Robust and Scalable Learning of Complex Intrinsic Dataset Geometry via ElPiGraph
Abstract
1. Introduction
2. Materials and Methods
2.1. General Objective of the ElPiGraph as a Data Approximation and Dimensionality Reduction Method
2.2. Elastic Graphs: Basic Definitions
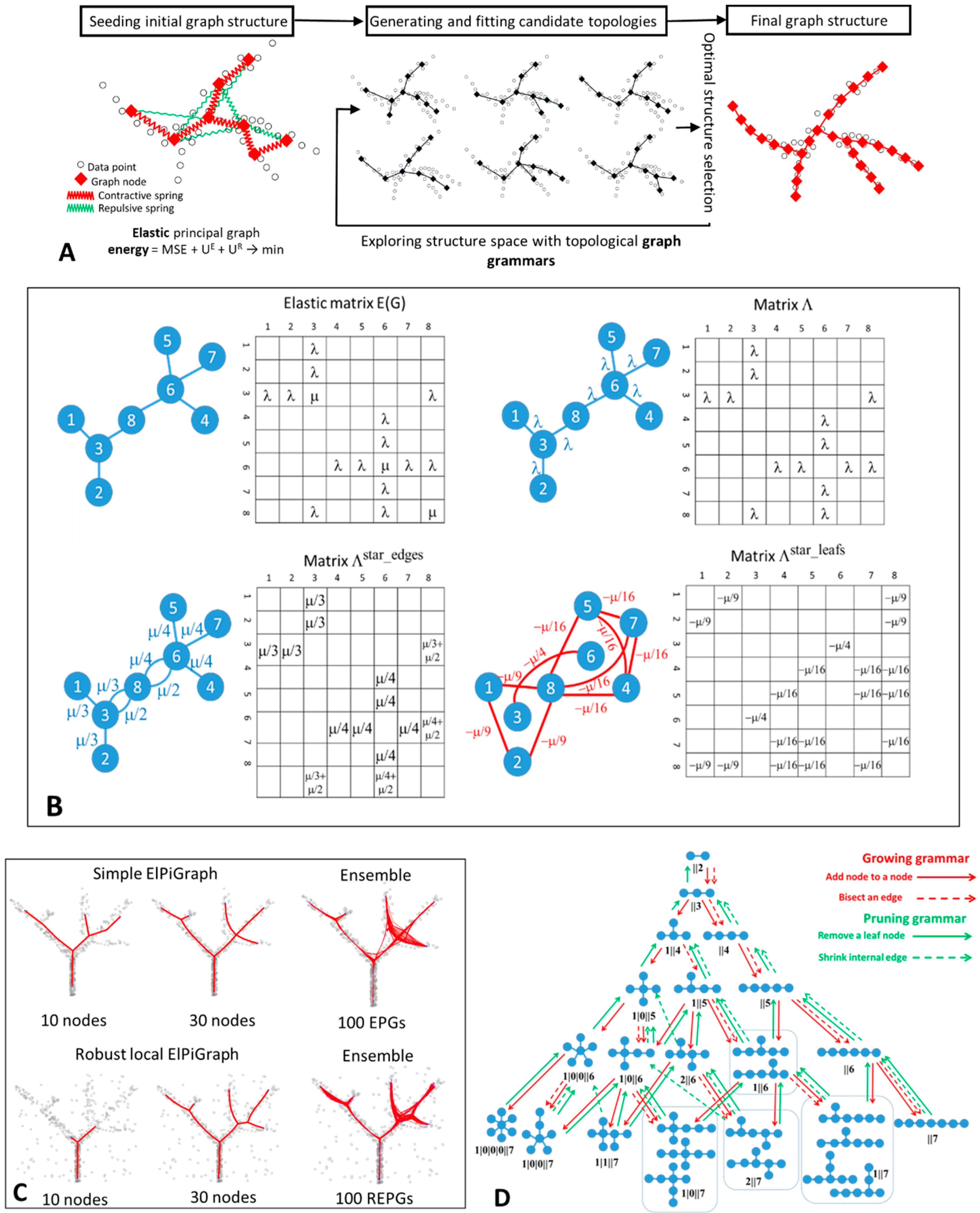
2.3. Elastic Energy Functional and Elastic Matrix
 graph is characterized by a contribution to the elastic energy. The graph with one star
graph is characterized by a contribution to the elastic energy. The graph with one star  will be characterized by a penalty,
will be characterized by a penalty,  by , and
by , and  by .
