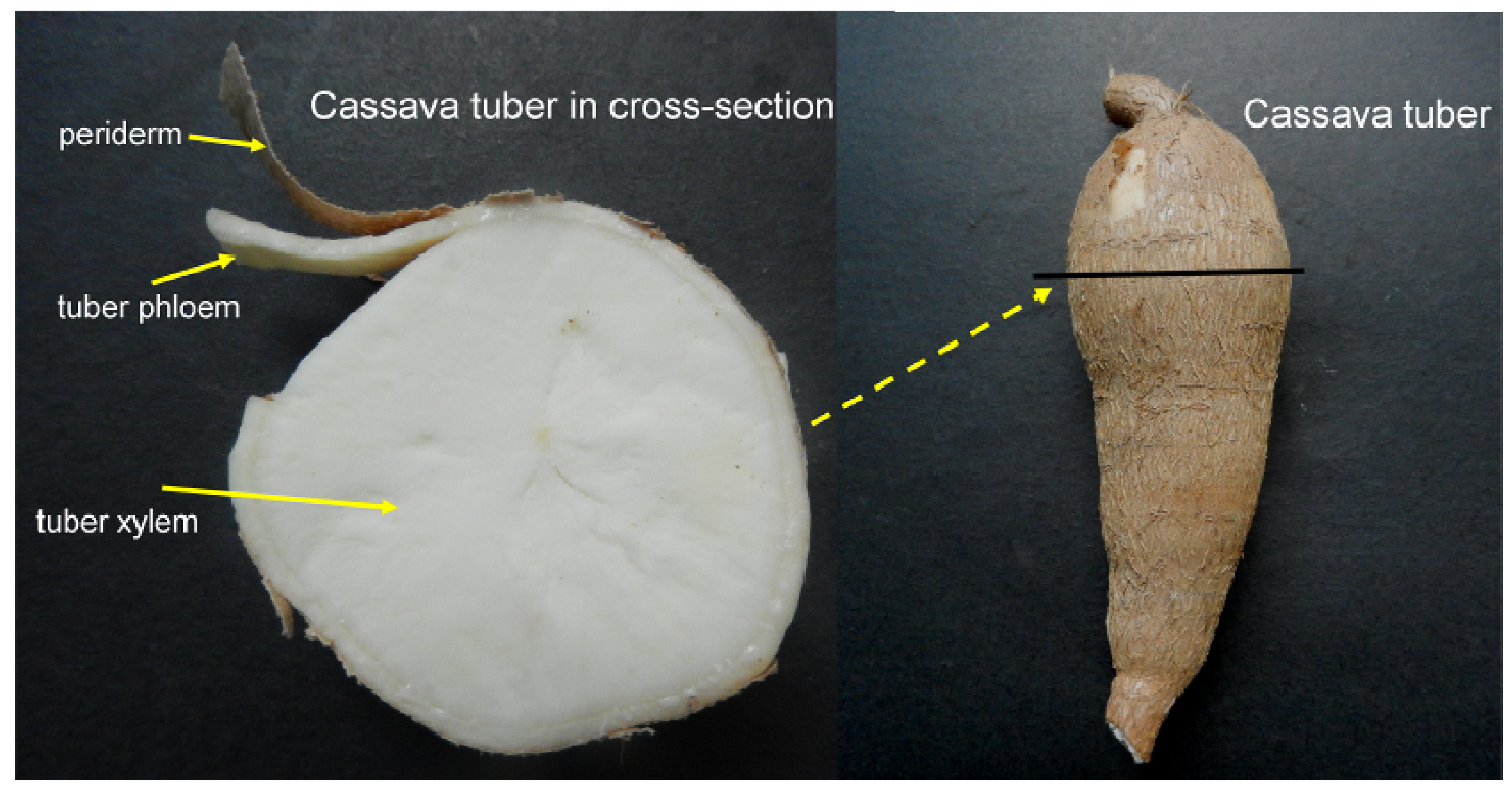
3.2. Molecular Cloning of MeVINVs
| Gene | Forward primer (5'–3') | Reverse primer (5'–3') |
|---|---|---|
| RT-PCR | ||
| MeCVINV1 | ATATCTAGAATGGCAGACCCCAACC | TATGTCGACAAAGATGAGTTTCACAGT |
| MeCVINV2 | ACCCTTCTTCCGTCCTTCTTC | AAATGAGGTAGACTTGGAAAGGTAAG |
| MeCVINV3 | TGAGGATCCCCAGCAAATACATGGACTAT | GCCGTCGACCAAAATAGGAGGTGTT |
| Real-time RT-PCR | ||
| MeCVINV1 | TTGAGACTAAGAGCCCGAGAAT | AGGACCAGAAGACCGAAGG |
| MeCVINV2 | CAGCCTGAGAAGAACTGGATG | GCCAGTCCATACACCATTTTG |
| MeCVINV3 | CACCTFTATTTTTCCGTCCTF | ATFCCCAACTTTCACCTTC |
3.3. Gene Bioinformatics Analysis
3.4. Real-time RT-PCR Analysis
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Hsieh, C.W.; Liu, L.K.; Yeh, S.H.; Chen, C.F.; Lin, H.I.; Sung, H.Y.; Wang, A.Y. Molecular cloning and functional identification of invertase isozymes from green bamboo Bmbusa oldhamii. J. Agric. Food Chem. 2006, 54, 3101–3107. [Google Scholar]
- Sturm, A. Invertases. Primary structures, functions, and roles in plant development and sucrose partitioning. Plant Physiol. 1999, 121, 1–7. [Google Scholar]
- Baroja-Fernández, E.; Muñoz, F.J.; Montero, M.; Etxeberria, E.; Sesma, M.T.; Ovecka, M.; Bahaji, A.; Ezquer, I.; Li, J.; Prat, S. Enhancing sucrose synthase activity in transgenic potato (Slanum tuberosum L) tubers results in increased levels of starch, adpglucose and udpglucose and total yield. Plant Cell Physiol. 2009, 50, 1651–1662. [Google Scholar]
- Sturm, A.; Tang, G.Q. The sucrose-cleaving enzymes of plants are crucial for development, growth and carbon partitioning. Trends Plant Sci. 1999, 4, 401–407. [Google Scholar]
- Flemetakis, E.; Efrose, R.C.; Ott, T.; Stedel, C.; Aivalakis, G.; Udvardi, M.K.; Katinakis, P. Satial and temporal organization of sucrose metabolism in lotus japonicus nitrogen-fixing nodules suggests a role for the elusive alkaline/neutral invertase. Plant Mol. Biol. 2006, 62, 53–69. [Google Scholar]
- Roitsch, T.; Ehneß, R.; Goetz, M.; Hause, B.; Hofmann, M.; Sinha, A.K. Regulation and function of extracellular invertase from higher plants in relation to assimilate partitioning, stress responses and sugar signalling. Funct. Plant Biol. 2000, 27, 815–825. [Google Scholar]
- Tymowska-Lalanne, Z.; Kreis, M. The plant invertases: Physiology, biochemistry and molecular biology. Adv. Bot. Res. 1998, 28, 71–117. [Google Scholar]
- Fotopoulos, V. Plant invertases: Structure, function and regulation of a diverse enzyme family. J. Biol. Res. 2005, 4, 127–137. [Google Scholar]
- Vargas, W.A.; Salerno, G.L. The cinderella story of sucrose hydrolysis: Alkaline/neutral invertases, from cyanobacteria to unforeseen roles in plant cytosol and organelles. Plant Sci. 2010, 178, 1–8. [Google Scholar]
