3.2. Preliminaries
Consider a dataset X with n objects, that is, , where each object is described by d categorical features, and the features belong to . Each feature has a finite set of values . Moreover, the values from different features has no intersection such that the number of total feature values is , denoted as m.
For better describing how to calculate the joint probability of two values
and
, we need to introduce some symbols. Let
denotes the feature that
belongs to, and let
denotes the value in feature
f of object
x. Let
denotes the probability of
that calculated by its occurrence frequency. Thus, the joint probability of
and
is
The normalized mutual information, denoted as NMI, is a measurement of the mutual dependence between two vectors [
25]. When we observe one vector, the information of the other vector that we can obtain can be quantified by NMI. Accordingly, the relation between two features
and
could defined as
where
is the relative entropy of joint distribution and marginal distribution, and it is written in
and
are the marginal entropies of feature
and
, respectively. The marginal entropy of the specific feature can be described by
3.3. Learning Value Couplings
The value couplings are learned to reflect the intrinsic relationship between feature values. As we used in the previous work [
5], which is proved effective and intuitional. The relation between values has two aspects: on the one hand, the occurrence frequency of one value is influenced by others; on the other hand, one value could be influenced by its pair value because of their co-occurrence relationship in one objects. For capturing the value couplings based on occurrence and co-occurrence, two coupling functions and their corresponding relation matrices (
) are constructed, respectively.
The occurrence-based value coupling function is
, which represents the occurrence frequency of
influenced by
. In this function, the NMI of two features works as a weight. After constructing the coupling function, the occurrence-based relationship matrix
is constructed by:
The co-occurrence-based value coupling function is
, which indicates the co-occurrence frequency of value
influenced by value
. Note that
and
will never be equal since it is impossible for two values owned by the same feature to co-occur in one object. Thus, the co-occurrence-based relationship matrix
is designed as follow:
The two matrices could be treated as new representations of value couplings based on occurrence and co-occurrence, respectively. Moreover, they could be applied in the following values clustering.
3.4. Hybrid Value Clustering
To capture the value clusters from different perspectives and semantics, we cluster the feature values in different granularities and use the new representation () as the input of the clustering algorithm. To make the cluster results more robust and reflect the data characteristics more precisely, we choose a hybrid clustering strategy, which combines the clustering results of DBSCAN (Density-Based Spatial Clustering of Applications with Noise) and HC (Hierarchical Clustering).
The motivation we use the hybrid clustering strategy is as follows: (i) The metric of DBSCAN is density-based, whereas HC is a partition-based method like K-means. So when we combine the cluster results of the two clustering methods, we can obtain the comprehensive value clusters, which is crucial for capturing the intrinsic data characteristics. (ii) DBSCAN has excellent performance for both convex data sets and non-convex data sets, whereas K-means is not suitable for non-convex data sets. HC can also solve the non-spherical datasets that K-means can not solve. (iii) DBSCAN is not sensitive to noisy points, which means DBSCAN is stable. Consequently, our hybrid clustering strategy suitable for majority data sets; meanwhile, it has a better clustering result.
DBSCAN contains a pairwise parameter , where represents the maximum radius of circles centered on cluster cores, and represents the minimum number of objects in the circle. HC only has one parameter K, which means the number of clusters likes K-means. Therefore, for clustering with different granularities, we set parameters and for and clustering with DBSCAN respectively. Likewise, we set parameters and for clustering with HC.
Parameter selection. In HC clustering, the strategy of choosing
K is demonstrated in Algorithm 1. Instead of giving a fixed value, we use another proportion factor
to decide the maximum cluster number as shown in Steps (3-12) of Algorithm 1. We remove those tiny clusters with only one value from the indicator matrix. When the number of removed clusters is larger than
, we stop increasing
K, whose initial value is 2. In DBSCAN clustering, for a specific
, the parameter
and
are selected based on k-distance graph. For a given
k, the k-distance function is mapping each point to its k-th nearest neighbor. We sort the points of the clustering database in descending order of their k-distance values. Furthermore, we set
to the first point in the first “valley” of the sorted k-distance graph, and we set
as value
k. The value
k is same to the parameter
K of HC. The parameter selection is following [
26].
