NMR Solution Structure of a Chymotrypsin Inhibitor from the Taiwan Cobra Naja naja atra
Abstract
:1. Introduction
2. Results and Discussion
2.1. Chemical Shift Assignment

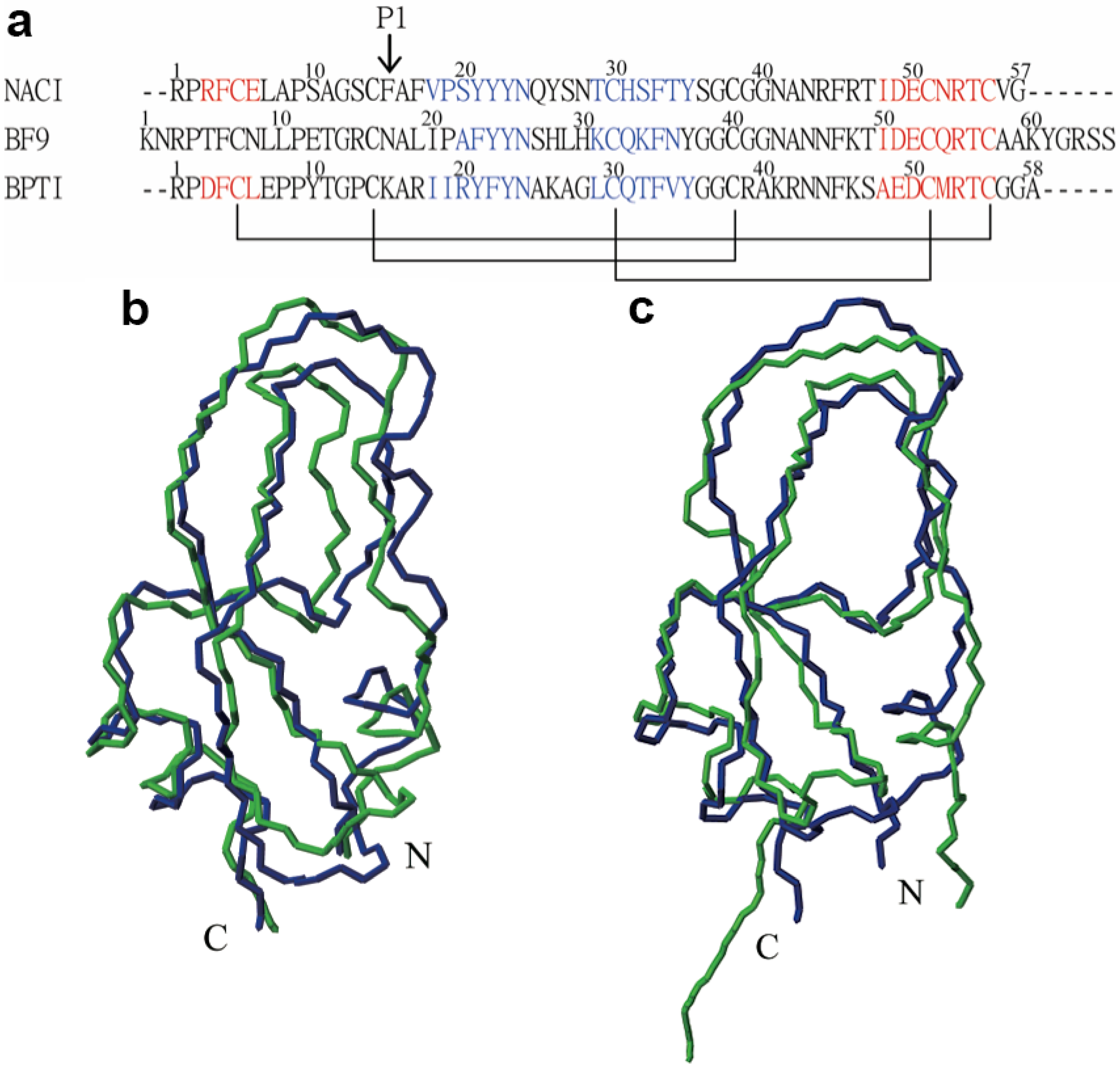
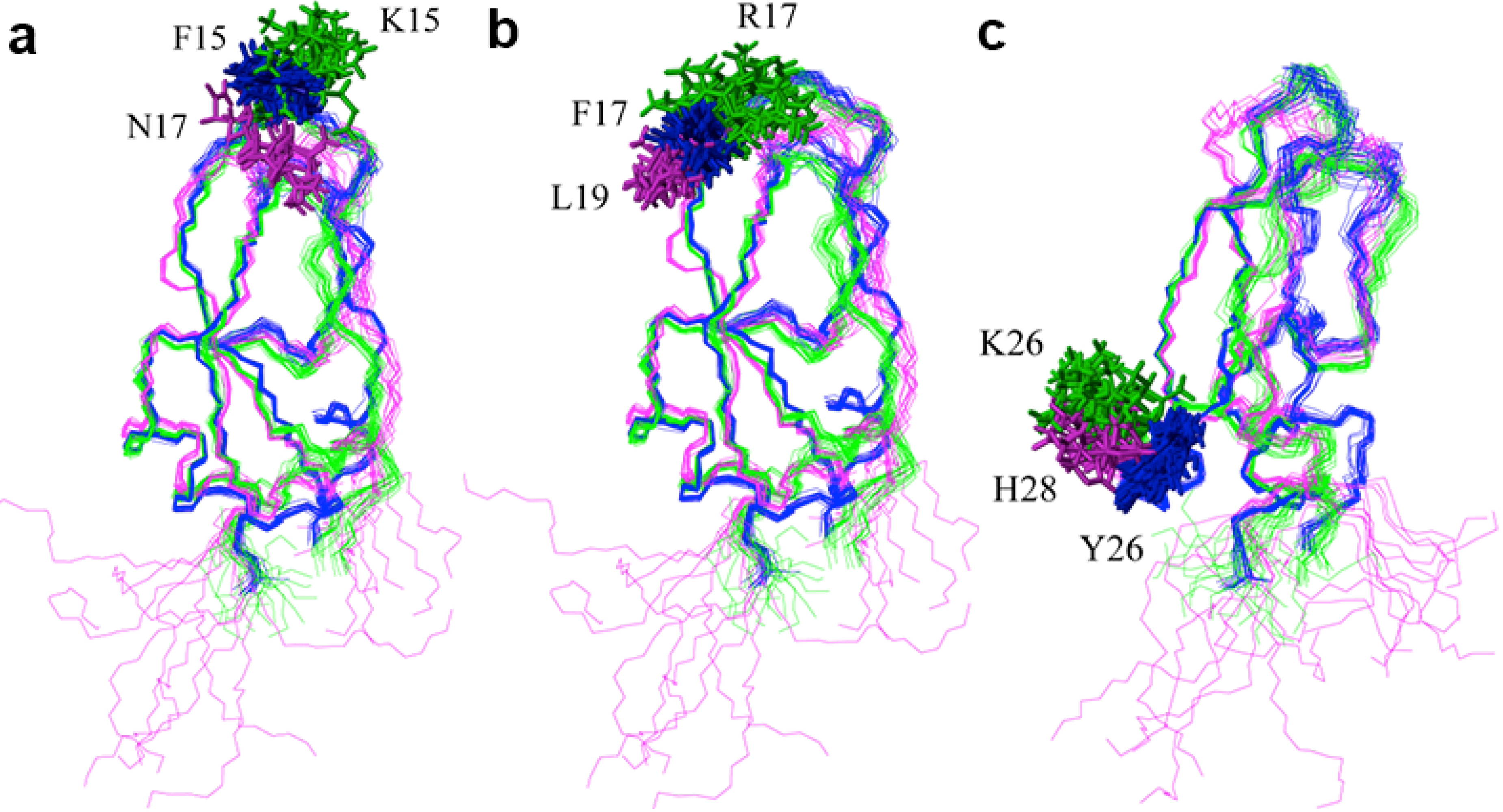
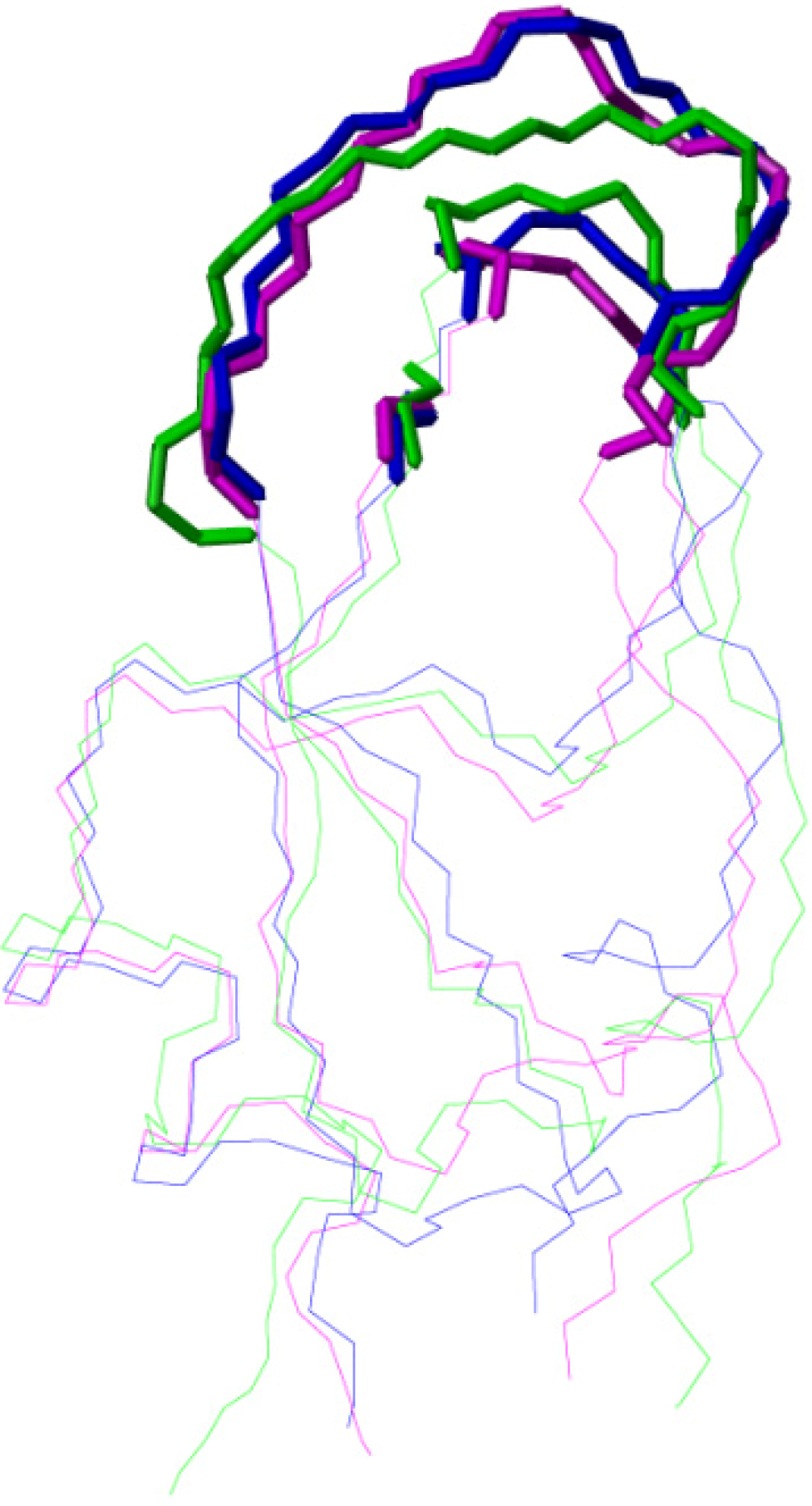
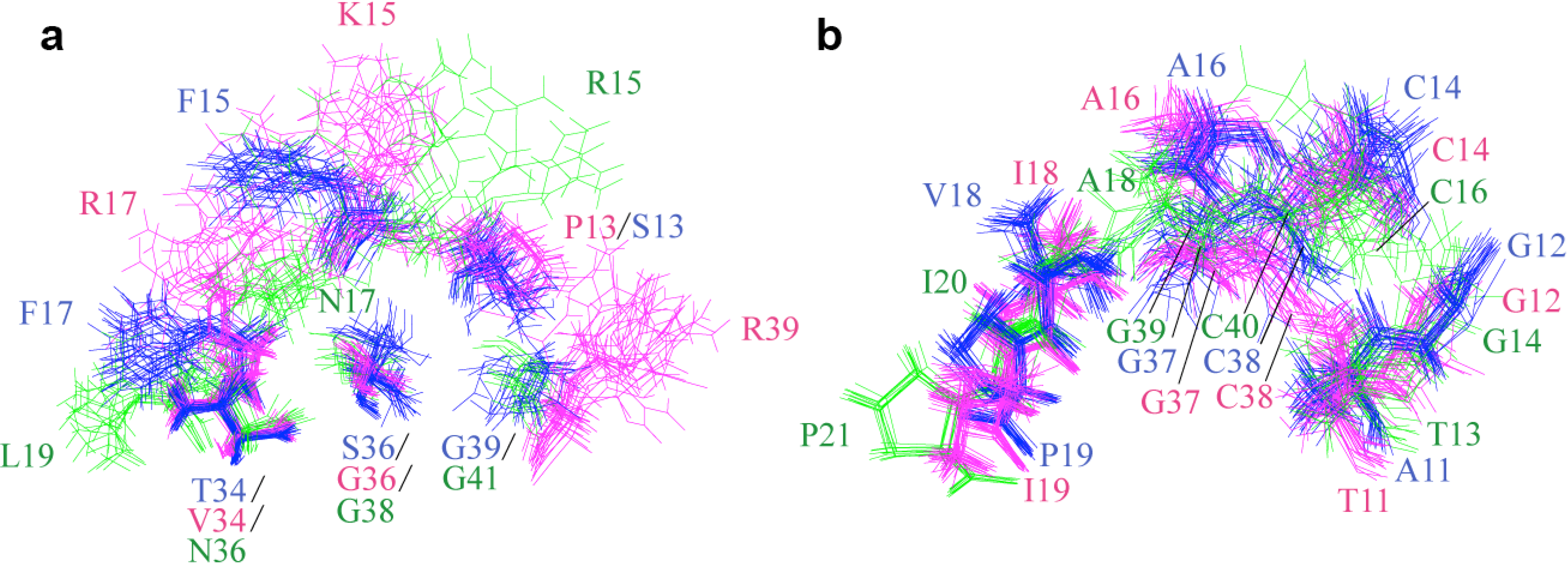
2.2. Structure Determination and Solution Structure of NaCl
| NOE distance restraints: | |
|---|---|
| Number | 1075 |
| Intraresidual, |i – j| = 0 | 121 |
| Sequential, |i – j| = 1 | 324 |
| Medium range, 1 < |i – j| < 5 | 253 |
| Long range, |i – j| >= 5 | 377 |
| Maximal violation | 0.10 ± 0.01 Å |
| Torsion angle restraints (ϕ/ψ): | |
| Number | 75 |
| Maximal violation | 2.94 ± 0.87° |
| Final CYANA target function value | 2.61 ± 1.61 Å2 |
| AMBER energy | −2208 ± 50 kcal/mol |
| RMSDs from ideal geometry: | |
| Bond lengths | 0.015 ± 0.001 Å |
| Bond angles | 1.93 ± 0.04° |
| RMSD to mean coordinates of residues 1–57: | |
| Backbone atoms N, Cα, C' | 0.37 ± 0.08Å |
| All heavy atoms | 0.73 ± 0.08Å |
| PROCHECK Ramachandran plot statistics: | |
| Most favorable regions | 78.5% |
| Additional allowed regions | 21.5% |
| Generously allowed regions | 0.0% |
| Disallowed regions | 0.0% |


3. Experimental