- Bocock, P.; Morse, A.; Dervinis, C.; Davis, J. Evolution and diversity of invertase genes in populus trichocarpa. Planta 2008, 227, 565–576. [Google Scholar]
- Vargas, W.; Cumino, A.; Salerno, G.L. Cyanobacterial alkaline/neutral invertases. Origin of sucrose hydrolysis in the plant cytosol? Planta 2003, 216, 951–960. [Google Scholar]
- Tang, G.Q.; Luscher, M.; Sturm, A. Antisense repression of vacuolar and cell wall invertase in transgenic carrot alters early plant development and sucrose partitioning. Plant Cell Online 1999, 11, 177–189. [Google Scholar]
- Schaarschmidt, S.; Roitsch, T.; Hause, B. Arbuscular mycorrhiza induces gene expression of the apoplastic invertase lin6 in tomato (Lcopersicon esculentum) roots. J. Exp. Bot. 2006, 57, 4015–4023. [Google Scholar]
- Jain, M.; Chourey, P.S.; Boote, K.J.; Allen, L.H., Jr. Short-term high temperature growth conditions during vegetative-to-reproductive phase transition irreversibly compromise cell wall invertase-mediated sucrose catalysis and microspore meiosis in grain sorghum (Srghum bicolor). J. Plant Physiol. 2010, 167, 578–582. [Google Scholar]
- Chourey, P.; Jain, M.; Li, Q.B.; Carlson, S. Genetic control of cell wall invertases in developing endosperm of maize. Planta 2006, 223, 159–167. [Google Scholar]
- Yu, X.; Wang, X.; Zhang, W.; Qian, T.; Tang, G.; Guo, Y.; Zheng, C. Antisense suppression of an acid invertase gene (MAI1) in muskmelon alters plant growth and fruit development. J. Exp. Bot. 2008, 59, 2969–2977. [Google Scholar]
- Qi, X.; Wu, Z.; Li, J.; Mo, X.; Wu, S.; Chu, J.; Wu, P. Atcyt-inv1, a neutral invertase, is involved in osmotic stress-induced inhibition on lateral root growth in Arabidopsis. Plant Mol. Biol. 2007, 64, 575–587. [Google Scholar]
- Kim, J.-Y.; Mahé, A.; Brangeon, J.; Prioul, J.-L. A maize vacuolar invertase, IVR2, is induced by water stress. Organ/tissue specificity and diurnal modulation of expression. Plant Physiol. 2000, 124, 71–84. [Google Scholar]
- Roitsch, T.; González, M.C. Function and regulation of plant invertases: Sweet sensations. Trends Plant Sci. 2004, 9, 606–613. [Google Scholar]
- Lou, Y.; Gou, J.Y.; Xue, H.W. PIP5K9, an arabidopsis phosphatidylinositol monophosphate kinase, interacts with a cytosolic invertase to negatively regulate sugar-mediated root growth. Plant Cell Online 2007, 19, 163–181. [Google Scholar]
- Vargas, W.A.; Pontis, H.G.; Salerno, G.L. New insights on sucrose metabolism: Evidence for an active A/N-inv in chloroplasts uncovers a novel component of the intracellular carbon trafficking. Planta 2008, 227, 795–807. [Google Scholar]