by .2.4. Elastic Principal Graph Optimization
| Algorithm 1 Base graph optimization for a fixed structure of the elastic graph |
Iterate 3–4 till the map φ does not change more than ε in some appropriate measure of difference between two consecutive iterations. |
2.5. Graph Grammar-Based Approach for Determining the Optimal Graph Topology
| Algorithm 2 graph grammar based optimization of graph structure and embedment |
Repeat 2–4 until the graph contains a required number of nodes. |
| GRAPH GRAMMAR OPERATION ‘BISECT AN EDGE’ | GRAPH GRAMMAR OPERATION ‘ADD NODE TO A NODE’ |
| Applicable to: any edge of the graph | Applicable to: any node of the graph |
| Update of the graph structure: for a given edge {A,B}, connecting nodes A and B, remove {A,B} from the graph, add a new node C, and introduce two new edges {A,C} and {C,B}. | Update of the graph structure: for a given node A, add a new node C, and introduce a new edge {A,C} |
| Update of the elasticity matrix: the elasticity of edges {A,C} and {C,B} equals elasticity of {A,B}. | Update of the elasticity matrix: if A is a leaf node (not a star center) then the elasticity of the edge {A,C} equals to the edge connecting A and its neighbor, the elasticity of the new star with the center in A equals to the elasticity of the star centered in the neighbor of A. If the graph contains only one edge then a predefined values is assigned. else the elasticity of the edge {A,C} is the mean elasticity of all edges in the star with the center in A, the elasticity of a star with the center in A does not change. |
| Update of the graph injection map: C is placed in the mean position between the positions of A and B. | Update of the graph injection map: if A is a leaf node (not a star center) then C is placed at the same distance and the same direction as the edge connecting A and its neighbor, else C is placed in the mean point of all data points for which A is the closest node |
| GRAPH GRAMMAR OPERATION ‘REMOVE A LEAF NODE’ | GRAPH GRAMMAR OPERATION ‘SHRINK INTERNAL EDGE’ |
| Applicable to: node A of the graph with deg(A) = 1 | Applicable to: any edge {A,B} such that deg(A) > 1 and deg(B) > 1. |
| Update of the graph structure: for a given edge {A,B}, connecting nodes A and B, remove edge {A,B} and node A from the graph | Update of the graph structure: for a given edge {A,B}, connecting nodes A and B, remove {A,B}, reattach all edges connecting A with its neighbors to B, remove A from the graph. |
| Update of the elasticity matrix: if B is the center of a 2-star then put zero for the elasticity of the star for B (B becomes a leaf) else do not change the elasticity of the star for B Remove the row and column corresponding to the vertex A | Update of the elasticity matrix: The elasticity of the new star with the center in B becomes the average elasticity of the previously existing stars with the centers in A and B Remove the row and column corresponding to the vertex A |
| Update of the graph injection map: all nodes besides A keep their positions. | Update of the graph injection map: B is placed in the mean position between A and B. |
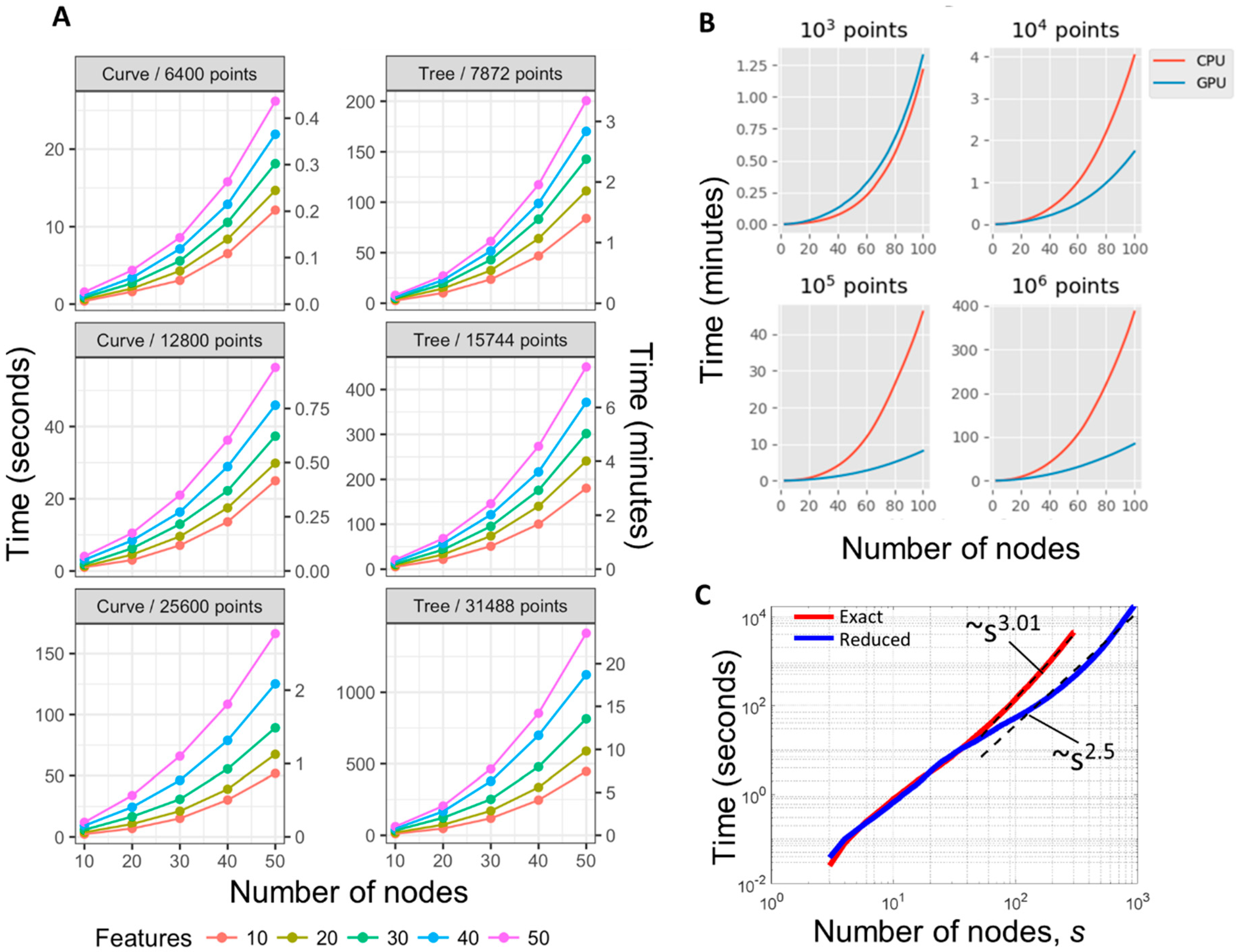
2.6. Computational Complexity of ElPiGraph
- (1)
- Effective reduction of the number of points in the dataset, e.g., by applying pre-clustering. The new datapoints represent the centroids of clusters weighted accordingly to the number of points in each cluster. Alternatively, ElPiGraph can use stochastic approximation approach, by exploiting a subsample of data at each step of growth.
