Profiling of Concanavalin A-Binding Glycoproteins in Human Hepatic Stellate Cells Activated with Transforming Growth Factor-β1
Abstract
:1. Introduction
2. Results and Discussion
2.1. Isolation of Con A-Binding Glycoproteins (CBGs) from LX-2 Cells
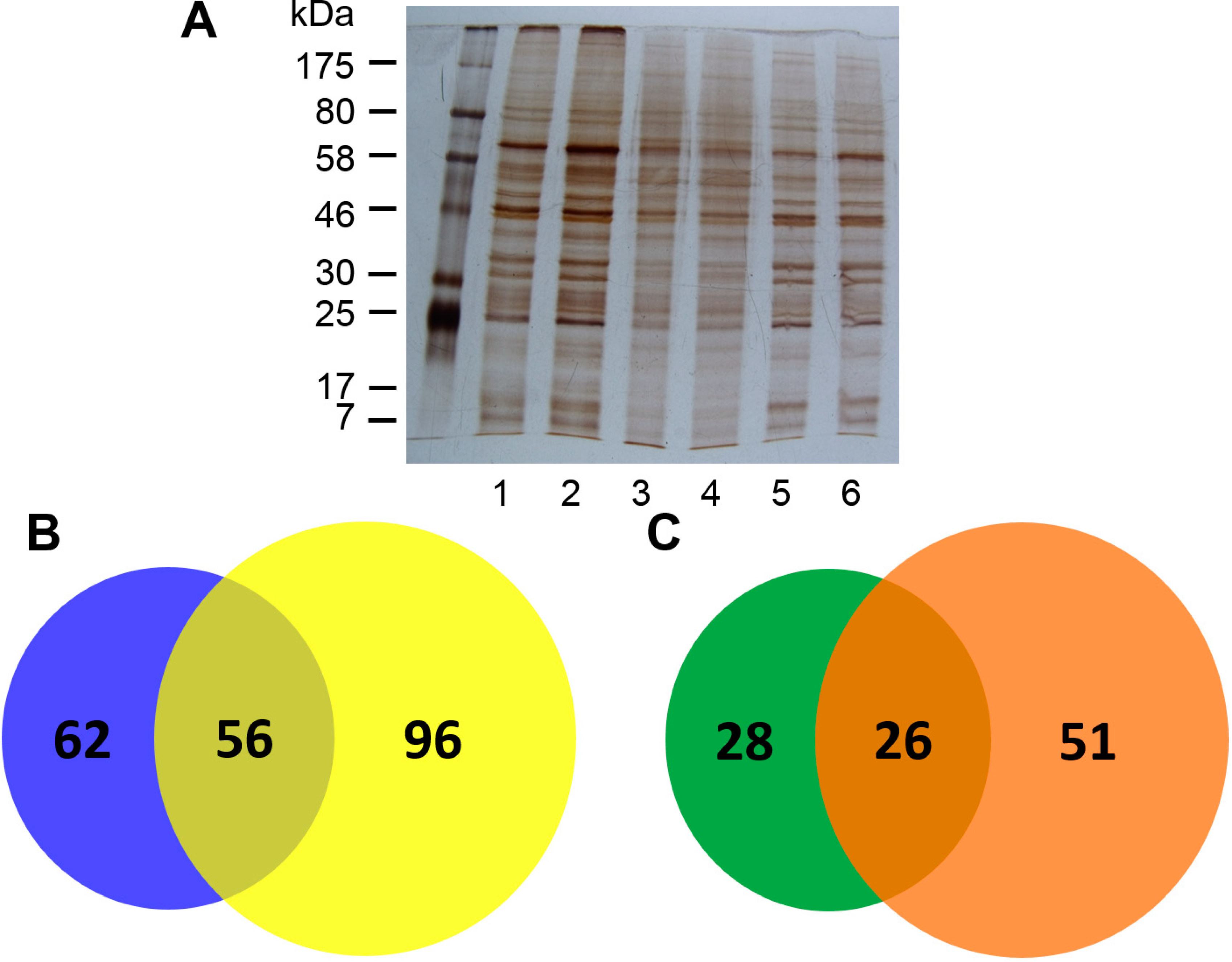
2.2. Identification of CBGs Isolated by CMPCs by LC-MS/MS
| No. | Accession | Description | Gene Name | Search Score | Coverage % | A/Q Ratio a | Q or A b | Known c |
|---|---|---|---|---|---|---|---|---|
| 1 | IPI00003865 | Isoform 1 of Heat shock cognate 71 kDa protein | HSPA8 | 375 | 26.8 | — | Q, A | PN |
| 2 | IPI00022434 | Putative uncharacterized protein ALB | ALB | 260 | 6.5 | — | Q, A | YN |
| 3 | IPI00021263 | 14-3-3 protein zeta/delta | YWHAZ | 164 | 15.1 | 0.16 | Q, A | PN |
| 4 | IPI00010796 | Protein disulfide-isomerase | P4HB | 109 | 17.5 | 0.5 | Q, A | PN |
| 5 | IPI00011134 | Putative heat shock 70 kDa protein 7 | HSPA7 | 96 | 6.5 | — | Q, A | PN |
| 6 | IPI00220327 | Keratin, type II cytoskeletal 1 | KRT1 | 92 | 3.4 | — | Q, A | PN |
| 7 | IPI00009865 | Keratin, type I cytoskeletal 10 | KRT10 | 75 | 14.6 | — | Q, A | PN |
| 8 | IPI00217966 | Isoform 1 of L-lactate dehydrogenase A chain | LDHA | 68 | 4.5 | — | Q, A | PN |
| 9 | IPI00021439 | Actin, cytoplasmic 1 | ACTB | 65 | 17.9 | — | Q, A | PN |
| 10 | IPI01969230 | LOC644914; H3F3B; LOC440926; H3F3A Histone H3;xx | Null | 60 | 8.9 | — | Q, A | Null |
| 11 | IPI00418471 | Vimentin | VIM | 56 | 2.1 | — | Q, A | Y° |
| 12 | IPI00020599 | Calreticulin | CALR | 53 | 10.8 | 2.44 | Q, A | YN |
| 13 | IPI00183968 | tropomyosin alpha-3 chain isoform 1 | TPM3 | 53 | 7.4 | — | Q, A | PN |
| 14 | IPI00218820 | Isoform 3 of Tropomyosin beta chain | TPM2 | 53 | 8.9 | — | Q, A | PN |
| 15 | IPI00293665 | Keratin, type II cytoskeletal 6B | KRT6B | 49 | 4.4 | — | Q, A | PN |
| 16 | IPI00453473 | Histone H4 | HIST1H4H | 36 | 29.1 | 3.13 | Q, A | P° |
| 17 | IPI00003269 | Beta-actin-like protein 2 | ACTBL2 | 32 | 6.6 | — | Q, A | PN |
| 18 | IPI00216783 | ubiquitin carboxyl-terminal hydrolase 2 isoform b | USP2 | 32 | 2.3 | — | Q, A | P° |
| 19 | IPI00219219 | Galectin-1 | LGALS1 | 29 | 8.9 | 2.8 | Q, A | PN |
| 20 | IPI00009636 | Membrane-spanning 4-domains subfamily A member 7 | MS4A7 | 28 | 7.1 | — | Q, A | N |
| 21 | IPI00375617 | Isoform 2 of Abhydrolase domain-containing protein 12B | ABHD12B | 26 | 2.1 | — | Q, A | P° |
| 22 | IPI00012048 | Isoform 1 of Nucleoside diphosphate kinase A | NME1-NME2 | 24 | 11.2 | 2.73 | Q, A | N |
| 23 | IPI00005685 | Paraneoplastic antigen Ma1 | PNMA1 | 22 | 3.7 | — | Q, A | PN |
| 24 | IPI00022774 | Transitional endoplasmic reticulum ATPase | VCP | 22 | 3 | — | Q, A | PN |
| 25 | IPI00470859 | Putative uncharacterized protein DKFZp686C04126 | MAN1B1 | 22 | 0.8 | — | Q, A | P° |
| 26 | IPI00888712 | actin, beta-like 3 | POTEE | 20 | 2.6 | — | Q, A | PN |
| 27 | IPI00000816 | 14-3-3 protein epsilon | YWHAE | 116 | 14.5 | 0.01 | Q | PN |
| 28 | IPI00385250 | Protease serine 4 isoform B | PRSS3 | 75 | 9.6 | 0.01 | Q | PN |
| 29 | IPI00013508 | Alpha-actinin-1 | ACTN1 | 61 | 1.3 | 0.01 | Q | PN |
| 30 | IPI00451401 | Triosephosphate isomerase | TPI1 | 17 | 7.4 | 0.01 | Q | PN |
| 31 | IPI00220642 | 14-3-3 protein gamma | YWHAG | 58 | 15 | 0.01 | Q | PN |
| 32 | IPI00465028 | TPI1 triosephosphate isomerase isoform 2 | TPI1P1 | 56 | 18.9 | 0.01 | Q | PN |
| 33 | IPI00479722 | Proteasome activator complex subunit 1 | PSME1 | 56 | 4.4 | 0.01 | Q | PN |