3.1. Sample Preparation
3.2. NMR Measurements
3.3. Structure Calculation
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Župunski, V.; Kordiš, D.; Gubenšek, F. Adaptive evolution in the snake venom Kunitz/BPTI protein family. FEBS Lett. 2003, 547, 131–136. [Google Scholar] [CrossRef]
- Gilquin, B.; Lecoq, A.; Desné, F.; Guenneugues, M.; Zinn-Justin, S.; Ménez, A. Conformational and functional variability supported by the BPTI fold: Solution structure of the Ca2+ channel blocker calcicludine. Proteins 1999, 34, 520–532. [Google Scholar] [CrossRef]
- Cheng, Y.C.; Yan, F.R.; Chang, L.S. Taiwan cobra chymotrypsin inhibitor: cloning, functional expression and gene organization. Biochim. Biophys. Acta 2005, 1747, 213–220. [Google Scholar] [CrossRef]
- Pritchard, L.; Dufton, M.J. Evolutionary trace analysis of the Kunitz/BPTI family of proteins: Functional divergence may have been based on conformational adjustment. J. Mol. Biol. 1999, 285, 1589–1607. [Google Scholar]
- Chen, C.P.; Hsu, C.H.; Su, N.Y.; Lin, Y.C.; Chiou, S.H.; Wu, S.H. Solution structure of a Kunitz-type chymotrypsin inhibitor isolated from the elapid snake Bungarus fasciatus. J. Biol. Chem. 2001, 276, 45079–45087. [Google Scholar]
- Branden, C.; Tooze, J. Introduction to Protein Structure, 2nd ed.; Garland: New York, NY, USA, 1999. [Google Scholar]