- (2)
- Application of accelerated strategies for the nearest neighbor search step (with the number of neighbors = 1) between graph node positions and data points. In relatively small dimensions and large graphs, the standard use of kdtree method can be already beneficial. The new partitioning can exploit the results of the previous partitioning in order to prevent recomputing all distances, similar to the fast versions of k-means algorithm [52,53]. Various strategies of approximate nearest neighbor search can be also exploited.
- (3)
- Reducing the number of tested candidate graph topologies using simple heuristics. For example, one can test at each application of ‘Bisect an edge’ grammar operation only k longest edges, which most probably will be selected as optimal, with small and fixed k. Similarly, ‘Add a node’ grammar operations can be applied to the several most charged with data points nodes.
2.7. Strategies for Graph Initialization
2.8. Fine-Tuning the Final Graph Structure
2.9. Choice of the Core ElPiGraph Parameters
2.10. Principal Graph Eensembles and Consensus Principal Graph
2.11. Code Availability
- MATLAB from https://github.com/sysbio-curie/ElPiGraph.M
- Python from https://github.com/sysbio-curie/ElPiGraph.P. Note that Python implementation provides support of GPU use.
3. Results
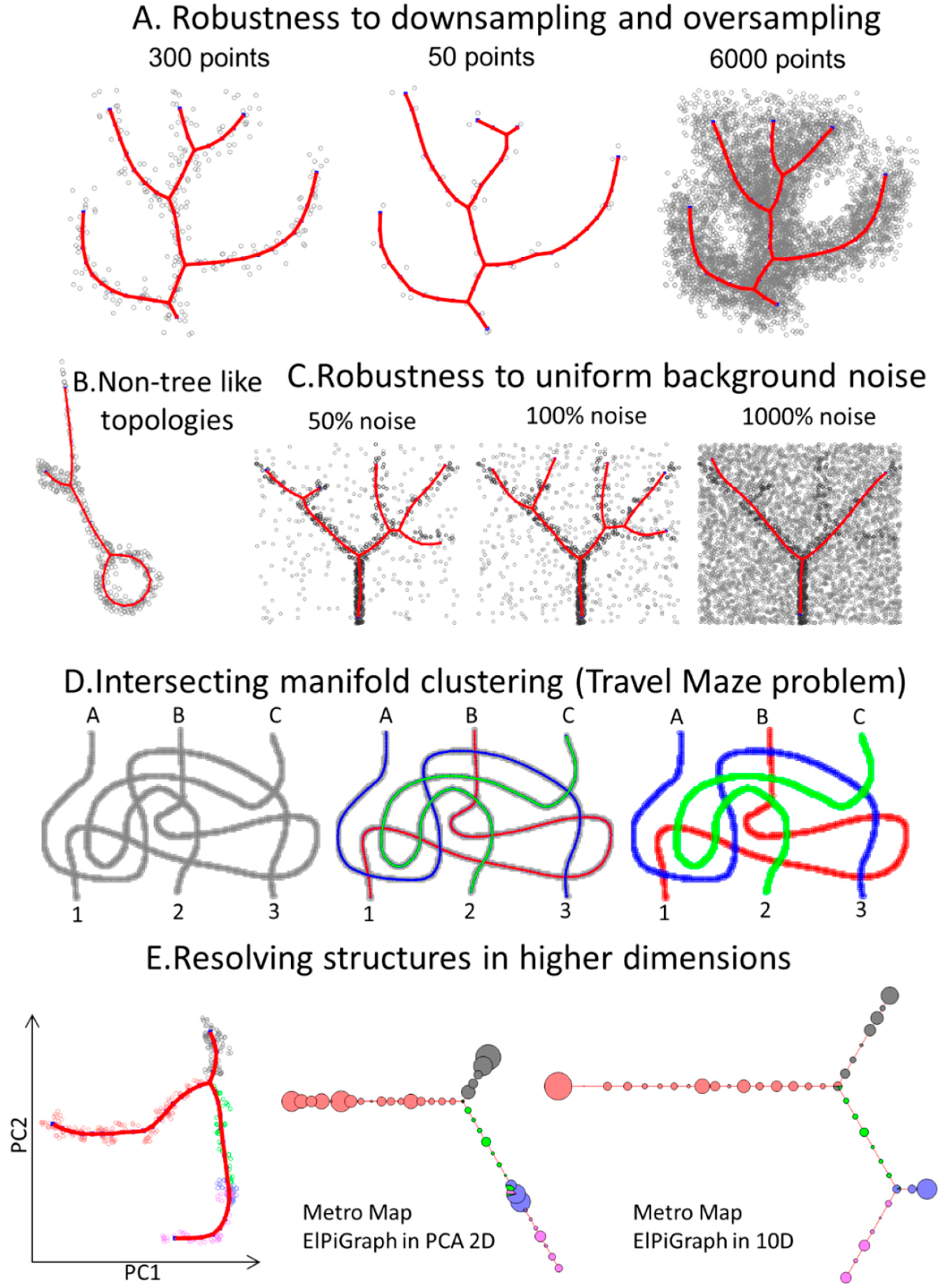
3.1. Approximating Complex Data Topologies in Synthetic Examples
3.2. Benchmarking ElPiGraph
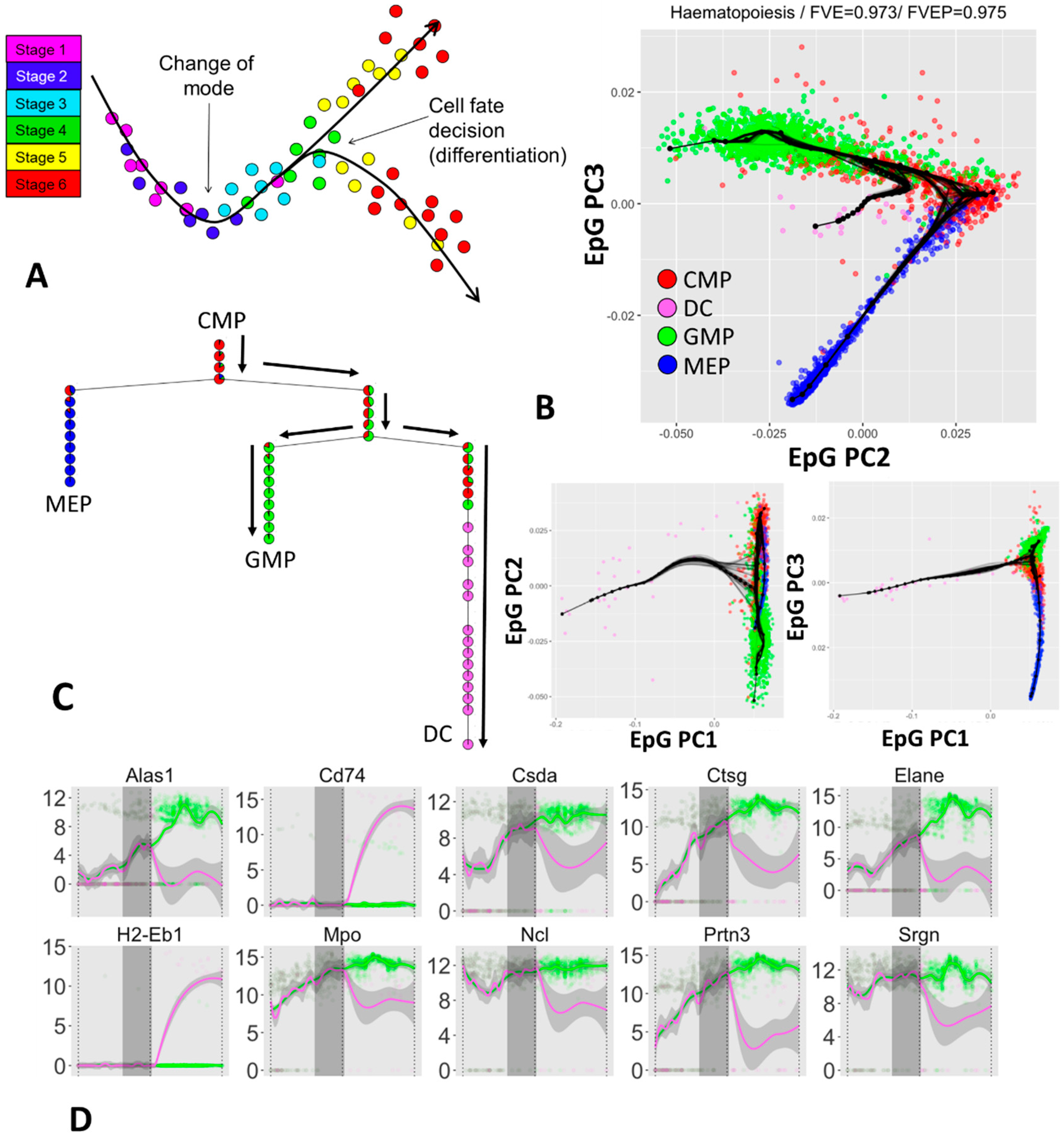
3.3. Inferring Branching Pseudo-Time Cell Trajectories from Single-Cell RNASeq Data via ElPiGraph
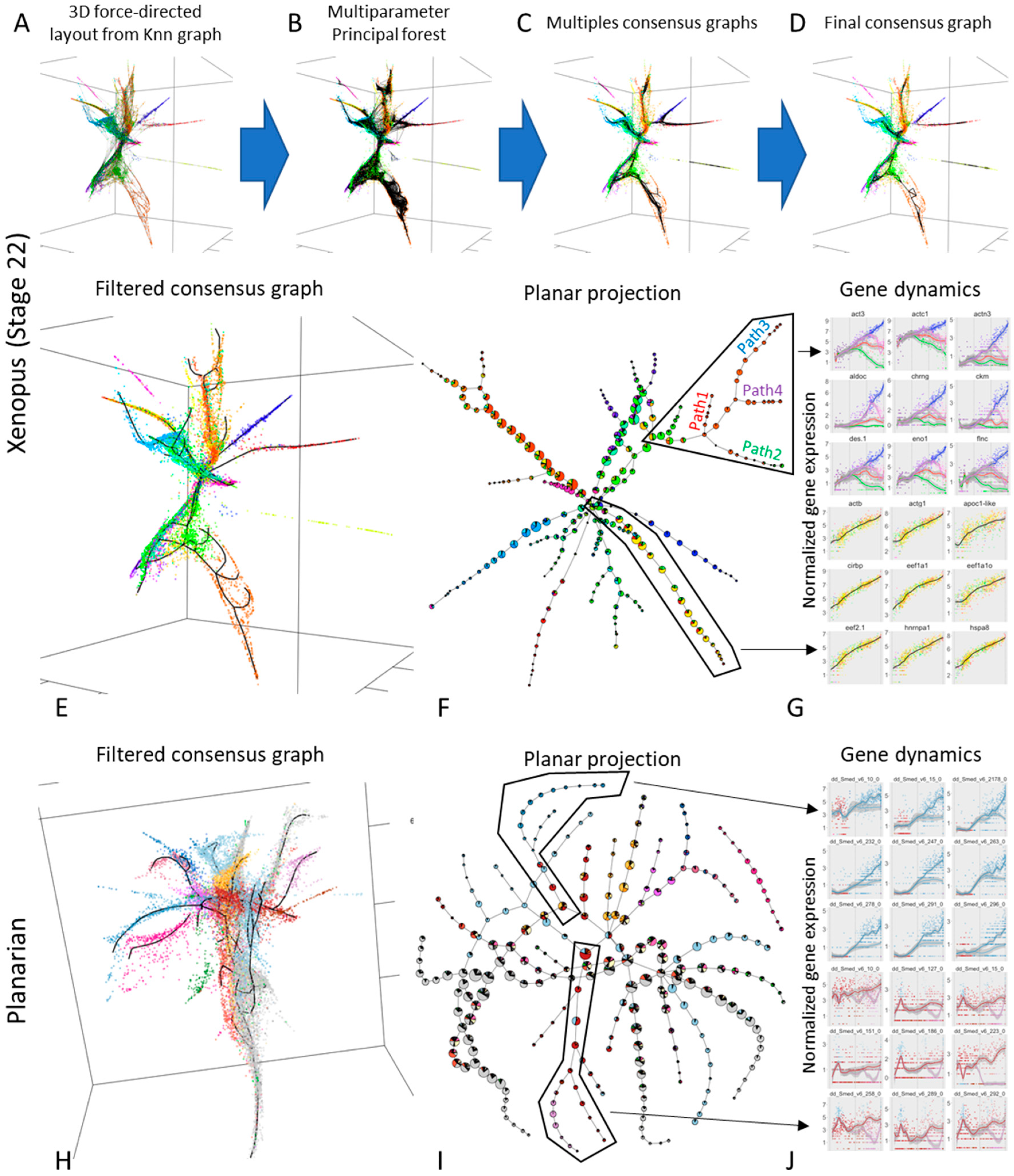
3.4. Approximating the Complex Structure of Development or Differentiation from Single Cell RNASeq Data
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Roux, B.L.; Rouanet, H. Geometric Data Analysis: From Correspondence Analysis to Structured Data Analysis; Springer: Dordrecht, The Netherlands, 2005; ISBN 1402022360. [Google Scholar]
- Gorban, A.; Kégl, B.; Wunch, D.; Zinovyev, A. Principal Manifolds for Data Visualisation and Dimension Reduction; Springer: Berlin/Heidelberg, Germany, 2008; ISBN 978-3-540-73749-0. [Google Scholar]
- Carlsson, G. Topology and data. Bull. Am. Math. Soc. 2009, 46, 255–308. [Google Scholar] [CrossRef]
- Nielsen, F. An elementary introduction to information geometry. arXiv Prepr. 2018, arXiv:1808.08271. [Google Scholar]
- Camastra, F.; Staiano, A. Intrinsic dimension estimation: Advances and open problems. Inf. Sci. 2016, 328, 26–41. [Google Scholar] [CrossRef]
- Albergante, L.; Bac, J.; Zinovyev, A. Estimating the effective dimension of large biological datasets using Fisher separability analysis. In Proceedings of the International Joint Conference on Neural Networks, Budapest, Hungary, 14–19 July 2019. [Google Scholar]
