Enhancement of Gene Silencing Effect and Membrane Permeability by Peptide-Conjugated 27-Nucleotide Small Interfering RNA
Abstract
:1. Introduction
2. Results and Discussion
2.1. Results
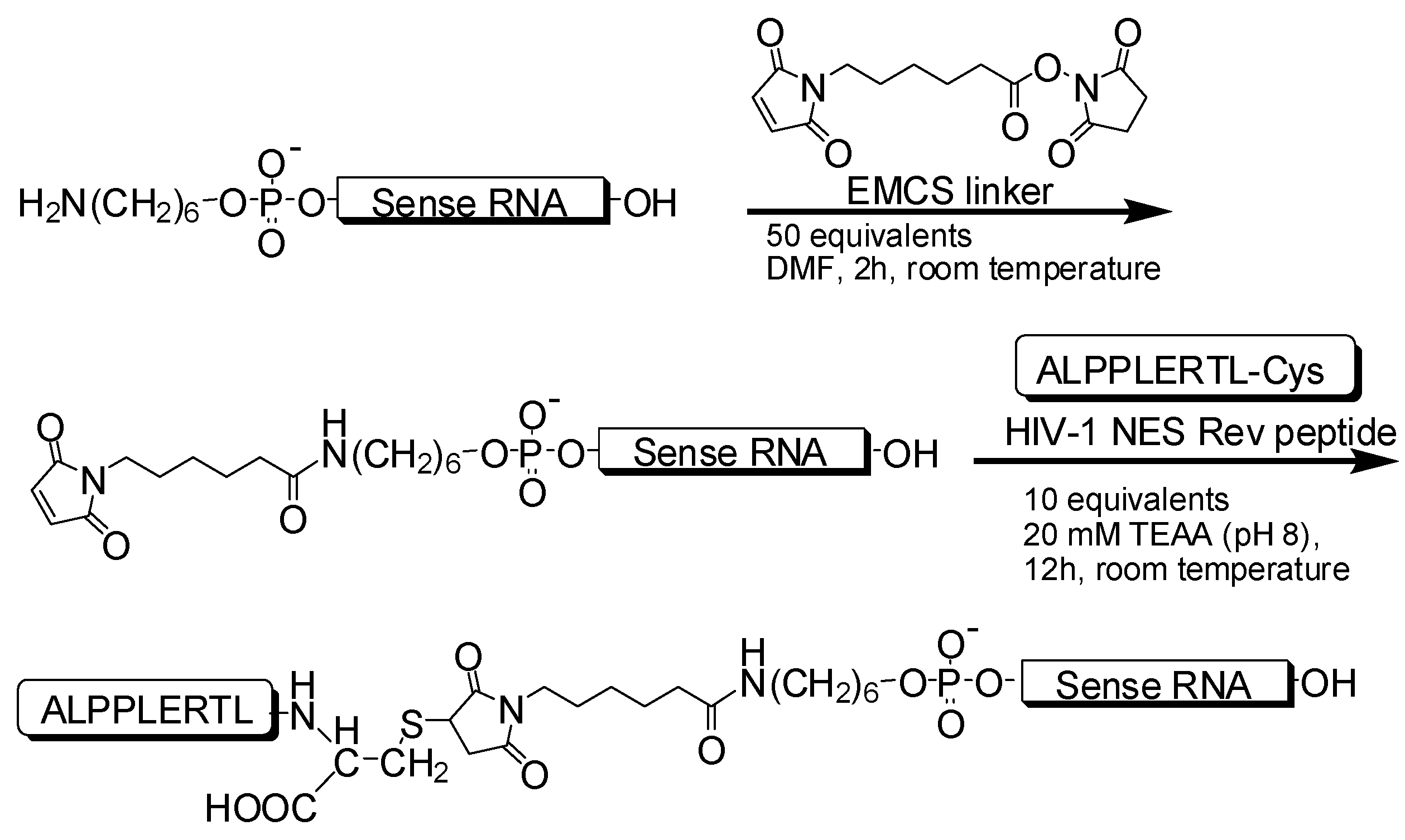
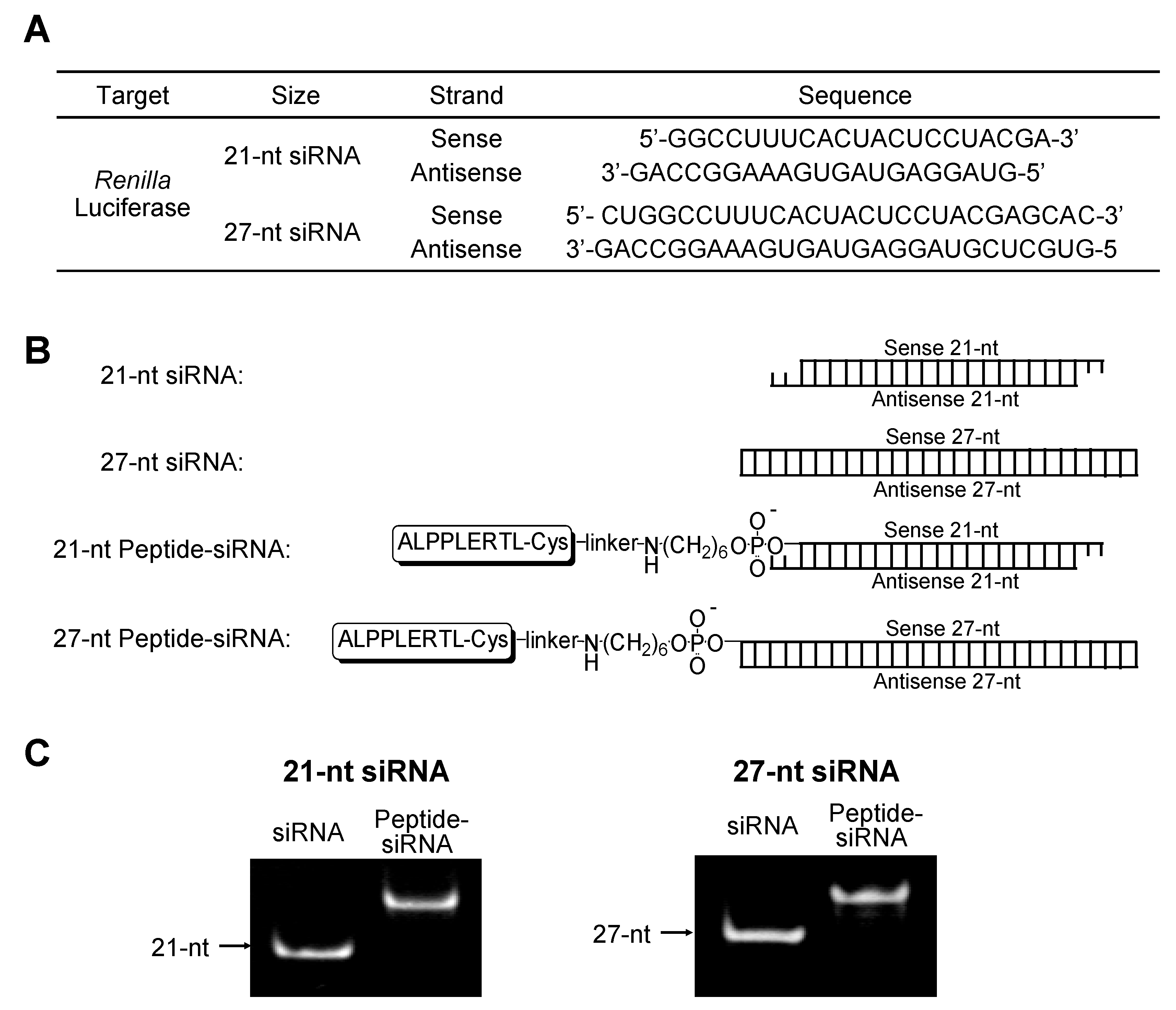
2.1.1. Synthesis of 21-nt and 27-nt siRNAs Conjugated with Peptide
| Size | Target gene | Conjugate | HPLC retention time a (min) | MALDI-TOF MS b Found/Calcd | Yield c (%) |
|---|---|---|---|---|---|
| 21-nt ssRNA | Luciferase | None | 6.5 | 6569.8/6569.9 | d |
| 21-nt ssRNA | Luciferase | HIV-1 Rev | 15.5 | 8154.5/8153.8 | 33.0 |
| 27-nt ssRNA | Luciferase | None | 6.9 | 8463.0/8465.1 | d |
| 27-nt ssRNA | Luciferase | HIV-1 Rev | 16.8 | 10,057.5/10,049.0 | 22.9 |
2.1.2. Processing of Peptide-siRNAs by Dicer
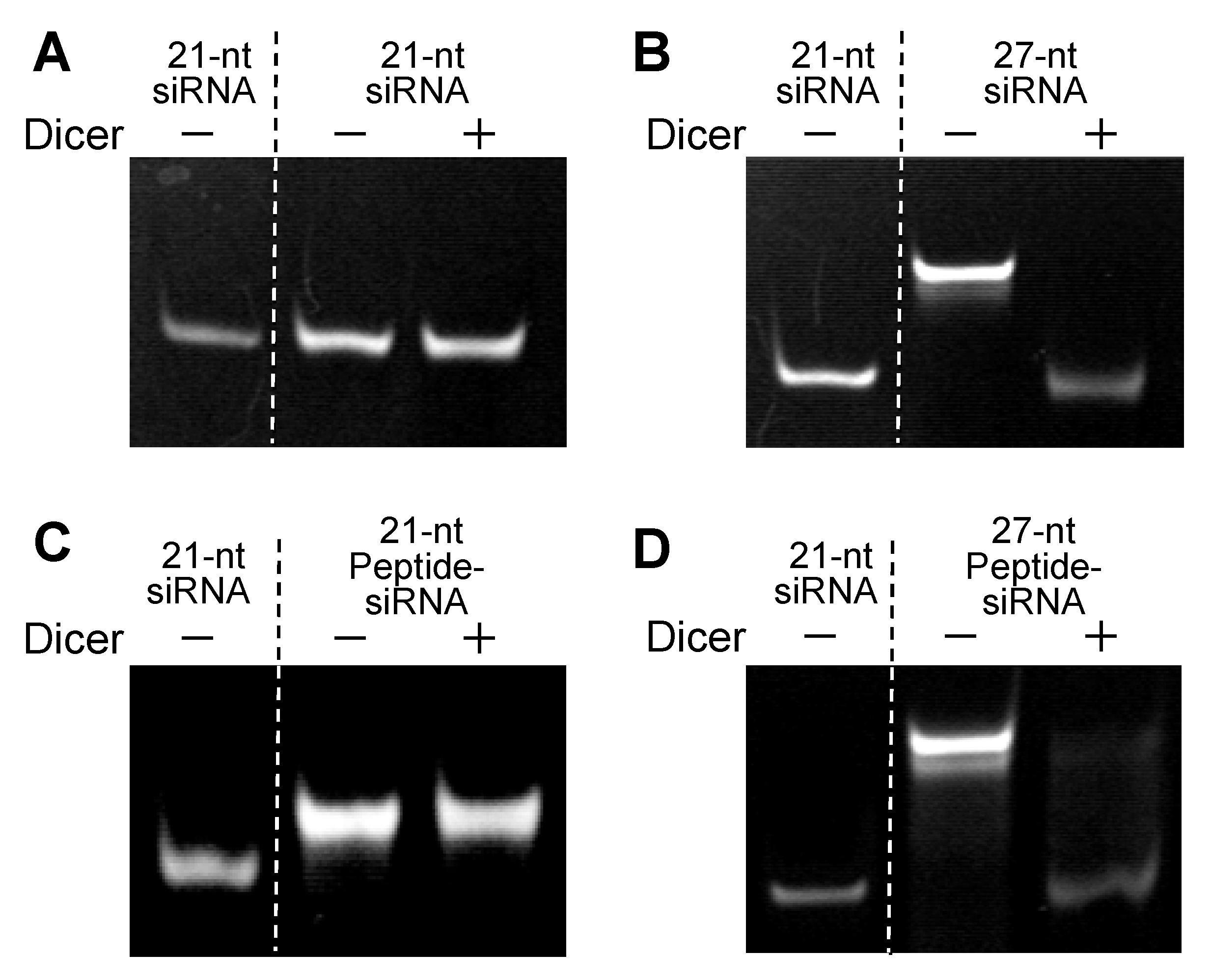
2.1.3. Gene Silencing of Peptide-siRNAs
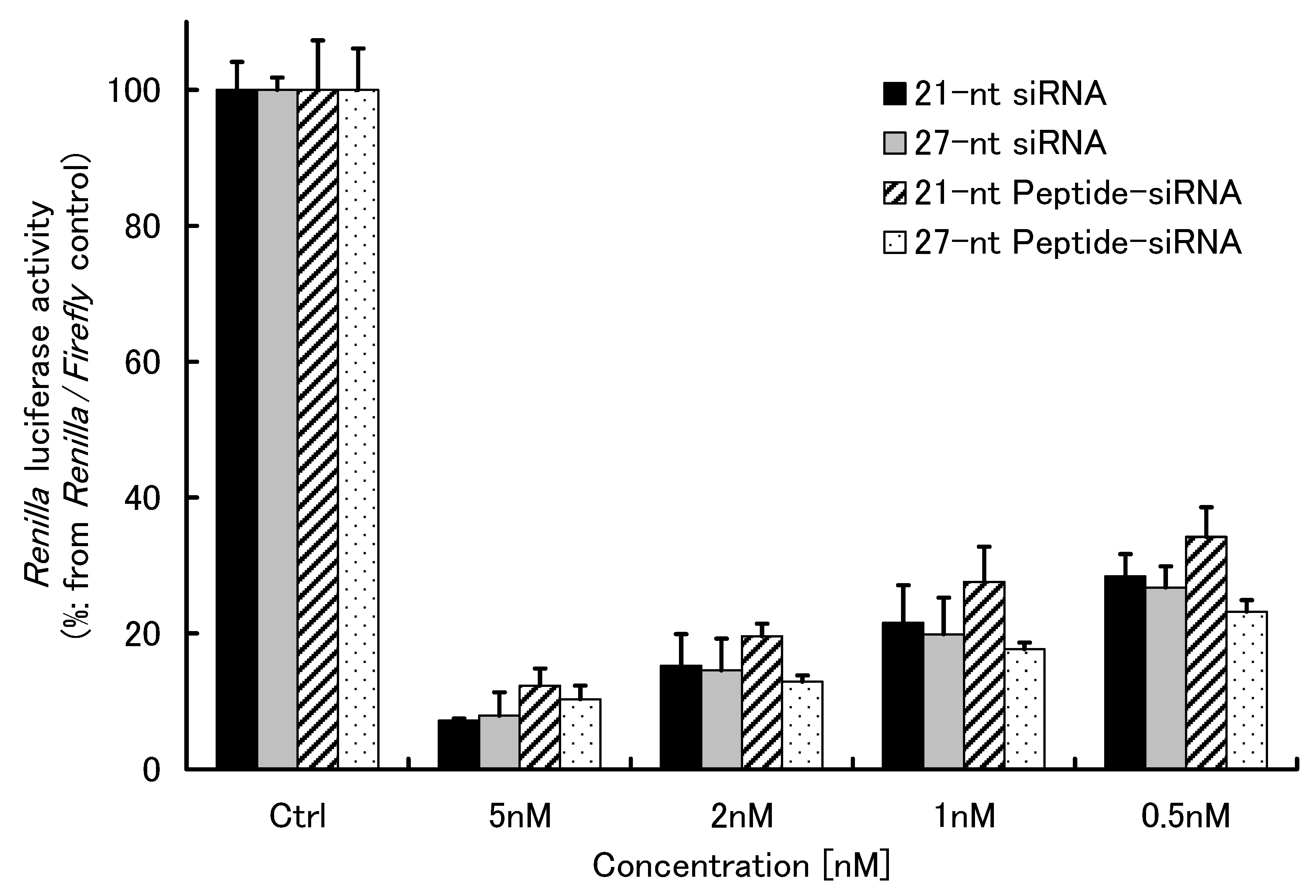
2.1.4. Cell Membrane Permeability of Peptide-siRNAs
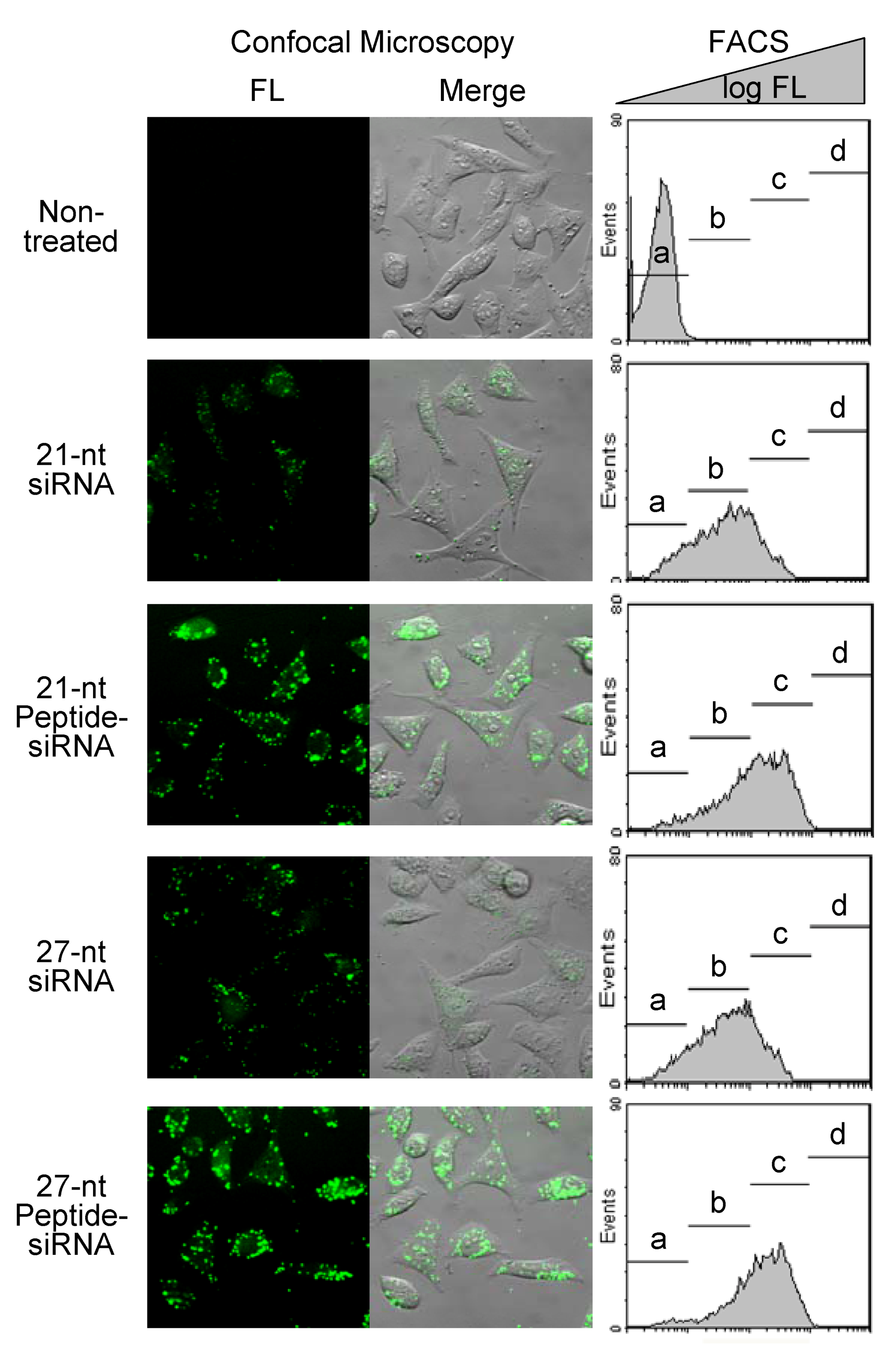
| siRNAs | Populations of the HeLa cells (%) | |||
|---|---|---|---|---|
| a | b | c | d | |
| Non-treated | 99.58 | 0.40 | 0.01 | 0.00 |
| 21-nt siRNA | 16.44 | 60.36 | 21.89 | 0.00 |
| 21-nt Peptide-siRNA | 7.35 | 36.57 | 55.00 | 0.31 |
| 27-nt siRNA | 14.37 | 62.29 | 22.30 | 0.00 |
| 27-nt Peptide-siRNA | 5.62 | 28.93 | 64.07 | 0.49 |
2.2. Discussion
3. Experimental
3.1. Synthesis of siRNAs Conjugated with NES Peptide
3.2. In Vitro Cleavage of Peptide-siRNAs by Recombinant Dicer
3.3. Cell Culture
3.4. Transfection of Peptide-siRNAs Targeting Renilla Luciferase mRNA
3.5. Luciferase Activity Assay
3.6. Confocal Microscopy and Flow Cytometry
4. Conclusions
