Homogeneous Adaboost Ensemble Machine Learning Algorithms with Reduced Entropy on Balanced Data
Abstract
1. Introduction
2. Related Works
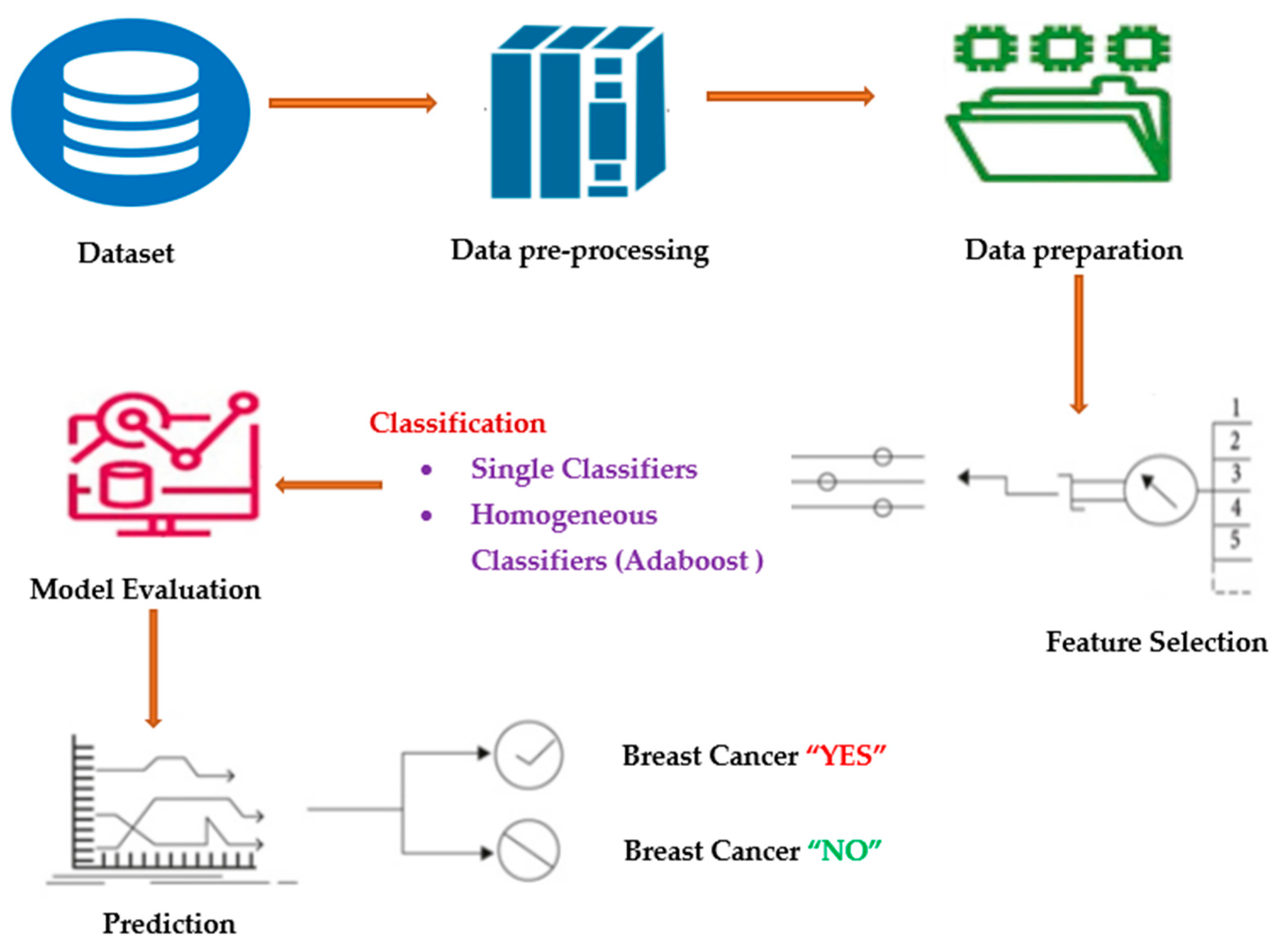
3. Proposed Methodology
3.1. Data-Preprocessing
3.2. Adaboost Classifier
| Algorithm 1: Adaboost classifier—pseudo code |
| Input: Let D be the dataset that includes {(a1,b1), (a2,b2), ….. (am, bm)}; |
| Let λ be the learning (base) algorithm |
| Let T be the total No. of learning rounds. |
| Process: |
| D1(i) = 1/m |
| for time = 1, …, T; |
| ht = λ (D, Dt); weak learner is trained with Distribution Dt |
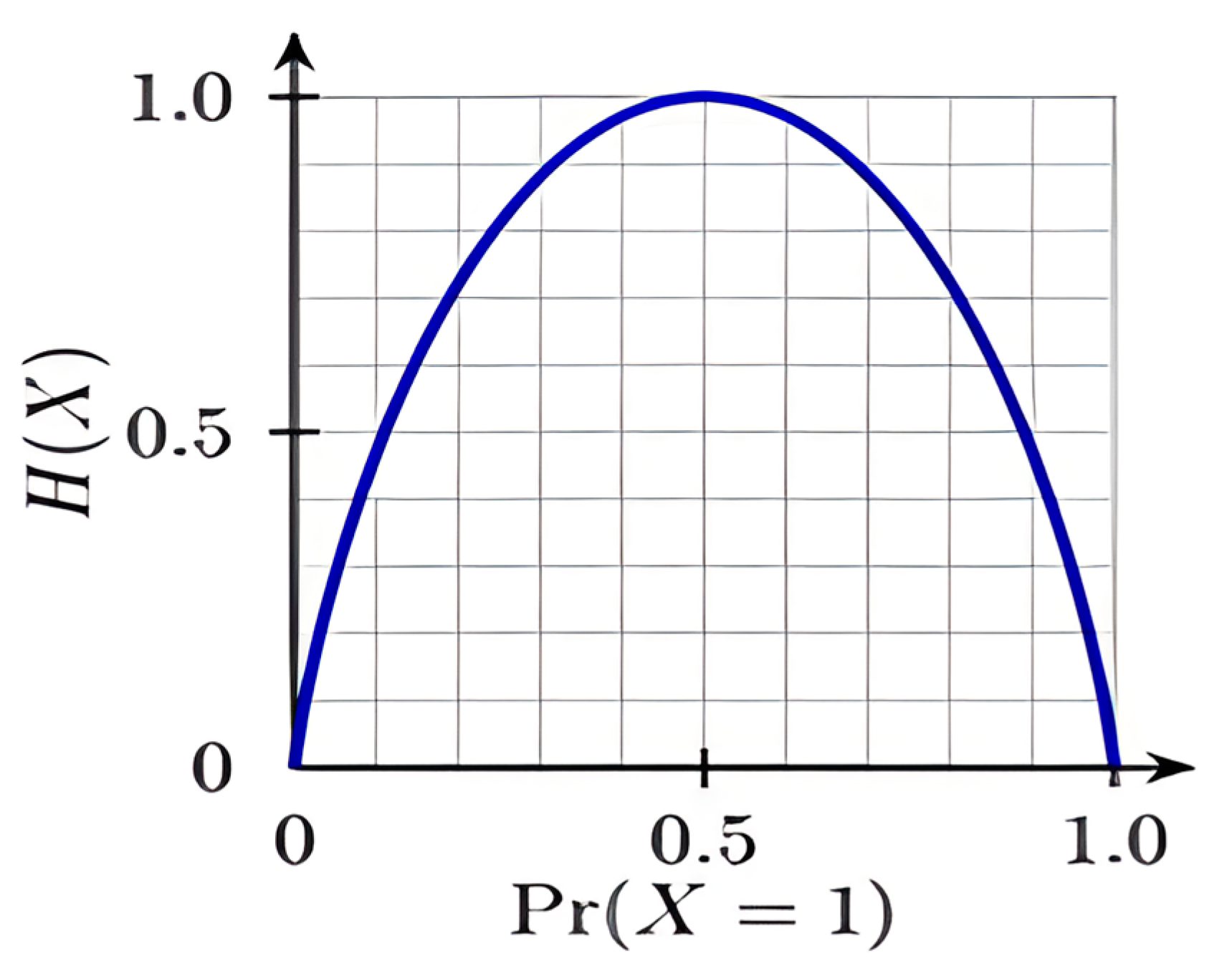
| t = [ (aii)]; Error measure (entropy) |
| = ln ( ht |
| Dt+1(i) = ∗ |
| = |
| Outcome: H(a) = sign ( |
| Algorithm 2: LogitBoost |
| Input: Dataset (Training) |
| 1. Initialize: = (b(i) + 1)/2, G() = 0, and p() = 0.5 |
| 2. For time = 1…..,T |
| 2.1a Calculate the working response as well as weights |
| = , |
| 2.1b Fit using weights |
| 2.1c Update G() ← G() + and |
| Output: G(a) = |
3.3. Alternating Decision-Tree (ADTree)
| Algorithm 3: ADTree with Adaboost |
| Input: Training Dataset -D |
| 1. Process of Initialization |
| 1.a Set = 0 = 1/n = {true} |
| 1.b First DT rule (x): { if (true) then = ln ( else 0} |
| 1.c Update = 1 = = 0 exp |
| 2. Do it again for boosting cycle t = 1:T |
| 2.1 For every pre-condition and each condition |
| Z + + W( |
| 2.1 Compute and for the selected that minimizes Z with = 1 |
| = ln (), = =l n() |
| 2.2 Update : { , |
| 2.3 Update = exp(- ( |
| Output: F(x) = |
3.4. Reduced Error Pruning Tree (REPTree)