- Laskowski, M.; Kato, I. Protein inhibitors of proteinases. Annu. Rev. Biochem. 1980, 49, 593–626. [Google Scholar] [CrossRef]
- Rühlmann, A.; Kukla, D.; Schwager, P.; Bartels, K.; Huber, R. Structure of the complex formed by bovine trypsin and bovine pancreatic trypsin inhibitor: Crystal structure determination and stereochemistry of the contact region. J. Mol. Biol. 1973, 77, 417–436. [Google Scholar] [CrossRef]
- Otting, G.; Liepinsh, E.; Wüthrich, K. Disulfide bond isomerization in BPTI and BPTI(G36S): An NMR study of correlated mobility in proteins. Biochemistry 1993, 32, 3571–3582. [Google Scholar] [CrossRef]
- Grey, M.J.; Wang, C.Y.; Palmer, A.G. Disulfide bond isomerization in basic pancreatic trypsin inhibitor: Multisite chemical exchange quantified by CPMG relaxation dispersion and chemical shift modeling. J. Am. Chem. Soc. 2003, 125, 14324–14335. [Google Scholar] [CrossRef]
- Herrmann, T.; Güntert, P.; Wüthrich, K. Protein NMR structure determination with automated NOE assignment using the new software CANDID and the torsion angle dynamics algorithm DYANA. J. Mol. Biol. 2002, 319, 209–227. [Google Scholar] [CrossRef]