Acknowledgments
References
- Tabara, H.; Sarkissian, M.; Kelly, W.G.; Fleenor, J.; Grishok, A.; Timmons, L.; Fire, A.; Mello, C.C. The rde-1 gene, RNA interference, and transposon silencing in C. elegans. Cell 1999, 99, 123–132. [Google Scholar] [CrossRef]
- Couzin, J. Small RNAs make big splash. Science 2002, 298, 2296–2297. [Google Scholar] [CrossRef]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar]
- Myers, J.W.; Jones, J.T.; Meyer, T.; Ferrell, J.E. Recombinant Dicer efficiently converts large dsRNAs into siRNAs suitable for gene silencing. Nat. Biotechnol. 2003, 21, 324–328. [Google Scholar] [CrossRef]
- Macrae, I.J.; Zhou, K.; Li, F.; Repic, A.; Brooks, A.N.; Cande, W.Z.; Adams, P.D.; Doudna, J.A. Structural basis for double-stranded RNA processing by Dicer. Science 2006, 311, 195–198. [Google Scholar] [CrossRef]
- Silhavy, D.; Molnar, A.; Lucioli, A.; Szittya, G.; Hornyik, C.; Tavazza, M.; Burgyan, J. A viral protein suppresses RNA silencing and binds silencing-generated, 21- to 25-nucleotide double-stranded RNAs. EMBO J. 2002, 21, 3070–3080. [Google Scholar] [CrossRef]
- Hornung, V.; Guenthner-Biller, M.; Bourquin, C.; Ablasser, A.; Schlee, M.; Uematsu, S.; Noronha, A.; Manoharan, M.; Akira, S.; de Fougerolles, A.; et al. Sequence-specific potent induction of IFN-alpha by short interfering RNA in plasmacytoid dendritic cells through TLR7. Nat. Med. 2005, 11, 263–270. [Google Scholar] [CrossRef]
- Elbashir, S.M.; Harborth, J.; Lendeckel, W.; Yalcin, A.; Weber, K.; Tuschl, T. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 2001, 411, 494–498. [Google Scholar]
- Schütze, N. siRNA technology. Mol. Cell. Endocrinol. 2004, 213, 115–119. [Google Scholar] [CrossRef]