3.5. Naïve Bayes (NB) Classifier
3.6. Random Forest (RF) Classifier
3.7. CART
3.8. Homogeneous Adaboost Technique
4. Results and Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Dhar, P. A Method to Detect Breast Cancer Based on Morphological Operation. Int. J. Educ. Manag. Eng. 2021, 11, 25–31. [Google Scholar] [CrossRef]
- Lu, Y.; Li, J.-Y.; Su, Y.-T.; Liu, A.-A. A Review of Breast Cancer Detection in Medical Images. In Proceedings of the 2018 IEEE Visual Communications and Image Processing (VCIP), Taichung, Taiwan, 9–12 December 2018; pp. 1–4. [Google Scholar]
- Shwetha, S.V.; Dharmanna, L. An Automatic Recognition, Identification and Classification of Mitotic Cells for the Diagnosis of Breast Cancer Stages. Int. J. Image Graph. Sign. Process 2021, 13, 1–11. [Google Scholar] [CrossRef]
- Khourdifi, Y.; Bahaj, M. Feature Selection with Fast Correlation-Based Filter for Breast Cancer Prediction and Classification Using Machine Learning Algorithms. In Proceedings of the 2018 International Symposium on Advanced Electrical and Communication Technologies (ISAECT), Rabat, Morocco, 21–23 November 2018; pp. 1–6. [Google Scholar]
- Chaudhuri, A.K.; Banerjee, D.K.; Das, A. A Dataset Centric Feature Selection and Stacked Model to Detect Breast Cancer. Int. J. Intell. Syst. Appl. 2021, 13, 24–37. [Google Scholar] [CrossRef]
- Neumayer, L.; Viscusi, R.K. 37—Assessment and Designation of Breast Cancer Stage. In The Breast, 5th ed.; Bland, K.I., Copeland, E.M., Klimberg, V.S., Gradishar, W.J., Eds.; Elsevier: Alpharetta, GA, USA, 2018; pp. 531–552.e6. ISBN 978-0-323-35955-9. [Google Scholar]