| 34 | IPI00453476 | 29 kDa protein | Null | 37 | 15.7 | 0.01 | Q | Null |
| 35 | IPI00021304 | Keratin, type II cytoskeletal 2 epidermal | KRT2 | 33 | 4.5 | 0.01 | Q | PN |
| 36 | IPI00008527 | 60S acidic ribosomal protein P1 | RPLP1 | 32 | 14 | 0.01 | Q | PN |
| 37 | IPI00021428 | Actin, alpha skeletal muscle | ACTA1 | 26 | 15.6 | 0.01 | Q | PN |
| 38 | IPI00479743 | Isoform 1 of POTE ankyrin domain family member E | POTEKP | 26 | 5.8 | 0.01 | Q | PN |
| 39 | IPI00009791 | Isoform WB of Plasma membrane calcium-transporting ATPase 2 | ATP2B2 | 25 | 0.7 | 0.01 | Q | PN |
| 40 | IPI00014845 | Isoform 1 of Dynein heavy chain 8, axonemal | DNAH8 | 25 | 0.9 | 0.01 | Q | PN |
| 41 | IPI00886949 | LOC100129520, similar to hCG2044193 | Null | 25 | 4.6 | 0.01 | Q | Null |
| 42 | IPI00002850 | Hepatic leukemia factor | HLF | 24 | 4.1 | 0.01 | Q | PN |
| 43 | IPI00550020 | Parathymosin | PTMS | 24 | 10.8 | 0.01 | Q | PN |
| 44 | IPI00064885 | Zinc finger protein 3 homolog | ZFP3 | 23 | 1.8 | 0.01 | Q | PN |
| 45 | IPI00925740 | ECT2 Protein | ECT2 | 20 | 47.8 | 0.01 | Q | PN |
| 46 | IPI00002349 | Nuclear fragile X mental retardation-interacting protein 2 | NUFIP2 | 19 | 2.6 | 0.01 | Q | PN |
| 47 | IPI00218667 | Stathmin-2 | STMN2 | 19 | 5 | 0.01 | Q | PN |
| 48 | IPI00018146 | 14-3-3 protein theta | YWHAQ | 18 | 9.8 | 0.01 | Q | PN |
| 49 | IPI00156689 | Synaptic vesicle membrane protein VAT-1 homolog | VAT1 | 18 | 6.4 | 0.01 | Q | PN |
| 50 | IPI00457114 | Isoform 1 of IQ motif and SEC7 domain-containing protein 1 | IQSEC1 | 18 | 3.7 | 0.01 | Q | PN |
| 51 | IPI00797750 | 11 kDa protein | Null | 17 | 16.7 | 0.01 | Q | Null |
| 52 | IPI00938247 | LOC100287408,hypothetical protein XP_002344194 | Null | 17 | 8.8 | 0.01 | Q | Null |
| 53 | IPI00966637 | Putative uncharacterized protein SDAD1 | SDAD1 | 17 | 23.9 | 0.01 | Q | PN |
| 54 | IPI00937642 | LOC100290701 hypothetical protein XP_002347764 | Null | 15 | 5.6 | 0.01 | Q | Null |
| 55 | IPI00304925 | Heat shock 70 kDa protein 1A/1B | HSPA1A; HSPA1B | 85 | 9.2 | 100 | A | PN |
| 56 | IPI00830052 | 62 kDa protein | Null | 85 | 10.5 | 100 | A | Null |
| 57 | IPI00015614 | Isoform A of Trypsin-3 | PRSS3 | 78 | 6.6 | 100 | A | P° |
| 58 | IPI00337495 | Isoform 2 of Procollagen-lysine,2-oxoglutarate 5-dioxygenase 2 | PLOD2 | 73 | 1.8 | 100 | A | YN |
| 59 | IPI00739539 | POTE ankyrin domain family member F | POTEKP | 63 | 3.6 | 100 | A | PN |
| 60 | IPI00000230 | tropomyosin alpha-1 chain isoform 2 | TPM1 | 59 | 7.7 | 100 | A | P° |
| 61 | IPI00013991 | Isoform 1 of Tropomyosin beta chain | TPM2 | 59 | 8.8 | 100 | A | P° |
| 62 | IPI00026260 | Isoform 1 of Nucleoside diphosphate kinase B | NME1-NME2 | 59 | 25 | 100 | A | N |
| 63 | IPI00879437 | Protein disulfide-isomerase A1 | P4HB | 54 | 16.5 | 100 | A | P° |
| 64 | IPI00937995 | Actin-like protein (Fragment) | ACTB | 50 | 19.4 | 100 | A | P° |
| 65 | IPI00024320 | Putative RNA-binding protein 3 | RBM3 | 40 | 20.4 | 100 | A | P° |
| 66 | IPI00171611 | Histone H3.2 | HIST2H3A | 40 | 11.8 | 100 | A | PN |
| 67 | IPI00798360 | SARNP 18 kDa protein | SARNP | 40 | 4.3 | 100 | A | PN |
| 68 | IPI00878173 | cDNA FLJ39583 fis, clone SKMUS2004897, highly similar to ACTIN, ALPHA SKELETAL MUSCLE | ACTA1 | 38 | 15.2 | 100 | A | P° |
| 69 | IPI00045396 | Calumenin isoform 4 | CALU | 34 | 2.2 | 100 | A | YN |
| 70 | IPI00016768 | L-lactate dehydrogenase A-like 6B | LDHAL6B | 31 | 4.5 | 100 | A | PN |