After clustering, we get four clustering indicator matrices to represent the clustering results. The clustering indicator matrix of is denoted as , the size of which is . Likewise, other indicator matrices are with size . Finally, we concatenate the four indicator matrices into one indicator matrix, denoted as C, which contains the comprehensive information of value clusters. Similar to and , C could also be regarded as a new representation of feature values based on value couplings and value clusters.
3.5. Embedding Values by Autoencoder
Deep Neural Network (DNN) is the hottest topic in machine learning because of its ability in feature extraction. Each middle layer in DNN has the ability of feature learning; it is a self-learning process without any prior knowledge.
After constructing the value clusters indicator matrix
C, which contains comprehensive information, we further learn the couplings between the value clusters. Meanwhile, it requires to build a concise but meaningful value representation. It is intuitional for us to use DNN for value clusters couplings learning, and we use Autoencoder to handle this in unsupervised circumstance. The simple function of Encoder and Decoder are as follows:
The Encoder is used to learn low-dimension representation of the input X. Each layer of the Encoder learns the feature and features couplings of Input X, therefore, contains the complete information of X. The Decoder is implemented to reconstruct X from its input, i.e., . The training process of Autoencoder is minimizing the loss function . After training, the will contain the feature couplings of X and convey similar information with X as well.
The Autoencoder makes it possible for us to capture the heterogeneous value clusters couplings and obtain a relatively low dimension values representation. In our method, we train the Autoencoder by using the value clusters indicator matrix