- Jee, J.; Güntert, P. Influence of the completeness of chemical shift assignments on NMR structures obtained with automated NOE assignment. J. Struct. Funct. Genom. 2003, 4, 179–189. [Google Scholar]
- Güntert, P.; Mumenthaler, C.; Wüthrich, K. Torsion angle dynamics for NMR structure calculation with the new program DYANA. J. Mol. Biol. 1997, 273, 283–298. [Google Scholar] [CrossRef]
- Güntert, P. Automated structure determination from NMR spectra. Eur. Biophys. J. 2009, 38, 129–143. [Google Scholar] [CrossRef]
- Laskowski, R.A.; Rullmann, J.A.C.; MacArthur, M.W.; Kaptein, R.; Thornton, J.M. AQUA and PROCHECK-NMR: Programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 1996, 8, 477–486. [Google Scholar]
- Koradi, R.; Billeter, M.; Wüthrich, K. MOLMOL: A program for display and analysis of macromolecular structures. J. Mol. Graphics 1996, 14, 51–55. [Google Scholar] [CrossRef]
- Ulrich, E.L.; Akutsu, H.; Doreleijers, J.F.; Harano, Y.; Ioannidis, Y.E.; Lin, J.; Livny, M.; Mading, S.; Maziuk, D.; Miller, Z.; et al. BioMagResBank. Nucleic Acids Res. 2008, 36, D402–D408. [Google Scholar]
- Berndt, K.D.; Güntert, P.; Orbons, L.P.M.; Wüthrich, K. Determination of a high-quality nuclear magnetic resonance solution structure of the bovine pancreatic trypsin inhibitor and comparison with three crystal structures. J. Mol. Biol. 1992, 227, 757–775. [Google Scholar] [CrossRef]