| 71 | IPI00784327 | Isoform 1 of 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase beta-2 | PLCB2 | 30 | 1.9 | 100 | A | PN |
| 72 | IPI00000874 | Peroxiredoxin-1 | PRDX1 | 29 | 10.1 | 100 | A | PN |
| 73 | IPI00216817 | HEAT repeat containing 7B1 | HEATR7B1 | 29 | 1.3 | 100 | A | PN |
| 74 | IPI00293975 | 16 kDa protein | Null | 29 | 6.2 | 100 | A | Null |
| 75 | IPI00298622 | Intestinal-type alkaline phosphatase | ALPI | 29 | 2.1 | 100 | A | YN |
| 76 | IPI00945706 | BOC Protein | BOC | 29 | 14.3 | 100 | A | PN |
| 77 | IPI00329801 | Annexin A5 | ANXA5 | 26 | 2.8 | 100 | A | P° |
| 78 | IPI00014424 | Elongation factor 1-alpha 2 | EEF1A2 | 25 | 3.9 | 100 | A | PN |
| 79 | IPI00030282 | Isoform 1 of Filensin | BFSP1 | 25 | 3.9 | 100 | A | P° |
| 80 | IPI00910689 | Polyserase-2 | PRSS36 | 25 | 3.2 | 100 | A | YN |
| 81 | IPI00023673 | Galectin-3-binding protein | LGALS3BP | 24 | 3.8 | 100 | A | YN |
| 82 | IPI00414717 | Isoform 2 of Golgi apparatus protein 1 | GLG1 | 24 | 2.1 | 100 | A | YN |
| 83 | IPI00007249 | Ectonucleotide pyrophosphatase/phosphodiesterase family member 4 | ENPP4 | 23 | 5.7 | 100 | A | YN |
| 84 | IPI00008692 | Isoform 1 of Keratin, type I cuticular Ha6 | KRT36 | 23 | 4.9 | 100 | A | PN |
| 85 | IPI00290077 | Keratin, type I cytoskeletal 15 | KRT15 | 23 | 4.4 | 100 | A | PN |
| 86 | IPI00384497 | Protein-tyrosine phosphatase-like member B | PTPLB | 23 | 7.1 | 100 | A | YN |
| 87 | IPI00908888 | cDNA FLJ57836, highly similar to Myb-binding protein 1A | MYBBP1A | 23 | 2.2 | 100 | A | PN |
| 88 | IPI00024658 | OTU domain-containing protein 7B | OTUD7B | 22 | 2.5 | 100 | A | P° |
| 89 | IPI00384972 | Isoform 1 of MLL1/MLL complex subunit KIAA1267 | KIAA1267 | 22 | 1.5 | 100 | A | PN |
| 90 | IPI00394814 | Serine protease 55 | PRSS55 | 22 | 4.8 | 100 | A | YN |
| 91 | IPI00216704 | Isoform 2 of Spectrin beta chain, erythrocyte | SPTB I | 21 | 1.7 | 100 | A | PN |
| 92 | IPI00455675 | Centrosomal protein of 192 kDa | CEP192 | 20 | 1.1 | 100 | A | PN |
| 93 | IPI00552749 | DNAH8 478 kDa protein | DNAH8 | 20 | 1.7 | 100 | A | PN |
| 94 | IPI00748715 | SEPT9 protein (Fragment) | Null | 19 | 3.5 | 100 | A | Null |
| 95 | IPI00020035 | Protein NipSnap homolog 3B | NIPSNAP3B | 18 | 12.6 | 100 | A | P° |
| 96 | IPI00217899 | E3 ubiquitin-protein ligase RNF168 | RNF168 | 18 | 1.9 | 100 | A | PN |
| 97 | IPI00010289 | D(1A) dopamine receptor | DRD1 | 17 | 4 | 100 | A | YN |
| 98 | IPI00478586 | Isoform 2 of Vacuolar protein sorting-associated protein 13A | VPS13A | 17 | 1.9 | 100 | A | PN |
| 99 | IPI00929137 | Conserved hypothetical protein | Null | 17 | 9.9 | 100 | A | Null |
| 100 | IPI00011500 | Isoform 1 of Testicular acid phosphatase | ACPT | 16 | 2.8 | 100 | A | YN |
| 101 | IPI00299507 | Condensin complex subunit 2 | NCAPH | 16 | 0.9 | 100 | A | PN |
| 102 | IPI00009101 | Isoform 2 of Helicase SRCAP | SRCAP | 15 | 1 | 100 | A | PN |
| 103 | IPI00442299 | Isoform 1 of Neurexin-1-alpha | NRXN1 | 15 | 3.3 | 100 | A | YN |
| 104 | IPI00947233 | PCOLCE2 Protein | PCOLCE2 | 14 | 42.6 | 100 | A | PN |
| 105 | IPI00294052 | Single-stranded DNA-binding protein 2 | SSBP2 | 13 | 3.6 | 100 | A | PN |
2.3. GO Classification of the Identified CBGs Using Blast2GO®
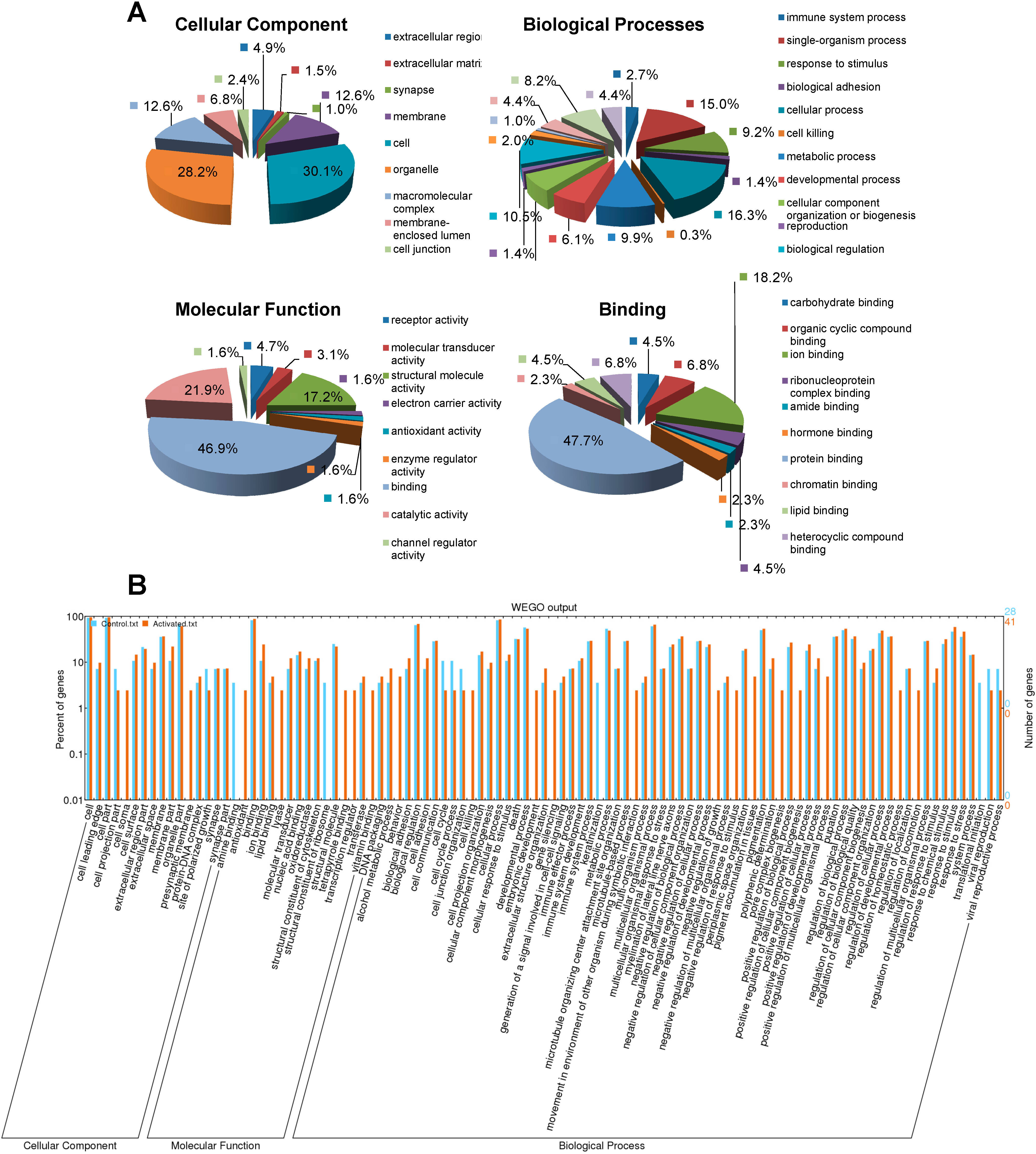
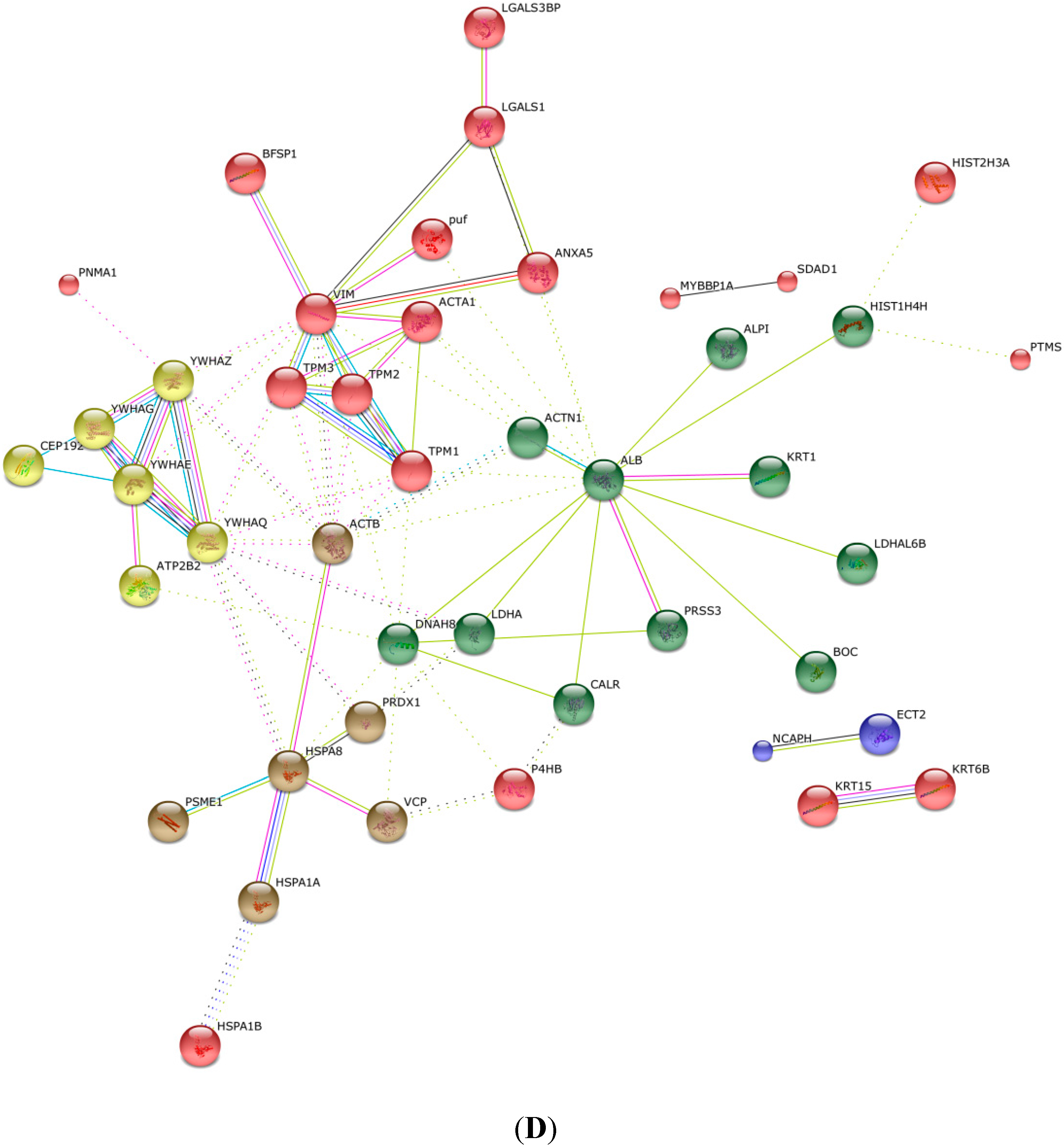
2.4. KEGG Pathway Analysis and Functional Protein Association Networks
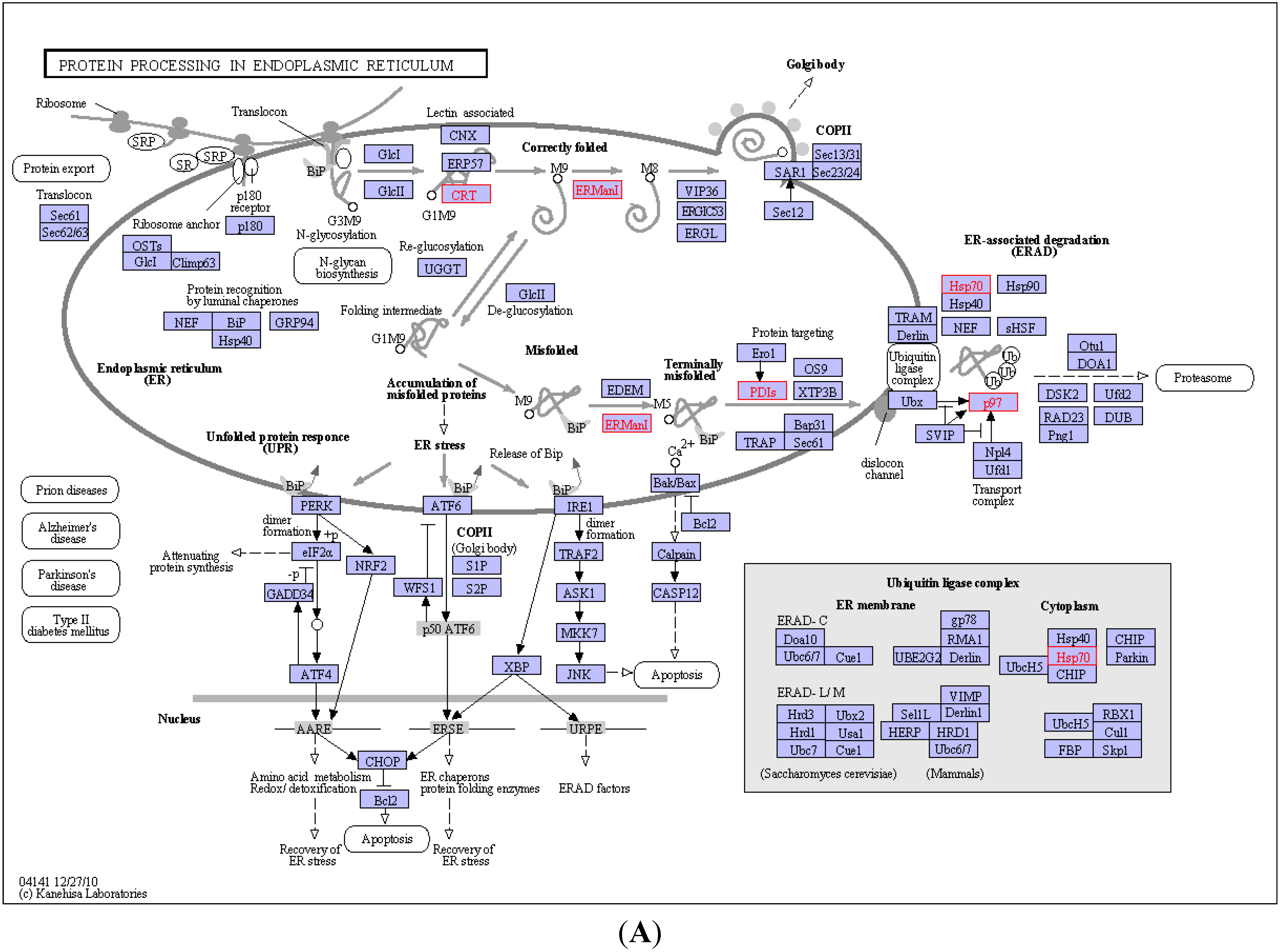
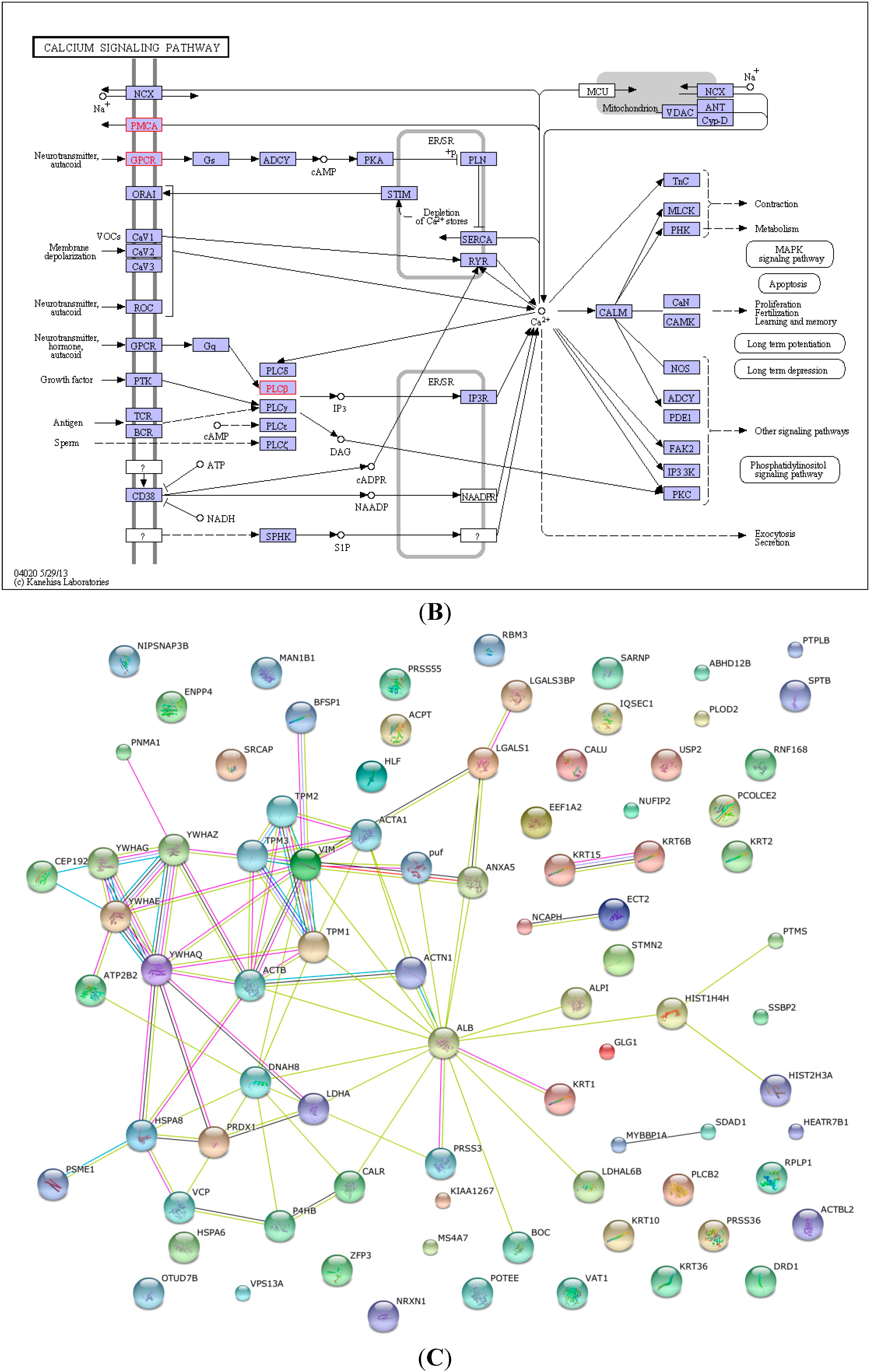
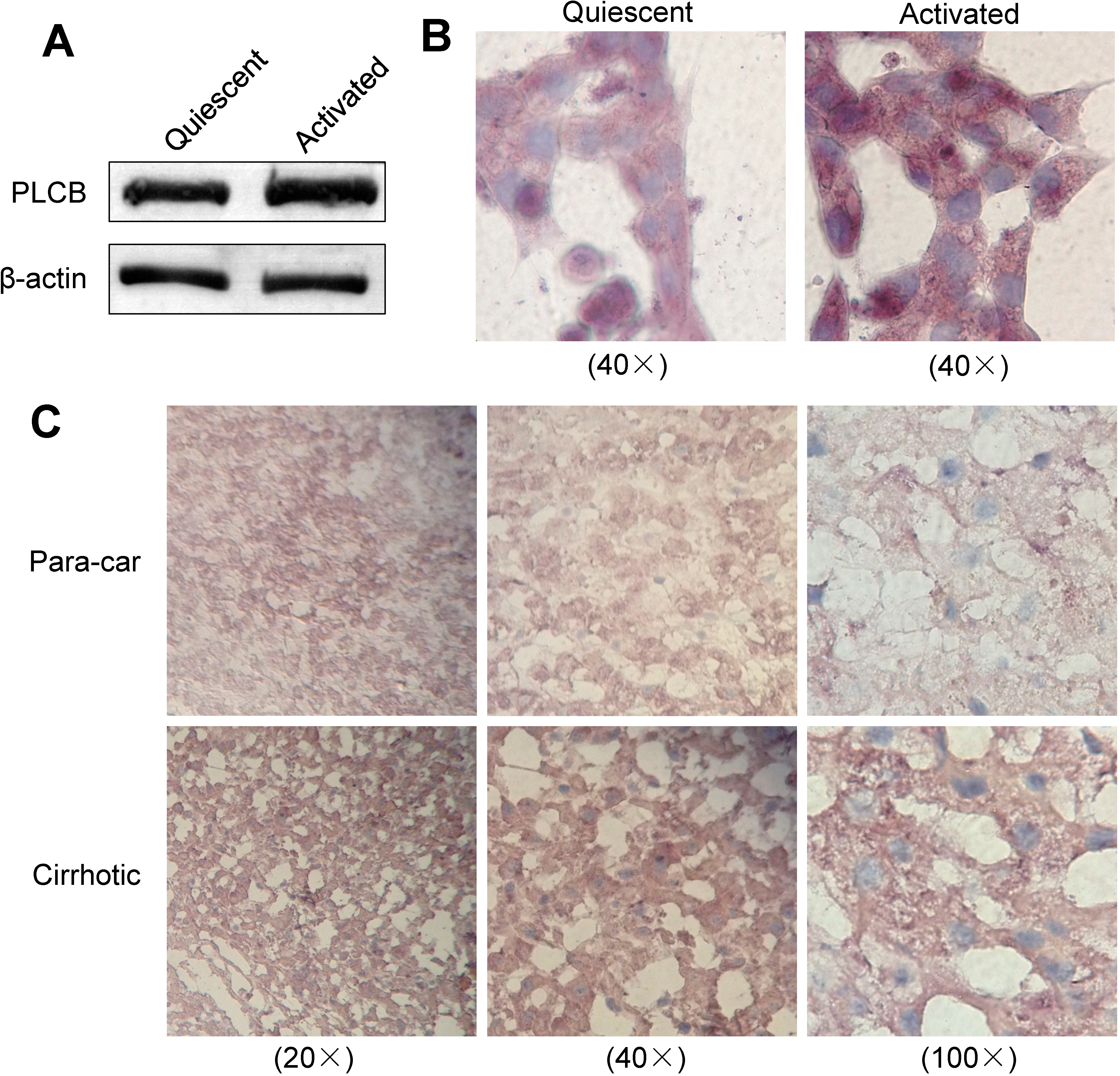