- John, M.; Constien, R.; Akinc, A.; Goldberg, M.; Moon, Y.A.; Spranger, M.; Hadwiger, P.; Soutschek, J.; Vornlocher, H.P.; Manoharan, M.; et al. Effective RNAi-mediated gene silencing without interruption ofthe endogenous microRNA pathway. Nature 2007, 449, 745–747. [Google Scholar]
- Leung, R.K.; Whittaker, P.A. RNA interference: From gene silencing to gene-specific therapeutics. Pharmacol. Ther. 2005, 107, 222–239. [Google Scholar] [CrossRef]
- Lu, D.; Xiao, Z.; Wang, W.; Xu, Y.; Gao, S.; Deng, L.; He, W.; Yang, Y.; Guo, X.; Wang, X. Down regulation of CIAPIN1 reverses multidrug resistance in human breast cancer cells by inhibiting MDR1. Molecules 2012, 17, 7595–7611. [Google Scholar] [CrossRef]
- Wang, Y.; Cao, Y.L.; Yang, F.; Zhang, Y.; Wang, S.H.; Liu, L. Small interfering RNA effectively inhibits the expression of SARS coronavirus membrane gene at two novel targeting sites. Molecules 2010, 15, 7197–7207. [Google Scholar] [CrossRef]
- Kubo, T.; Zhelev, Z.; Bakalova, R.; Ohba, H.; Doi, K.; Fujii, M. Controlled intracellular localization and enhanced antisense effect of oligonucleotides by chemical conjugation. Org. Biomol. Chem. 2005, 21, 3257–3259. [Google Scholar]
- Kubo, T.; Takamori, K.; Kanno, K.; Bakalova, R.; Ohba, H.; Matsukisono, M.; Akebiyama, Y.; Fujii, M. Efficient cleavage of RNA, enhanced cellular uptake, and controlled intracellular localization of conjugate DNAzymes. Bioorg. Med. Chem. Lett. 2005, 15, 167–170. [Google Scholar] [CrossRef]
- Kubo, T.; Morikawa, M.; Ohba, H.; Fujii, M. Synthesis of DNA-peptide conjugates by solid-phase fragment condensation. Org. Lett. 2003, 5, 2623–2626. [Google Scholar] [CrossRef]