- Gorban, A.N.; Tyukin, I.Y. Blessing of dimensionality: Mathematical foundations of the statistical physics of data. Philos. Trans. R. Soc. A Math. Phys. Eng. Sci. 2018, 376, 20170237. [Google Scholar] [CrossRef]
- Pearson, K.; Pearson, K. On lines and planes of closest fit to systems of points in space. Philos. Mag. 1901, 2, 559–572. [Google Scholar] [CrossRef]
- Kohonen, T. The self-organizing map. Proc. IEEE 1990, 78, 1464–1480. [Google Scholar] [CrossRef]
- Gorban, A.; Zinovyev, A. Elastic principal graphs and manifolds and their practical applications. Computing 2005, 75, 359–379. [Google Scholar] [CrossRef]
- Gorban, A.N.; Zinovyev, A. Principal manifolds and graphs in practice: From molecular biology to dynamical systems. Int. J. Neural Syst. 2010, 20, 219–232. [Google Scholar] [CrossRef]
- Smola, A.J.; Williamson, R.C.; Mika, S.; Scholkopf, B. Regularized Principal Manifolds. Comput. Learn. Theory 1999, 1572, 214–229. [Google Scholar]
- Tenenbaum, J.B.; De Silva, V.; Langford, J.C. A global geometric framework for nonlinear dimensionality reduction. Science 2000, 290, 2319–2323. [Google Scholar] [CrossRef]
- Roweis, S.T.; Saul, L.K. Nonlinear dimensionality reduction by locally linear embedding. Science 2000, 290, 2323–2326. [Google Scholar] [CrossRef]
- Van Der Maaten, L.J.P.; Hinton, G.E. Visualizing high-dimensional data using t-sne. J. Mach. Learn. Res. 2008, 9, 2579–2605. [Google Scholar]
- McInnes, L.; Healy, J.; Saul, N.; Großberger, L. UMAP: Uniform Manifold Approximation and Projection. J. Open Source Softw. 2018, 3, 861. [Google Scholar] [CrossRef]
- Gorban, A.N.; Zinovyev, A.Y. Principal graphs and manifolds. In Handbook of Research on Machine Learning Applications and Trends: Algorithms, Methods, and Techniques; Information Science Reference: Hershey, PA, USA, 2009; pp. 28–59. ISBN 9781605667669. [Google Scholar]
- Zinovyev, A.; Mirkes, E. Data complexity measured by principal graphs. Comput. Math. Appl. 2013, 65, 1471–1482. [Google Scholar] [CrossRef]
- Mao, Q.; Wang, L.; Tsang, I.W.; Sun, Y. Principal Graph and Structure Learning Based on Reversed Graph Embedding. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 2227–2241. [Google Scholar] [CrossRef] [PubMed]
- Gorban, A.N.; Sumner, N.R.; Zinovyev, A.Y. Topological grammars for data approximation. Appl. Math. Lett. 2007, 20, 382–386. [Google Scholar] [CrossRef]
- Gorban, A.N.; Sumner, N.R.; Zinovyev, A.Y. Beyond the concept of manifolds: Principal trees, metro maps, and elastic cubic complexes. In Principal Manifolds for Data Visualization and Dimension Reduction; Lecture Notes in Computational Science and Engineering; Springer: Berlin/Heidelberg, Germany, 2008; Volume 58, pp. 219–237. [Google Scholar]
- Mao, Q.; Yang, L.; Wang, L.; Goodison, S.; Sun, Y. SimplePPT: A simple principal tree algorithm. In Proceedings of the SIAM International Conference on Data Mining, Vancouver, BC, Canada, 30 April–2 May 2015; Society for Industrial and Applied Mathematics Publications: Philadelphia, PA, USA, 2015; pp. 792–800. [Google Scholar]
