Influence of Ectopic Beats on Heart Rate Variability Analysis
Abstract
1. Introduction
2. Methods
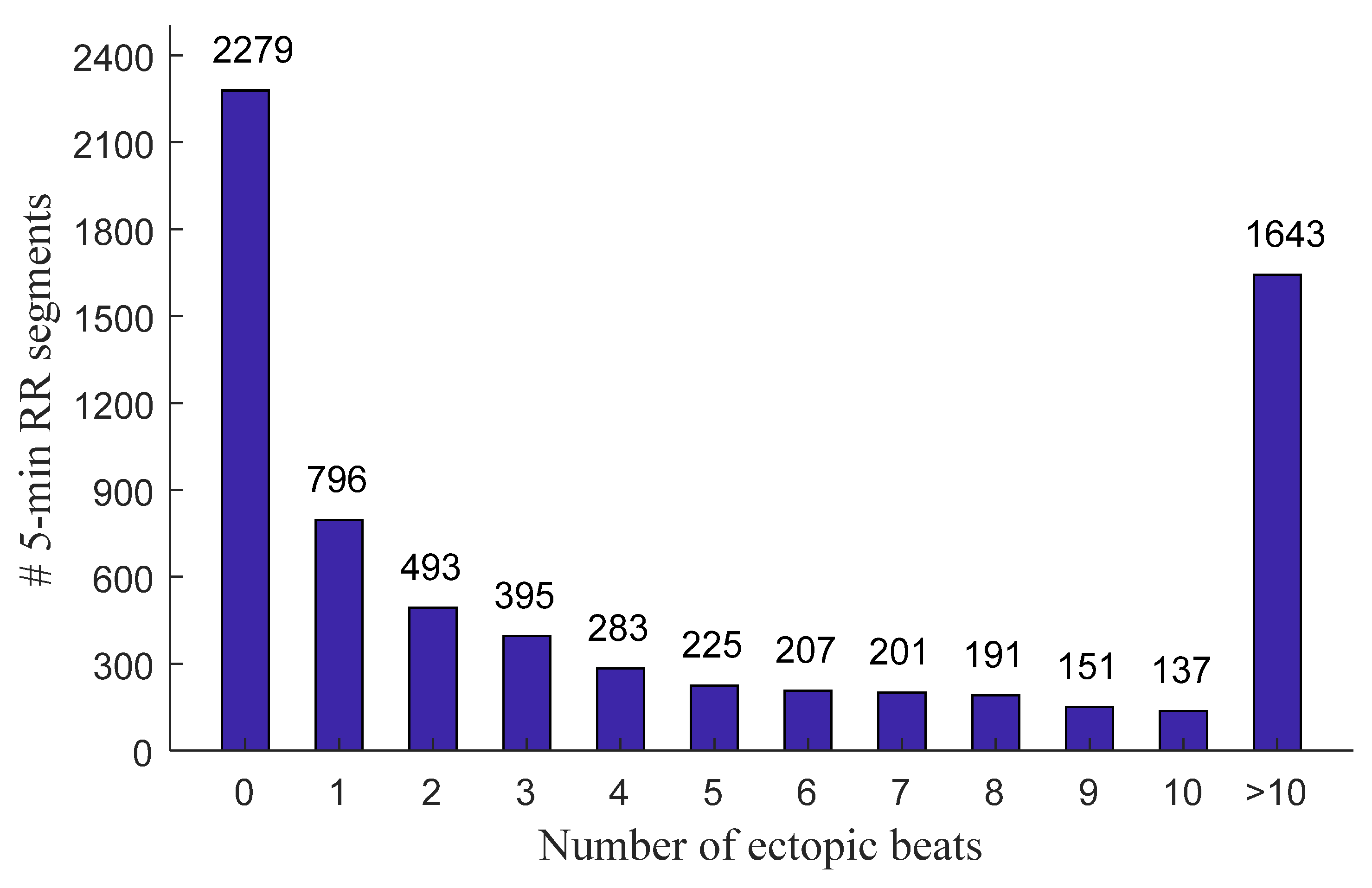
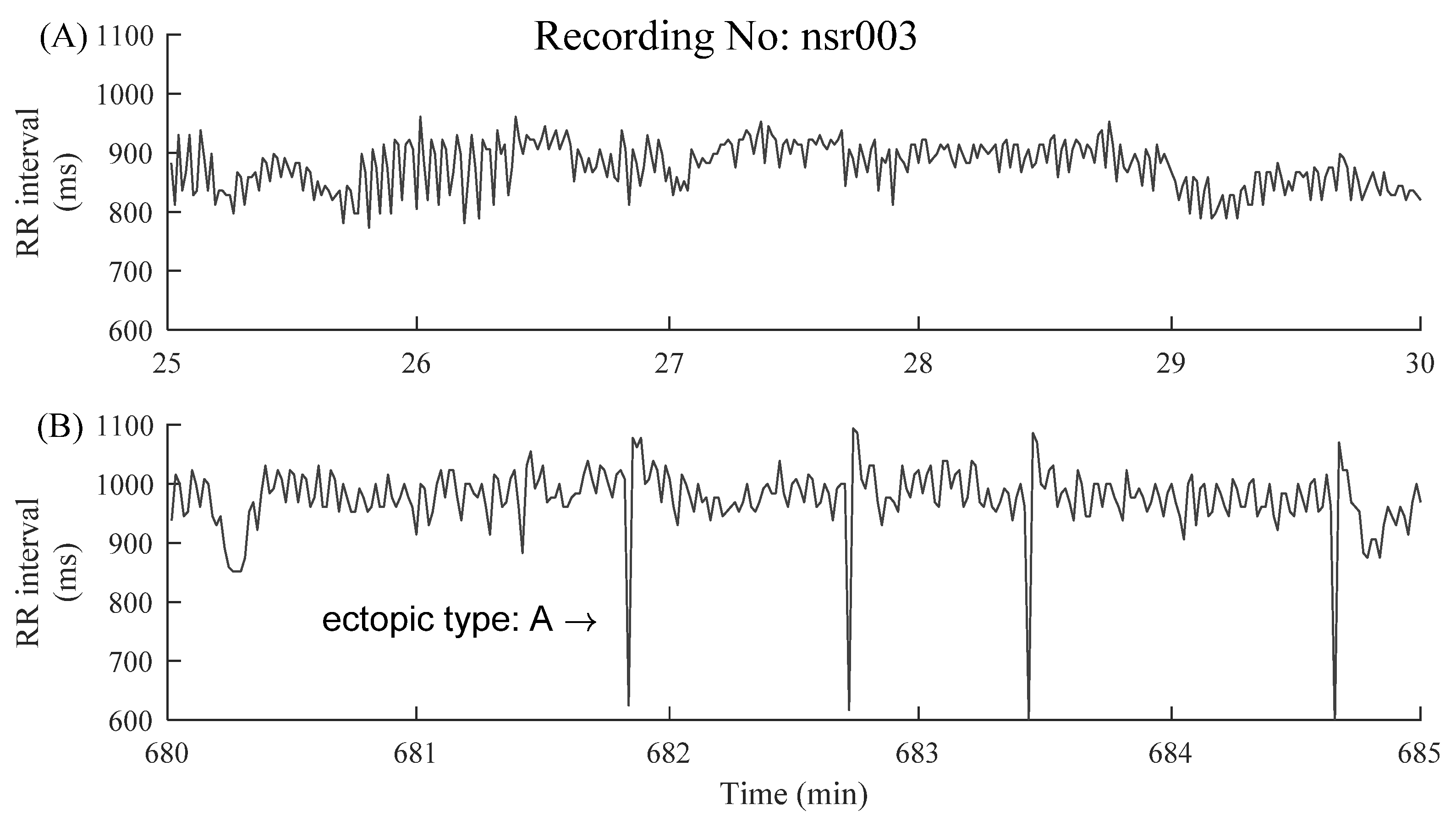
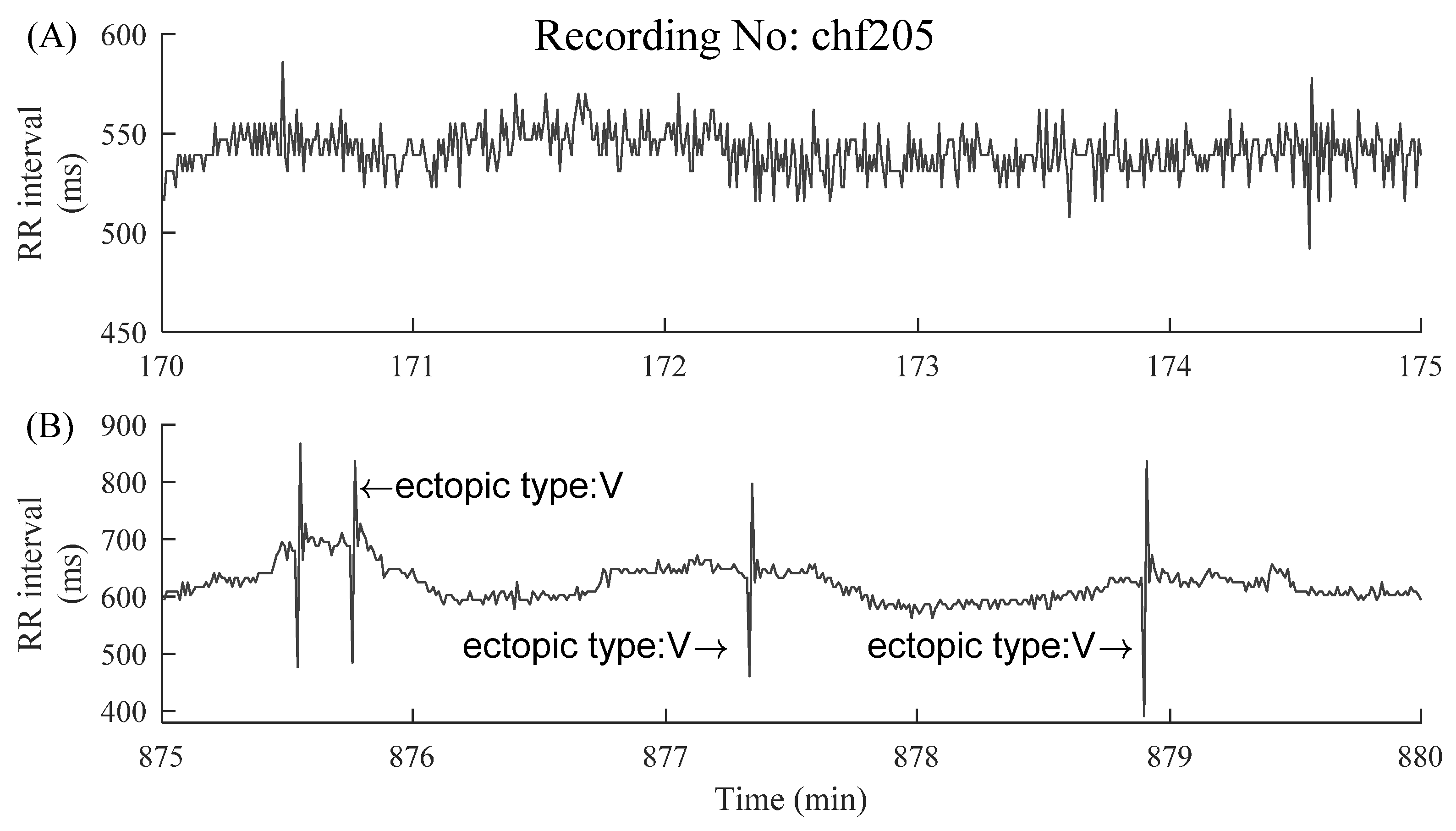
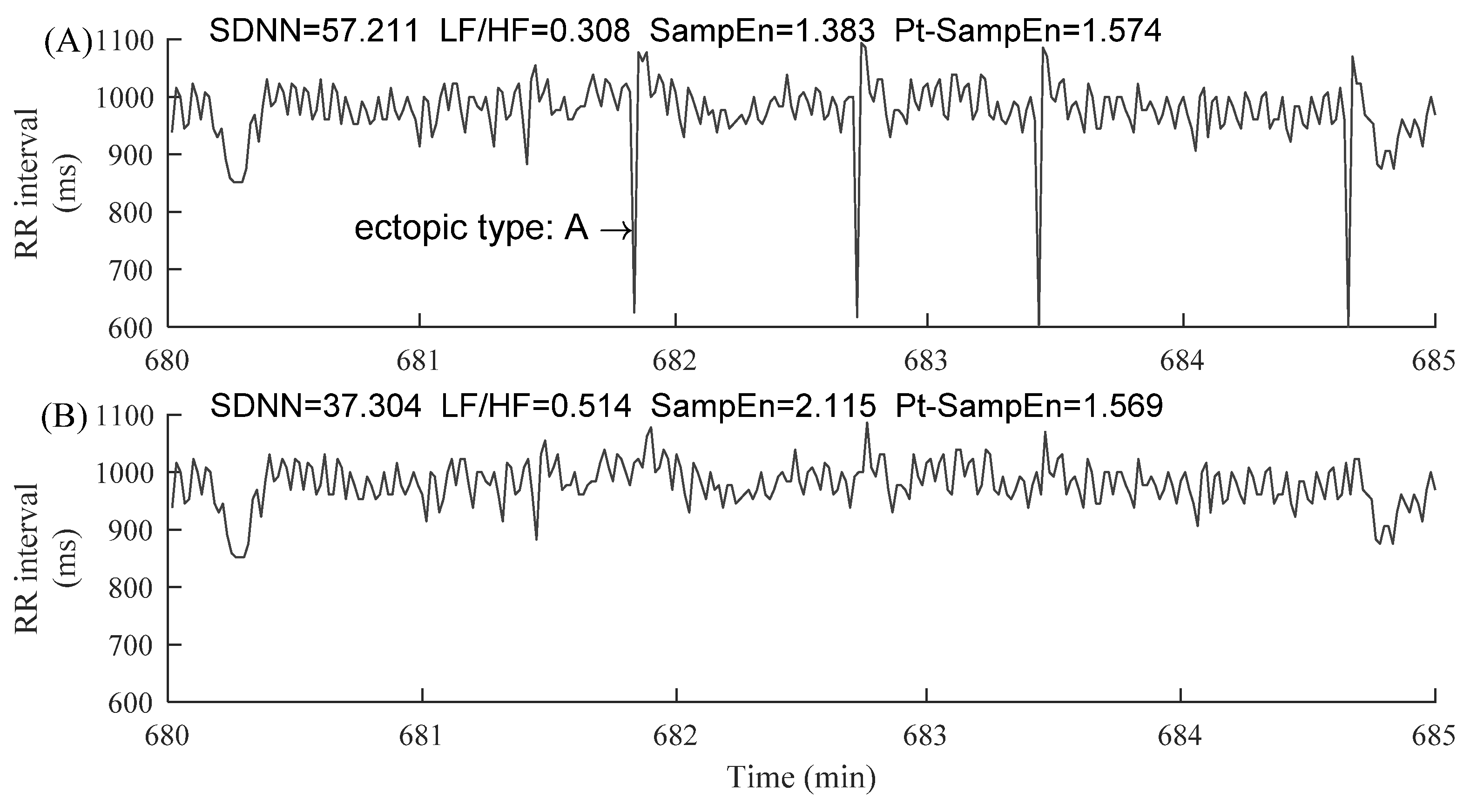
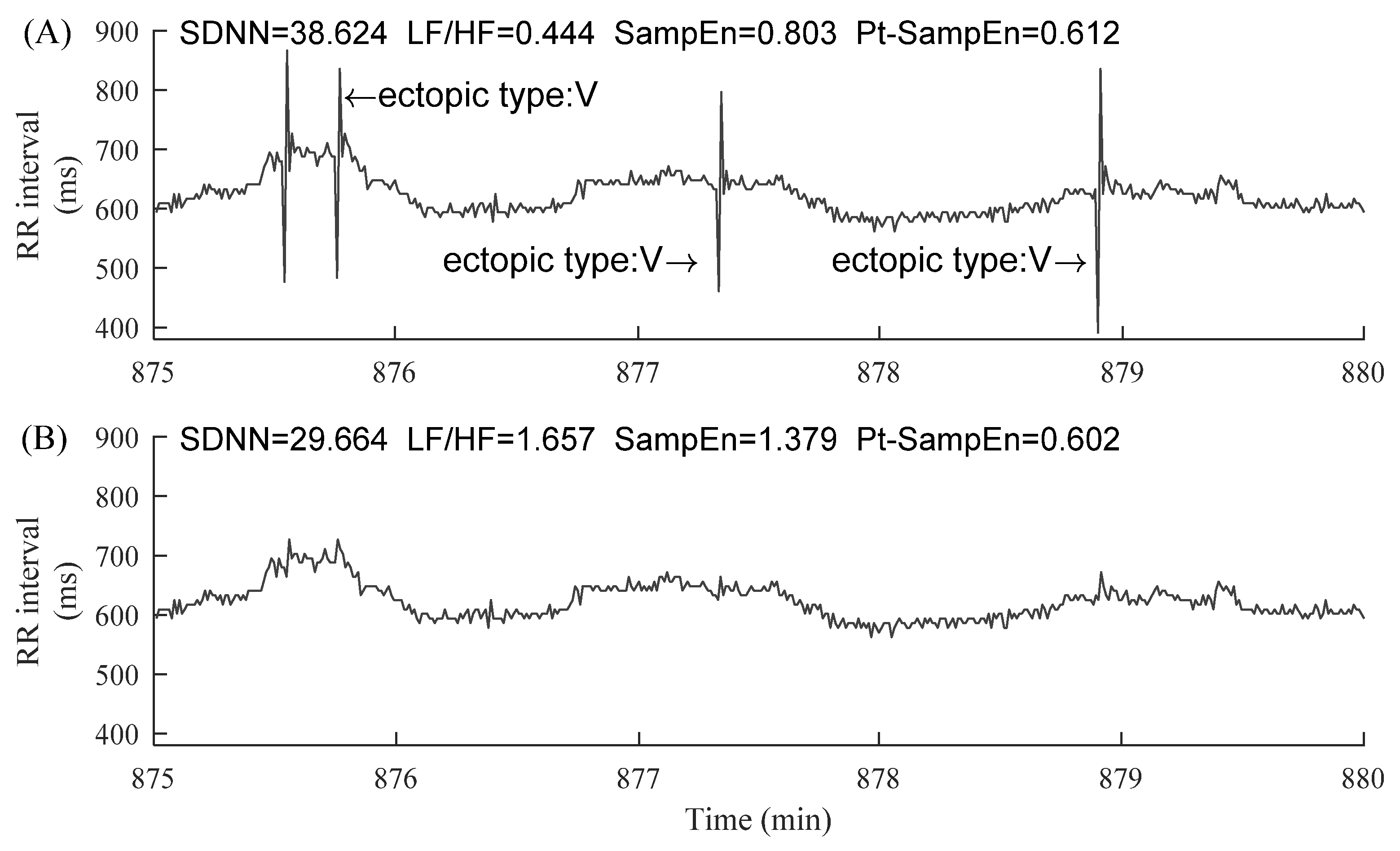
2.1. Data and Preprocessing
2.2. HRV Indices
- SDNN: SDNN is the standard deviation of a 5 min RR segment, and was calculated as follows:where is the ith RR interval, is the mean value of all RR intervals, N is the number of RR intervals.
- LF/HF: The frequency-domain features were calculated using power spectral density. Prior to the frequency-domain analysis, spline interpolation was used to resample the RR intervals time series evenly at 4 Hz. HRV spectrum was produced using Burg’s method with an order of 10 [24]. It was then decomposed into two separate frequency bands: a low-frequency band (LF, 0.04 to 0.15 Hz) and a high-frequency band (HF, 0.15 to 0.4 Hz). The ratio of low-frequency power to high-frequency power (LF/HF) was calculated, which reflected the balance between the sympathetic and parasympathetic (or vagal) activity.