- Luo, D.; Saltzman, W.M. Synthetic DNA delivery systems. Nat. Biotechnol. 2000, 18, 33–37. [Google Scholar] [CrossRef]
- Pungente, M.D.; Jubeli, E.; Øpstad, C.L.; Al-Kawaz, M.; Barakat, N.; Ibrahim, T.; Abdul Khalique, N.; Raju, L.; Jones, R.; et al. Synthesis and preliminary investigations of the siRNA delivery potential of novel, single-chain rigid cationic carotenoid lipids. Molecules 2012, 17, 3484–3500. [Google Scholar]
- Houra, K.; Turčić, P.; Gabričević, M.; Weitner, T.; Konjevoda, P.; Stambuk, N. Interaction of α-melanocortin and its pentapeptide antisense LVKAT: Effects on hepatoprotection in male CBA mice. Molecules 2011, 26, 7331–7343. [Google Scholar]
- Takei, Y.; Kadomatsu, K.; Yuzawa, Y.; Matsuo, S.; Muramatsu, T. A small interfering RNA targeting vascular endothelial growth factor as cancer therapeutics. Cancer Res. 2004, 64, 3365–3370. [Google Scholar] [CrossRef]
- Czauderna, F.; Fechtner, M.; Dames, S.; Aygun, H.; Klippel, A.; Pronk, G.J.; Giese, K.; Kaufmann, J.F. Structural variations and stabilising modifications of synthetic siRNAs in mammalian cells. Nucleic Acids Res. 2003, 31, 2705–2716. [Google Scholar] [CrossRef]
- Elmén, J.; Thonberg, H.; Ljungberg, K.; Frieden, M.; Westergaard, M.; Xu, Y.; Wahren, B.; Liang, Z.; Ørum, H.; Koch, T.; et al. Locked nucleic acid (LNA) mediated improvements in siRNA stability and functionality. Nucleic Acids Res. 2005, 33, 439–447. [Google Scholar] [CrossRef]
- Amarzguioui, M.; Holen, T.; Babaie, E.; Prydz, H. Tolerance for mutations and chemical modifications in a siRNA. Nucleic Acids Res. 2003, 31, 589–595. [Google Scholar]