- Bala, N. Strategies for elimination of cyanogens from cassava for reducing toxicity and improving food safety. Food Chem. Toxicol. 2011, 49, 690–693. [Google Scholar]
- Ceballos, H.; Iglesias, C.A.; Pérez, J.C.; Dixon, A.G. Cassava breeding: Opportunities and challenges. Plant Mol. Biol. 2004, 56, 503–516. [Google Scholar]
- Ihemere, U.; Arias-Garzon, D.; Lawrence, S.; Sayre, R. Genetic modification of cassava for enhanced starch production. Plant Biotechnol. J. 2006, 4, 453–465. [Google Scholar]
- Nagele, T.; Henkel, S.; Hormiller, I.; Sauter, T.; Sawodny, O.; Ederer, M.; Heyer, A.G. Mathematical modeling of the central carbohydrate metabolism in Arabidopsis reveals a substantial regulatory influence of vacuolar invertase on whole plant carbon metabolism. Plant Physiol. 2010, 153, 260–272. [Google Scholar]
- Huber, S.C. Biochemical mechanism for regulation of sucrose accumulation in leaves during photosynthesis. Plant Physiol. 1989, 91, 656–662. [Google Scholar]
- Paul, M.J.; Foyer, C.H. Sink regulation of photosynthesis. J. Exp. Bot. 2001, 52, 1383–1400. [Google Scholar]
- Xiang, L.; van den Ende, W. Trafficking of plant vacuolar invertases: From a membrane-anchored to a soluble status. Understanding sorting information in their complex n-terminal motifs. Plant Cell Physiol. 2013, 54, 1263–1277. [Google Scholar]
- Obenland, D.M.; Simmen, U.; Boller, T.; Wiemken, A. Purification and characterization of three soluble invertases from barley (Hordeum vulgare L.) leaves. Plant Physiol. 1993, 101, 1331–1339. [Google Scholar]
- Lammens, W.; le Roy, K.; Yuan, S.; Vergauwen, R.; Rabijns, A.; van Laere, A.; Strelkov, S.V.; van den Ende, W. Crystal structure of 6-SST/6-SFT from pachysandra terminalis, a plant fructan biosynthesizing enzyme in complex with its acceptor substrate 6-kestose. Plant J. 2012, 70, 205–219. [Google Scholar]
- Luo, X.; Chi, M.; Huang, X.; Xie, H.; Lu, F. Studies on the relationship between soluble sugar content in the leaves and the starch accumulation in the root tuber of cassava. Chin. Agric. Sci. Bull. 2006, 22, 289. (In Chinese) [Google Scholar]
- Wang, L.T.; Wang, A.Y.; Hsieh, C.W.; Chen, C.Y.; Sung, H.Y. Vacuolar invertases in sweet potato: Molecular cloning, characterization, and analysis of gene expression. J. Agric. Food Chem. 2005, 53, 3672–3678. [Google Scholar]
- Ji, X.; van den Ende, W.; van Laere, A.; Cheng, S.; Bennett, J. Structure, evolution, and expression of the two invertase gene families of rice. J. Mol. Evolut. 2005, 60, 615–634. [Google Scholar]