- Sample entropy: SampEn is a widely used index for HRV analysis and can reflect the inherent complexity or regularity of RR interval time series. The calculation for SampEn was summarized as follows [20]:For a 5 min RR segment , given the parameters m and r, first form the vector sequences , which represented m consecutive values:The distance between and was defined using the maximum absolute difference:For each , denote as (N-m)-1 times the number of (, ) that meets . Similarly, set is times the number of that meets for all .Then SampEn is defined by:In the current study, the parameter settings for SampEn used the recommendation from our previous study [22], i.e., embedding dimension m = 2 and tolerance threshold r = 0.10 times the standard deviation (SD) of the time series.
- Physical threshold-based SampEn (Pt-SampEn): For entropy analysis, three intrinsic parameters (embedding dimension 𝑚, tolerance threshold r, and time-series length N) needed to be initialized. SampEn was reported not sensitive to the time series length if but sensitive to the parameter tolerance threshold. Since parameter m is based on the length N (), SampEn was also not sensitive to m. Tolerance threshold r was difficult to be determined and was recommended between 0.10 and 0.25 times the SD of the time series. However, in practice, if the r value was too small, the number of matched vectors was small, and, on the contrary, if the r value was too large, detailed information within the time series was ignored. RR interval time series usually have variable SD values, and it is not easy to find an appropriate r value to achieve an optimal result. Hence, simply use the suggested range of 0.10 to 0.25 times the SD. Herein, a physical threshold r was used to form a unified comparison baseline for determining the vector similarity, and, thus, the Pt-SampEn was developed in our previous study [28].
2.3. Evaluation Method
2.4. Statistical Analysis
3. Results
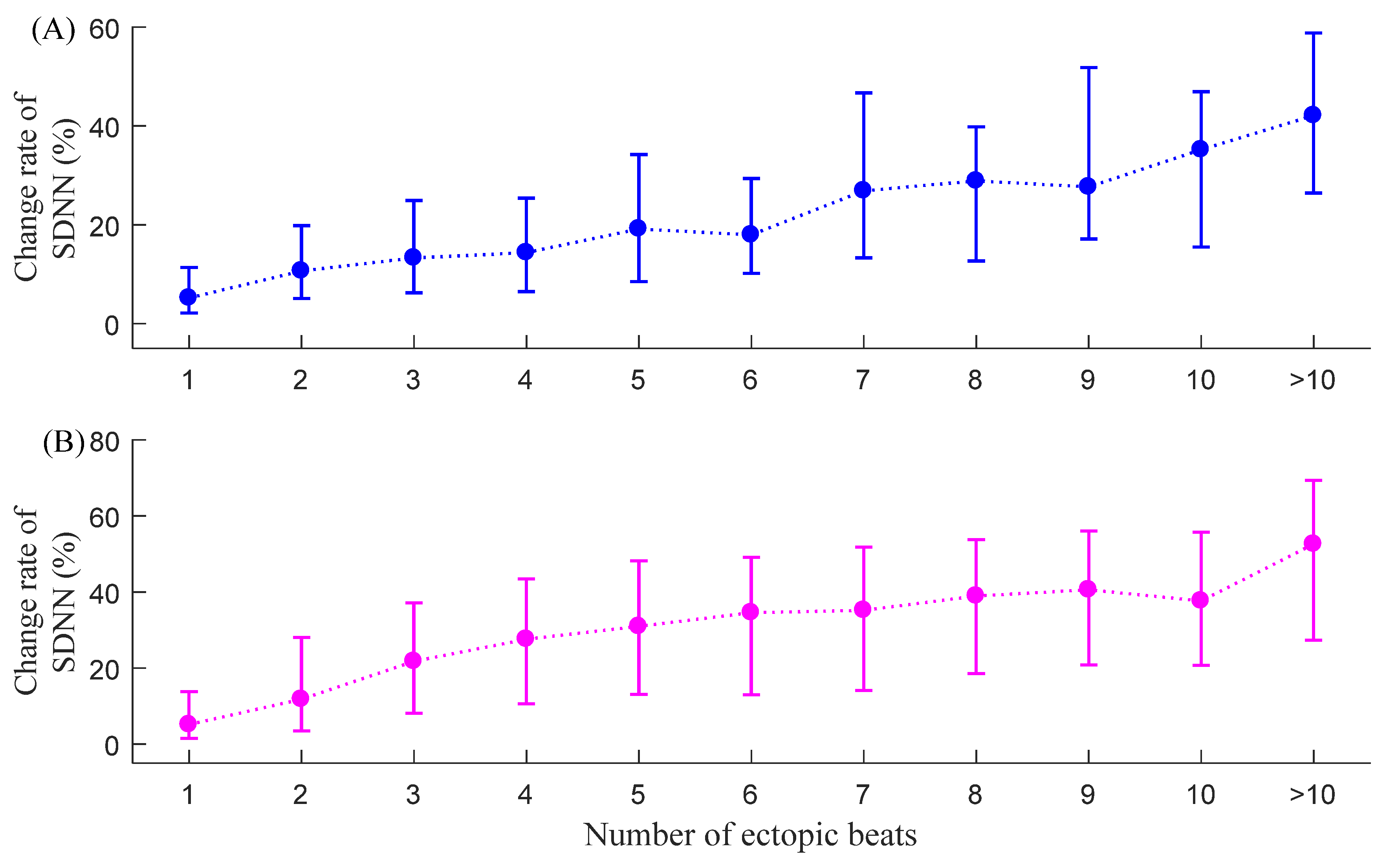
3.1. CSDNN
3.2. CLF/HF
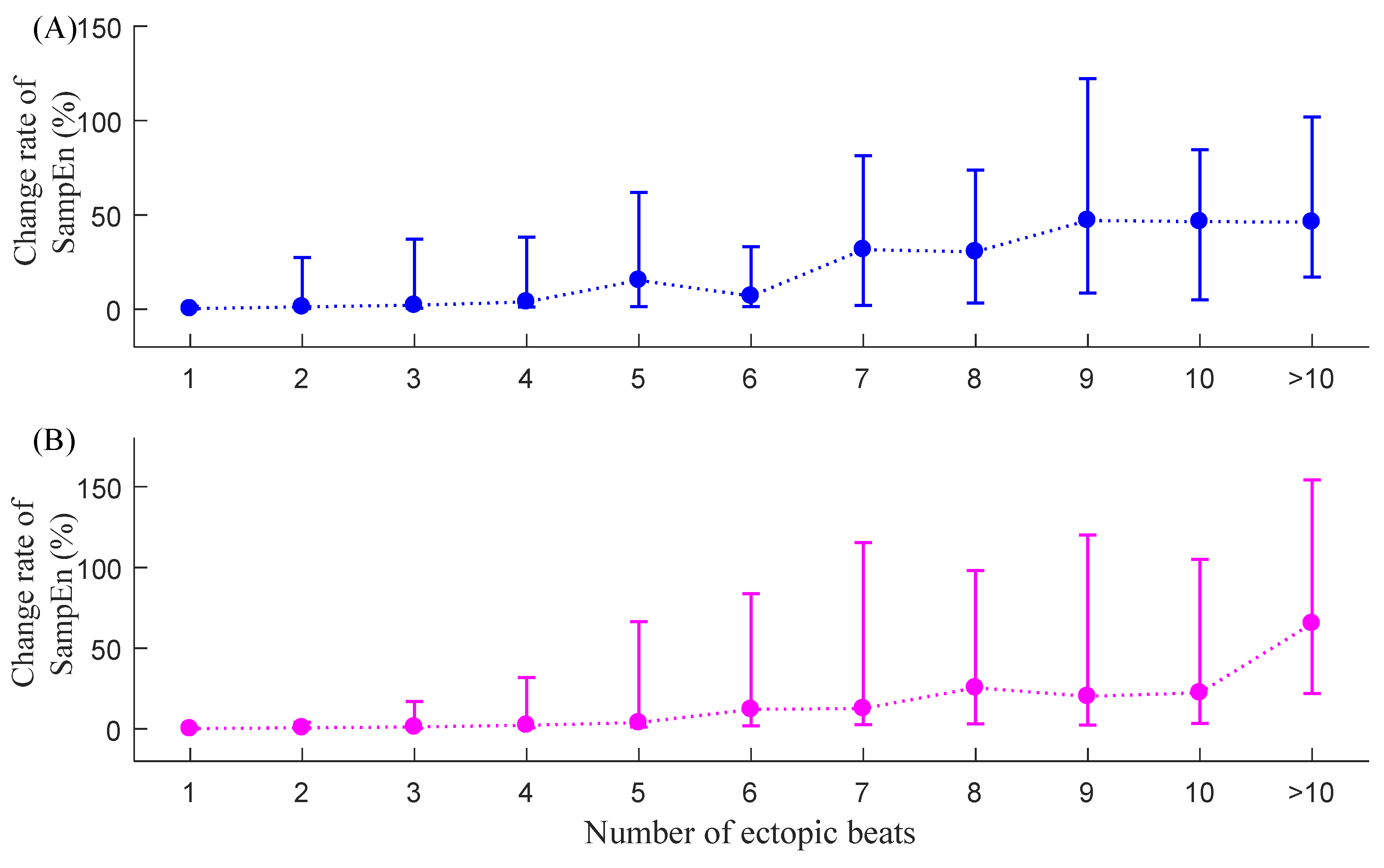
3.3. CSampEn
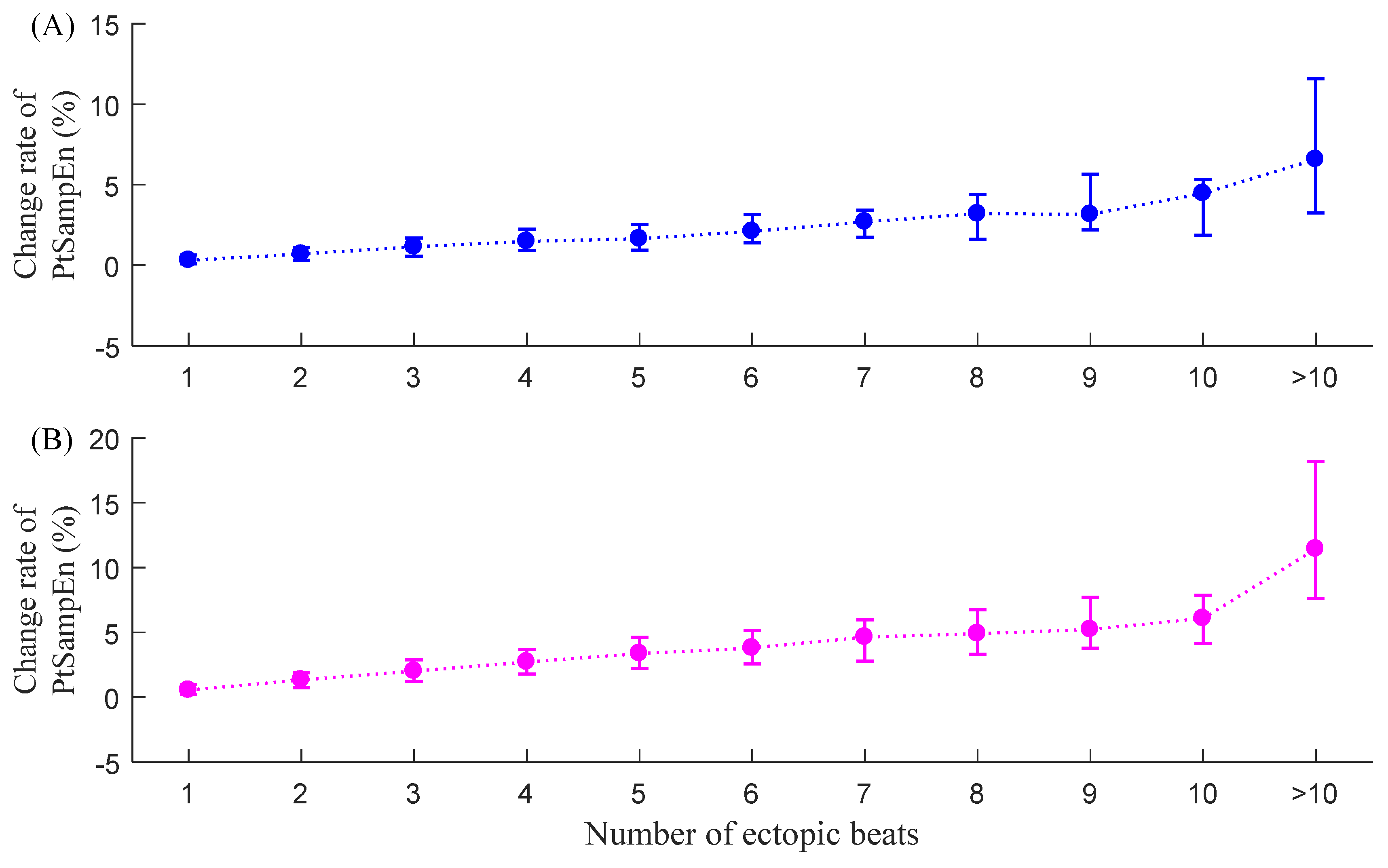
3.4. CPT-SampEn
3.5. Comparison of Variances of the Change Rate Index
4. Discussion and Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Hon, E.H.; Lee, S.T. Electronic evaluation of the fetal heart rate patterns preceeding fetal death: Further observations. Am. J. Obstet. Gynecol. 1965, 87, 814–826. [Google Scholar]
- Kobayashi, H.; Ishibashi, K.; Noguchi, H. Heart Rate Variability; An Index for Monitoring and Analyzing Human Autonomic Activities. Appl. Hum. Sci. J. Physiol. Anthr. 1999, 18, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Joseph, K.P.; Kannathal, N.; Lim, C.M.; Suri, J.S. Heart rate variability: A review. Med. Biol. Eng. Comput. 2006, 44, 1031–1051. [Google Scholar] [CrossRef] [PubMed]