C as the input. Furthermore, we use the Encoder to calculate a new values representation matrix
in
. The column size
q is determined by
(denoted by
) and hidden factor
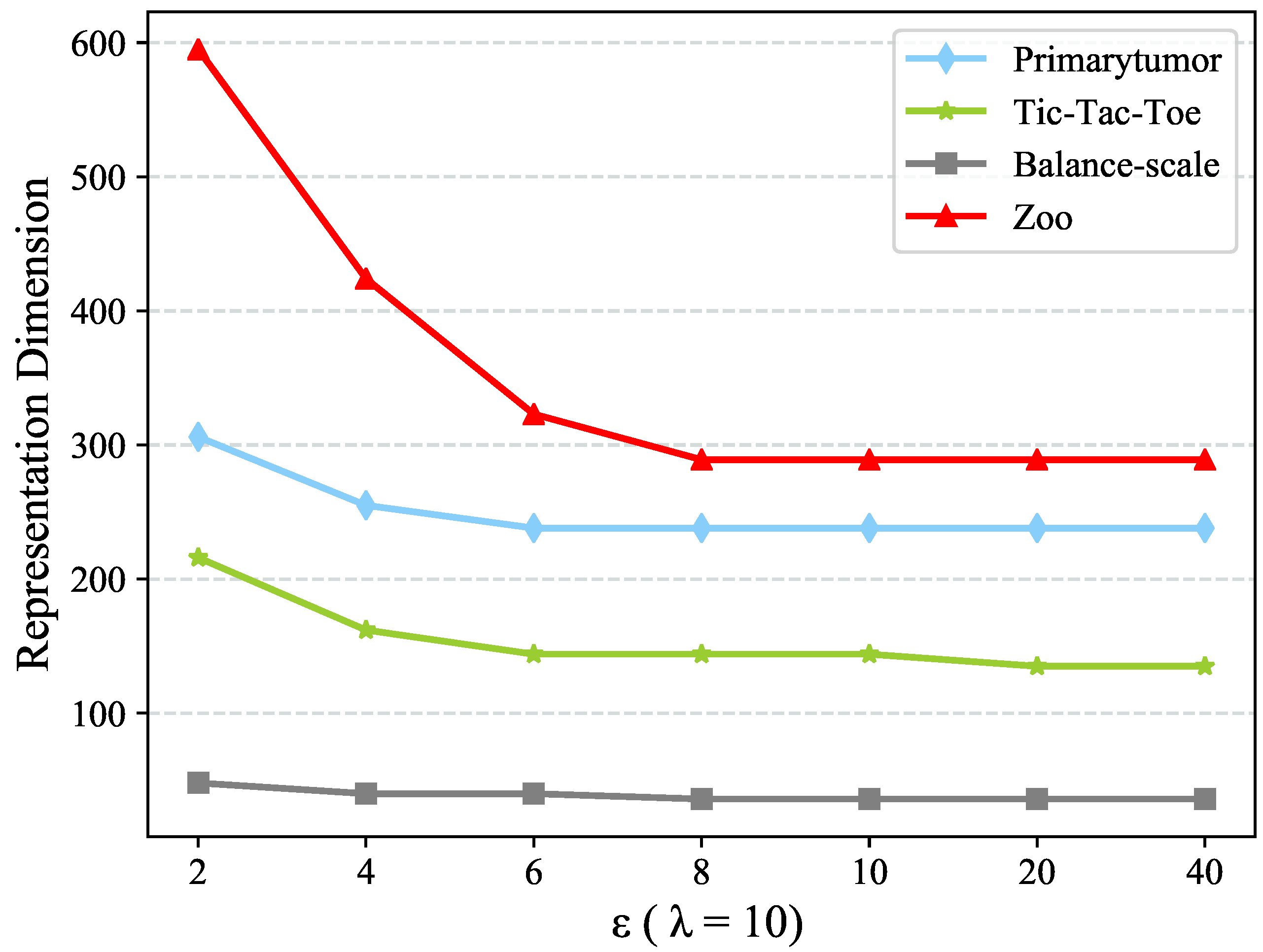
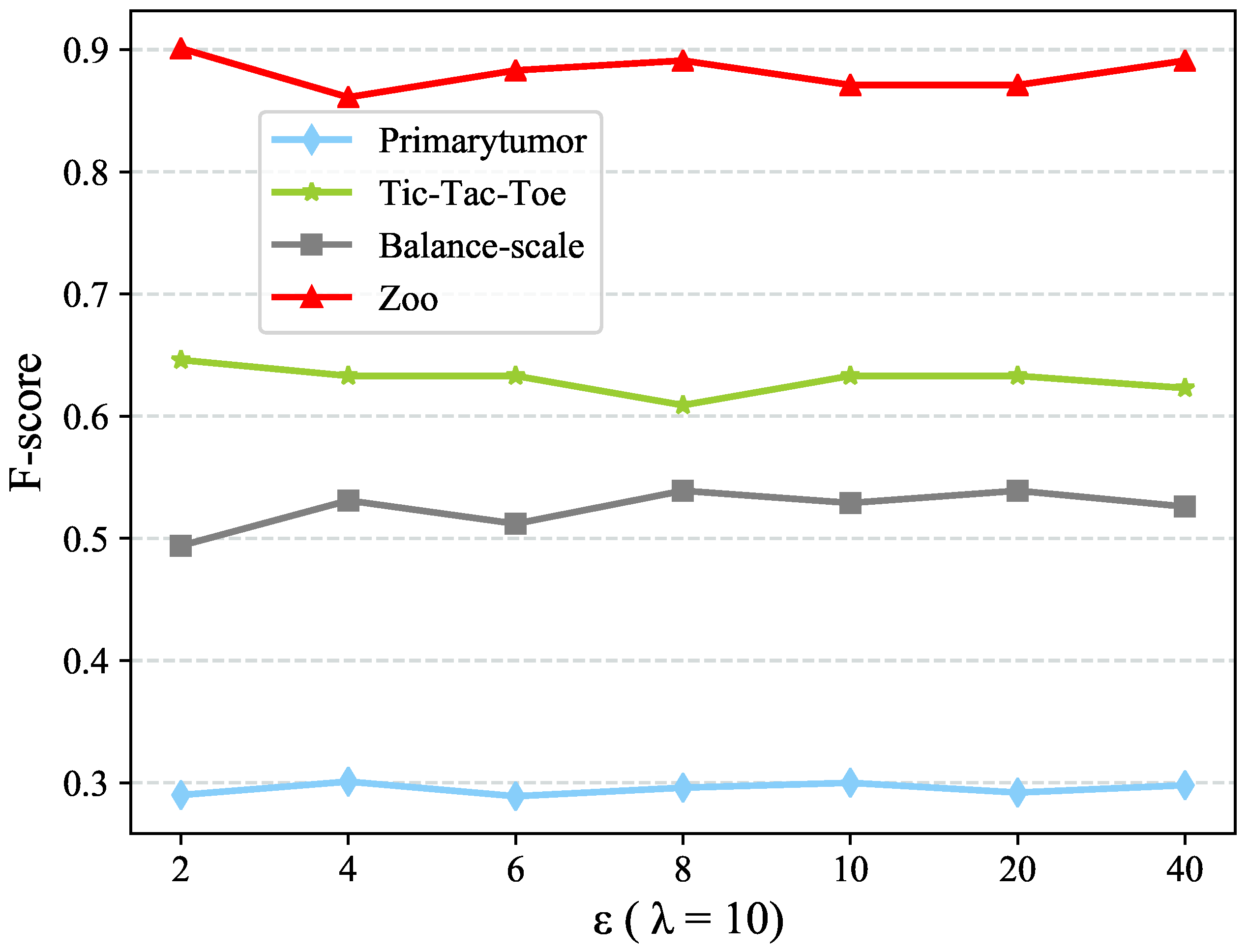
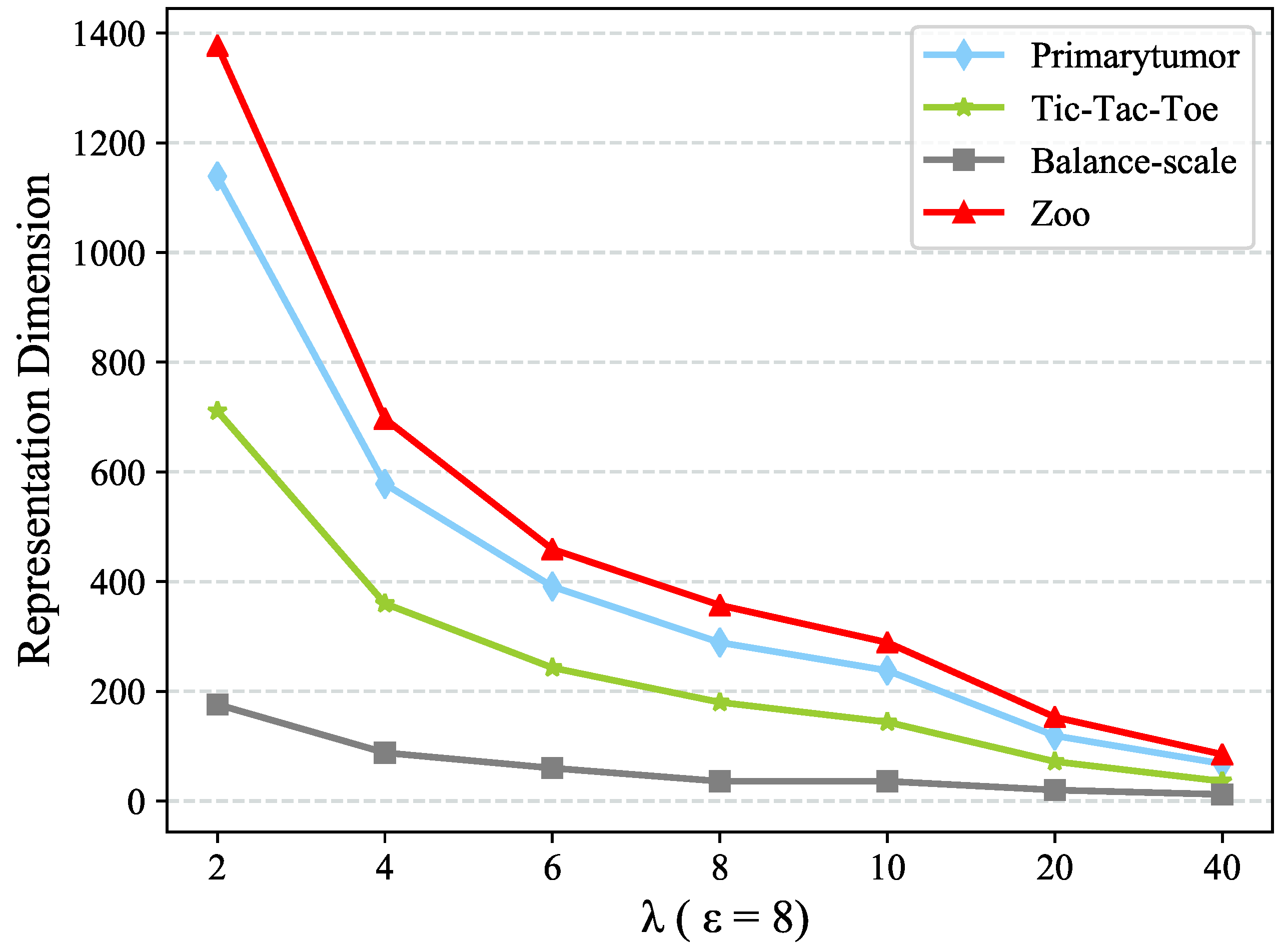
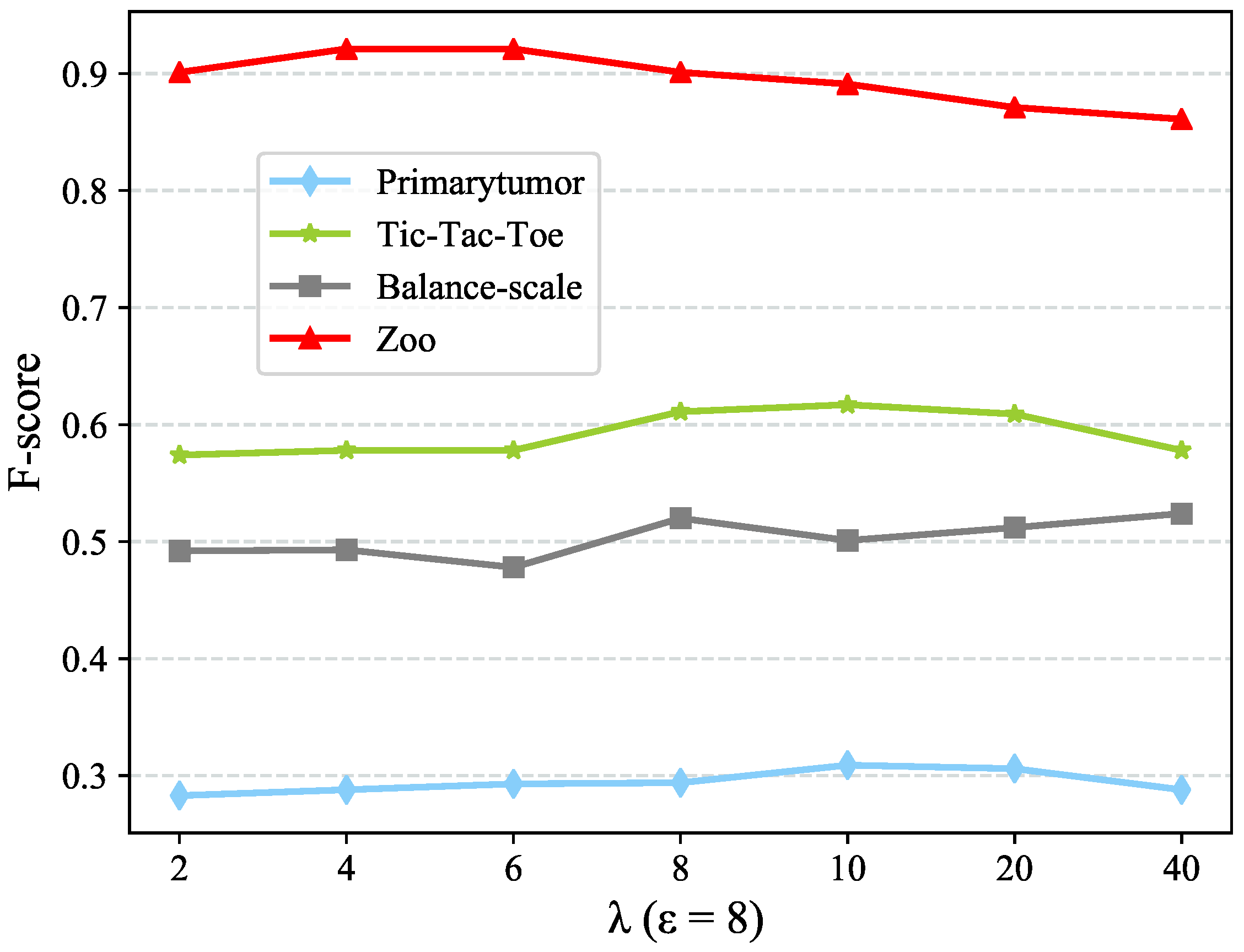
which will be discussed in
Section 4.5. The new value representation
would convey the information of value clusters
C as well as the clusters couplings, which is considered as a concise but meaningful value representation.
3.6. The Embedding for Objects
The final step is to model the objects embedding after we get values representation from Autoencoder. The general function is presented as
The function
in Equation (
7) could be customized to suit for learning task in the following. We concatenate the new values from
to generate the new objects embedding.
The main procedures of CDE++ are presented in Algorithm 1. Algorithm 1 has three inputs, that is, the data set
