Computational Complexity Reduction of Neural Networks of Brain Tumor Image Segmentation by Introducing Fermi–Dirac Correction Functions
Abstract
1. Introduction
2. Datasets and Methods
2.1. Datasets
2.2. Theoretical Scheme of Fermi–Dirac Correction Function
2.3. Experimental Framework
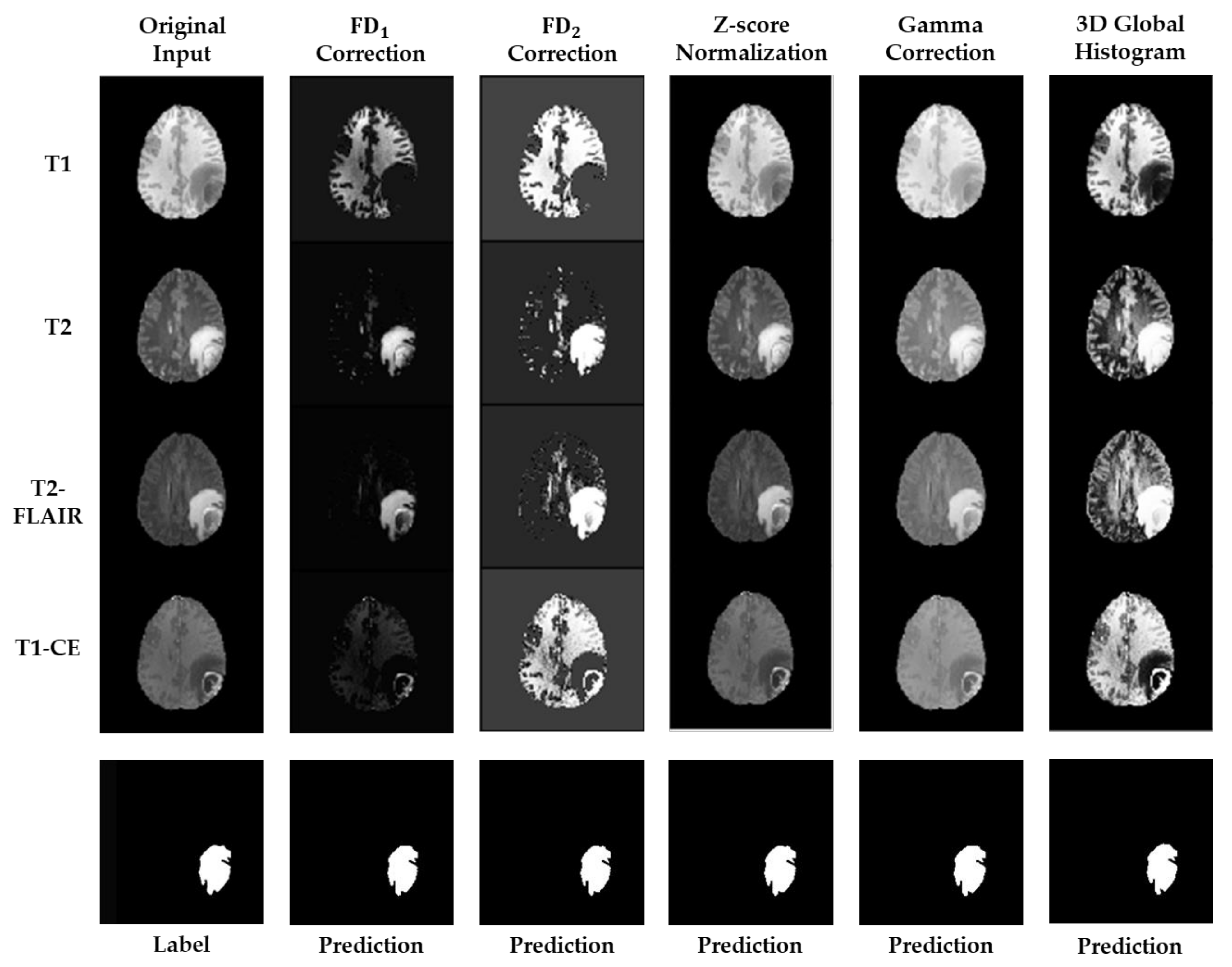
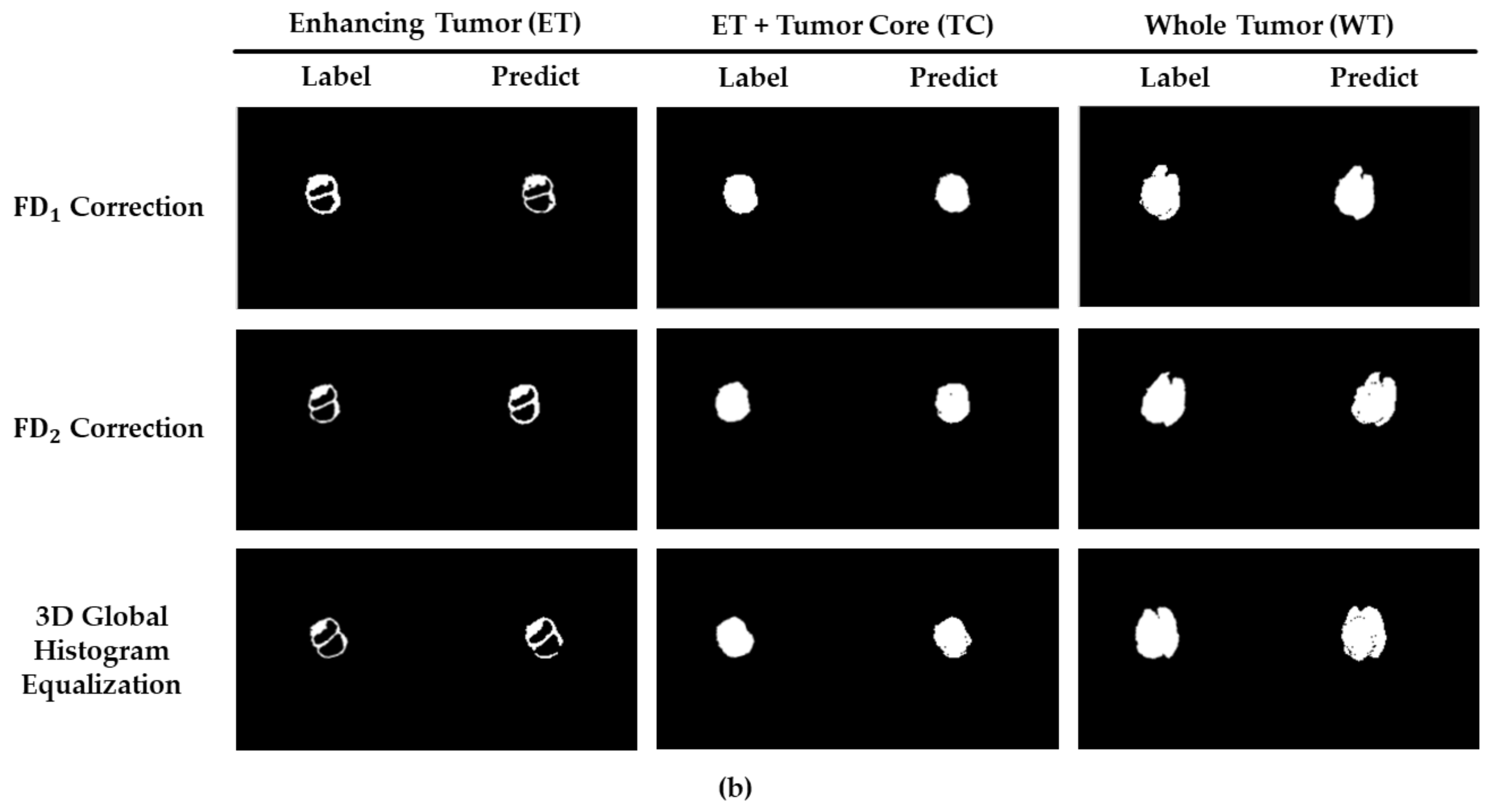
3. Results
4. Discussion and Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bauer, S.; Wiest, R.; Nolte, L.; Reyes, M. A survey of MRI-based medical image analysis for brain tumor studies. Phys. Med. Biol. 2013, 58, R97–R129. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Li, R.; Deng, H.; Wang, L.; Lin, W.; Ji, S.; Shen, D. Deep convolutional neural networks for multi-modality isointense infant brain image segmentation. Neuroimage 2015, 108, 214–224. [Google Scholar] [CrossRef] [PubMed]
- Gurusamy, R.; Subramaniam, D.V. A Machine Learning Approach for MRI Brain Tumor Classification. CMC Comput. Mat. Contin. 2017, 53, 91–108. [Google Scholar]
- Wang, S.; Zhang, Y.; Liu, G.; Phillip, P.; Yuan, T.F. Detection of Alzheimer’s Disease by Three-Dimensional Displacement Field Estimation in Structural Magnetic Resonance Imaging. J. Alzheimers Dis. 2016, 50, 233–248. [Google Scholar] [CrossRef]
- Qiu, A.; Brown, T.; Fischl, B.; Ma, J.; Miller, M.I. Atlas Generation for Subcortical and Ventricular Structures with Its Applications in Shape Analysis. IEEE Trans. Image Process. 2010, 19, 1539–1547. [Google Scholar] [CrossRef]
- Kim, J.; Duchin, Y.; Shamir, R.R.; Patriat, R.; Vitek, J.; Harel, N.; Sapiro, G. Automatic localization of the subthalamic nucleus on patient-specific clinical MRI by incorporating 7 T MRI and machine learning: Application in deep brain stimulation. Hum. Brain Mapp. 2019, 40, 679–698. [Google Scholar] [CrossRef]
- Kreshuk, A.; Straehle, C.N.; Sommer, C.; Koethe, U.; Knott, G.; Hamprecht, F.A. Automated segmentation of synapses in 3D EM data. In Proceedings of the 2011 IEEE International Symposium on Biomedical Imaging: From Nano to Macro, Chicago, IL, USA, 30 March–2 April 2011; pp. 220–223. [Google Scholar]
- Vasilkoski, Z.; Stepanyants, A. Detection of the optimal neuron traces in confocal microscopy images. J. Neurosci. Methods 2009, 178, 197–204. [Google Scholar] [CrossRef]