2.5. Expression of PLCB2 in Activated LX-2 Cells and Human Liver Cirrhosis Tissue
3. Experimental Section
3.1. Materials
3.2. Cell Culture and Stimulation
3.3. Non-Denaturing Extraction of Total Proteins
3.4. Isolation of Glycoproteins from LX-2 Cells Using ConA-Magnetic Particle Conjugates (CMPCs)
3.5. Identification of Peptides by LC-MS/MS
3.6. Label-Free Relative Quantification of Identified Glycoproteins by emPAI
3.7. Data Mining and Bioinformatics
3.8. SDS-PAGE and Western Blotting
3.9. Immunohistochemistry
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Muhanna, N.; Doron, S.; Wald, O.; Horani, A.; Eid, A.; Pappo, O.; Friedman, S.L.; Safadi, R. Activation of hepatic stellate cells after phagocytosis of lymphocytes: A novel pathway of fibrogenesis. Hepatology 2008, 48, 963–977. [Google Scholar] [CrossRef] [PubMed]
- Troeger, J.S.; Mederacke, I.; Gwak, G.Y.; Dapito, D.H.; Mu, X.; Hsu, C.C.; Pradere, J.P.; Friedman, R.A.; Schwabe, R.F. Deactivation of hepatic stellate cells during liver fibrosis resolution in mice. Gastroenterology 2012, 143, 1073–1083. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Zhang, L.; Yin, Z.; Su, W.; Ren, G.; Zhou, C.; You, J.; Fan, J.; Wang, X. Activated hepatic stellate cells promote hepatocellular carcinoma development in immunocompetent mice. Int. J. Cancer 2011, 129, 2651–2661. [Google Scholar] [CrossRef] [PubMed]
- Inagaki, Y.; Okazaki, I. Emerging insights into transforming growth factor beta Smad signal in hepatic fibrogenesis. Gut 2007, 56, 284–292. [Google Scholar] [CrossRef] [PubMed]
- Friedman, S.L. Mechanisms of hepatic fibrogenesis. Gastroenterology 2008, 134, 1655–1669. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Trevijano, E.R.; Iraburu, M.J.; Fontana, L. Transforming growth factor beta1 induces the expression of alpha1 (I) procollagen mRNA by a hydrogen peroxide-C/EBPbeta-dependent mechanism in rat hepatic stellate cells. Hepatology 1999, 29, 960–970. [Google Scholar] [CrossRef] [PubMed]
- Varki, A. Essentials of Glycobiology; Cold Spring Harbor Laboratory Press: New York, NY, USA, 2009. [Google Scholar]
- Gerald, W.H.; Ronald, J. Glycomics hits the big time. Cell 2010, 143, 672–676. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Mallorya, T.; Satomura, S. AFP-L3: A new generation of tumor marker for hepatocellular carcinoma. Clin. Chim. Acta 2001, 313, 15–19. [Google Scholar] [CrossRef] [PubMed]
- Lee, L.Y.; Thaysen-Andersen, M.; Baker, M.S.; Packer, N.H.; Hancock, W.S.; Fanayan, S. Comprehensive N-glycome profiling of cultured human epithelial breast cells identifies unique secretome N-glycosylation signatures enabling tumorigenic subtype classification. J. Proteome Res. 2014, in press. [Google Scholar]
- Goto, Y.; Obata, T.; Kunisawa, J.; Sato, S.; Ivanov, I.; Lamichhane, A.; Takeyama, N. Innate lymphoid cells regulate intestinal epithelial cell glycosylation. Science 2014, 345, 1254009. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.X.; Weng, Z.H.; Zhang, S.L. Notch3 regulates the activation of hepatic stellate cells. World J. Gastroenterol. 2012, 18, 1397–1403. [Google Scholar] [CrossRef] [PubMed]
- Moloney, D.J.; Panin, V.M.; Johnston, S.H.; Chen, J.; Shao, L.; Wilson, R.; Wang, Y.; Stanley, P.; Irvine, K.D.; Haltiwanger, R.S.; et al. Fringe is a glycosyltransferase that modifies Notch. Nature 2000, 406, 369–375. [Google Scholar] [CrossRef] [PubMed]
- Muchmore, A.V.; Decker, J.M. The lectin-like interaction between recombinant tumor necrosis factor and uromodulin. J. Immunol. 1987, 138, 2541–2546. [Google Scholar] [PubMed]