- Braasch, D.A.; Jensen, S.; Liu, Y.; Kaur, K.; Arar, K.; White, M.A.; Corey, D.R. RNA interference in mammalian cells by chemically-modified RNA. Biochemistry 2003, 42, 7967–7975. [Google Scholar] [CrossRef]
- Kenski, D.M.; Cooper, A.J.; Li, J.J.; Willingham, A.T.; Haringsma, J.J; Young, T.A.; Kuklin, N.A.; Jones, J.J.; Cancilla, M.T.; McMasters, D.R.; et al. Analysis of acyclic nucleoside modifications in siRNAs finds sensitivity at position 1 that is restored by 5'-terminal phosphorylation both in vitro and in vivo. Nucleic Acids Res. 2010, 38, 660–671. [Google Scholar] [CrossRef]
- Grünweller, A.; Wyszko, E.; Bieber, B.; Jahnel, R.; Erdmann, V.A.; Kurreck, J. Comparison of different antisense strategies in mammalian cells using locked nucleic acids, 2'-O-methyl RNA, Phosphorothioates and small interfering RNA. Nucleic Acids Res. 2003, 12, 3185–3193. [Google Scholar]
- Hall, A.H.; Wan, J.; Spesock, A.; Sergueeva, Z.; Shaw, B.R.; Alexander, K.A. High potency silencing by single-stranded boranophosphate siRNA. Nucleic Acids Res. 2006, 34, 2773–2781. [Google Scholar] [CrossRef]
- Jung, S.; Lee, S.H.; Mok, H.; Chung, H.J.; Park, T.G. Gene silencing efficiency of siRNA-PEG conjugates: Effect of PEGylation site and PEG molecular weight. J. Control. Release 2010, 144, 306–313. [Google Scholar] [CrossRef]
- Oishi, M.; Nagasaki, Y.; Itaka, K.; Nishiyama, N.; Kataoka, K. Lactosylated poly(ethylene glycol)-siRNA conjugate through acid-labile beta-thiopropionate linkage to construct pH-sensitive polyion complex micelles achieving enhanced gene silencing in hepatoma cells. J. Am. Chem. Soc. 2005, 127, 1624–1625. [Google Scholar]