- Wang, Y.; Narayanaswamy, A.; Tsai, C.L.; Roysam, B. A Broadly Applicable 3-D Neuron Tracing Method Based on Open-Curve Snake. Neuroinformatics 2011, 9, 193–217. [Google Scholar] [CrossRef]
- Park, S.H.; Gao, Y.; Shen, D. Multiatlas-Based Segmentation Editing with Interaction-Guided Patch Selection and Label Fusion. IEEE Trans. Biomed. Eng. 2016, 63, 1208–1219. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Wang, Y.; Yuan, J.; Cheng, Q.; Wang, X.; Carson, P.L. Medical breast ultrasound image segmentation by machine learning. Ultrasonics 2019, 91, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Zarpalas, D.; Gkontra, P.; Daras, P.; Maglaveras, N. Gradient-Based Reliability Maps for ACM-Based Segmentation of Hippocampus. IEEE Trans. Biomed. Eng. 2014, 61, 1015–1026. [Google Scholar] [CrossRef]
- Rajinikanth, V.; Satapathy, S.C.; Fernandes, S.L.; Nachiappan, S. Entropy based segmentation of tumor from brain MR images—A study with teaching learning based optimization. Pattern Recognit. Lett. 2017, 94, 87–95. [Google Scholar] [CrossRef]
- Ma, C.; Luo, G.; Wang, K. Concatenated and Connected Random Forests with Multiscale Patch Driven Active Contour Model for Automated Brain Tumor Segmentation of MR Images. IEEE Trans. Med. Imaging 2018, 37, 1943–1954. [Google Scholar] [CrossRef]
- Milletari, F.; Navab, N.; Ahmadi, S. V-Net: Fully Convolutional Neural Networks for Volumetric Medical Image Segmentation. In Proceedings of the Fourth International Conference on 3D Vision (3DV), Stanford, CA, USA, 25–28 October 2016; pp. 565–571. [Google Scholar]
- Smistad, E.; Falch, T.L.; Bozorgi, M.; Elster, A.C.; Lindseth, F. Medical image segmentation on GPUs—A comprehensive review. Med. Image Anal. 2015, 1, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Papandreou, G.; Kokkinos, I.; Murphy, K.; Yuille, A.L. DeepLab: Semantic Image Segmentation with Deep Convolutional Nets, Atrous Convolution, and Fully Connected CRFs. IEEE Trans. Pattern Anal. Mach. Intell. 2018, 40, 834–848. [Google Scholar] [CrossRef] [PubMed]
- Dasgupta, A.; Singh, S. A fully convolutional neural network based structured prediction approach towards the retinal vessel segmentation. In Proceedings of the IEEE 14th International Symposium on Biomedical Imaging, Melbourne, Australia, 18–21 April 2017; pp. 248–251. [Google Scholar]