- Anonymous. Task Force of the European Society of Cardiology and the North American Society of Pacing and Electrophysiology, Heart Rate Variability Standards of measurement, physiological interpretation, and clinical use. Circulation 1996, 93, 1043–1065. [Google Scholar] [CrossRef]
- Lanza, G.A.; Guido, V.; Galeazzi, M.; Mustilli, M.; Natali, R.; Ierardi, C.; Milici, C.; Burzotta, F.; Pasceri, V.; Tomassini, F.; et al. Prognostic role of heart rate variability in patients with a recent acute myocardial infarction. Am. J. Cardiol. 1998, 82, 1323–1328. [Google Scholar] [CrossRef]
- Liao, D.; Carnethon, M.; Evans, G.W.; Cascio, W.E.; Heiss, G. Lower Heart Rate Variability Is Associated With the Development of Coronary Heart Disease in Individuals With Diabetes: The Atherosclerosis Risk in Communities (ARIC) Study. Diabetes 2002, 51, 3524–3531. [Google Scholar] [CrossRef]
- Ding, L.F.; Wang, L. Relationship between LVMI and HRV in patients with primary hypertension. J. Clin. Med. Pract. 2016, 1, 29–31. [Google Scholar]
- Salo, M.A.; Huikuri, H.V.; Seppanen, T. Ectopic Beats in Heart Rate Variability Analysis: Effects of Editing on Time and Frequency Domain Measures. Ann. Noninvasive Electrocardiol. 2001, 6, 5–17. [Google Scholar] [CrossRef]
- Singh, B.; Singh, D.; Jaryal, A.K.; Deepak, K.K. Ectopic beats in approximate entropy and sample entropy-based HRV assess-ment. Int. J. Syst. Sci. 2012, 43, 884–893. [Google Scholar] [CrossRef]
- Liu, C.; Zhao, L.; Cai, Z.; Liu, F.; Li, Y.; Wei, S.; Li, J.; Murray, A. Effect of Ectopic Beats on Heart Rate Variability Indices in Heart Failure Patients. In Proceedings of the World Congress on Medical Physics and Biomedical Engineering, Prague, Czech Republic, 3–8 June 2018; pp. 361–365. [Google Scholar]
- Albrecht, P.; Cohen, R.J. Estimation of Heart Rate Power Spectrum Bands from Real-World Data: Dealing with Ectopic Beats and Noisy Data. In Proceedings of the Computers in Cardiology 1988, Washington, DC, USA, 25–28 September 1988; pp. 311–314. [Google Scholar]
- Lippman, N.; Stein, K.M.; Lerman, B.B. Comparison of methods for removal of ectopy in measurement of heart rate variability. Am. J. Physiol. Circ. Physiol. 1994, 267, H411–H418. [Google Scholar] [CrossRef]
- Wen, F.; He, F.-T. An efficient method of addressing ectopic beats: New insight into data preprocessing of heart rate variability analysis. J. Zhejiang Univ. Sci. B 2011, 12, 976–982. [Google Scholar] [CrossRef] [PubMed]
- Mateo, J.; Laguna, P. Analysis of heart rate variability in the presence of ectopic beats using the heart timing signal. IEEE Trans. Biomed. Eng. 2003, 50, 334–343. [Google Scholar] [CrossRef] [PubMed]