- Reddy, P.C.; Chandra, R.; Vadiraj, P.; Reddy, M.A.; Mahesh, T.; Madhuri, G.S. Detection of Plant Leaf-based Diseases Using Machine Learning Approach. In Proceedings of the 2021 IEEE International Conference on Computation System and Information Technology for Sustainable Solutions (CSITSS), Bangalore, India, 16–18 December 2021; pp. 1–4. [Google Scholar] [CrossRef]
- Makrariya, A.; Kumar Shandilya, S.; Shandilya, S.; Gregus, M.; Izonin, I.; Makrariya, R. Mathematical Simulation of Behavior of Female Breast Consisting Malignant Tumor During Hormonal Changes. IEEE Access 2022, 10, 91346–91355. [Google Scholar] [CrossRef]
- Islam, M.; Haque, R.; Iqbal, H.; Hasan, M.; Hasan, M.; Kabir, M.N. Breast Cancer Prediction: A Comparative Study Using Machine Learning Techniques. SN Comput. Sci. 2020, 1, 290. [Google Scholar] [CrossRef]
- Kaur, P.; Pruthi, Y.; Bhatia, V.; Singh, J. Empirical Analysis of Cervical and Breast Cancer Prediction Systems Using Classification. Int. J. Educ. Manag. Eng. 2019, 9, 1–15. [Google Scholar] [CrossRef]
- Jha, K.K.; Jha, R.; Jha, A.; Hassan, M.; Yadav, S.; Mahesh, T. A Brief Comparison on Machine Learning Algorithms Based on Various Applications: A Comprehensive Survey. In Proceedings of the 2021 IEEE International Conference on Computation System and Information Technology for Sustainable Solutions (CSITSS), Bangalore, India, 16–18 December 2021; pp. 1–5. [Google Scholar] [CrossRef]
- Li, J.; Liu, H.; Ng, S.K.; Wong, L. Discovery of Significant Rules for Classifying Cancer Diagnosis Data. Bioinformatics 2003, 19 (Suppl. 2), ii93–ii102. [Google Scholar] [CrossRef]