- Markland, W.; Ley, A.C.; Lee, S.W.; Ladner, R.C. Iterative optimization of high-affinity protease inhibitors using phage display .1. Plasmin. Biochemistry 1996, 35, 8045–8057. [Google Scholar] [CrossRef]
- Goddard, T.D.; Kneller, D.G. Sparky 3; University of California: San Francisco, CA, USA, 2001. [Google Scholar]
- Güntert, P. Automated NMR protein structure calculation. Prog. Nucl. Magn. Reson. Spectrosc. 2003, 43, 105–125. [Google Scholar] [CrossRef]
- Wishart, D.S.; Sykes, B.D. The 13C chemical-shift index: a simple method for the identification of protein secondary structure using 13C chemical-shift data. J. Biomol. NMR 1994, 4, 171–180. [Google Scholar]
- Cornilescu, G.; Delaglio, F.; Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 1999, 13, 289–302. [Google Scholar] [CrossRef]
- Cornell, W.D.; Cieplak, P.; Bayly, C.I.; Gould, I.R.; Merz, K.M.; Ferguson, D.M.; Spellmeyer, D.C.; Fox, T.; Caldwell, J.W.; Kollman, P.A. A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J. Am. Chem. Soc. 1995, 117, 5179–5197. [Google Scholar] [CrossRef]
- Koradi, R.; Billeter, M.; Güntert, P. Point-centered domain decomposition for parallel molecular dynamics simulation. Comput. Phys. Commun. 2000, 124, 139–147. [Google Scholar] [CrossRef]