- Kim, S.; Luna, M.; Chikontwe, P.; Park, S.H. Two-Step U-Nets for Brain Tumor Segmentation and Random Forest with Radiomics for Survival Time Prediction. In Proceedings of the International MICCAI Brainlesion Workshop, Shenzhen, China, 17 October 2019; pp. 200–209. [Google Scholar]
- Pei, L.; Vidyaratne, L.; Rahman, M.M.; Iftekharuddin, K.M. Context aware deep learning for brain tumor segmentation, subtype classification, and survival prediction using radiology images. Sci. Rep. 2020, 10, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Pei, L.; Vidyaratne, L.; Rahman, M.M.; Shboul, Z.A.; Iftekharuddin, K.M. Multimodal Brain Tumor Segmentation and Survival Prediction Using Hybrid Machine Learning. In Proceedings of the International MICCAI Brainlesion Workshop, Shenzhen, China, 17 October 2019; pp. 73–81. [Google Scholar]
- Rehman, M.U.; Cho, S.; Kim, J.H.; Chong, K.T. BU-Net: Brain Tumor Segmentation Using Modified U-Net Architecture. Electronics 2020, 9, 2203. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Lecture Notes in Computer Science; Navab, N., Panizo, L., Islam, M.M.M., Varela, M.L.R., Ali Mohamad, N., Xiao, Z., Eds.; Springer: Cham, Switzerland, 2015; Volume 9351, pp. 234–241. [Google Scholar]
- Li, X.; Wang, W.; Hu, X.; Yang, J. Selective Kernel Networks. In Proceedings of the 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition, Long Beach, CA, USA, 15–20 June 2019; pp. 510–519. [Google Scholar]
- Hu, J.; Shen, L.; Sun, G. Squeeze-and-Excitation Networks. In Proceedings of the 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–22 June 2018; pp. 7132–7141. [Google Scholar]
- Woo, S.; Park, J.; Lee, J.Y.; Kweon, I.S. CBAM: Convolutional block attention module. In Proceedings of the 15th the European Conference on Computer Vision (ECCV), Munich, Germany, 8–14 September 2018; pp. 3–19. [Google Scholar]
- Abraham, N.; Khan, N.M. A Novel Focal Tversky Loss Function with Improved Attention U-Net for Lesion Segmentation. In Proceedings of the 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019), Venice, Italy, 8–11 April 2019; pp. 683–687. [Google Scholar]