- Owoseni, A.T.; Olabode, O.; Akintola, K.G. Comparative Descriptive Analysis of Breast Cancer Tissues Using K-Means and SelfOrganizing Map. Int. J. Inf. Technol. Comput. Sci. 2018, 10, 46–55. [Google Scholar] [CrossRef]
- Reddy, A.M.; Reddy, K.S.; Jayaram, M.; Venkata Maha Lakshmi, N.; Aluvalu, R.; Mahesh, T.R.; Kumar, V.V.; Stalin Alex, D. An Efficient Multilevel Thresholding Scheme for Heart Image Segmentation Using a Hybrid Generalized Adversarial Network. J. Sens. 2022, 2022, 4093658. [Google Scholar] [CrossRef]
- Akbugday, B. Classification of Breast Cancer Data Using Machine Learning Algorithms. In Proceedings of the 2019 Medical Technologies Congress (TIPTEKNO), Izmir, Turkey, 3–5 October 2019; pp. 1–4. [Google Scholar]
- Alwidian, J.; Hammo, B.H.; Obeid, N. WCBA: Weighted Classification Based on Association Rules Algorithm for Breast Cancer Disease. Appl. Soft Comput. 2018, 62, 536–549. [Google Scholar] [CrossRef]
- Reddy, K.H.K.; Luhach, A.; Kumar, V.; Pratihar, S.; Kumar, D.; Roy, D.S. Towards energy efficient Smart city services: A software defined resource management scheme for data centers. Sustain. Comput. Inform. Syst. 2022, 35, 100776. [Google Scholar] [CrossRef]
- Senthilkumar, B.; Zodinpuii, D.; Pachuau, L.; Chenkual, S.; Zohmingthanga, J.; Kumar, N.S.; Hmingliana, L. Ensemble Modelling for Early Breast Cancer Prediction from Diet and Lifestyle. IFAC-PapersOnLine 2022, 55, 429–435. [Google Scholar] [CrossRef]