X, the factor of drop redundancy value clusters
, the hidden factor of Autoencoder
. The algorithm mainly consists of four steps. The first step is to calculate
and
based on occurrence and co-occurrence value coupling function. Then CDE++ utilizes hybrid clustering strategy to cluster values with
and
. The parameter
is used to control the clustering results and determines the time to terminate the clustering process. In the third step, the algorithm uses Autoencoder to learn the couplings of value clusters and generates the concise but meaningful value embedding. The parameter
is a hidden factor of the input dimension and output dimension of Encoder, which indicates the ratio of dimension compression. Finally, CDE++ embeds objects in the data set by concatenating the value embedding.
| Algorithm 1 Object Embedding |
| Input: Dataset X, Parameters and |
| Output: the new representation of X () |
- 1:
Generate and - 2:
Initialize - 3:
fordo - 4:
for do - 5:
Initialize - 6:
Initialize - 7:
while do - 8:
- 9:
Remove the clusters containing only one value and count as r - 10:
end while - 11:
end for - 12:
end for - 13:
Train Autoencoder - 14:
- 15:
- 16:
Return
|
Complexity Analysis. (1) Generating value couplings matrix incurs the complexity of ; (2) Values clustering needs the complexity of ; (3) The complexity of Autoencoder is , where and are total value clusters and the iteration times respectively. (4) Generating numerical object embedding by value embeddings has the complexity of . Accordingly, the total complexity of CDE++ is . In real data sets, the number of values in one feature is generally small, thus, is a little larger than . Meanwhile, is not comparable with . The total number of value clusters is much smaller than m and is iteration times which is manual setting. Therefore, the approximate time complexity of CDE++ is simplified as ).