- Schlemper, J.; Oktay, O.; Schaap, M.; Heinrich, M.; Kainz, B.; Glocker, B.; Rueckert, D. Attention gated networks: Learning to leverage salient regions in medical images. Med. Image Anal. 2019, 53, 197–207. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Huang, W.; Dong, P.; Xia, Y.; Wang, S. D-UNet: A dimension-fusion U shape network for chronic stroke lesion segmentation. IEEE/ACM Trans. Comput. Biol. Bioinform. 2019, 1. [Google Scholar] [CrossRef] [PubMed]
- Myronenko, A. 3D MRI Brain Tumor Segmentation Using Autoencoder Regularization. In Lecture Notes in Computer Science; Crimiet, A., Navab, N., Panizo, L., Islam, M.M.M., Varela, M.L.R., Ali Mohamad, N., Eds.; Springer: Cham, Switzerland, 2019; Volume 11384, pp. 311–320. [Google Scholar]
- Dawngliana, M.; Deb, D.; Handique, M.; Roy, S. Automatic brain tumor segmentation in MRI: Hybridized multilevel thresholding and level set. In Proceedings of the 2015 International Symposium on Advanced Computing and Communication (ISACC), Silchar, India, 14–15 September 2015; pp. 219–223. [Google Scholar]
- Cai, H.; Yang, Z.; Cao, X.; Xia, W.; Xu, X. A New Iterative Triclass Thresholding Technique in Image Segmentation. IEEE Trans. Image Process. 2014, 23, 1038–1046. [Google Scholar] [CrossRef]
- Feng, Y.; Zhao, H.; Li, X.; Zhang, X.; Li, H. A multi-scale 3D Otsu thresholding algorithm for medical image segmentation. Digit. Signal Prog. 2017, 60, 186–199. [Google Scholar] [CrossRef]
- Li, Y.; Bai, X.; Jiao, L.; Xue, Y. Partitioned-cooperative quantum-behaved particle swarm optimization based on multilevel thresholding applied to medical image segmentation. Appl. Soft. Comput. 2017, 56, 345–356. [Google Scholar] [CrossRef]
- Dong, X.; Shen, J.; Shao, L.; Gool, L.V. Sub-Markov Random Walk for Image Segmentation. IEEE Trans. Image Process. 2016, 25, 516–527. [Google Scholar] [CrossRef]
- Li, C.; Ding, Z.; Gatenby, J.C.; Metaxas, D.N.; Gore, J.C. A Level Set Method for Image Segmentation in the Presence of Intensity Inhomogeneities with Application to MRI. IEEE Trans. Image Process. 2011, 20, 2007–2016. [Google Scholar]
- Pratondo, A.; Chui, C.-K.; Ong, S.-H. Integrating machine learning with region-based active contour models in medical image segmentation. J. Vis. Commun. Image R. 2017, 43, 1–9. [Google Scholar] [CrossRef]