- Wang, L.; Mao, Q. Probabilistic Dimensionality Reduction via Structure Learning. IEEE Trans. Pattern Anal. Mach. Intell. 2019, 41, 205–219. [Google Scholar] [CrossRef]
- Briggs, J.; Weinreb, C.; Wagner, D.; Megason, S.; Peshki, L.; Kirschner, M.; Klein, A. The dynamics of gene expression in vertebrate embryogenesis at single-cell resolution. Science 2018, 360, eaar5780. [Google Scholar] [CrossRef]
- Wagner, D.; Weinreb, C.; Collins, Z.; Briggs, J.; Megason, S.; Klein, A. Single-cell mapping of gene expression landscapes and lineage in the zebrafish embryo. Science 2018, 360, 981–987. [Google Scholar] [CrossRef]
- Plass, M.; Solana, J.; Wolf, F.A.; Ayoub, S.; Misios, A.; Glažar, P.; Obermayer, B.; Theis, F.J.; Kocks, C.; Rajewsky, N. Cell type atlas and lineage tree of a whole complex animal by single-cell transcriptomics. Science 2018, 360, eaaq1723. [Google Scholar] [CrossRef]
- Furlan, A.; Dyachuk, V.; Kastriti, M.E.; Calvo-Enrique, L.; Abdo, H.; Hadjab, S.; Chontorotzea, T.; Akkuratova, N.; Usoskin, D.; Kamenev, D.; et al. Multipotent peripheral glial cells generate neuroendocrine cells of the adrenal medulla. Science 2017, 357, eaal3753. [Google Scholar] [CrossRef] [PubMed]
- Trapnel, C.; Cacchiarelli, D.; Grimsby, J.; Pokharel, P.; Li, S.; Morse, M.; Lennon, N.J.; Livak, K.J.; Mikkelsen, T.S.; Rinn, J.L. Pseudo-temporal ordering of individual cells reveals dynamics and regulators of cell fate decisions. Nat. Biotechnol. 2012, 29, 997–1003. [Google Scholar]
- Athanasiadis, E.I.; Botthof, J.G.; Andres, H.; Ferreira, L.; Lio, P.; Cvejic, A. Single-cell RNA-sequencing uncovers transcriptional states and fate decisions in hematopoiesis. Nat. Commun. 2017, 8, 2045. [Google Scholar] [CrossRef] [PubMed]
- Velten, L.; Haas, S.F.; Raffel, S.; Blaszkiewicz, S.; Islam, S.; Hennig, B.P.; Hirche, C.; Lutz, C.; Buss, E.C.; Nowak, D.; et al. Human hematopoietic stem cell lineage commitment is a continuous process. Nat. Cell Biol. 2017, 19, 271–281. [Google Scholar] [CrossRef]
- Tirosh, I.; Venteicher, A.S.; Hebert, C.; Escalante, L.E.; Patel, A.P.; Yizhak, K.; Fisher, J.M.; Rodman, C.; Mount, C.; Filbin, M.G.; et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 2016, 539, 309–313. [Google Scholar] [CrossRef]
- Cannoodt, R.; Saelens, W.; Saeys, Y. Computational methods for trajectory inference from single-cell transcriptomics. Eur. J. Immunol. 2016, 46, 2496–2506. [Google Scholar] [CrossRef] [PubMed]
- Moon, K.R.; Stanley, J.; Burkhardt, D.; van Dijk, D.; Wolf, G.; Krishnaswamy, S. Manifold learning-based methods for analyzing single-cell RNA-sequencing data. Curr. Opin. Syst. Biol. 2017, 7, 36–46. [Google Scholar] [CrossRef]
- Saelens, W.; Cannoodt, R.; Todorov, H.; Saeys, Y. A comparison of single-cell trajectory inference methods. Nat. Biotechnol. 2019, 37, 547–554. [Google Scholar] [CrossRef]