- Cebo, C.; Dambrouck, E.; Maes, C.; Laden, G.; Strecker, J.C.; Michalski, J.P.; Zanetta, J.P. Recombinant human interleukins IL-1alpha, IL-1beta, IL-4, IL-6, and IL-7 show different and specific calcium-independent carbohydrate-binding properties. J. Biol. Chem. 2001, 276, 5685–5691. [Google Scholar] [CrossRef] [PubMed]
- Hellerbrand, C.; Jobin, C.; Licato, L.L.; Sartor, R.B.; Brenner, D.A. Cytokines induce NF-kappaB in activated but not in quiescent rat hepatic stellate cells. Am. J. Physiol. 1998, 275, G269–G278. [Google Scholar] [PubMed]
- Spratte, J.; Meyer, Z.S.H.; Endlich, N.; Zygmunt, M.; Fluhr, H. Heparin inhibits TNF-α signaling in human endometrial stromal cells by interaction with NF-κB. Mol. Hum. Reprod. 2013, 19, 227–236. [Google Scholar] [CrossRef] [PubMed]
- Qin, Y.; Zhong, Y.; Dang, L.; Zhu, M.; Yu, H.; Chen, W.; Cui, J.; Bian, H.; Li, Z. Alteration of protein glycosylation in human hepatic stellate cells activated with transforming growth factor-β1. J. Proteomics 2012, 75, 4114–4123. [Google Scholar] [CrossRef] [PubMed]
- Spicer, S.S.; Schulte, B.A. Diversity of cell glycoconjugates shown histochemically: A perspective. J. Histochem. Cytochem. 1992, 40, 1–38. [Google Scholar] [CrossRef] [PubMed]
- Yue, T.; Goldstein, I.J.; Hollingsworth, M.A.; Kaul, K.; Brand, R.E.; Haab, B.B. The prevalence and nature of glycan alterations on specific proteins in pancreatic cancer patients revealed using antibody-lectin Sandwich arrays. Mol. Cell Proteomics 2009, 8, 1697–1707. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Li, X.J.; Martin, D.B.; Aebersold, R. Identification and quantification of N-linked glycoproteins using hydrazide chemistry, stable isotope labeling and mass spectrometry. Nat. Biotechnol. 2003, 21, 660–666. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.A.; Zhou, Y.; Elliott, S.; Aebersold, R.; Zhang, H. Solid-phase extraction of N-linked glycopeptides. Nat. Protoc. 2007, 2, 334–339. [Google Scholar] [CrossRef] [PubMed]
- Kaji, H.; Saito, H.; Yamauchi, Y.; Shinkawa, T.; Taoka, M.; Hirabayashi, J. Lectin affinity capture, isotope-coded tagging and mass spectrometry to identify N-linked glycoproteins. Nat. Biotechnol. 2003, 21, 667–672. [Google Scholar] [CrossRef] [PubMed]
- Kaji, H.; Yamauchi, Y.; Takahashi, N.; Isobe, T. Mass spectrometric identification of N-linked glycopeptides using lectin-mediated affinity capture and glycosylation site-specific stable isotope tagging. Nat. Protoc. 2007, 1, 3019–3027. [Google Scholar] [CrossRef]
- Sturiale, L.; Barone, R.; Palmigiano, A.; Ndosimao, C.N.; Briones, P.; Adamowicz, M. Multiplexed glycoproteomic analysis of glycosylation disorders by sequential yolk immunoglobulins immunoseparation and MALDI-TOF MS. Proteomics 2008, 8, 3822–3832. [Google Scholar] [CrossRef] [PubMed]
- Cho, W.; Jung, K.; Regnier, F.E. Use of glycan targeting antibodies to identify cancer-associated glycoproteins in plasma of breast cancer patients. Anal. Chem. 2008, 80, 5286–5292. [Google Scholar] [CrossRef] [PubMed]
- Sparbier, K.; Asperger, A.; Resemann, A.; Kessler, I.; Koch, S.; Wenzel, T. Analysis of glycoproteins in human serum by means of glycospecific magnetic bead separation and LC-MALDI-TOF/TOF analysis with automated glycopeptide detection. J. Biomol. Tech. 2007, 18, 252–258. [Google Scholar] [PubMed]