- Luginbühl, P.; Güntert, P.; Billeter, M.; Wüthrich, K. The new program OPAL for molecular dynamics simulations and energy refinements of biological macromolecules. J. Biomol. NMR 1996, 8, 136–146. [Google Scholar]
- Sample Availability: Not available.
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lin, Y.-J.; Ikeya, T.; Güntert, P.; Chang, L.-S. NMR Solution Structure of a Chymotrypsin Inhibitor from the Taiwan Cobra Naja naja atra. Molecules 2013, 18, 8906-8918. https://doi.org/10.3390/molecules18088906
Lin Y-J, Ikeya T, Güntert P, Chang L-S. NMR Solution Structure of a Chymotrypsin Inhibitor from the Taiwan Cobra Naja naja atra. Molecules. 2013; 18(8):8906-8918. https://doi.org/10.3390/molecules18088906
Chicago/Turabian StyleLin, Yi-Jan, Teppei Ikeya, Peter Güntert, and Long-Sen Chang. 2013. "NMR Solution Structure of a Chymotrypsin Inhibitor from the Taiwan Cobra Naja naja atra" Molecules 18, no. 8: 8906-8918. https://doi.org/10.3390/molecules18088906
APA StyleLin, Y.-J., Ikeya, T., Güntert, P., & Chang, L.-S. (2013). NMR Solution Structure of a Chymotrypsin Inhibitor from the Taiwan Cobra Naja naja atra. Molecules, 18(8), 8906-8918. https://doi.org/10.3390/molecules18088906