- Moschos, S.A.; Jones, S.W.; Perry, M.N.; Williams, A.E.; Erjefalt, J.S.; Turner, J.J.; Barnes, P.J.; Sproat, B.S.; Gait, M.J.; Lindsay, M.A. Lung delivery studies using siRNA conjugated to TAT(48–60) and penetratin reveal peptide induced reduction in gene expression and induction of innate immunity. Bioconjug. Chem. 2007, 18, 1450–1459. [Google Scholar] [CrossRef]
- Muratovska, A.; Eccles, M.R. Conjugate for efficient delivery of short interfering RNA (siRNA) into mammalian cells. FEBS Lett. 2004, 58, 63–68. [Google Scholar] [CrossRef]
- Turner, J.J.; Jones, S.; Fabani, M.M.; Ivanova, G.; Arzumanov, A.A.; Gait, M.J. RNA targeting with peptide conjugates of oligonucleotides, siRNA and PNA. Blood Cells Mol. Dis. 2007, 38, 1–7. [Google Scholar] [CrossRef]
- Detzer, A.; Overhoff, M.; Wunsche, W.; Rompf, M.; Turner, J.J.; Ivanova, G.D.; Gait, M.J.; Sczakiel, G. Increased RNAi is related to intracellular release of siRNA via a covalently attached signal peptide. RNA 2009, 15, 627–636. [Google Scholar] [CrossRef]
- Soutschek, J.; Akinc, A.; Bramlage, B.; Charisse, K.; Constien, R.; Donoghue, M.; Elbashir, S.; Geick, A.; Hadwiger, P.; Harborth, J.; et al. Therapeutic silencing of an endogenous gene by systemic administration of modified siRNAs. Nature 2004, 432, 173–178. [Google Scholar]
- Kubo, T.; Takei, Y.; Mihara, K.; Yanagihara, K.; Seyama, T. Amino-modified and lipid-conjugated Dicer-substrate siRNA enhances RNAi efficacy. Bioconjug. Chem. 2012, 23, 164–173. [Google Scholar] [CrossRef]
- Kubo, T.; Yanagihara, K.; Takei, Y.; Mihara, K.; Morita, Y.; Seyama, T. Palmitic acid-conjugated 21-nt siRNA enhances gene-silencing activity. Mol. Pharm. 2011, 8, 2193–2203. [Google Scholar] [CrossRef]