- Storck, N.; Ericson, M.; Lindblad, L.E.; Urstad, J.M. Automatic computerized analysis of heart rate variability with digital fil-tering of ectopic beats. Clin. Physiol. 2001, 21, 15–24. [Google Scholar] [CrossRef]
- Keenan, D.B. Detection and correction of ectopic beats for HRV analysis applying discrete wavelet transforms. Int. J. Inf. Technol. 2005, 2, 54–60. [Google Scholar]
- Ahyoung, C.; Hangsik, S. Quantitative analysis of the effect of an ectopic beat on the heart rate variability in the resting con-dition. Front. Physiol. 2018, 9, 922. [Google Scholar]
- Abásolo, D.; Hornero, R.; Espino, P.; Poza, J.; Sánchez, C.I.; de la Rosa, R. Analysis of regularity in the EEG background activity of Alzheimer’s disease patients with Approximate Entropy. Clin. Neurophysiol. 2005, 116, 1826–1834. [Google Scholar] [CrossRef] [PubMed]
- Holzinger, A.; Stocker, C.; Bruschi, M.; Auinger, A.; Silva, H.; Gamboa, H.; Fred, A. On Applying Approximate Entropy to ECG Signals for Knowledge Discovery on the Example of Big Sensor Data. In Proceedings of the International Conference on Active Media Technology, Macau, China, 4–7 December 2012; pp. 646–657. [Google Scholar]
- Richman, J.S.; Moorman, J.R. Physiological time-series analysis using approximate entropy and sample entropy. Am. J. Physiol. Circ. Physiol. 2000, 278, H2039–H2049. [Google Scholar] [CrossRef]
- Pincus, S.M. Approximate entropy as a measure of system complexity. Proc. Natl. Acad. Sci. USA 1991, 88, 2297–2301. [Google Scholar] [CrossRef]
- Sunkaria, R.; Kumar, V.; Saxena, S.C. Sample entropy-based HRV characterisation. Int. J. Med. Eng. Inform. 2012, 4, 398. [Google Scholar] [CrossRef]
- Bolea, J.; Bailón, R.; Pueyo, E. On the Standardization of Approximate Entropy: Multidimensional Approximate Entropy Index Evaluated on Short-Term HRV Time Series. Complexity 2018, 2018, 4953273. [Google Scholar] [CrossRef]
- Bolea, J.; Pueyo, E.; Orini, M.; Bailón, R. Influence of Heart Rate in Non-linear HRV Indices as a Sampling Rate Effect Evaluated on Supine and Standing. Front. Physiol. 2016, 7, 501. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.N.; Li, J.Q.; Xiong, J.L.; Liang, X.Y.; Liu, C.Y. Suppressing the influence of ectopic beats by applying a physical thresh-old-based sample entropy. Entropy 2020, 22, 411. [Google Scholar] [CrossRef] [PubMed]
- Goldberger, A.L.; Amaral, L.A.N.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.-K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet. Circulation 2000, 101, e215–e220. [Google Scholar] [CrossRef]
- Zhao, L.; Wei, S.; Zhang, C.; Zhang, Y.; Jiang, X.; Liu, F.; Liu, C. Determination of Sample Entropy and Fuzzy Measure Entropy Parameters for Distinguishing Congestive Heart Failure from Normal Sinus Rhythm Subjects. Entropy 2015, 17, 6270–6288. [Google Scholar] [CrossRef]