- Qiu, X.; Mao, Q.; Tang, Y.; Wang, L.; Chawla, R.; Pliner, H.A.; Trapnell, C. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 2017, 14, 979–982. [Google Scholar] [CrossRef]
- Drier, Y.; Sheffer, M.; Domany, E. Pathway-based personalized analysis of cancer. Proc. Natl. Acad. Sci. USA 2013, 110, 6388–6393. [Google Scholar] [CrossRef]
- Welch, J.D.; Hartemink, A.J.; Prins, J.F. SLICER: Inferring branched, nonlinear cellular trajectories from single cell RNA-seq data. Genome Biol. 2016, 17, 106. [Google Scholar] [CrossRef]
- Setty, M.; Tadmor, M.D.; Reich-Zeliger, S.; Angel, O.; Salame, T.M.; Kathail, P.; Choi, K.; Bendall, S.; Friedman, N.; Pe’Er, D. Wishbone identifies bifurcating developmental trajectories from single-cell data. Nat. Biotechnol. 2016, 34, 637–645. [Google Scholar] [CrossRef]
- Kégl, B.; Krzyzak, A. Piecewise linear skeletonization using principal curves. IEEE Trans. Pattern Anal. Mach. Intell. 2002, 24, 59–74. [Google Scholar] [CrossRef]
- Hastie, T.; Stuetzle, W. Principal curves. J. Am. Stat. Assoc. 1989, 84, 502–516. [Google Scholar] [CrossRef]
- Kégl, B.; Krzyzak, A.; Linder, T.; Zeger, K. A polygonal line algorithm for constructing principal curves. In Proceedings of the Advances in Neural Information Processing Systems, Denver, CO, USA, 29 November–4 December 1999. [Google Scholar]
- Gorban, A.N.; Rossiev, A.A.; Wunsch, D.C.; Gorban, A.A.; Rossiev, D.C. Wunsch II. In Proceedings of the International Joint Conference on Neural Networks, Washington, DC, USA, 10–16 July 1999. [Google Scholar]
- Zinovyev, A. Visualization of Multidimensional Data; Krasnoyarsk State Technical Universtity: Krasnoyarsk, Russia, 2000. [Google Scholar]
- Gorban, A.N.; Zinovyev, A. Method of elastic maps and its applications in data visualization and data modeling. Int. J. Comput. Anticip. Syst. Chaos 2001, 12, 353–369. [Google Scholar]
- Delicado, P. Another Look at Principal Curves and Surfaces. J. Multivar. Anal. 2001, 77, 84–116. [Google Scholar] [CrossRef]
- Gorban, A.N.; Mirkes, E.; Zinovyev, A.Y. Robust principal graphs for data approximation. Arch. Data Sci. 2017, 2, 1–16. [Google Scholar]
- Chen, H.; Albergante, L.; Hsu, J.Y.; Lareau, C.A.; Lo Bosco, G.; Guan, J.; Zhou, S.; Gorban, A.N.; Bauer, D.E.; Aryee, M.J.; et al. Single-cell trajectories reconstruction, exploration and mapping of omics data with STREAM. Nat. Commun. 2019, 10, 1–14. [Google Scholar] [CrossRef]
- Parra, R.G.; Papadopoulos, N.; Ahumada-Arranz, L.; Kholtei, J.E.; Mottelson, N.; Horokhovsky, Y.; Treutlein, B.; Soeding, J. Reconstructing complex lineage trees from scRNA-seq data using MERLoT. Nucleic Acids Res. 2019, 47, 8961–8974. [Google Scholar] [CrossRef]
- Wolf, F.A.; Hamey, F.; Plass, M.; Solana, J.; Dahlin, J.S.; Gottgens, B.; Rajewsky, N.; Simon, L.; Theis, F.J. PAGA: Graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 2019, 20, 59. [Google Scholar] [CrossRef]