- Roopashree, S.; Anitha, J.; Mahesh, T.; Kumar, V.V.; Viriyasitavat, W.; Kaur, A. An IoT based authentication system for therapeutic herbs measured by local descriptors using machine learning approach. Measurement 2022, 200, 111484, ISSN 0263-2241. [Google Scholar] [CrossRef]
- Patel, A. Benign vs Malignant Tumors. JAMA Oncol. 2020, 6, 1488. [Google Scholar] [CrossRef] [PubMed]
- Qiu, M.; Shi, S.; Chen, Z.; Yu, H.; Sheng, H.; Jin, Y.; Wang, D.; Wang, F.; Li, Y.; Xie, D.; et al. Frequency and Clinicopathological Features of Metastasis to Liver, Lung, Bone, and Brain from Gastric Cancer: A SEER-based Study. Cancer Med. 2018, 7, 3662–3672. [Google Scholar] [CrossRef]
- Hussain, L.; Saeed, S.; Awan, I.A.; Idris, A.; Nadeem, M.S.A.; Chaudhry, Q.-U.-A. Detecting Brain Tumor Using Machines Learning Techniques Based on Different Features Extracting Strategies. Curr. Med. Imaging Rev. 2019, 15, 595–606. [Google Scholar] [CrossRef]
- Schneble, E.J.; Graham, L.J.; Shupe, M.P.; Flynt, F.L.; Banks, K.P.; Kirkpatrick, A.D.; Nissan, A.; Henry, L.; Stojadinovic, A.; Shumway, N.M.; et al. Current Approaches and Challenges in Early Detection of Breast Cancer Recurrence. J. Cancer 2014, 5, 281–290. [Google Scholar] [CrossRef]
- Mahesh, T.R.; Vinoth Kumar, V.; Muthukumaran, V.; Shashikala, H.K.; Swapna, B.; Guluwadi, S. Performance Analysis of XGBoost Ensemble Methods for Survivability with the Classification of Breast Cancer. J. Sens. 2022, 2022, e4649510. [Google Scholar] [CrossRef]
- Chakraborty, V.; Sundaram, M. An Efficient Smote-Based Model for Dyslexia Prediction. J. Inf. Eng. Electron. Bus. 2021, 13, 13–21. [Google Scholar] [CrossRef]