- Alberto, F.; Bignon, C.; Sulzenbacher, G.; Henrissat, B.; Czjzek, M. The three-dimensional structure of invertase (β-fructosidase) from Thermotoga maritima reveals a bimodular arrangement and an evolutionary relationship between retaining and inverting glycosidases. J. Biol. Chem. 2004, 279, 18903–18910. [Google Scholar]
- Chen, T.H.; Huang, Y.C.; Yang, C.S.; Yang, C.C.; Wang, A.Y.; Sung, H.Y. Insights into the catalytic properties of bamboo vacuolar invertase through mutational analysis of active site residues. Phytochemistry 2009, 70, 25–31. [Google Scholar]
- Bournay, A.S.; Hedley, P.E.; Maddison, A.; Waugh, R.; Machray, G.C. Exon skipping induced by cold stress in a potato invertase gene transcript. Nucleic Acids Res. 1996, 24, 2347–2351. [Google Scholar]
- Andersen, M.N.; Asch, F.; Wu, Y.; Jensen, C.R.; Næsted, H.; Mogensen, V.O.; Koch, K.E. Soluble invertase expression is an early target of drought stress during the critical, abortion-sensitive phase of young ovary development in maize. Plant Physiol. 2002, 130, 591–604. [Google Scholar]
- Elliott, K.J.; Butler, W.O.; Dickinson, C.D.; Konno, Y.; Vedvick, T.S.; Fitzmaurice, L.; Mirkov, T.E. Isolation and characterization of fruit vacuolar invertase genes from two tomato species and temporal differences in mrna levels during fruit ripening. Plant Mol. Biol. 1993, 21, 515–524. [Google Scholar]
- Li, Z.; Palmer, W.M.; Martin, A.P.; Wang, R.; Rainsford, F.; Jin, Y.; Patrick, J.W.; Yang, Y.; Ruan, Y.L. High invertase activity in tomato reproductive organs correlates with enhanced sucrose import into, and heat tolerance of, young fruit. J. Exp. Bot. 2012, 63, 1155–1166. [Google Scholar]
- Jansson, C.; Westerbergh, A.; Zhang, J.; Hu, X.; Sun, C. Cassava, a potential biofuel crop in (the) People’s Republic of China. Appl. Energy 2009, 86, S95–S99. [Google Scholar]
- Baguma, Y.; Sun, C.; Borén, M.; Olsson, H.; Rosenqvist, S.; Mutisya, J.; Rubaihayo, P.R.; Jansson, C. Sugar-mediated semidian oscillation of gene expression in the cassava storage root regulates starch synthesis. Plant Signal. Behav. 2008, 3, 439–445. [Google Scholar]
- Alves, A.A.C. Cassava botany and physiology. In Cassava: Biology, Production and Utilization; Hillocks, R.J., Thresh, J.M., Bellotti, A.C., Eds.; CABI Publishing: New York, NY, USA, 2002; Chapter 5; pp. 67–90. [Google Scholar]
- McKenzie, M.J.; Chen, R.K.Y.; Harris, J.C.; Ashworth, M.J.; Brummell, D.A. Post-translational regulation of acid invertase activity by vacuolar invertase inhibitor affects resistance to cold-induced sweetening of potato tubers. Plant Cell Environ. 2013, 36, 176–185. [Google Scholar]
- Liu, X.; Lin, Y.; Liu, J.; Song, B.; Ou, Y.; Zhang, H.; Li, M.; Xie, C. Stinvinh2 as an inhibitor of StvacINV1 regulates the cold-induced sweetening of potato tubers by specifically capping vacuolar invertase activity. Plant Biotechnol. J. 2013, 11, 640–647. [Google Scholar]