- Cuesta-Albertos, J.A.; Gordaliza, A.; Matrán, C. Trimmed k-means: An attempt to robustify quantizers. Ann. Stat. 1997, 25, 553–576. [Google Scholar] [CrossRef]
- Gorban, A.N.; Mirkes, E.M.; Zinovyev, A. Piece-wise quadratic approximations of arbitrary error functions for fast and robust machine learning. Neural Netw. 2016, 84, 28–38. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Elkan, C. Using the Triangle Inequality to Accelerate k-Means. In Proceedings of the Twentieth International Conference on Machine Learning, Washington, DC, USA, 21–24 August 2003. [Google Scholar]
- Hamerly, G. Making k-means even faster. In Proceedings of the 10th SIAM International Conference on Data Mining, Columbus, OH, USA, 29 April–1 May 2010. [Google Scholar]
- Politis, D.; Romano, J.; Wolf, M. Subsampling; Springer: New York, NY, USA, 1999. [Google Scholar]
- Babaeian, A.; Bayestehtashk, A.; Bandarabadi, M. Multiple manifold clustering using curvature constrained path. PLoS ONE 2015, 10, e0137986. [Google Scholar] [CrossRef] [PubMed]
- Bac, J.; Zinovyev, A. Lizard Brain: Tackling Locally Low-Dimensional Yet Globally Complex Organization of Multi-Dimensional Datasets. Front. Neurorobot. 2020, 13, 110. [Google Scholar] [CrossRef] [PubMed]
- Mao, Q.; Wang, L.; Goodison, S.; Sun, Y. Dimensionality Reduction Via Graph Structure Learning. In Proceedings of the 21th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Sydney, Australia, 10–13 August 2015; pp. 765–774. [Google Scholar]
- Aynaud, M.M.; Mirabeau, O.; Gruel, N.; Grossetête, S.; Boeva, V.; Durand, S.; Surdez, D.; Saulnier, O.; Zaïdi, S.; Gribkova, S.; et al. Transcriptional Programs Define Intratumoral Heterogeneity of Ewing Sarcoma at Single-Cell Resolution. Cell Rep. 2020, 30, 1767–1779. [Google Scholar] [CrossRef]
- Paul, F.; Arkin, Y.; Giladi, A.; Jaitin, D.A.; Kenigsberg, E.; Keren-Shaul, H.; Winter, D.; Lara-Astiaso, D.; Gury, M.; Weiner, A.; et al. Transcriptional Heterogeneity and Lineage Commitment in Myeloid Progenitors. Cell 2015, 163, 1663–1677. [Google Scholar] [CrossRef]
- Guo, G.; Pinello, L.; Han, X.; Lai, S.; Shen, L.; Lin, T.W.; Zou, K.; Yuan, G.C.; Orkin, S.H. Serum-Based Culture Conditions Provoke Gene Expression Variability in Mouse Embryonic Stem Cells as Revealed by Single-Cell Analysis. Cell Rep. 2016, 14, 956–965. [Google Scholar] [CrossRef]
- Zhang, Z.; Wang, J. MLLE: Modified Locally Linear Embedding Using Multiple Weights. Adv. Neural Inf. Process. Syst. 2007, 19(19), 1593–1600. [Google Scholar]
- Weinreb, C.; Wolock, S.; Klein, A.M. SPRING: A kinetic interface for visualizing high dimensional single-cell expression data. Bioinformatics 2017, 34, 1246–1248. [Google Scholar] [CrossRef]
- Gorban, A.N.; Zinovyev, A. Visualization of Data by Method of Elastic Maps and its Applications in Genomics, Economics and Sociology. IHES Preprints. 2001. IHES/M/01/36. Available online: http://cogprints.org/3088/ (accessed on 11 March 2011).