- Wang, L.; Pan, C. Robust level set image segmentation via a local correntropy-based K-means clustering. Pattern Recognit. 2014, 47, 1917–1925. [Google Scholar] [CrossRef]
- Norouzi, A.; Rahim, M.S.M.; Altameem, A.; Saba, T.; Rad, A.E.; Rehman, A.; Uddin, M. Medical Image Segmentation Methods, Algorithms, and Applications. IETE Tech. Rev. 2014, 31, 199–213. [Google Scholar] [CrossRef]
- Singh, P.; Bhadauria, H.S.; Singh, A. Automatic brain MRI image segmentation using FCM and LSM. In Proceedings of the 3rd International Conference on Reliability, Infocom Technologies and Optimization, Noida, India, 8–10 October 2014; pp. 1–6. [Google Scholar]
- Natarajan, P.; Krishnan, N.; Kenkre, N.S.; Nancy, S.; Singh, B.P. Tumor detection using threshold operation in MRI brain images. In Proceedings of the 2012 IEEE International Conference on Computational Intelligence and Computing Research, Coimbatore, India, 18–20 December 2012; pp. 1–4. [Google Scholar]
- Chen, C.-C.; Juan, H.-H.; Tsai, M.-Y.; Lu, H.H.-S. Unsupervised Learning and Pattern Recognition of Biological Data Structures with Density Functional Theory and Machine Learning. Sci. Rep. 2018, 8, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-C.; Tsai, M.-Y.; Kao, M.-Z.; Lu, H.H.-S. Medical Image Segmentation with Adjustable Computational Complexity Using Data Density Functionals. Appl. Sci. 2019, 9, 1718. [Google Scholar] [CrossRef]
- Menze, B.H.; Jakab, A.; Bauer, S.; Kalpathy-Cramer, J.; Farahani, K.; Kirby, J.; Burren, Y.; Porz, N.; Slotboom, J.; Wiest, R.; et al. The Multimodal Brain Tumor Image Segmentation Benchmark (BRATS). IEEE Trans. Med. Imaging. 2015, 34, 1993–2024. [Google Scholar] [CrossRef] [PubMed]
- Bakas, S.; Akbari, H.; Sotiras, A.; Bilello, M.; Rozycki, M.; Kirby, J.S.; Freymann, J.B.; Farahani, K.; Davatzikos, C. Advancing The Cancer Genome Atlas glioma MRI collections with expert segmentation labels and radiomic features. Nat. Sci. Data 2017, 4, 170117. [Google Scholar] [CrossRef]
- Huang, S.-J.; Wu, C.-J.; Chen, C.-C. Pattern Recognition of Human Postures Using Data Density Functional Method. Appl. Sci. 2018, 8, 1615. [Google Scholar] [CrossRef]
- Amorim, P.; Moraes, T.; Silva, J.; Pedrini, H. 3D Adaptive Histogram Equalization Method for Medical Volumes. In Proceedings of the 13th International Joint Conference on Computer Vision, Imaging and Computer Graphics Theory and Applications, Funchal, Portugal, 27–29 January 2018; Volume 4, pp. 363–370. [Google Scholar]