- Xiong, J.; Liang, X.; Zhu, T.; Zhao, L.; Li, J.; Liu, C. A New Physically Meaningful Threshold of Sample Entropy for Detecting Cardiovascular Diseases. Entropy 2019, 21, 830. [Google Scholar] [CrossRef]
- Lu, S.; Chen, X.; Kanters, J.K.; Solomon, I.C.; Chon, K.H. Automatic Selection of the Threshold Value R for Approximate Entropy. IEEE Trans. Biomed. Eng. 2008, 55, 1966–1972. [Google Scholar] [CrossRef] [PubMed]
- Castiglioni, P.; Di Rienzo, M. How the threshold “r” influences approximate entropy analysis of heart-rate variability. In Proceedings of the Computers in Cardiology 2016, Vancouver, BC, Canada, 11–14 September 2016; pp. 561–564. [Google Scholar]
- Lake, D.E.; Moorman, J.R. Accurate estimation of entropy in very short physiological time series: The problem of atrial fibrillation detection in implanted ventricular devices. Am. J. Physiol. Circ. Physiol. 2011, 300, H319–H325. [Google Scholar] [CrossRef] [PubMed]
- Magagnin, V.; Bassani, T.; Bari, V.; Turiel, M.; Maestri, R.; Pinna, G.D.; Porta, A. Non-stationarities significantly distort short-term spectral, symbolic and entropy heart rate variability indices. Physiol. Meas. 2011, 32, 1775–1786. [Google Scholar] [CrossRef]
- Porta, A.; De Maria, B.; Bari, V.; Marchi, A.; Faes, L. Are Nonlinear Model-Free Conditional Entropy Approaches for the Assessment of Cardiac Control Complexity Superior to the Linear Model-Based One? IEEE Trans. Biomed. Eng. 2016, 64, 1287–1296. [Google Scholar] [CrossRef]
- Porta, A.; Castiglioni, P.; Bari, V.; Bassani, T.; Marchi, A.; Cividjian, A.; Quintin, L.; Di Rienzo, M. K-nearest-neighbor condi-tional entropy approach for the assessment of the short-term complexity of cardiovascular control. Physiol. Meas. 2013, 34, 17–33. [Google Scholar] [CrossRef]
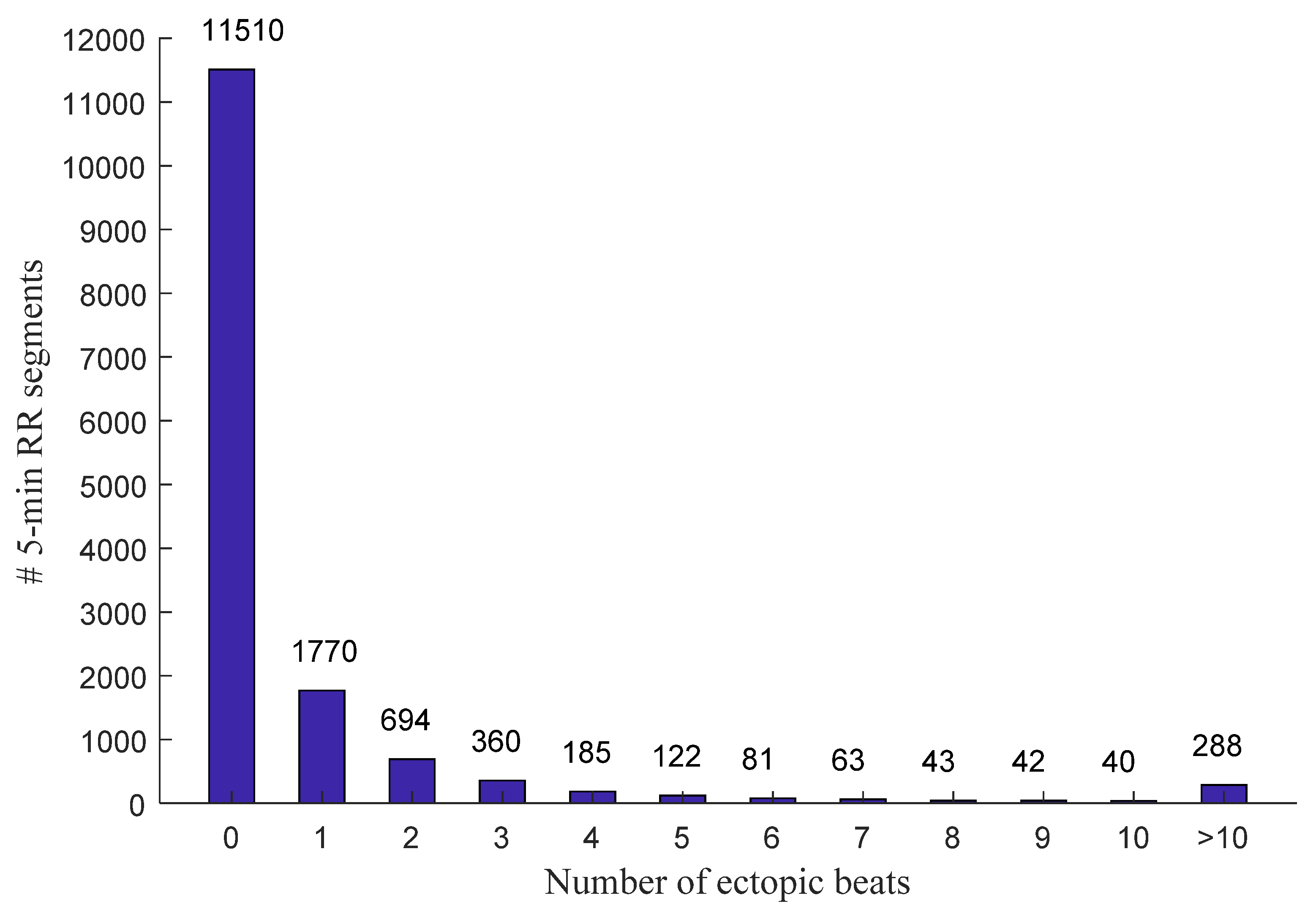

| Record | # Ectopic-Free 5 min Segments | # Ectopic 5 min Segments | Record | # Ectopic-Free 5 min Segments | # Ectopic 5 min Segments |
|---|---|---|---|---|---|
| NSR001 | 212 | 58 | NSR028 | 190 | 95 |
| NSR002 | 134 | 146 | NSR029 | 269 | 18 |
| NSR003 | 247 | 37 | NSR030 | 229 | 58 |
| NSR004 | 245 | 33 | NSR031 | 97 | 191 |
| NSR005 | 89 | 198 | NSR032 | 99 | 188 |
| NSR006 | 239 | 40 | NSR033 | 261 | 14 |
| NSR007 | 203 | 81 | NSR034 | 265 | 18 |
| NSR008 | 237 | 50 | NSR035 | 258 | 29 |
| NSR009 | 262 | 25 | NSR036 | 254 | 28 |
| NSR010 | 168 | 107 | NSR037 | 246 | 29 |
| NSR011 | 195 | 92 | NSR038 | 251 | 4 |
| NSR012 | 247 | 40 | NSR039 | 200 | 87 |
| NSR013 | 249 | 32 | NSR040 | 266 | 17 |
| NSR014 | 175 | 112 | NSR041 | 253 | 29 |
| NSR015 | 263 | 24 | NSR042 | 277 | 10 |
| NSR016 | 245 | 42 | NSR043 | 159 | 123 |
| NSR017 | 22 | 265 | NSR044 | 17 | 270 |
| NSR018 | 73 | 213 | NSR045 | 135 | 149 |
| NSR019 | 254 | 33 | NSR046 | 181 | 94 |
| NSR020 | 172 | 108 | NSR047 | 266 | 22 |