- Wolfrum, C.; Shi, S.; Jayaprakash, K.N.; Jayaraman, M.; Wang, G.; Pandey, R.K.; Rajeev, K.G.; Nakayama, T.; Charrise, K.; Ndungo, E.M.; et al. Mechanisms and optimization of in vivo delivery of lipophilic siRNAs. Nat. Biotechnol. 2007, 25, 1149–1157. [Google Scholar] [CrossRef]
- Nishina, K.; Unno, T.; Uno, Y.; Kubodera, T.; Kanouchi, T.; Mizusawa, H.; Yokota, T. Efficient in vivo delivery of siRNA to the liver by conjugation of alpha-tocopherol. Mol. Ther. 2008, 16, 734–740. [Google Scholar] [CrossRef]
- Kim, D.H.; Behlke, M.A.; Rose, S.D.; Chang, M.S.; Choi, S.; Rossi, J.J. Synthetic dsRNA Dicer substrates enhance RNAi potency and efficacy. Nat. Biotechnol. 2005, 23, 222–226. [Google Scholar] [CrossRef]
- Marques, J.T.; Devosse, T.; Wang, D.; Zamanian-Daryoush, M.; Serbinowski, P.; Hartmann, R.; Fujita, T.; Behlke, M.A.; Williams, B.R. A structural basis for discriminating between self and nonself double-stranded RNAs in mammalian cells. Nat. Biotechnol. 2006, 24, 559–565. [Google Scholar]
- Kubo, T.; Zhelev, Z.; Ohba, H.; Bakalova, R. Modified 27-nt dsRNAs with dramatically enhanced stability in serum and long-term RNAi activity. Oligonucleotides 2007, 17, 445–464. [Google Scholar] [CrossRef]
- Kubo, T.; Zhelev, Z.; Bakalova, R.; Ohba, H. Highly efficient gene suppression by chemically modified 27 nucleotide double-stranded RNAs. Jpn. J. Appl. Phys. 2008, 47, 1346–1350. [Google Scholar] [CrossRef]
- Kubo, T.; Zhelev, Z.; Bakalova, R.; Ohba, H. Enhancement of gene silencing potency and nuclease stability by chemically modified duplex RNA. Nucleic Acids Symp. Ser. 2007, 51, 407–408. [Google Scholar] [CrossRef]