- Rahman, M.; Zhou, Y.; Wang, S.; Rogers, J. Wart Treatment Decision Support Using Support Vector Machine. Int. J. Intell. Syst. Appl. 2020, 12, 1–11. [Google Scholar] [CrossRef]
- Mahesh, T.R.; Kumar, D.; Vinoth Kumar, V.; Asghar, J.; Mekcha Bazezew, B.; Natarajan, R.; Vivek, V. Blended Ensemble Learning Prediction Model for Strengthening Diagnosis and Treatment of Chronic Diabetes Disease. Comput. Intell. Neurosci. 2022, 2022, e4451792. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.; Ivashchenko, M.; Lyushenko, L.; Klyushnyk, D. Artificial Neural Network Training Criterion Formulation Using Error Continuous Domain. Int. J. Mod. Educ. Comput. Sci. 2021, 13, 13–22. [Google Scholar] [CrossRef]
- Hu, Z.; Tereykovski, I.A.; Tereykovska, L.O.; Pogorelov, V.V. Determination of Structural Parameters of Multilayer Perceptron Designed to Estimate Parameters of Technical Systems. Int. J. Intell. Syst. Appl. 2017, 9, 57–62. [Google Scholar] [CrossRef]
- Mahesh, T.R.; Vinoth Kumar, V.; Vivek, V.; Karthick Raghunath, K.M.; Sindhu Madhuri, G. Early predictive model for breast cancer classification using blended ensemble learning. Int. J. Syst. Assur. Eng. Manag. 2022. [CrossRef]
- Chaurasia, V.; Pal, S.; Tiwari, B.B. Prediction of benign and malignant breast cancer using data mining techniques. J. Algorithm Comput. Technol. 2018, 12, 119–126. [Google Scholar] [CrossRef]
- Islam, M.; Iqbal, H.; Haque, R.; Hasan, K. Prediction of breast cancer using support vector machine and K-nearest neighbors. In Proceedings of the IEEE Region 10 Humanitarian Technology Conf (R10-HTC), Dhaka, Bangladesh, 21–23 December 2017. [Google Scholar]