- Xu, Y.; Wu, Z.; Zhang, L.; Lu, H.; Yang, P.; Webley, P.A. Highly specific enrichment of glycopeptides using boronic acid-functionalized mesoporous silica. Anal. Chem. 2008, 81, 503–508. [Google Scholar] [CrossRef]
- Alvarez-Manilla, G.A.; Guo, Y.; Warren, N.L.; Orlando, R.; Pierce, M. Tools for glycoproteomic analysis: size exclusion chromatography facilitates identification of tryptic glycopeptides with N-linked glycosylation sites. J. Proteome Res. 2006, 5, 701–708. [Google Scholar] [CrossRef] [PubMed]
- Jia, W.; Lu, Z.; Fu, Y.; Wang, H.P.; Wang, L.H.; Chi, H. A strategy for precise and large scale identification of core fucosylated glycoproteins. Mol. Cell Proteomics 2009, 8, 913–923. [Google Scholar] [CrossRef] [PubMed]
- Lewandrowski, U.; Lohrig, K.; Zahedi, R.; Wolters, D.; Sickmann, A. Glycosylation site analysis of human platelets by electrostatic repulsion hydrophilic interaction chromatography. Clin. Proteomics 2008, 4, 25–36. [Google Scholar] [CrossRef]
- Zhang, H.; Guo, T.; Li, X.; Datta, A.; Park, J.E.; Yang, J. Simultaneous characterization of glyco- and phosphoproteomes of mouse brain membrane proteome with electrostatic repulsion hydrophilic interaction chromatography. Mol. Cell. Proteomics 2010, 9, 635–647. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.; Jung, E.; Brunak, S. Prediction of N-glycosylation Sites in Human Proteins. In Preparation. 2004. Available online: http://www.cbs.dtu.dk/services/NetNGlyc/ (accessed on 25 September 2014).
- Steentoft, C.; Vakhrushev, S.Y.; Joshi, H.J.; Kong, Y.; Vester-Christensen, M.B.; Schjoldager, K.T.; Lavrsen, K.; Dabelsteen, S.; Pedersen, N.B.; Marcos-Silva, L.; et al. Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO J. 2013, 32, 1478–1488. [Google Scholar] [CrossRef] [PubMed]
- Goldstein, I.J.; Poretz, R.D. Isolation, physicochemical characterization and carbohydrate-binding specificity of lectins. In The Lectins: Properties, Functions, and Applications in Biology and Medicine; Liener, I.E., Sharon, N., Goldstein, I.J., Eds.; Academic Press: New York, NY, USA, 1986; pp. 33–247. [Google Scholar]
- Apweiler, R.; Jesus Martin, M.; O’onovan, C.; Magrane, M.; Alam-Faruque, Y.; Antunes, R. The UniProt Consortium: Reorganizing the protein space at the Universal Protein Resource (UniProt). Nucl. Acids Res. 2012, 40, D71–D75. [Google Scholar] [CrossRef] [PubMed]
- Aebi, M.; Bernasconi, R.; Clerc, S.; Molinari, M. N-glycan structures: Recognition and processing in the ER. Cell 2010, 35, 74–82. [Google Scholar]
- Helenius, A.; Aebi, M. Intracellular functions of N-Linked glycans. Science 2001, 291, 2364–2369. [Google Scholar] [CrossRef] [PubMed]
- Naismith, J.H.; Field, R.A. Structural basis of trimannoside recognition by concanavalin A. J. Biol. Chem. 1996, 271, 972–976. [Google Scholar] [CrossRef] [PubMed]
- Ahn, J.; Febbraio, M.; Silverstein, R.L. A novel isoform of human Golgi complex-localized glycoprotein-1 (also known as E-selectin ligand-1, MG-160 and cysteine-rich fibroblast growth factor receptor) targets differential subcellular localization. J. Cell Sci. 2005, 118, 1725–1731. [Google Scholar] [CrossRef] [PubMed]
- Kowalewski, B.; Lübke, T.; Kollmann, K.; Braulke, T.; Reinheckel, T.; Dierks, T.; Damme, M. Molecular Characterization of Arylsulfatase G: Expression, processing, glycosylation, transport, and activity. J. Biol. Chem. 2014, 289, 27992–28005. [Google Scholar] [CrossRef] [PubMed]
- Reily, C.; Ueda, H.; Huang, Z.Q.; Mestecky, J.; Julian, B.A.; Willey, C.D.; Novak, J. Cellular signaling and production of galactose-deficient IgA1 in IgA nephropathy, an autoimmune disease. J. Immunol. Res. 2014, 2014. [Google Scholar] [CrossRef] [PubMed]