- Tauzin, A.S.; Sulzenbacher, G.; Lafond, M.; Desseaux, V.; Reca, I.B.; Perrier, J.; Bellincampi, D.; Fourquet, P.; Lévêque, C.; Giardina, T. Functional characterization of a vacuolar invertase from Solanum lycopersicum: Post-translational regulation by n-glycosylation and a proteinaceous inhibitor. Biochimie 2014, 101, 39–49. [Google Scholar]
- Geng, M.T.; Yao, Y.; Wu, X.H.; Min, Y.; Fu, S.P.; Hu, X.W.; Guo, J.C. Cloning and sequence analysis of two cDNA encoding invertase inhibitors from cassava (Manihot esculenta Crantz). Adv. Mater. Res. 2013, 726, 4326–4329. [Google Scholar]
- Phytozome. Available online: http://www.phytozome.net/cassava (accessed on 15 April 2011).
- TMHMM. Available online: http://www.cbs.dtu.dk/services/TMHMM (accessed on 10 September 2013).
- Gene Structure Display Server. Available online: http://gsds.cbi.pku.edu.cn/index.php (accessed on 25 April 2013).
- SWISS-MODEL. Available online: http://swissmodel.expasy.org (accessed on 15 January 2014).
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔ Ct method. Methods 2001, 25, 402–408. [Google Scholar]
- Narayanan, N.N.; Ihemere, U.; Ellery, C.; Sayre, R.T. Overexpression of hydroxynitrile lyase in cassava roots elevates protein and free amino acids while reducing residual cyanogen levels. PLoS One 2011, 6, e21996. [Google Scholar]
- Sample Availability: Not available.
© 2014 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Yao, Y.; Wu, X.-H.; Geng, M.-T.; Li, R.-M.; Liu, J.; Hu, X.-W.; Guo, J.-C. Cloning, 3D Modeling and Expression Analysis of Three Vacuolar Invertase Genes from Cassava (Manihot Esculenta Crantz). Molecules 2014, 19, 6228-6245. https://doi.org/10.3390/molecules19056228
Yao Y, Wu X-H, Geng M-T, Li R-M, Liu J, Hu X-W, Guo J-C. Cloning, 3D Modeling and Expression Analysis of Three Vacuolar Invertase Genes from Cassava (Manihot Esculenta Crantz). Molecules. 2014; 19(5):6228-6245. https://doi.org/10.3390/molecules19056228
Chicago/Turabian StyleYao, Yuan, Xiao-Hui Wu, Meng-Ting Geng, Rui-Mei Li, Jiao Liu, Xin-Wen Hu, and Jian-Chun Guo. 2014. "Cloning, 3D Modeling and Expression Analysis of Three Vacuolar Invertase Genes from Cassava (Manihot Esculenta Crantz)" Molecules 19, no. 5: 6228-6245. https://doi.org/10.3390/molecules19056228
APA StyleYao, Y., Wu, X.-H., Geng, M.-T., Li, R.-M., Liu, J., Hu, X.-W., & Guo, J.-C. (2014). Cloning, 3D Modeling and Expression Analysis of Three Vacuolar Invertase Genes from Cassava (Manihot Esculenta Crantz). Molecules, 19(5), 6228-6245. https://doi.org/10.3390/molecules19056228