- Jayasuruthi, L.; Shalini, A.; Kumar, V.V. Application of Rough Set Theory in Data Mining Market Analysis Using Rough Sets Data Explorer. J. Comput. Theor. Nanosci. 2018, 15, 2126–2130. [Google Scholar] [CrossRef]
- Asri, H.; Mousannif, H.; Moatassime, H.A.; Noel, T. Using machine learning algorithms for breast cancer risk prediction and diagnosis. In Proceedings of the 6th International Symposium on Frontiers in Ambient and Mobile Systems (FAMS), Madrid, Spain, 23–26 May 2016. [Google Scholar]
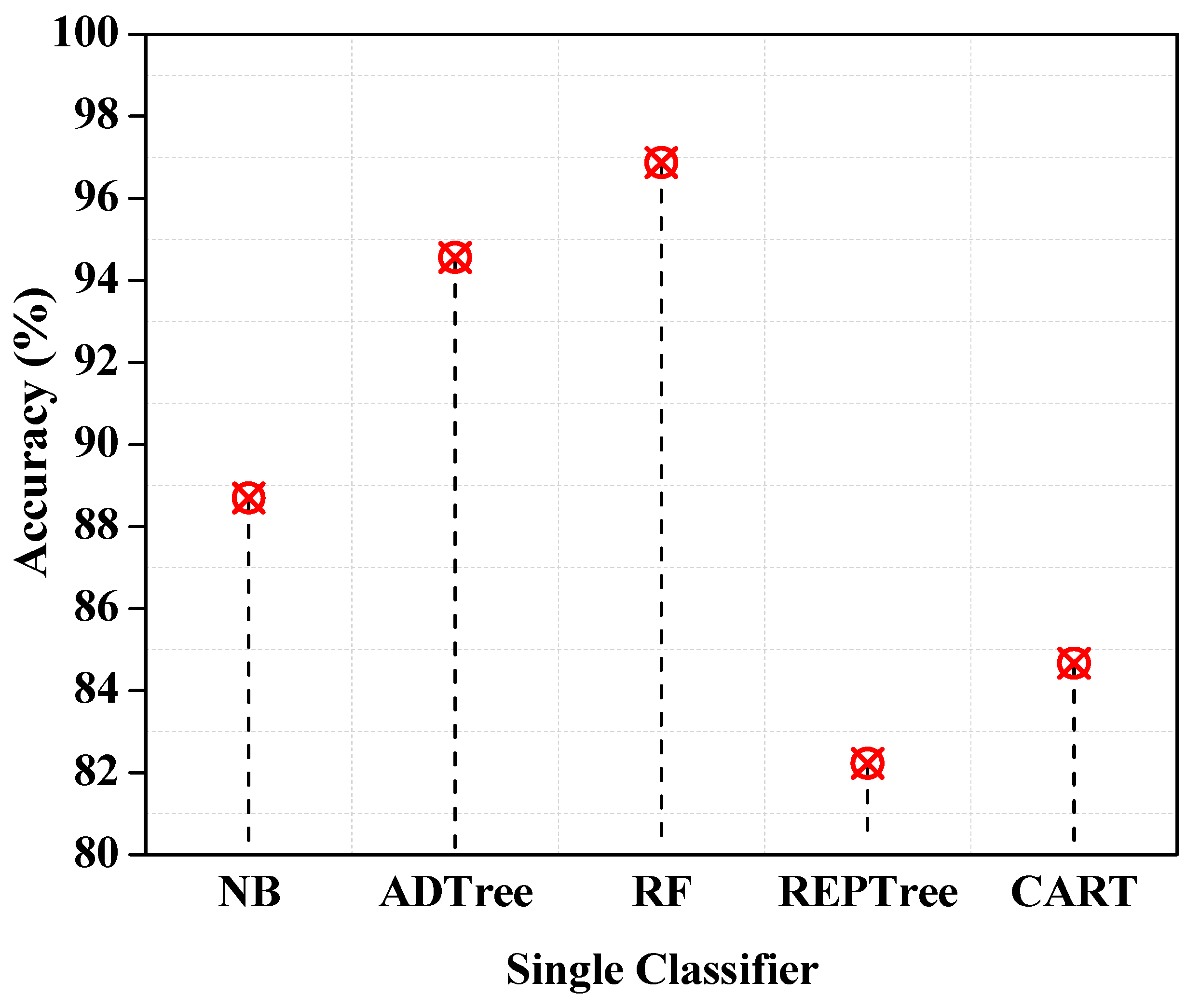


| Performance Metrics | NB | ADTree | RF | REPTree | CART |
|---|---|---|---|---|---|
| TTBM (s) | 5.77 | 58.28 | 2.11 | 11.76 | 55.55 |
| Accuracy (%) | 88.7 | 94.56 | 96.87 | 82.23 | 84.67 |
| F1-Score | 0.3 | 0.85 | 0.84 | 0.83 | 0.81 |
| RAE | 120 | 56.71 | 76.12 | 76.92 | 65.77 |
| MAE | 0.65 | 0.24 | 0.24 | 0.7 | 0.26 |
| RRSE | 137.51 | 94.33 | 80.72 | 96. 79 | 96.44 |
| RMSE | 0.72 | 0.36 | 0.37 | 0.42 | 0.55 |
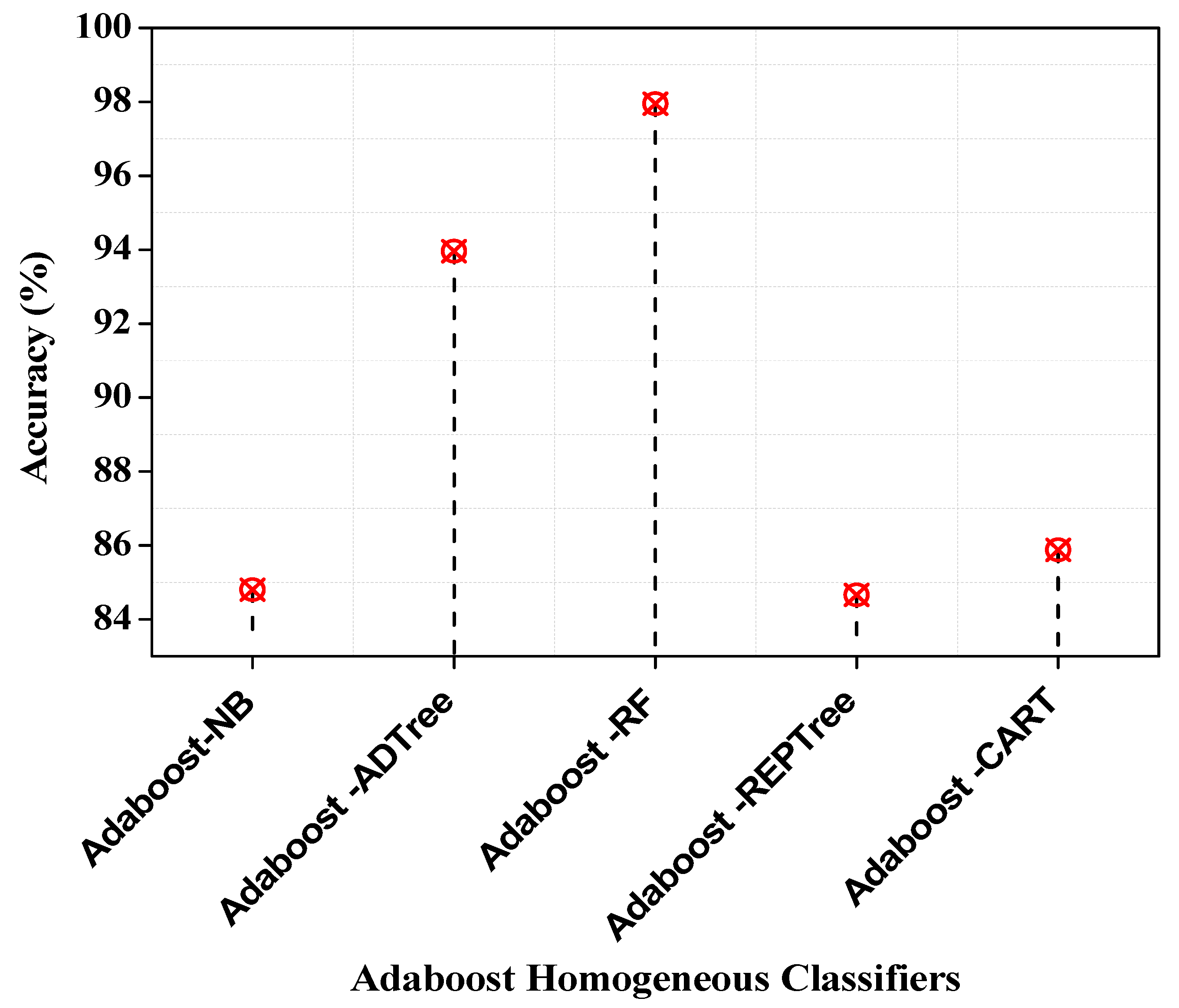
| Performance Metrics | Adaboost-NB | Adaboost-ADTree | Adaboost-RF | Adaboost-REPTree | Adaboost-CART |
|---|---|---|---|---|---|
| TTBM (s) | 18.32 | 30.21 | 8.52 | 61.44 | 200.12 |
| Accuracy (%) | 84.8 | 93.96 | 97.95 | 84.66 | 85.88 |
| F1-Score | 0.70 | 0.87 | 0.98 | 0.94 | 0.79 |
| MAE | 0.51 | 0.18 | 0.11 | 0.18 | 0.17 |
| RMSE | 0.69 | 0.39 | 0.33 | 0.38 | 0.37 |
| RAE | 107.44 | 55.64 | 33.77 | 44.29 | 42.48 |