- Lin, M.C.; Huang, M.J.; Liu, C.H.; Yang, T.L.; Huang, M.C. GALNT2 enhances migration and invasion of oral squamous cell carcinoma by regulating EGFR glycosylation and activity. Oral Oncol. 2014, 50, 478–484. [Google Scholar] [CrossRef] [PubMed]
- Zachara, N.E.; Hart, G.W. Cell signaling, the essential role of O-GlcNAc! Biochim. Biophys. Acta 2006, 1761, 599–617. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, D.B.; Kawada, N.; Imamura, K.; Miyamoto, Y.; Tateno, C.; Seki, S.; Kuroki, T.; Yoshizato, K. Proteome analysis of rat hepatic stellate cells. Hepatology 2000, 32, 268–277. [Google Scholar] [CrossRef] [PubMed]
- Santucci, L.; Fiorucci, S.; Cammilleri, F.; Servillo, G.; Federici, B.; Morelli, A. Galectin-1 exerts immunomodulatory and protective effects on concanavalin A-induced hepatitis in mice. Hepatology 2000, 31, 399–406. [Google Scholar] [CrossRef] [PubMed]
- Hsu, D.K.; Dowling, C.A.; Jeng, K.C.; Chen, J.T.; Yang, R.Y.; Liu, F.T. Galectin-3 expression is induced in cirrhotic liver and hepatocellular carcinoma. Int. J. Cancer 1999, 81, 519–526. [Google Scholar] [CrossRef] [PubMed]
- Maeda, N.; Kawada, N.; Seki, S.; Arakawa, T.; Ikeda, K.; Iwao, H.; Okuyama, H.; Hirabayashi, J.; Kasai, K.; Yoshizato, K. Stimulation of proliferation of rat hepatic stellate cells by galectin-1 and galectin-3 through different intracellular signaling pathways. J. Biol. Chem. 2003, 278, 18938–18944. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Li, X.; Wang, Y.; Wang, H.; Huang, C.; Li, J. Endoplasmic reticulum stress-induced hepatic stellate cell apoptosis through calcium-mediated JNK/P38 MAPK and Calpain/Caspase-12 pathways. Mol. Cell Biochem. 2014, 394, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Berridge, M.J.; Peter, B.M.D. The versatility and universality of calcium signalling. Nat. Rev. Mol. Cell Biol. 2000, 1, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Clapham, D.E. Calcium signaling. Cell 2007, 131, 1047–1058. [Google Scholar] [CrossRef] [PubMed]
- Verma, A.K.; Filoteo, A.G.; Stanford, D.R.; Wieben, E.D.; Penniston, J.T. Complete primary structure of a human plasma membrane Ca2+ pump. J. Biol. Chem. 1988, 263, 14152–14159. [Google Scholar] [PubMed]
- Singer, A.U.; Waldo, G.L.; Harden, T.K.; Sondek, J. A unique fold of phospholipase C-beta mediates dimerization and interaction with G alpha q. Nat. Struct. Biol. 2002, 9, 32–36. [Google Scholar] [CrossRef]
- Wu, F.R.; Pan, C.X.; Rong, C.; Xia, Q.; Yuan, F.L.; Tang, J.; Wang, X.Y.; Wang, N.; Ni, W.L.; Chen, F.H. Inhibition of acid-sensing ion channel 1a in hepatic stellate cells attenuates PDGF-induced activation of HSCs through MAPK pathway. Mol. Cell. Biochem. 2014, 395, 199–209. [Google Scholar] [CrossRef] [PubMed]
- Ding, Z.Y.; Jin, G.N.; Liang, H.F.; Wang, W.; Chen, W.X.; Datta, P.K.; Zhang, M.Z.; Zhang, B.; Chen, X.P. Transforming growth factor β induces expression of connective tissue growth factor in hepatic progenitor cells through Smad independent signaling. Cell Signal 2013, 25, 1981–1992. [Google Scholar] [CrossRef] [PubMed]
- Friedman, S.L. Cytokines and fibrogenesis. Semin. Liver Dis. 1999, 19, 129–140. [Google Scholar] [PubMed]