- Kang, D.; Park, S.; Paik, J. SdBAN: Salient Object Detection Using Bilateral Attention Network with Dice Coefficient Loss. IEEE Access 2020, 8, 104357–104370. [Google Scholar] [CrossRef]
- Eelbode, T.; Bertels, J.; Berman, M.; Vandermeulen, D.; Maes, F.; Bisschops, R.; Blaschko, M.B. Optimization for Medical Image Segmentation: Theory and Practice When Evaluating With Dice Score or Jaccard Index. IEEE Trans. Med. Imaging 2020, 39, 3679–3690. [Google Scholar] [CrossRef] [PubMed]
- Source Codes of D-Unet and Dice Score. Available online: https://github.com/SZUHvern/D-UNet (accessed on 21 October 2020).
- Fawcett, T. An Introduction to ROC Analysis. Pattern Recognit. Lett. 2006, 27, 861–874. [Google Scholar] [CrossRef]
- Chicco, D.; Jurman, G. The advantages of the Matthews correlation coefficient (MCC) over F1 score and accuracy in binary classification evaluation. BMC Genom. 2020, 21, 6-1–6-13. [Google Scholar] [CrossRef] [PubMed]



| Parameter/Function | Value/Method |
|---|---|
| Number of epochs | 30 |
| Batch size | 32 |
| Learning rate (Initial value) | 0.00015 |
| Loss function | 3D soft dice loss function |
| Optimizer | Adam |
| Preprocessing Method | Dice Score (WT only) | Computational Time (min) |
|---|---|---|
| Null | 0.7183 | 91 |
| z-score Normalization | 0.9296 | 142 |
| 3D Global Histogram Equalization | 0.9148 | 84 |
| Gamma Correction 1 | 0.9319 | 93 |
| correction | 0.9431 | 88 |
| correction | 0.9347 | 90 |
| Preprocessing Method | Validation Stage (WT Only) | ||||||
|---|---|---|---|---|---|---|---|
| TP | TN | FP | FN | Accuracy | Recall | Precision | |
| Null | 15,306 | 789,342 | 2003 | 5886 | 0.9901 | 0.71 | 0.85 |
| z-score Normalization | 19,512 | 789,570 | 1776 | 1680 | 0.9957 | 0.90 | 0.90 |
| 3D Global Histogram Equalization | 19,293 | 789,400 | 1947 | 1899 | 0.9952 | 0.89 | 0.89 |
| Gamma Correction 1 | 18,603 | 790,100 | 1248 | 2562 | 0.9953 | 0.86 | 0.92 |
| correction | 19,180 | 790,010 | 1339 | 2012 | 0.9959 | 0.89 | 0.91 |
| correction | 19,471 | 789,500 | 1841 | 1721 | 0.9956 | 0.90 | 0.89 |
| Preprocessing Method | Dice Score | Computational Time (min) | ||