| RRSE | 135.62 | 91.12 | 61.33 | 91.33 | 89.66 |
| Study and Year | Sampling Strategy | Accuracy |
|---|---|---|
| Alzubaidi A et al. 2016 [30] | 70–30% training–testing | 97.0% |
| Chaurasia V et al. 2018 [31] | 10-fold cross validation | 97.36% |
| Islam et al. 2017 [32] | 10-fold cross validation | 97.0% |
| Walid Theib Mohammad et. 2022 [33] | 70–30% training–testing | 97.7% |
| Asri et al. 2016 [34] | 10-fold cross validation | 97.13% |
| Proposed Method (Adaboost-RF) | 70–30% training–testing | 97.95% |
| 95% CI | 96.5–98.6% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramakrishna, M.T.; Venkatesan, V.K.; Izonin, I.; Havryliuk, M.; Bhat, C.R. Homogeneous Adaboost Ensemble Machine Learning Algorithms with Reduced Entropy on Balanced Data. Entropy 2023, 25, 245. https://doi.org/10.3390/e25020245
Ramakrishna MT, Venkatesan VK, Izonin I, Havryliuk M, Bhat CR. Homogeneous Adaboost Ensemble Machine Learning Algorithms with Reduced Entropy on Balanced Data. Entropy. 2023; 25(2):245. https://doi.org/10.3390/e25020245
Chicago/Turabian StyleRamakrishna, Mahesh Thyluru, Vinoth Kumar Venkatesan, Ivan Izonin, Myroslav Havryliuk, and Chandrasekhar Rohith Bhat. 2023. "Homogeneous Adaboost Ensemble Machine Learning Algorithms with Reduced Entropy on Balanced Data" Entropy 25, no. 2: 245. https://doi.org/10.3390/e25020245
APA StyleRamakrishna, M. T., Venkatesan, V. K., Izonin, I., Havryliuk, M., & Bhat, C. R. (2023). Homogeneous Adaboost Ensemble Machine Learning Algorithms with Reduced Entropy on Balanced Data. Entropy, 25(2), 245. https://doi.org/10.3390/e25020245