- Pasquinelli, A.E.; Powers, M.A.; Lund, E.; Forbes, D.; Dahlberg, J.E. Inhibition of mRNA export in vertebrate cells by nuclear export signal conjugates. Proc. Natl. Acad. Sci. USA 1997, 94, 14394–14399. [Google Scholar]
- Chiu, Y.L.; Rana, T.M. RNAi in human cells: Basic structural and functional features of small interfering RNA. Mol. Cell. 2002, 10, 549–561. [Google Scholar] [CrossRef]
- Jeong, J.H.; Mok, H.; Oh, Y.K.; Park, T.G. siRNA conjugate delivery systems. Bioconjug. Chem. 2009, 20, 5–14. [Google Scholar] [CrossRef]
- Sample Availability: Not available.
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kubo, T.; Yanagihara, K.; Sato, Y.; Morita, Y.; Seyama, T. Enhancement of Gene Silencing Effect and Membrane Permeability by Peptide-Conjugated 27-Nucleotide Small Interfering RNA. Molecules 2012, 17, 11089-11102. https://doi.org/10.3390/molecules170911089
Kubo T, Yanagihara K, Sato Y, Morita Y, Seyama T. Enhancement of Gene Silencing Effect and Membrane Permeability by Peptide-Conjugated 27-Nucleotide Small Interfering RNA. Molecules. 2012; 17(9):11089-11102. https://doi.org/10.3390/molecules170911089
Chicago/Turabian StyleKubo, Takanori, Kazuyoshi Yanagihara, Yuichiro Sato, Yasuhiro Morita, and Toshio Seyama. 2012. "Enhancement of Gene Silencing Effect and Membrane Permeability by Peptide-Conjugated 27-Nucleotide Small Interfering RNA" Molecules 17, no. 9: 11089-11102. https://doi.org/10.3390/molecules170911089
APA StyleKubo, T., Yanagihara, K., Sato, Y., Morita, Y., & Seyama, T. (2012). Enhancement of Gene Silencing Effect and Membrane Permeability by Peptide-Conjugated 27-Nucleotide Small Interfering RNA. Molecules, 17(9), 11089-11102. https://doi.org/10.3390/molecules170911089