- Gorban, A.N.; Zinovyev, A.Y.; Wunsch, D.C. Application of the method of elastic maps in analysis of genetic texts. In Proceedings of the International Joint Conference on Neural Networks, Portland, OR, USA, 20–24 July 2003; p. 3. [Google Scholar]
- Failmezger, H.; Jaegle, B.; Schrader, A.; Hülskamp, M.; Tresch, A. Semi-automated 3D Leaf Reconstruction and Analysis of Trichome Patterning from Light Microscopic Images. PLoS Comput. Biol. 2013, 9. [Google Scholar] [CrossRef] [PubMed]
- Cohen, D.P.A.; Martignetti, L.; Robine, S.; Barillot, E.; Zinovyev, A.; Calzone, L. Mathematical Modelling of Molecular Pathways Enabling Tumour Cell Invasion and Migration. PLoS Comput. Biol. 2015, 11, e1004571. [Google Scholar] [CrossRef] [PubMed]
| Element | Initial Publication | Principal Advances |
|---|---|---|
| Principal curves | Hastie and Stuelze, 1989 [24] | Definition of principal curves based on self-consistency |
| Piece-wise linear principal curves | Kégl et al., 1999 [25] | Length constrained principal curves, polygonal line algorithm |
| Elastic energy functional for elastic principal curves and manifolds | Gorban, Rossiev, Wunsch II, 1999 [26] | Fast iterative splitting algorithm to minimize the elastic energy, based on sequence of solutions of simple quadratic minimization problems |
| Method of elastic maps | Zinovyev, 2000 [27], Gorban and Zinovyev, 2001 [28] | Construction of principal manifold approximations possessing various topologies |
| Principal Oriented Points | Delicado, 2001 [29] | Principal curves passing through a set of principal oriented points |
| Principal graphs specialized for image skeletonization | Kégl and Krzyzak, 2002 [30] | Coining the term principal graph, an algorithm extending the polygonal line algorithm, specialized on image skeletonization |
| Self-assembling principal graphs | Gorban and Zinovyev, 2005 [10] | Simple principal graph algorithm, based on application of elastic map method, specialized on image skeletonization |
| General purpose elastic principal graphs | Gorban, Sumner, Zinovyev, 2007 [20] | Suggesting the principle of (pluri-)harmonic graph embedding, coining the terms ‘principal tree’ and ‘principal cubic complex’ with algorithms for their construction |
| Topological grammars | Gorban, Sumner, Zinovyev, 2007 [20] | Exploring multiple principal graph topologies via gradient descent-like search in the space of admissible structures |
| Explicit control of principal graph complexity | Gorban and Zinovyev, 2009 [17] | Introducing three types of principal graph complexity and ways to constrain it |
| Regularized principal graphs | Mao et al., 2015 [22] | Formulating reverse graph embedding problem. Suggesting SimplePPT algorithm. Further development in Mao et al., 2017 [19] |
| Robust principal graphs | Gorban, Mirkes, Zinovyev, 2015 [31] | Using trimmed version of the mean squared error, resulting in the ‘local’ growth of the principal graphs and robustness to background noise |
| Domain-specific adaptations of principal graphs | Qiu et al., 2017 [32] Chen et al., 2019 [33] Para et al., 2019 [34] | Use of principal graphs in single cell data analysis as a part of pipelines Monocle, STREAM, MERLoT. Introducing heuristics for problem-specific initializations of principal graphs. Benchmarked in Saelens et al., 2019 [35]. |
| Partition-based graph abstraction | Wolf et al., 2019 [36] | Dealing with non-tree like topologies, large-scale and multi-scale data analysis using graphs |
| Excessive branching control | This publication | Introducing a penalty for excessive branching of elastic principal graphs (α parameter) |
| Principal graph ensemble approach | This publication | Estimating confidence of branching point positions, constructing consensus principal graphs |
| Reducing the computational complexity of elastic principal graphs | This publication | Accelerated procedures and heuristics in order to enable constructing large principal graphs |
| Use of GPUs for constructing elastic principal graphs | This publication | Scalable Python and R implementations of elastic principal graphs (ElPiGraph), improved scalability and introducing various plotting functions |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Albergante, L.; Mirkes, E.; Bac, J.; Chen, H.; Martin, A.; Faure, L.; Barillot, E.; Pinello, L.; Gorban, A.; Zinovyev, A. Robust and Scalable Learning of Complex Intrinsic Dataset Geometry via ElPiGraph. Entropy 2020, 22, 296. https://doi.org/10.3390/e22030296
Albergante L, Mirkes E, Bac J, Chen H, Martin A, Faure L, Barillot E, Pinello L, Gorban A, Zinovyev A. Robust and Scalable Learning of Complex Intrinsic Dataset Geometry via ElPiGraph. Entropy. 2020; 22(3):296. https://doi.org/10.3390/e22030296
Chicago/Turabian StyleAlbergante, Luca, Evgeny Mirkes, Jonathan Bac, Huidong Chen, Alexis Martin, Louis Faure, Emmanuel Barillot, Luca Pinello, Alexander Gorban, and Andrei Zinovyev. 2020. "Robust and Scalable Learning of Complex Intrinsic Dataset Geometry via ElPiGraph" Entropy 22, no. 3: 296. https://doi.org/10.3390/e22030296
APA StyleAlbergante, L., Mirkes, E., Bac, J., Chen, H., Martin, A., Faure, L., Barillot, E., Pinello, L., Gorban, A., & Zinovyev, A. (2020). Robust and Scalable Learning of Complex Intrinsic Dataset Geometry via ElPiGraph. Entropy, 22(3), 296. https://doi.org/10.3390/e22030296