|---|---|---|---|---|
| Training Stage | Validation Stage | |||
| 3D Global Histogram Equalization | WT: | 0.9220 | 0.8002 | 138 |
| TC: | 0.9419 | 0.7688 | ||
| ET: | 0.9142 | 0.6365 | ||
| correction | WT: | 0.9491 | 0.8337 | 140 |
| TC: | 0.9757 | 0.7976 | ||
| ET: | 0.9559 | 0.6802 | ||
| correction | WT: | 0.9336 | 0.8433 | 141 |
| TC: | 0.9773 | 0.8041 | ||
| ET: | 0.9606 | 0.6848 | ||
| Preprocessing Method | Validation Stage | |||||||
|---|---|---|---|---|---|---|---|---|
| TP | TN | FP | FN | Accuracy | Recall | Precision | ||
| 3D Global Histogram Equalization | WT: | 18,910 | 789,880 | 1465 | 2281 | 0.9953 | 0.89 | 0.89 |
| TC: | 7818 | 801,309 | 1490 | 1918 | 0.9958 | 0.80 | 0.84 | |
| ET: | 3141 | 807,914 | 700 | 780 | 0.9981 | 0.80 | 0.82 | |
| correction | WT: | 18,949 | 789,874 | 1470 | 2242 | 0.9954 | 0.89 | 0.93 |
| TC: | 7627 | 801,691 | 1107 | 2110 | 0.9960 | 0.78 | 0.87 | |
| ET: | 3007 | 808,156 | 459 | 914 | 0.9983 | 0.77 | 0.87 | |
| correction | WT: | 19,240 | 789,955 | 1340 | 1952 | 0.9959 | 0.91 | 0.93 |
| TC: | 7966 | 801,841 | 958 | 1772 | 0.9966 | 0.82 | 0.89 | |
| ET: | 3153 | 808,200 | 415 | 768 | 0.9998 | 0.80 | 0.88 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tai, Y.-L.; Huang, S.-J.; Chen, C.-C.; Lu, H.H.-S. Computational Complexity Reduction of Neural Networks of Brain Tumor Image Segmentation by Introducing Fermi–Dirac Correction Functions. Entropy 2021, 23, 223. https://doi.org/10.3390/e23020223
Tai Y-L, Huang S-J, Chen C-C, Lu HH-S. Computational Complexity Reduction of Neural Networks of Brain Tumor Image Segmentation by Introducing Fermi–Dirac Correction Functions. Entropy. 2021; 23(2):223. https://doi.org/10.3390/e23020223
Chicago/Turabian StyleTai, Yen-Ling, Shin-Jhe Huang, Chien-Chang Chen, and Henry Horng-Shing Lu. 2021. "Computational Complexity Reduction of Neural Networks of Brain Tumor Image Segmentation by Introducing Fermi–Dirac Correction Functions" Entropy 23, no. 2: 223. https://doi.org/10.3390/e23020223
APA StyleTai, Y.-L., Huang, S.-J., Chen, C.-C., & Lu, H. H.-S. (2021). Computational Complexity Reduction of Neural Networks of Brain Tumor Image Segmentation by Introducing Fermi–Dirac Correction Functions. Entropy, 23(2), 223. https://doi.org/10.3390/e23020223