- Pinzani, M.; Marra, F. Cytokine receptors and signaling in hepatic stellate cells. Semin. Liver Dis. 2001, 21, 397–416. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.L.; Wu, J.; Wei, D.; Zhang, D.W.; Feng, H.; Chen, Z.N. Newcastle disease virus represses the activation of human hepatic stellate cells and reverses the development of hepatic fibrosis in mice. Liver Int. 2009, 29, 593–602. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Cui, T.; Chen, Q.; Ma, T.; Li, Z. Isolation and identification of native membrane glycoproteins from living cell by concanavalin A—Magnetic particle conjugates. Anal. Biochem. 2012, 421, 339–341. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Cui, T.; Wang, Y.; Sun, S.; Ma, T.; Wang, T.; Chen, Q.; Li, Z. Selective isolation and analysis of glycoprotein fractions and their glycomes from hepatocellular carcinoma sera. Proteomics 2013, 13, 1481–1498. [Google Scholar] [CrossRef] [PubMed]
- Ishihama, Y.; Oda, Y.; Tabata, T.; Sato, T.; Nagasu, T.; Rappsilber, J.; Mann, M. Exponentially modified protein abundance index (emPAI) for estimation of absolute protein amount in proteomics by the number of sequenced peptides per protein. Mol. Cell Proteomics 2005, 4, 1265–1272. [Google Scholar] [CrossRef]
- Conesa, A.; Götz, S.; Garcia-Gomez, J.M.; Terol, J.; Talon, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef] [PubMed]
- Ye, J.; Fang, L.; Zheng, H.; Zhang, Y.; Chen, J.; Zhang, Z.; Wang, J.; Li, S.; Li, R.; Bolund, L.; et al. WEGO: A web tool for plotting GO annotations. Nucl. Acids Res. 2006, 34, W293–W297. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S.; Sato, Y.; Furumichi, M.; Tanabe, M. KEGG for integration and interpretation of large-scale molecular datasets. Nucl. Acids Res. 2012, 40, D109–D114. [Google Scholar] [CrossRef] [PubMed]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID Bioinformatics Resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Franceschini, A.; Kuhn, M.; Simonovic, M.; Roth, A.; Minguez, P. The STRING database in 2011: Functional interaction networks of proteins, globally integrated and scored. Nucl. Acids Res. 2011, 39, D561–D568. [Google Scholar] [CrossRef] [PubMed]
- Gomes, T.S.; Oshima, C.T.; Forones, N.M.; de Oliveira Lima, F.; Ribeiro, D.A. Expression of galectin-3 in gastric adenocarcinoma. Indian J. Med. Res. 2014, 140, 69–76. [Google Scholar] [PubMed]
- Sample Availability: Samples of the compounds upon request are available from the authors.
© 2014 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qin, Y.; Zhong, Y.; Yang, G.; Ma, T.; Jia, L.; Huang, C.; Li, Z. Profiling of Concanavalin A-Binding Glycoproteins in Human Hepatic Stellate Cells Activated with Transforming Growth Factor-β1. Molecules 2014, 19, 19845-19867. https://doi.org/10.3390/molecules191219845
Qin Y, Zhong Y, Yang G, Ma T, Jia L, Huang C, Li Z. Profiling of Concanavalin A-Binding Glycoproteins in Human Hepatic Stellate Cells Activated with Transforming Growth Factor-β1. Molecules. 2014; 19(12):19845-19867. https://doi.org/10.3390/molecules191219845
Chicago/Turabian StyleQin, Yannan, Yaogang Zhong, Ganglong Yang, Tianran Ma, Liyuan Jia, Chen Huang, and Zheng Li. 2014. "Profiling of Concanavalin A-Binding Glycoproteins in Human Hepatic Stellate Cells Activated with Transforming Growth Factor-β1" Molecules 19, no. 12: 19845-19867. https://doi.org/10.3390/molecules191219845
APA StyleQin, Y., Zhong, Y., Yang, G., Ma, T., Jia, L., Huang, C., & Li, Z. (2014). Profiling of Concanavalin A-Binding Glycoproteins in Human Hepatic Stellate Cells Activated with Transforming Growth Factor-β1. Molecules, 19(12), 19845-19867. https://doi.org/10.3390/molecules191219845