| NSR021 | 275 | 12 | NSR048 | 268 | 21 |
| NSR022 | 236 | 47 | NSR049 | 285 | 3 |
| NSR023 | 253 | 34 | NSR050 | 285 | 3 |
| NSR024 | 15 | 272 | NSR051 | 281 | 6 |
| NSR025 | 166 | 120 | NSR052 | 274 | 10 |
| NSR026 | 243 | 44 | NSR053 | 269 | 1 |
| NSR027 | 280 | 5 | NSR054 | 271 | 8 |
| Record | # Ectopic-Free 5 min Segments | # Ectopic 5 min Segments | Record | # Ectopic-Free 5 min Segments | # Ectopic 5 min Segments |
|---|---|---|---|---|---|
| CHF201 | 240 | 36 | CHF216 | 250 | 14 |
| CHF202 | 97 | 150 | CHF217 | 53 | 228 |
| CHF203 | 75 | 187 | CHF218 | 47 | 217 |
| CHF204 | 0 | 247 | CHF219 | 236 | 28 |
| CHF205 | 31 | 245 | CHF220 | 138 | 143 |
| CHF206 | 11 | 240 | CHF221 | 0 | 276 |
| CHF207 | 1 | 249 | CHF222 | 1 | 274 |
| CHF208 | 31 | 257 | CHF223 | 0 | 274 |
| CHF209 | 70 | 156 | CHF224 | 137 | 150 |
| CHF210 | 16 | 258 | CHF225 | 97 | 121 |
| CHF211 | 275 | 11 | CHF226 | 18 | 257 |
| CHF212 | 0 | 205 | CHF227 | 0 | 275 |
| CHF213 | 7 | 281 | CHF228 | 71 | 204 |
| CHF214 | 0 | 204 | CHF229 | 267 | 20 |
| CHF215 | 110 | 166 |
| # Ectopic Beat | Variances in NSR Subjects (%), std | Variances in CHF Patients (%), std | ||||||
|---|---|---|---|---|---|---|---|---|
| CSDNN | CLF/HF | CSampEn | CPt-SampEn | CSDNN | CLF/HF | CSampEn | CPt-SampEn | |
| 1 | 9.3 | 222.5 | 29.5 | 0.4 | 13.3 | 177.7 | 26.7 | 0.6 |
| 2 | 12.1 | 415.3 | 40.2 | 0.6 | 16.9 | 262.2 | 41.8 | 1.1 |
| 3 | 14.5 | 547.9 | 49.2 | 1.0 | 18.6 | 334.2 | 61.6 | 1.6 |
| 4 | 16.3 | 483.5 | 55.5 | 1.2 | 20.3 | 385.4 | 68.2 | 2.1 |
| 5 | 16.4 | 742.1 | 59.7 | 1.7 | 20.8 | 354.4 | 71.4 | 2.4 |
| 6 | 15.1 | 520.7 | 47.9 | 1.7 | 21.0 | 338.2 | 79.1 | 2.2 |
| 7 | 18.5 | 679.5 | 55.7 | 1.3 | 22.0 | 356.1 | 71.2 | 2.5 |
| 8 | 18.2 | 1001.8 | 65.9 | 1.9 | 21.0 | 375.3 | 91.2 | 2.8 |
| 9 | 21.1 | 818.6 | 77.0 | 2.4 | 22.0 | 571.1 | 86.5 | 2.7 |
| 10 | 19.3 | 1037.3 | 84.8 | 3.2 | 22.2 | 524.1 | 92.1 | 3.4 |
| >10 | 19.7 | 1566.0 | 75.4 | 9.4 | 25.3 | 589.7 | 134.0 | 15.0 |
| Mean | 16.4 | 730.5 | 58.3 | 2.3 | 20.3 | 388.0 | 74.9 | 3.3 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, L.; Li, P.; Li, J.; Liu, C. Influence of Ectopic Beats on Heart Rate Variability Analysis. Entropy 2021, 23, 648. https://doi.org/10.3390/e23060648
Zhao L, Li P, Li J, Liu C. Influence of Ectopic Beats on Heart Rate Variability Analysis. Entropy. 2021; 23(6):648. https://doi.org/10.3390/e23060648
Chicago/Turabian StyleZhao, Lina, Peng Li, Jianqing Li, and Chengyu Liu. 2021. "Influence of Ectopic Beats on Heart Rate Variability Analysis" Entropy 23, no. 6: 648. https://doi.org/10.3390/e23060648
APA StyleZhao, L., Li, P., Li, J., & Liu, C. (2021). Influence of Ectopic Beats on Heart Rate Variability Analysis. Entropy, 23(6), 648. https://doi.org/10.3390/e23060648







